Heterogeneity of the Abnormal Prion Protein (PrPSc) of the Chandler Scrapie Strain
Abstract
:1. Introduction
2. Results and Discussion
2.1. Size Distribution of PrPSc Aggregates
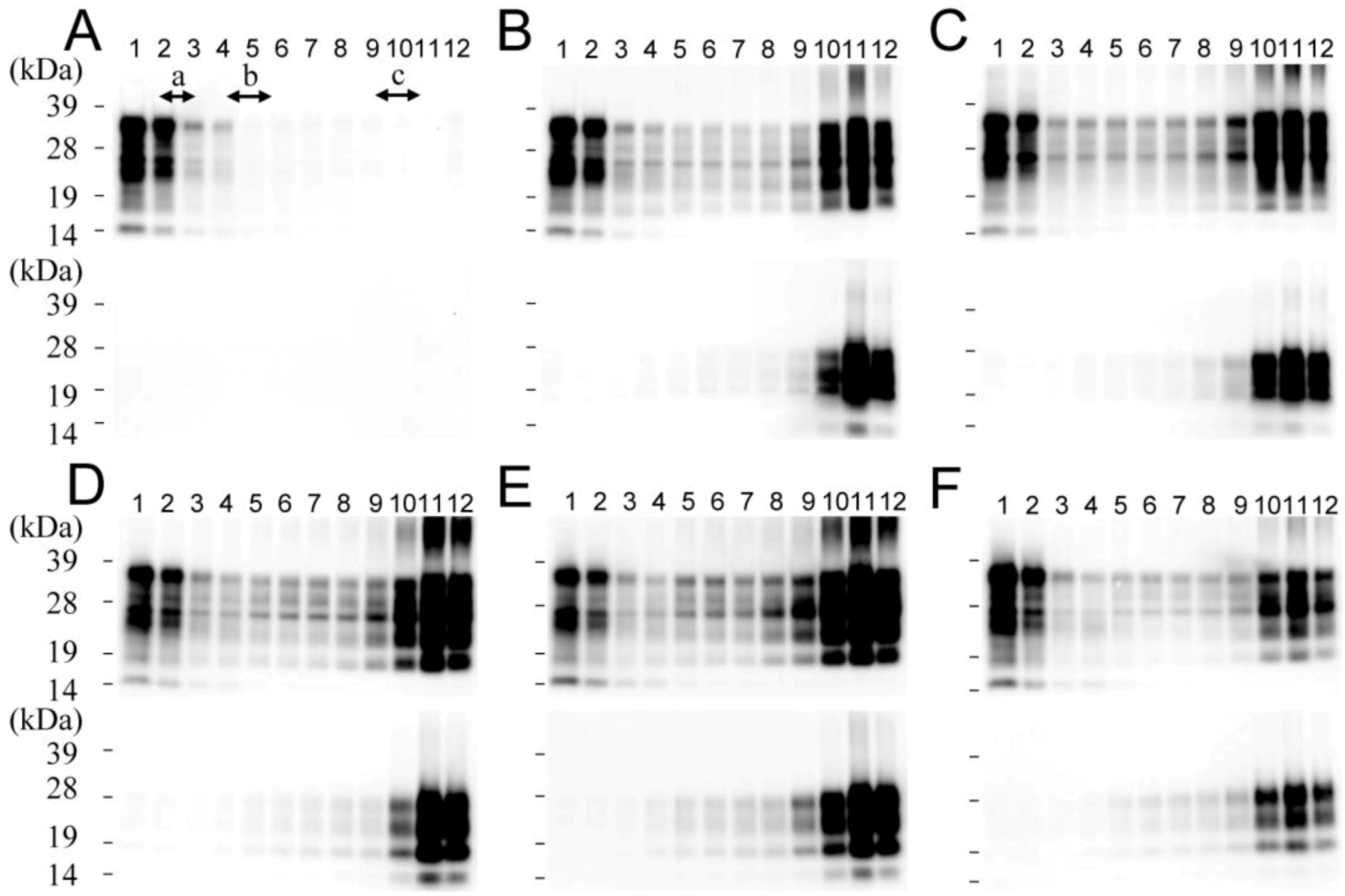
2.2. Conformational Stability of Small and Large PrPSc Aggregates against GdnHCl
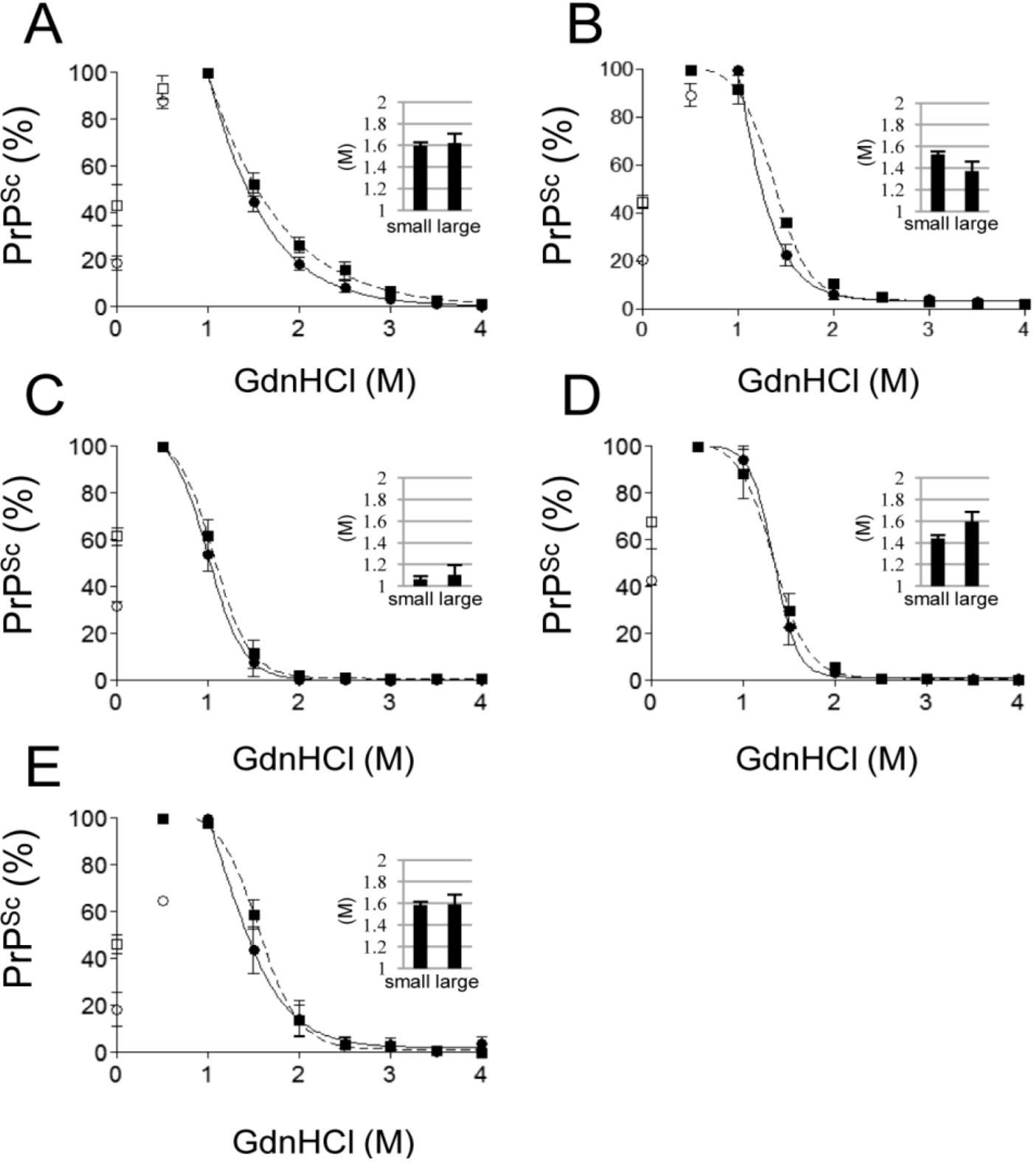
2.3. Aggregation Stability of Small and Large PrPSc Aggregates against SDS

2.4. Comparison of the PK Sensitivity of Small and Large PrPSc Aggregates

2.5. Transmissibility of Small and Large PrPSc Aggregates in the Chandler Strain
| Fraction No. | Chandler | mBSE | |||||
| Incubation period 1 | n/n02 | PrP 3 | Incubation period | n/n0 | PrP | ||
| 5 | 176.5 ± 4.6 | 4/4 | 0.11 | 241.8 ± 13.0* | 4/4 | 0.16 | |
| 7 | 174.5 ± 0.3 | 4/4 | 0.10 | 222.8 ± 6.3* | 4/4 | 0.19 | |
| 9 | 178.5 ± 1.8 | 4/4 | 0.26 | 216.0 ± 12.7 | 4/4 | 0.54 | |
| 11 | 175.3 ± 0.6 | 4/4 | 1.00 | 196.0 ± 12.7 | 4/4 | 1.00 | |
2.6. Discussion
3. Experimental Section
3.1. Animals and Prions
3.2. Velocity Sedimentation in Sucrose Gradients
3.3. Conformational Stability Assays
3.4. PK-Sensitivity Assay
3.5. Incubation Period Assay
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Prusiner, S.B. Novel proteinaceous infectious particles cause scrapie. Science 1982, 216, 136–144. [Google Scholar]
- Bolton, D.C.; McKinley, M.P.; Prusiner, S.B. Identification of a protein that purifies with the scrapie prion. Science 1982, 218, 1309–1311. [Google Scholar]
- Meyer, R.K.; McKinley, M.P.; Bowman, K.A.; Braunfeld, M.B.; Barry, R.A.; Prusiner, S.B. Separation and properties of cellular and scrapie prion proteins. Proc. Natl. Acad. Sci. USA 1986, 83, 2310–2314. [Google Scholar] [CrossRef]
- Pan, K.M.; Baldwin, M.; Nguyen, J.; Gasset, M.; Serban, A.; Groth, D.; Mehlhorn, I.; Huang, Z.; Fletterick, R.J.; Cohen, F.E.; et al. Conversion of alpha-helices into beta-sheets features in the formation of the scrapie prion proteins. Proc. Natl. Acad. Sci. USA 1993, 90, 10962–10966. [Google Scholar]
- Fraser, H.; Dickinson, A.G. The sequential development of the brain lesion of scrapie in three strains of mice. J. Comp. Pathol. 1968, 78, 301–311. [Google Scholar] [CrossRef]
- Bessen, R.A.; Marsh, R.F. Biochemical and physical properties of the prion protein from two strains of the transmissible mink encephalopathy agent. J.Virol. 1992, 66, 2096–2101. [Google Scholar]
- Collinge, J.; Sidle, K.C.; Meads, J.; Ironside, J.; Hill, A.F. Molecular analysis of prion strain variation and the aetiology of 'new variant' CJD. Nature 1996, 383, 685–690. [Google Scholar] [CrossRef]
- Korth, C.; Streit, P.; Oesch, B. Monoclonal antibodies specific for the native, disease-associated isoform of the prion protein. Methods Enzymol. 1999, 309, 106–122. [Google Scholar] [CrossRef]
- Ushiki-Kaku, Y.; Endo, R.; Iwamaru, Y.; Shimizu, Y.; Imamura, M.; Masujin, K.; Yamamoto, T.; Hattori, S.; Itohara, S.; Irie, S.; Yokoyama, T. Tracing conformational transition of abnormal prion proteins during interspecies transmission by using novel antibodies. J. Biol. Chem. 2010, 285, 11931–11936. [Google Scholar]
- Tixador, P.; Herzog, L.; Reine, F.; Jaumain, E.; Chapuis, J.; Le Dur, A.; Laude, H.; Beringue, V. The physical relationship between infectivity and prion protein aggregates is strain-dependent. PLoS Pathog. 2010, 6, e1000859. [Google Scholar] [CrossRef]
- Silveira, J.R.; Raymond, G.J.; Hughson, A.G.; Race, R.E.; Sim, V.L.; Hayes, S.F.; Caughey, B. The most infectious prion protein particles. Nature 2005, 437, 257–261. [Google Scholar]
- Tzaban, S.; Friedlander, G.; Schonberger, O.; Horonchik, L.; Yedidia, Y.; Shaked, G.; Gabizon, R.; Taraboulos, A. Protease-sensitive scrapie prion protein in aggregates of heterogeneous sizes. Biochemistry 2002, 41, 12868–12875. [Google Scholar]
- Come, J.H.; Fraser, P.E.; Lansbury, P.T., Jr. A kinetic model for amyloid formation in the prion diseases: importance of seeding. Proc. Natl. Acad. Sci. USA 1993, 90, 5959–5963. [Google Scholar] [CrossRef]
- Safar, J.; Wille, H.; Itri, V.; Groth, D.; Serban, H.; Torchia, M.; Cohen, F.E.; Prusiner, S.B. Eight prion strains have PrPSc molecules with different conformations. Nat. Med. 1998, 4, 1157–1165. [Google Scholar] [CrossRef]
- Ayers, J.I.; Schutt, C.R.; Shikiya, R.A.; Aguzzi, A.; Kincaid, A.E.; Bartz, J.C. The strain-encoded relationship between PrP replication, stability and processing in neurons is predictive of the incubation period of disease. PLoS Pathog. 2011, 7, e1001317. [Google Scholar] [CrossRef] [Green Version]
- Peretz, D.; Scott, M.R.; Groth, D.; Williamson, R.A.; Burton, D.R.; Cohen, F.E.; Prusiner, S.B. Strain-specified relative conformational stability of the scrapie prion protein. Protein Sci. 2001, 10, 854–863. [Google Scholar] [CrossRef]
- Legname, G.; Nguyen, H.O.; Baskakov, I.V.; Cohen, F.E.; Dearmond, S.J.; Prusiner, S.B. Strain-specified characteristics of mouse synthetic prions. Proc. Natl. Acad. Sci. USA 2005, 102, 2168–2173. [Google Scholar]
- Legname, G.; Nguyen, H.O.; Peretz, D.; Cohen, F.E.; DeArmond, S.J.; Prusiner, S.B. Continuum of prion protein structures enciphers a multitude of prion isolate-specified phenotypes. Proc. Natl. Acad. Sci. USA 2006, 103, 19105–19110. [Google Scholar]
- Bett, C.; Joshi-Barr, S.; Lucero, M.; Trejo, M.; Liberski, P.; Kelly, J.W.; Masliah, E.; Sigurdson, C.J. Biochemical properties of highly neuroinvasive prion strains. PLoS Pathog. 2012, 8, e1002522. [Google Scholar] [CrossRef]
- Anaya, Z.E.; Savistchenko, J.; Massonneau, V.; Lacroux, C.; Andreoletti, O.; Vilette, D. Recovery of small infectious PrPres aggregates from prion-infected cultured cells. J. Biol. Chem. 2011, 286, 8141–8148. [Google Scholar]
- Li, J.; Browning, S.; Mahal, S.P.; Oelschlegel, A.M.; Weissmann, C. Darwinian evolution of prions in cell culture. Science 2010, 327, 869–872. [Google Scholar] [CrossRef]
- Mahal, S.P.; Browning, S.; Li, J.; Suponitsky-Kroyter, I.; Weissmann, C. Transfer of a prion strain to different hosts leads to emergence of strain variants. Proc. Natl. Acad. Sci. USA 2010, 107, 22653–22658. [Google Scholar]
- Weissmann, C.; Li, J.; Mahal, S.P.; Browning, S. Prions on the move. EMBO Rep. 2011, 12, 1109–1117. [Google Scholar] [CrossRef]
- Puoti, G.; Giaccone, G.; Rossi, G.; Canciani, B.; Bugiani, O.; Tagliavini, F. Sporadic Creutzfeldt-Jakob disease: co-occurrence of different types of PrPSc in the same brain. Neurology 1999, 53, 2173–2176. [Google Scholar]
- Masujin, K.; Shu, Y.; Okada, H.; Matsuura, Y.; Iwamaru, Y.; Imamura, M.; Mohri, S.; Yokoyama, T. Isolation of two distinct prion strains from a scrapie-affected sheep. Arch. Virol. 2009, 154, 1929–1932. [Google Scholar] [CrossRef]
- Yokoyama, T.; Masujin, K.; Schmerr, M.J.; Shu, Y.; Okada, H.; Iwamaru, Y.; Imamura, M.; Matsuura, Y.; Murayama, Y.; Mohri, S. Intraspecies prion transmission results in selection of sheep scrapie strains. PLoS ONE 2010, 5, e15450. [Google Scholar]
- Dickinson, A.G. Scrapie in sheep and goats. Front Biol. 1976, 44, 209–241. [Google Scholar]
- Oelschlegel, A.M.; Fallahi, M.; Ortiz-Umpierre, S.; Weissmann, C. The extended cell panel assay characterizes the relationship of prion strains RML, 79A, and 139A and reveals conversion of 139A to 79A-like prions in cell culture. J. Virol. 2012, 86, 5297–5303. [Google Scholar] [CrossRef]
- Hayashi, H.K.; Yokoyama, T.; Takata, M.; Iwamaru, Y.; Imamura, M.; Ushiki, Y.K.; Shinagawa, M. The N-terminal cleavage site of PrPSc from BSE differs from that of PrPSc from scrapie. Biochem. Biophys. Res. Commun. 2005, 328, 1024–1027. [Google Scholar] [CrossRef]
- Yokoyama, T.; Masujin, K.; Iwamaru, Y.; Imamura, M.; Mohri, S. Alteration of the biological and biochemical characteristics of bovine spongiform encephalopathy prions during interspecies transmission in transgenic mice models. J. Gen. Virol. 2009, 90, 261–268. [Google Scholar] [CrossRef]
- Iwamaru, Y.; Takenouchi, T.; Ogihara, K.; Hoshino, M.; Takata, M.; Imamura, M.; Tagawa, Y.; Hayashi-Kato, H.; Ushiki-Kaku, Y.; Shimizu, Y.; Okada, H.; Shinagawa, M.; Kitani, H.; Yokoyama, T. Microglial cell line established from prion protein-overexpressing mice is susceptible to various murine prion strains. J. Virol. 2007, 81, 1524–1527. [Google Scholar]
- Hirogari, Y.; Kubo, M.; Kimura, K.M.; Haritani, M.; Yokoyama, T. Two different scrapie prions isolated in Japanese sheep flocks. Microbiol. Immunol. 2003, 47, 871–876. [Google Scholar]
- Yokoyama, T.; Kimura, K.M.; Ushiki, Y.; Yamada, S.; Morooka, A.; Nakashiba, T.; Sassa, T.; Itohara, S. In vivo conversion of cellular prion protein to pathogenic isoforms, as monitored by conformation-specific antibodies. J. Biol. Chem. 2001, 276, 11265–11271. [Google Scholar]
- Kasai, K.; Hirata, A.; Ohyama, T.; Nokihara, K.; Yokoyama, T.; Mohri, S. Novel assay with fluorescence-labelled PrP peptides for differentiating L-type atypical and classical BSEs, and scrapie. FEBS Lett. 2012, 586, 325–329. [Google Scholar] [CrossRef]
- Shimizu, Y.; Kaku-Ushiki, Y.; Iwamaru, Y.; Muramoto, T.; Kitamoto, T.; Yokoyama, T.; Mohri, S.; Tagawa, Y. A novel anti-prion protein monoclonal antibody and its single-chain fragment variable derivative with ability to inhibit abnormal prion protein accumulation in cultured cells. Microbiol. Immunol. 2010, 54, 112–121. [Google Scholar]
- Pirisinu, L.; Di Bari, M.; Marcon, S.; Vaccari, G.; D'Agostino, C.; Fazzi, P.; Esposito, E.; Galeno, R.; Langeveld, J.; Agrimi, U.; Nonno, R. A new method for the characterization of strain-specific conformational stability of protease-sensitive and protease-resistant PrP. PLoS ONE 2010, 5, e12723. [Google Scholar]
Supplementary Figures
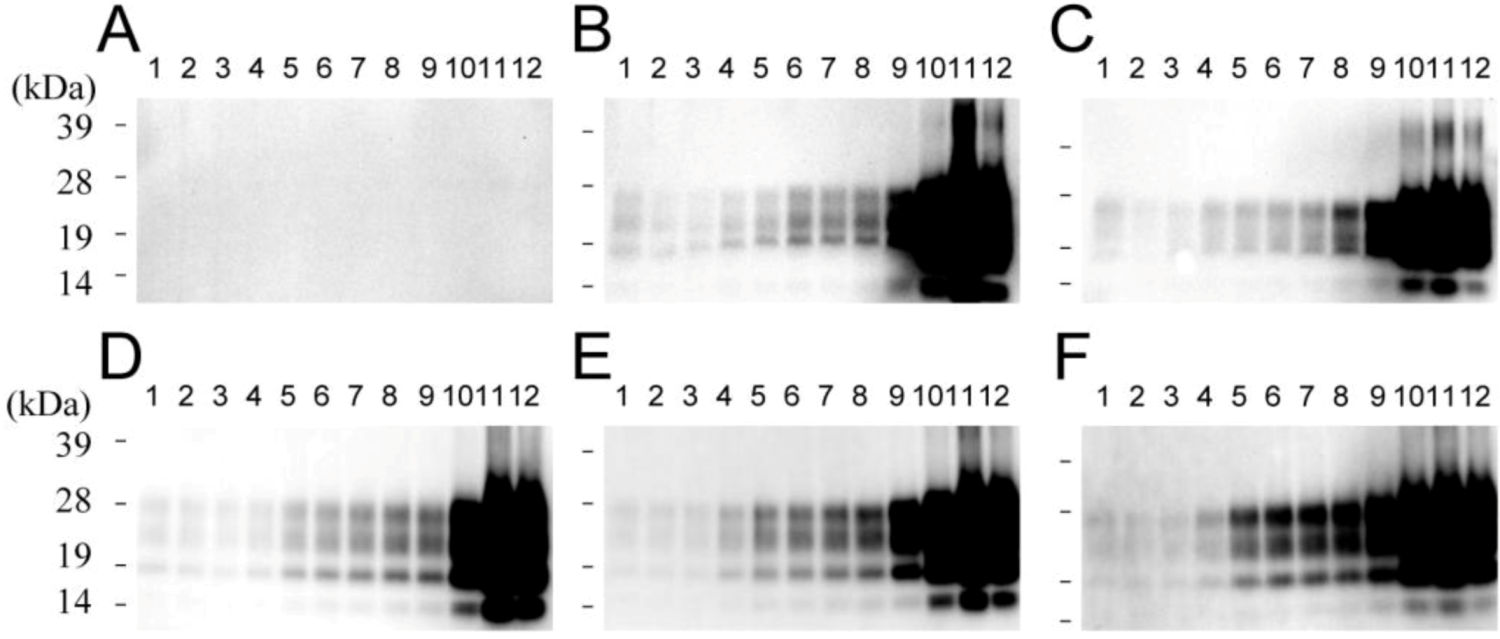

© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kasai, K.; Iwamaru, Y.; Masujin, K.; Imamura, M.; Mohri, S.; Yokoyama, T. Heterogeneity of the Abnormal Prion Protein (PrPSc) of the Chandler Scrapie Strain. Pathogens 2013, 2, 92-104. https://doi.org/10.3390/pathogens2010092
Kasai K, Iwamaru Y, Masujin K, Imamura M, Mohri S, Yokoyama T. Heterogeneity of the Abnormal Prion Protein (PrPSc) of the Chandler Scrapie Strain. Pathogens. 2013; 2(1):92-104. https://doi.org/10.3390/pathogens2010092
Chicago/Turabian StyleKasai, Kazuo, Yoshifumi Iwamaru, Kentaro Masujin, Morikazu Imamura, Shirou Mohri, and Takashi Yokoyama. 2013. "Heterogeneity of the Abnormal Prion Protein (PrPSc) of the Chandler Scrapie Strain" Pathogens 2, no. 1: 92-104. https://doi.org/10.3390/pathogens2010092




