Clinical Utility of Gene Expression Profiling Data for Clinical Decision-Making Regarding Adjuvant Therapy in Early Stage, Node-Negative Breast Cancer: A Case Report
Abstract
:1. Introduction
2. Case Presentation
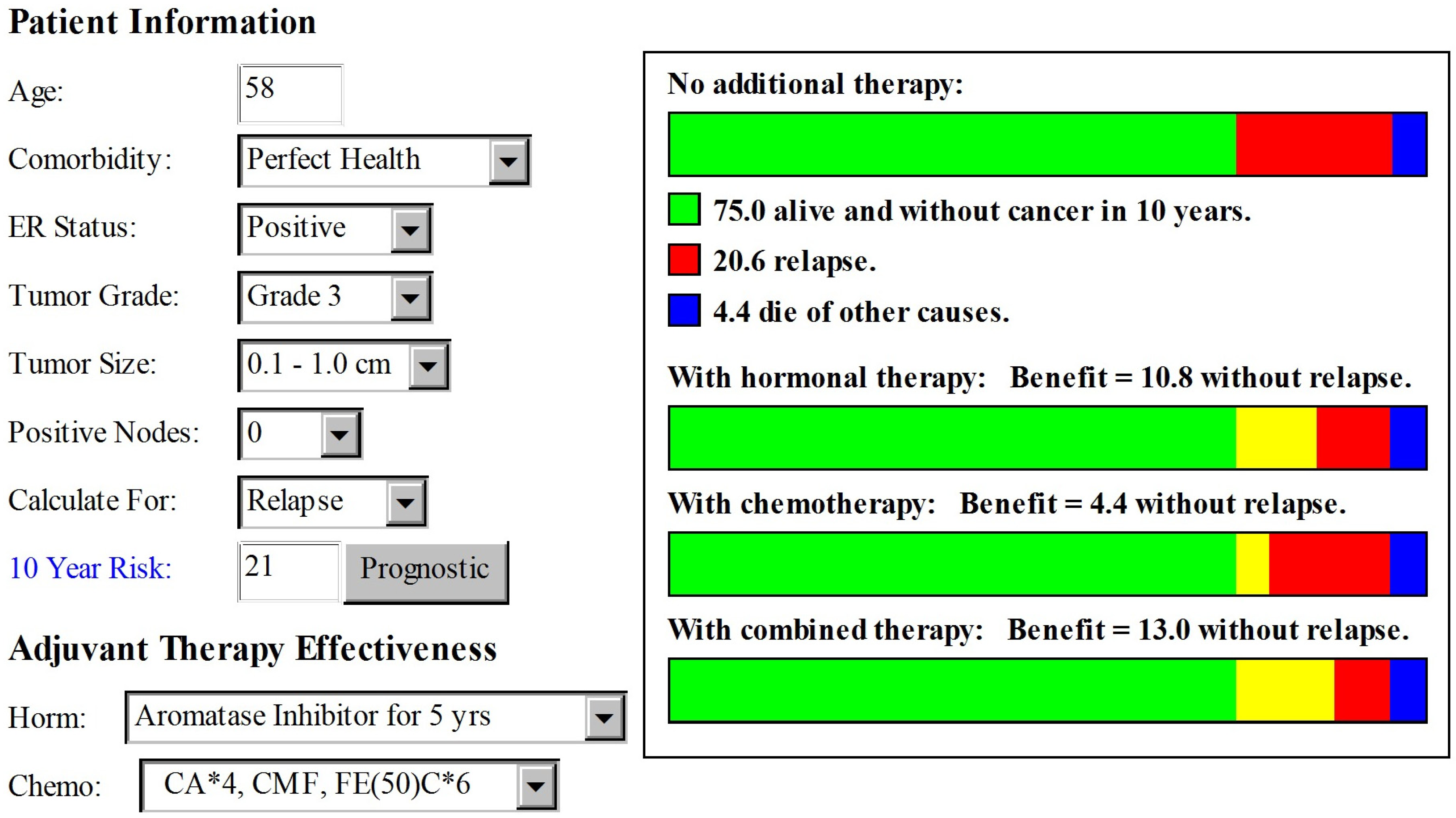

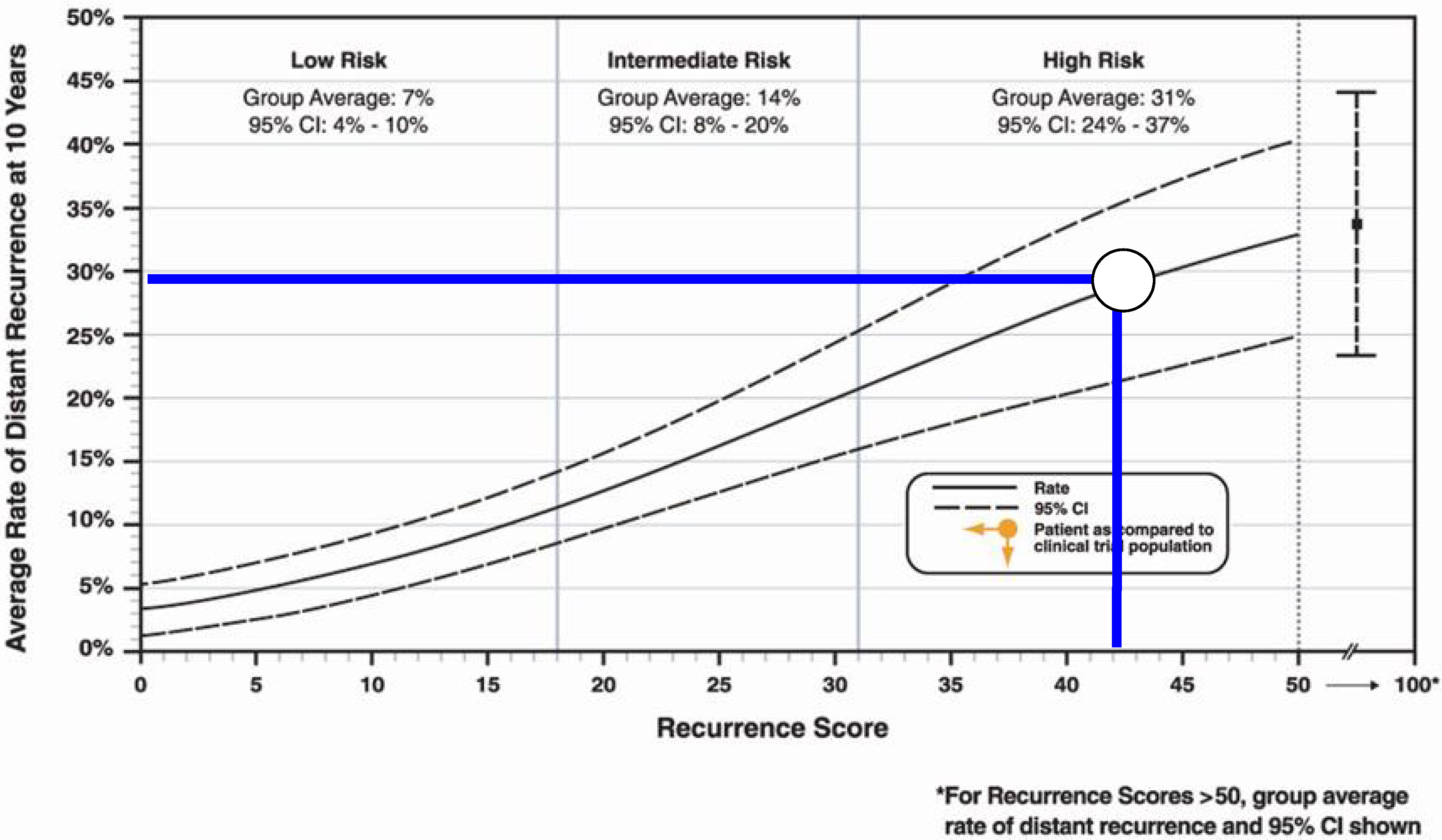
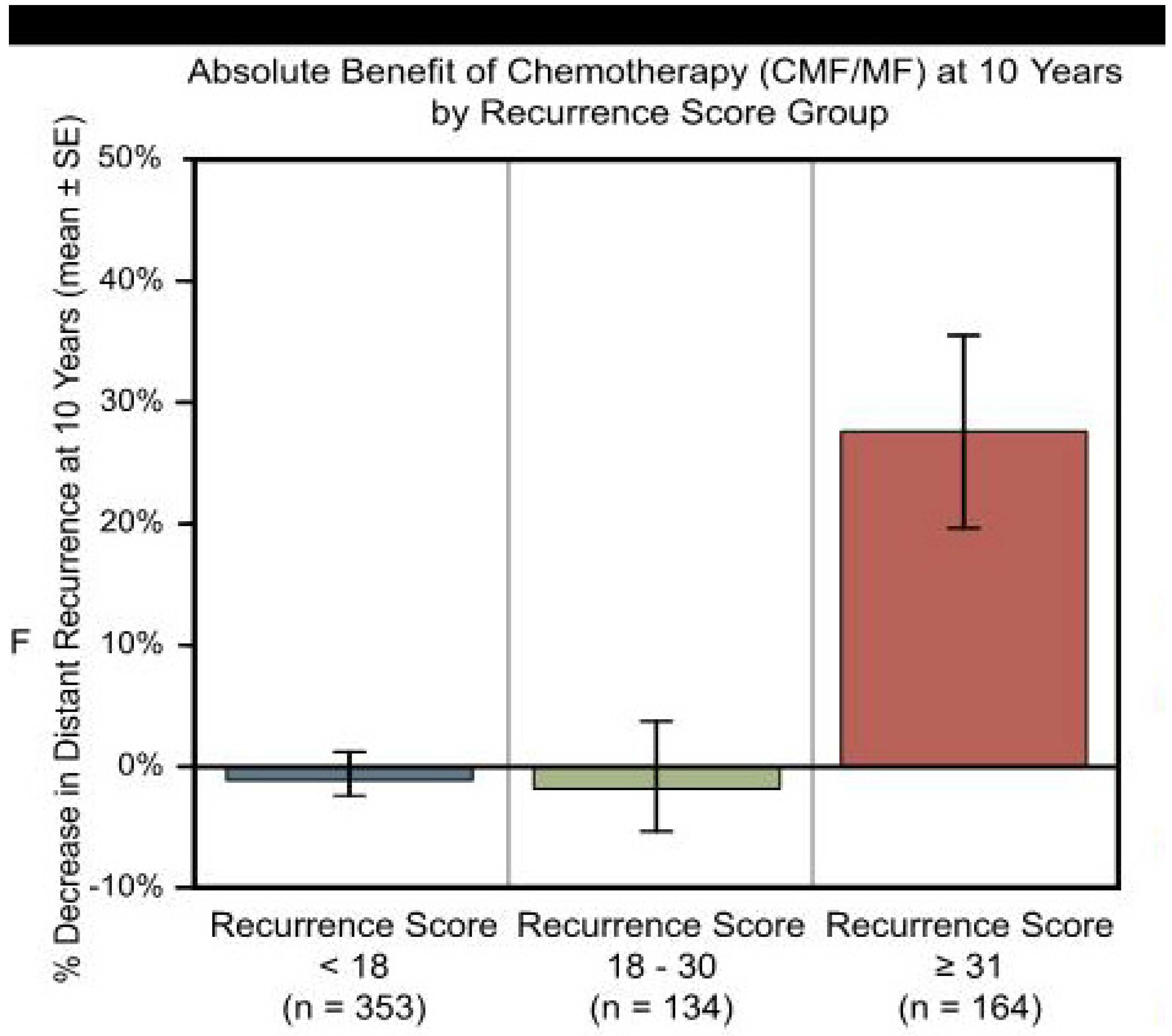
3. Discussion
Conflict of Interest
References
- Jemal, A.; Slegal, R.; Ward, E.; Hao, Y.; Xu, J.; Thun, M.J. Cancer statistics, 2009. CA Cancer J. Clin. 2009, 59, 225–249. [Google Scholar] [CrossRef]
- Adjuvant! Breast Cancer Help Files. Available online: http://www.adjuvantonline.com/breasthelp0306/breastindex.html (accessed on 25 March 2010).
- Cronin, M.; Sangli, C.; Liu, M.-L.; Pho, M.; Dutta, D.; Nguyen, A.; Jeong, J.; Wu, J.; Langone, K.C.; Watson, D. Analytical Validation of the Oncotype DX genomic diagnostic test for recurrence prognosis and therapeutic response prediction in node-negative, estrogen-receptor-positive breast cancer. Clin. Chem. 2007, 53, 1084–1091. [Google Scholar]
- Adjuvant! Online 2012. Available online: http://www.adjuvantonline.com/ (accessed on 01 August 2012).
- Paik, S.; Shak, S.; Tang, G.; Kim, C.; Baker, J.; Cronin, M.; Baehner, F.L.; Walker, M.G.; Watson, D.; Park, T.; et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N. Engl. J. Med. 2004, 351, 2817–2826. [Google Scholar]
- Paik, S.; Tang, G.; Shak, S.; Kim, C.; Baker, J.; Kim, W.; Cronin, M.; Baehner, F.L.; Watson, D.; Bryant, J.; et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 2006, 24, 3726–3734. [Google Scholar]
- Habel, L.A.; Shak, S.; Jacobs, M.K.; Capra, A.; Alexander, C.; Pho, M.; Baker, J.; Walker, M.; Watson, D.; Hackett, J.; et al. A population-based study of tumor gene expression and risk of breast cancer death among lymph node-negative patients. Breast Cancer Res. 2006, 8, R25. [Google Scholar]
- Henry, L.R.; Stojadinovic, A.; Swain, S.M.; Prindiville, S.; Cordes, R.; Soballe, P.W. The influence of a gene expression profile on breast cancer decisions. J. Surg. Oncol. 2009, 99, 319–323. [Google Scholar] [CrossRef]
- Asad, J.; Jacobson, A.F.; Estabrook, A.; Smith, S.R.; Boolbol, S.K.; Feldman, S.M.; Osborne, M.P.; Boachie-Adjei, K.; Twardzik, W.; Tartter, P.I. Does oncotype DX recurrence score affect the management of patients with early-stage breast cancer? Am. J. Surg. 2008, 196, 527–529. [Google Scholar]
- Sparano, J.A. The TAILORx trial: Individualized options for treatment. Commun. Oncol. 2006, 3, 494–496. [Google Scholar] [CrossRef]
- Whelan, T.J.; Loprinzi, C. Physician/patient decision aids for adjuvant therapy. J. Clin. Oncol. 2005, 23, 1627–1630. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Schuster, S.R.; Pockaj, B.A.; Bothe, M.R.; David, P.S.; Northfelt, D.W. Clinical Utility of Gene Expression Profiling Data for Clinical Decision-Making Regarding Adjuvant Therapy in Early Stage, Node-Negative Breast Cancer: A Case Report. J. Pers. Med. 2012, 2, 71-76. https://doi.org/10.3390/jpm2030071
Schuster SR, Pockaj BA, Bothe MR, David PS, Northfelt DW. Clinical Utility of Gene Expression Profiling Data for Clinical Decision-Making Regarding Adjuvant Therapy in Early Stage, Node-Negative Breast Cancer: A Case Report. Journal of Personalized Medicine. 2012; 2(3):71-76. https://doi.org/10.3390/jpm2030071
Chicago/Turabian StyleSchuster, Steven R., Barbara A. Pockaj, Mary R. Bothe, Paru S. David, and Donald W. Northfelt. 2012. "Clinical Utility of Gene Expression Profiling Data for Clinical Decision-Making Regarding Adjuvant Therapy in Early Stage, Node-Negative Breast Cancer: A Case Report" Journal of Personalized Medicine 2, no. 3: 71-76. https://doi.org/10.3390/jpm2030071



