MicroRNAs as Biomarkers in Cancer
Abstract
:1. Introduction
2. MicroRNAs and Cancer
2.1. MicroRNAs as Oncogenes
2.2. MicroRNAs as Tumor Suppressors
3. MicroRNAs as Potential Biomarkers in Cancer
4. Circulating MicroRNAs
| Cancer | Sample type | Comparison criteria | Methods | miRNAs | Target/mechanisms | Ref. |
|---|---|---|---|---|---|---|
| Epithelial cancers | ||||||
| Breast | Tissue cancer cells | ER+, ER− tissues | Microarray, qPCR | miR-34b (↓) | miR-34b target cyclin D1 and JAG-1 | [66] |
| Whole blood | Tumor vs. normal | qPCR | miR-195 (↑)
let-7a (↑) miR-155 (↑) | let-7a target KRAS and miR-155 target RhoA transforming growth factor and induces EMT | [67] | |
| Cancer cells | Cancer cell growth | Microarray | miR-21 (↑) | PDCD4 | [68] | |
| Colorectal | Tissue & serum | Tumor vs. normal | qPCR microarray | miR-17-3p (↑)
miR-92 (↑) | miR-92 is elevated in plasma and can be used as a non-invasive molecular marker for screening | [53] |
| Plasma | Tumor vs. normal | qPCR | miR-29a (↑)
miR-92a (↑) | Promote cell proliferation, suppressed apoptosis, induce tumor angiogenesis and accelerated tumor progression | [69] | |
| Bladder | Tissue & cancer cell lines | Tumor vs. normal | qPCR | miR-145 (↓)
miR-30a-3p (↓) miR-133a (↓) miR-133b (↓) miR-195 (↓) miR-125b (↓) miR-199a (↓) | These miRNAs are downregulated and play the role of tumor suppressors by targeting KRT7, a common target with oncogenic function | [70] |
| Glioblastoma | Tissue | Tumor vs. normal, Different grades of malignancy | qPCR Northern blotting | miR-21 (↑)
miR-221 (↑) miR-128 (↓) miR-181b (↓) | Knockdown of miR-21 triggered the activation of caspases leading to apoptosis. miR-221 & miR-222 repress expression p27Kip1. miR-181b triggered growth inhibition, apoptosis, and inhibited invasion | [71] |
| Gastric | Tissue & cell lines | Tumor vs. normal | qPCR | miR-409 (↓) | miR-409 target RDX and suppresses cell invasion and metastasis | [72] |
| Lung | Serum | Overall survival in NSCLC | sequencingqPCR | miR-486 (↑)
miR-30d (↑) miR-1 (↓) miR-499 (↓) | miR-1 downregulates MET oncogene. Facilitates activation of Caspase 3, Caspase 7, and PARP-1 as well as depletion of MCL-1 | [73] |
| Plasma | Tumor vs. normal | qPCR | let-7f (↓)
miR-20b (↓) miR-30e-3p (↓) | let-7 targets cMyc. miR-30e regulates of Ubc9 | [74] | |
| Oral & squamous cell | Tissue | Tumor vs. normal | qPCR | miR-184 (↑) | miR-184 alters cMyc expression and affects anti-apoptosis and proliferation of tongue SCC cells | [58] |
| Tissue & saliva | Tumor vs. normal | qPCR | miR-31 (↑) | - | [75] | |
| Ovarian | Serum | Tumor vs. normal | qPCR | miR-21 (↑)
miR-29a (↑) miR-92 (↑) miR-93 (↑) miR-99b (↓) miR-126 (↓) miR-127 (↓)) miR-155 (↓)) | miR-21 regulates PDCD4 and maspin miR-92 and miR-93 regulates TGFβ miR-29a potentially targets PTEN miR-127 regulates BCL6 | [39] |
| Prostate | Plasma, serum, murine | Tumor vs. normal | qPCR | miR-141 (↑)
miR-375 (↑) miR-107 (↑) miR-574-3p (↑) | induces abnormal cell division and proliferation and the development of aggressive prostate cancer | [76] |
| Tissue & Serum | Metastatic, localized tumors vs. normal | qPCR | miR-141 (↑)
miR-375 (↑) | regulates genes controlling cellular growth and proliferation | [77] | |
| Pancreatic | Tissue | Tumor vs. normal | qPCR | miR-155 (↑)
miR-203 (↑) miR-210 (↑) miR-222 (↑) | - | [78] |
| Plasma | Tumor vs. normal | qPCR | miR-21 (↑)
miR-155 (↑) miR-196a (↑) miR-210 (↑) | miR-21 targets PTEN and PDCD4 miR-210 affect DNA repair and genomic instability miR-155 target TP53INP1 | [79] | |
| Hepatocellular | Tissue & cell cultures | Tumor vs. normal | qPCR | miR-519d (↑) | miR-519d has inhibitory effect on CDKN1A/p21, PTEN and TIMP2 expression | [80] |
| Endometrial | Tissue | Tumor vs. hyperplasia vs. normal | qPCR | miR-200 family (↑) | Negatively regulates ZEB1 and ZEB2 and implicated in EMT | [81] |
| Renal cell | Tissue | Tumor vs. normal | qPCR Microarray | miR-122 (↑)
miR-155 (↑) miR-210 (↑) miR-200c (↓) miR-335 (↓) miR-218 (↓) | - | [82] |
| Melanoma | Tissue & cell lines | Normal vs. cancer cell lines | Microarray | miR-193a (↓)
miR-338 (↓) miR-565 (↓) miR-191 (↓) miR-193b (↑) | miR-193 is regulated by HNF-1a and p53; predicted targets for miR-191 include FZD5 and BDNF | [83] |
| Thyroid | Tissue | Tumor vs. Normal | qPCR | miR-187 (↑)
miR-221 (↑) miR-222 (↑) miR-146b (↑) miR-155 (↑) miR-224 (↑) miR-197 (↑) | The oncogenic mutations in PCs, RET/PTC, BRAF, and RAS are all capable of activation of the MAPK pathway | [84] |
| SARCOMAS | ||||||
| Osteosarcoma | Tissue & cancer cell lines | Tumor vs. normal | qPCR | miR-135b (↑)
miR-150 (↑) miR-542-5p (↑) miR-652 (↑) | Pro-apoptotic EGR2 and P2X7 are targets of miR-150 | [85] |
| Tissue | Tumor vs. normal | qPCR Microarray | miR-17-92 (↓) | 14q32 miRNAs (miR-544, miR-369-3p, miR-134 and miR-382) act cooperatively to destabilize cMYC and in turn, control expression of miR-17-92 miRNAs | [86] | |
| Leiomyosarcoma | Tissue | Tumor vs. normal | qPCR Microarray CGH | miR-21 (↑)
let7 (↑) miR- 27a (↑) miR-30a (↑) miR-23b (↑) miR-29b (↓) miR-32 (↓) miR-144 (↓) miR-212 (↓) miR-197 (↓) | Targets MAPK pathway genes | [87] |
| Rhabdomyo-sarcoma | Cell lines, Tissue & Serum | Tumor vs. normal | qPCR | miR-206 (↑) | expression of miR-206 in RMS cells promoted myogenic differentiation and blocked tumor growth | [88] |
| Gastrointestinal Stromal Tumor | Tissue | Tumor vs. normal | qPCR | miR-221 (↓)
miR-222 (↓) | Regulates cKIT | [89] |
| Ewing's Sarcoma | Cell lines | Primary sarcoma vs. Progenitor cells | miRNA Profiling | miR-145 (↓) | miR-145 inhibits stem cell transcription factors Oct4, Sox2, Klf4 and Myc | [90] |
| Schwannoma | Tissue, cell lines | Tumor vs. Normal | Microarray, qPCR | miR-7 (↓) | Inhibited expression of Ack1, Pak1, and EGFR | [91] |
| MPNST | Tissue | MPNST vs. neurofibroma | Microarray, qPCR | miR-34a (↓) | Partly due to p53 inactivation | [93] |
| LEUKEMIA/ LYMPHOMA | ||||||
| Adult T Cell Leukemia | Cells | primary ATL cells vs. normal CD4+ T cells | Microarray | miR-31 (↓) | miR-31 is a suppressor of NIK and pathway involving polycomb-mediated miRNA silencing and NF-kB activation | [94] |
| Acute promyelocytic leukemia | Cells | Leukemia vs. Normal Promyelocytes | qPCR | miR-15b (↓)
miR-16 (↓) miR-107 (↓) miR-223 (↓) miR-342 (↓) and let-7c (↓) | PML/RARa binds the regulatory sequences of the intragenic miR-342 and let-7c | [95] |
| AML | Cell lines | AML, Human myeloid, CLL cell lines | qPCR microarray | miR-34b (↓) | Cyclic AMP-Responsive Element Binding Protein down-regulation | [96] |
| CLL | Peripheral blood mononuclear cells | Cancer cells vs. normal cells | qPCR | miR-92 (↑) | Abnormal elevation of HIF-1α, the key upstream regulator of VEGF | [97] |
| Peripheral Blood CD19+ cells | Prognostic factors | qPCR Western blot | miR-29c (↓)
miR-223 (↓) | Regulates the Tcl1 oncogene Down-regulation of miR-29 inversely correlates with DNMT expression | [98] | |
| Hodgkins lymphoma | Cancer cell lines | Hodgkins vs. B cell non Hodgkins | qPCR Microarray | miR-155 (↑) | IKBKE, ZNF537, ZIC3, FGF7, and AGTR1 are functional targets of miR-155 | [34] |
| Diffuse Large B-cell lymphoma | Serum | Tumor vs. Normal | qPCR | miR-15a (↑)
miR-16-1 (↑) miR-29c (↑) miR-155 (↑) miR-34a (↓) | miR-155 directly down regulates one of the MYC antagonists like MAD1, MXI1, ROX/MNT | [99] |
5. miRNA Detection in Body Fluids and Stability
5.1. miRNA Extraction and Quantifying Methods
5.2. Potential Pitfalls in Developing miRNAs as Circulating Biomarkers
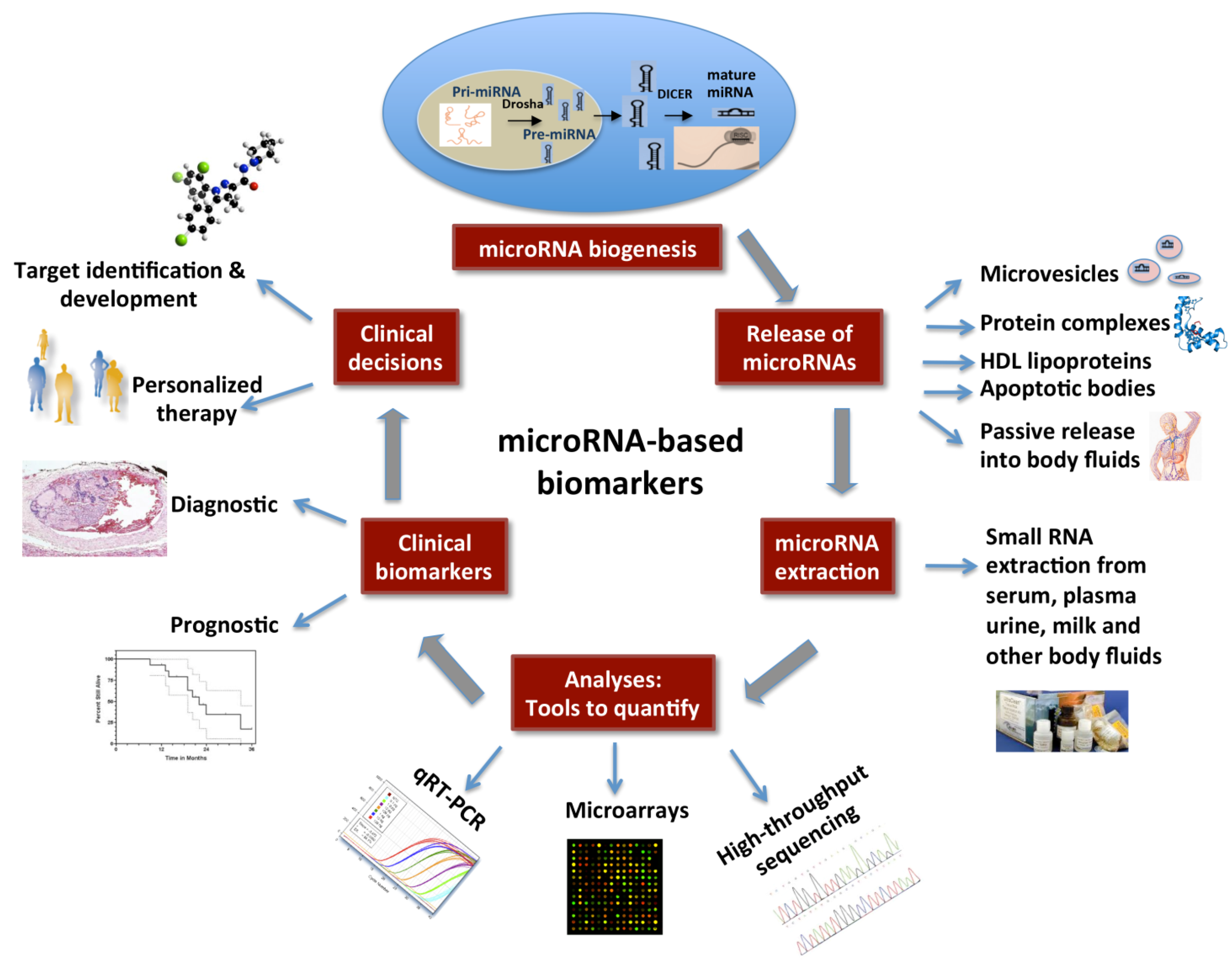
6. Future Directions
Acknowledgments
References
- Cho, W.C. OncomiRs: The discovery and progress of microRNAs in cancers. Mol. Cancer 2007, 6. [Google Scholar] [CrossRef]
- Junn, E.; Mouradian, M.M. MicroRNAs in neurodegenerative diseases and their therapeutic potential. Pharmacol. Ther. 2012, 133, 142–150. [Google Scholar] [CrossRef]
- Maccani, M.A.; Padbury, J.F.; Marsit, C.J. miR-16 and miR-21 expression in the placenta is associated with fetal growth. PLoS ONE 2011, 6. [Google Scholar] [CrossRef]
- Zidar, N.; Bostjancic, E.; Glavac, D.; Stajer, D. MicroRNAs, innate immunity and ventricular rupture in human myocardial infarction. Dis. Markers 2011, 31, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Rothschild, S.I.; Tschan, M.P.; Federzoni, E.A.; Jaggi, R.; Fey, M.F.; Gugger, M.; Gautschi, O. MicroRNA-29b is involved in the Src-ID1 signaling pathway and is dysregulated in human lung adenocarcinoma. Oncogene 2012, 31, 4221–4232. [Google Scholar] [CrossRef]
- Tavazoie, S.F.; Alarcon, C.; Oskarsson, T.; Padua, D.; Wang, Q.; Bos, P.D.; Gerald, W.L.; Massague, J. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 2008, 451, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Porkka, K.P.; Pfeiffer, M.J.; Waltering, K.K.; Vessella, R.L.; Tammela, T.L.; Visakorpi, T. MicroRNA expression profiling in prostate cancer. Cancer Res. 2007, 67, 6130–6135. [Google Scholar] [CrossRef] [PubMed]
- Roccaro, A.M.; Sacco, A.; Jia, X.; Azab, A.K.; Maiso, P.; Ngo, H.T.; Azab, F.; Runnels, J.; Quang, P.; Ghobrial, I.M. microRNA-dependent modulation of histone acetylation in Waldenstrom macroglobulinemia. Blood 2010, 116, 1506–1514. [Google Scholar] [CrossRef] [PubMed]
- Razumilava, N.; Bronk, S.F.; Smoot, R.L.; Fingas, C.D.; Werneburg, N.W.; Roberts, L.R.; Mott, J.L. miR-25 targets TNF-related apoptosis inducing ligand (TRAIL) death receptor-4 and promotes apoptosis resistance in cholangiocarcinoma. Hepatology 2012, 55, 465–475. [Google Scholar] [CrossRef]
- Bartel, D.P.; Chen, C.Z. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat. Rev. Genet. 2004, 5, 396–400. [Google Scholar] [CrossRef]
- Hobert, O. miRNAs play a tune. Cell 2007, 131, 22–24. [Google Scholar] [CrossRef]
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef]
- Li, X.; Cassidy, J.J.; Reinke, C.A.; Fischboeck, S.; Carthew, R.W. A microRNA imparts robustness against environmental fluctuation during development. Cell 2009, 137, 273–282. [Google Scholar] [CrossRef]
- Mendell, J.T. miRiad roles for the miR-17–92 cluster in development and disease. Cell 2008, 133, 217–222. [Google Scholar] [CrossRef]
- Croce, C.M. Causes and consequences of microRNA dysregulation in cancer. Nat. Rev. Genet. 2009, 10, 704–714. [Google Scholar] [CrossRef]
- Kim, V.N. Small RNAs: Classification, biogenesis, and function. Mol. Cells 2005, 19, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Flynt, A.S.; Lai, E.C. Biological principles of microRNA-mediated regulation: Shared themes amid diversity. Nat. Rev. Genet. 2008, 9, 831–842. [Google Scholar] [CrossRef]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- miRBase: The MicroRNA Database. Available online: http://www.mirbase.org (accessed on 15 November 2012).
- Lujambio, A.; Lowe, S.W. The microcosmos of cancer. Nature 2012, 482, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Hwang, H.W.; Mendell, J.T. MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br. J. Cancer 2006, 94, 776–780. [Google Scholar] [CrossRef]
- Conne, B.; Stutz, A.; Vassalli, J.D. The 3' untranslated region of messenger RNA: A molecular “hotspot” for pathology? Nat. Med. 2000, 6, 637–641. [Google Scholar] [CrossRef]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef] [PubMed]
- Chan, E.; Patel, R.; Nallur, S.; Ratner, E.; Bacchiocchi, A.; Hoyt, K.; Szpakowski, S.; Godshalk, S.; Ariyan, S.; Sznol, M.; et al. MicroRNA signatures differentiate melanoma subtypes. Cell Cycle 2011, 10, 1845–1852. [Google Scholar] [CrossRef]
- Wurz, K.; Garcia, R.L.; Goff, B.A.; Mitchell, P.S.; Lee, J.H.; Tewari, M.; Swisher, E.M. MiR-221 and MiR-222 alterations in sporadic ovarian carcinoma: Relationship to CDKN1B, CDKNIC and overall survival. GCC (Genes Chromosomes Cancer) 2010, 49, 577–584. [Google Scholar]
- Guttilla, I.K.; White, B.A. Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. J. Biol. Chem. 2009, 284, 23204–23216. [Google Scholar] [CrossRef] [PubMed]
- Van Der Heide, L.P.; Hoekman, M.F.; Smidt, M.P. The ins and outs of FoxO shuttling: Mechanisms of FoxO translocation and transcriptional regulation. Biochem. J. 2004, 380, 297–309. [Google Scholar] [CrossRef]
- Kefas, B.; Godlewski, J.; Comeau, L.; Li, Y.; Abounader, R.; Hawkinson, M.; Lee, J.; Fine, H.; Chiocca, E.A.; Lawler, S.; Purow, B. microRNA-7 inhibits the epidermal growth factor receptor and the Akt pathway and is down-regulated in glioblastoma. Cancer Res. 2008, 68, 3566–3572. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.; Lorenz, P.; Gross, G.; Ibrahim, S.; Kunz, M. MicroRNA let-7b targets important cell cycle molecules in malignant melanoma cells and interferes with anchorage-independent growth. Cancer Res. 2008, 18, 549–557. [Google Scholar]
- Tam, W.; Ben-Yehuda, D.; Hayward, W.S. Bic, a novel gene activated by proviral insertions in avian leukosis virus-induced lymphomas, is likely to function through its noncoding RNA. Mol. Cell Biol. 1997, 17, 1490–1502. [Google Scholar] [CrossRef] [PubMed]
- Cano, C.E.; Gommeaux, J.; Pietri, S.; Culcasi, M.; Garcia, S.; Seux, M.; Barelier, S.; Vasseur, S.; Spoto, R.P.; Pebusque, M.J.; Dusetti, N.J.; Iovanna, J.L.; Carrier, A. Tumor protein 53-induced nuclear protein 1 is a major mediator of p53 antioxidant function. Cancer Res. 2009, 69, 219–226. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J.; Hammond, S.M. A microRNA polycistron as a potential human oncogene. Nature 2005, 435, 828–833. [Google Scholar] [CrossRef] [PubMed]
- Dong, F.; Lou, D. MicroRNA-34b/c suppresses uveal melanoma cell proliferation and migration through multiple targets. Mol. Vis. 2012, 18, 537–546. [Google Scholar] [PubMed]
- Gibcus, J.H.; Tan, L.P.; Harms, G.; Schakel, R.N.; de Jong, D.; Blokzijl, T.; Moller, P.; Poppema, S.; Kroesen, B.J.; van den Berg, A. Hodgkin lymphoma cell lines are characterized by a specific miRNA expression profile. Neoplasia 2009, 11, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Taulli, R.; Bersani, F.; Foglizzo, V.; Linari, A.; Vigna, E.; Ladanyi, M.; Tuschl, T.; Ponzetto, C. The muscle-specific microRNA miR-206 blocks human rhabdomyosarcoma growth in xenotransplanted mice by promoting myogenic differentiation. J. Clin. Invest. 2009, 119, 2366–2378. [Google Scholar] [PubMed]
- Heneghan, H.M.; Miller, N.; Kelly, R.; Newell, J.; Kerin, M.J. Systemic miRNA-195 differentiates breast cancer from other malignancies and is a potential biomarker for detecting noninvasive and early stage disease. Oncologist 2010, 15, 673–682. [Google Scholar] [CrossRef]
- White, N.M.; Bao, T.T.; Grigull, J.; Youssef, Y.M.; Girgis, A.; Diamandis, M.; Fatoohi, E.; Metias, M.; Honey, R.J.; Stewart, R.; Pace, K.T.; Bjarnason, G.A.; Yousef, G.M. miRNA profiling for clear cell renal cell carcinoma: biomarker discovery and identification of potential controls and consequences of miRNA dysregulation. J. Urol. 2011, 186, 1077–1083. [Google Scholar] [CrossRef]
- Liu, J.; Gao, J.; Du, Y.; Li, Z.; Ren, Y.; Gu, J.; Wang, X.; Gong, Y.; Wang, W.; Kong, X. Combination of plasma microRNAs with serum CA19-9 for early detection of pancreatic cancer. Int. J. Cancer 2012, 131, 683–691. [Google Scholar] [CrossRef] [PubMed]
- Resnick, K.E.; Alder, H.; Hagan, J.P.; Richardson, D.L.; Croce, C.M.; Cohn, D.E. The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol. Oncol. 2009, 112, 55–59. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Zheng, T.; Wang, J.; Chen, X.; Liu, L. Role of microRNA in anticancer drug resistance. Int. J. Cancer 2010, 126, 2–10. [Google Scholar] [CrossRef]
- Ryu, J.K.; Matthaei, H.; Dal Molin, M.; Hong, S.M.; Canto, M.I.; Schulick, R.D.; Wolfgang, C.; Goggins, M.G.; Hruban, R.H.; Cope, L.; Maitra, A. Elevated microRNA miR-21 levels in pancreatic cyst fluid are predictive of mucinous precursor lesions of ductal adenocarcinoma. Pancreatology 2011, 11, 343–350. [Google Scholar] [CrossRef]
- Ryu, J.K.; Hong, S.M.; Karikari, C.A.; Hruban, R.H.; Goggins, M.G.; Maitra, A. Aberrant MicroRNA-155 expression is an early event in the multistep progression of pancreatic adenocarcinoma. Pancreatology 2010, 10, 66–73. [Google Scholar] [CrossRef]
- Russo, F.; Di Bella, S.; Nigita, G.; Macca, V.; Lagana, A.; Giugno, R.; Pulvirenti, A.; Ferro, A. miRandola: Extracellular circulating microRNAs database. PloS ONE 2012, 7. [Google Scholar] [CrossRef]
- Mo, M.H.; Chen, L.; Fu, Y.; Wang, W.; Fu, S.W. Cell-free Circulating miRNA Biomarkers in Cancer. Int. J. Cancer 2012, 3, 432–448. [Google Scholar] [CrossRef]
- Sun, Y.; Wang, M.; Lin, G.; Sun, S.; Li, X.; Qi, J.; Li, J. Serum microRNA-155 as a potential biomarker to track disease in breast cancer. PloS ONE 2012, 7. [Google Scholar] [CrossRef]
- Zhao, A.; Li, G.; Peoc’h, M.; Genin, C.; Gigante, M. Serum miR-210 as a novel biomarker for molecular diagnosis of clear cell renal cell carcinoma. Exp. Mol. Pathol. 2012, 94, 115–120. [Google Scholar] [PubMed]
- Dieckmann, K.P.; Spiekermann, M.; Balks, T.; Flor, I.; Loning, T.; Bullerdiek, J.; Belge, G. MicroRNAs miR-371–3 in serum as diagnostic tools in the management of testicular germ cell tumours. Br. J. Cancer 2012, 107, 1754–1760. [Google Scholar] [CrossRef]
- Yu, Z.; Kastenmuller, G.; He, Y.; Belcredi, P.; Moller, G.; Prehn, C.; Mendes, J.; Wahl, S.; Roemisch-Margl, W.; Ceglarek, U.; et al. Differences between human plasma and serum metabolite profiles. PLoS ONE 2011, 6. [Google Scholar] [CrossRef]
- Lawrie, C.H.; Gal, S.; Dunlop, H.M.; Pushkaran, B.; Liggins, A.P.; Pulford, K.; Banham, A.H.; Pezzella, F.; Boultwood, J.; Wainscoat, J.S.; Hatton, C.S.; Harris, A.L. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br. J. Haematol. 2008, 141, 672–675. [Google Scholar] [CrossRef]
- Tanaka, M.; Oikawa, K.; Takanashi, M.; Kudo, M.; Ohyashiki, J.; Ohyashiki, K.; Kuroda, M. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PloS ONE 2009, 4. [Google Scholar] [CrossRef]
- Zhu, W.; Qin, W.; Atasoy, U.; Sauter, E.R. Circulating microRNAs in breast cancer and healthy subjects. BMC Res. Notes 2009, 2. [Google Scholar] [CrossRef]
- Ng, E.K.; Chong, W.W.; Jin, H.; Lam, E.K.; Shin, V.Y.; Yu, J.; Poon, T.C.; Ng, S.S.; Sung, J.J. Differential expression of microRNAs in plasma of patients with colorectal cancer: A potential marker for colorectal cancer screening. Gut 2009, 58, 1375–1381. [Google Scholar] [CrossRef]
- Tsujiura, M.; Ichikawa, D.; Komatsu, S.; Shiozaki, A.; Takeshita, H.; Kosuga, T.; Konishi, H.; Morimura, R.; Deguchi, K.; Fujiwara, H.; Okamoto, K.; Otsuji, E. Circulating microRNAs in plasma of patients with gastric cancers. Br. J. Cancer 2010, 102, 1174–1179. [Google Scholar] [CrossRef]
- Skog, J.; Wurdinger, T.; van Rijn, S.; Meijer, D.H.; Gainche, L.; Sena-Esteves, M.; Curry, W.T., Jr.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef]
- Yamamoto, Y.; Kosaka, N.; Tanaka, M.; Koizumi, F.; Kanai, Y.; Mizutani, T.; Murakami, Y.; Kuroda, M.; Miyajima, A.; Kato, T.; Ochiya, T. MicroRNA-500 as a potential diagnostic marker for hepatocellular carcinoma. Biomarkers 2009, 14, 529–538. [Google Scholar] [CrossRef]
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006. [Google Scholar] [CrossRef]
- Wong, T.S.; Liu, X.B.; Wong, B.Y.; Ng, R.W.; Yuen, A.P.; Wei, W.I. Mature miR-184 as potential oncogenic microRNA of squamous cell carcinoma of tongue. Clin. Cancer Res. 2008, 14, 2588–2592. [Google Scholar] [CrossRef]
- Baraniskin, A.; Nopel-Dunnebacke, S.; Ahrens, M.; Jensen, S.G.; Zollner, H.; Maghnouj, A.; Wos, A.; Mayerle, J.; Munding, J.; Kost, D.; et al. Circulating U2 small nuclear RNA fragments as a novel diagnostic biomarker for pancreatic and colorectal adenocarcinoma. Int. J. Cancer. 2012, 132, E48–E57. [Google Scholar] [PubMed]
- Bhat, K.; Wang, F.; Ma, Q.; Li, Q.; Mallik, S.; Hsieh, T.C.; Wu, E. Advances in biomarker research for pancreatic cancer. Curr. Pharm. Design 2012, 18, 2439–2451. [Google Scholar] [CrossRef]
- Liu, R.; Chen, X.; Du, Y.; Yao, W.; Shen, L.; Wang, C.; Hu, Z.; Zhuang, R.; Ning, G.; Zhang, C.; et al. Serum microRNA expression profile as a biomarker in the diagnosis and prognosis of pancreatic cancer. Clin. Chem. 2012, 58, 610–618. [Google Scholar] [CrossRef]
- Morimura, R.; Komatsu, S.; Ichikawa, D.; Takeshita, H.; Tsujiura, M.; Nagata, H.; Konishi, H.; Shiozaki, A.; Ikoma, H.; Okamoto, K.; Ochiai, T.; Taniguchi, H.; Otsuji, E. Novel diagnostic value of circulating miR-18a in plasma of patients with pancreatic cancer. Br. J. Cancer 2011, 105, 1733–1740. [Google Scholar] [CrossRef]
- Zheng, J.; Dong, P.; Gao, S.; Wang, N.; Yu, F. High expression of serum miR-17-5p associated with poor prognosis in patients with hepatocellular carcinoma. Hepato-Gastroenterology 2012, 60. [Google Scholar] [CrossRef]
- Qi, J.; Wang, J.; Katayama, H.; Sen, S.; Liu, S.M. Circulating microRNAs (cmiRNAs) as novel potential biomarkers for hepatocellular carcinoma. Neoplasma 2013, 60, 135–142. [Google Scholar] [PubMed]
- Kirschner, M.B.; Kao, S.C.; Edelman, J.J.; Armstrong, N.J.; Vallely, M.P.; van Zandwijk, N.; Reid, G. Haemolysis during sample preparation alters microRNA content of plasma. PLoS ONE 2011, 6, e24145. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.M.; Lee, J.Y.; Ho, C.C.; Hong, Q.S.; Yu, S.L.; Tzeng, C.R.; Yang, P.C.; Chen, H.W. miRNA-34b as a tumor suppressor in estrogen-dependent growth of breast cancer cells. Breast Cancer Res. 2011, 13, R166. [Google Scholar]
- Heneghan, H.M.; Miller, N.; Kelly, R.; Newell, J.; Kerin, M.J. Systemic miRNA-195 differentiates breast cancer from other malignancies and is a potential biomarker for detecting noninvasive and early stage disease. Oncologist 2010, 15, 673–682. [Google Scholar] [CrossRef]
- Frankel, L.B.; Christoffersen, N.R.; Jacobsen, A.; Lindow, M.; Krogh, A.; Lund, A.H. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J. Biol. Chem. 2008, 283, 1026–1033. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Huang, D.; Ni, S.; Peng, Z.; Sheng, W.; Du, X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int. J. Cancer 2010, 127, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Ichimi, T.; Enokida, H.; Okuno, Y.; Kunimoto, R.; Chiyomaru, T.; Kawamoto, K.; Kawahara, K.; Toki, K.; Kawakami, K.; Nishiyama, K.; Tsujimoto, G.; Nakagawa, M.; Seki, N. Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int. J. Cancer 2009, 125, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Conti, A.; Aguennouz, M.; La Torre, D.; Tomasello, C.; Cardali, S.; Angileri, F.F.; Maio, F.; Cama, A.; Germano, A.; Vita, G.; Tomasello, F. miR-21 and 221 upregulation and miR-181b downregulation in human grade II-IV astrocytic tumors. J. Neuro-Oncology 2009, 93, 325–332. [Google Scholar] [CrossRef]
- Zheng, B.; Liang, L.; Huang, S.; Zha, R.; Liu, L.; Jia, D.; Tian, Q.; Wang, Q.; Wang, C.; Long, Z.; et al. MicroRNA-409 suppresses tumour cell invasion and metastasis by directly targeting radixin in gastric cancers. Oncogene 2012, 31, 4509–4516. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, X.; Zhao, Y.; Tian, T.; Jin, G.; Shu, Y.; Chen, Y.; Xu, L.; Zen, K.; Zhang, C.; Shen, H. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J. Clin. Ooncol. 2010, 28, 1721–1726. [Google Scholar] [CrossRef]
- Silva, J.; Garcia, V.; Zaballos, A.; Provencio, M.; Lombardia, L.; Almonacid, L.; Garcia, J.M.; Dominguez, G.; Pena, C.; Diaz, R.; et al. Vesicle-related microRNAs in plasma of nonsmall cell lung cancer patients and correlation with survival. Eur. Respir. J. 2011, 37, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.J.; Kao, S.Y.; Tu, H.F.; Tsai, M.M.; Chang, K.W.; Lin, S.C. Increase of microRNA miR-31 level in plasma could be a potential marker of oral cancer. Oral Dis. 2010, 16, 360–364. [Google Scholar] [CrossRef] [PubMed]
- Bryant, R.J.; Pawlowski, T.; Catto, J.W.; Marsden, G.; Vessella, R.L.; Rhees, B.; Kuslich, C.; Visakorpi, T.; Hamdy, F.C. Changes in circulating microRNA levels associated with prostate cancer. Br. J. Cancer 2012, 106, 768–774. [Google Scholar] [CrossRef] [PubMed]
- Brase, J.C.; Johannes, M.; Schlomm, T.; Falth, M.; Haese, A.; Steuber, T.; Beissbarth, T.; Kuner, R.; Sultmann, H. Circulating miRNAs are correlated with tumor progression in prostate cancer. Int. J. Cancer 2011, 128, 608–616. [Google Scholar] [CrossRef] [PubMed]
- Greither, T.; Grochola, L.F.; Udelnow, A.; Lautenschlager, C.; Wurl, P.; Taubert, H. Elevated expression of microRNAs 155, 203, 210 and 222 in pancreatic tumors is associated with poorer survival. Int. J. Cancer 2010, 126, 73–80. [Google Scholar] [CrossRef]
- Wang, J.; Chen, J.; Chang, P.; LeBlanc, A.; Li, D.; Abbruzzesse, J.L.; Frazier, M.L.; Killary, A.M.; Sen, S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev. Res. (Phila) 2009, 2, 807–813. [Google Scholar] [CrossRef]
- Fornari, F.; Milazzo, M.; Chieco, P.; Negrini, M.; Marasco, E.; Capranico, G.; Mantovani, V.; Marinello, J.; Sabbioni, S.; Callegari, E.; et al. In hepatocellular carcinoma miR-519d is up-regulated by p53 and DNA hypomethylation and targets CDKN1A/p21, PTEN, AKT3 and TIMP2. J. Pathol. 2012, 227, 275–285. [Google Scholar] [CrossRef]
- Snowdon, J.; Zhang, X.; Childs, T.; Tron, V.A.; Feilotter, H. The microRNA-200 family is upregulated in endometrial carcinoma. PLoS ONE 2011, 6. [Google Scholar] [CrossRef]
- White, N.M.; Bao, T.T.; Grigull, J.; Youssef, Y.M.; Girgis, A.; Diamandis, M.; Fatoohi, E.; Metias, M.; Honey, R.J.; Stewart, R.; et al. miRNA profiling for clear cell renal cell carcinoma: Biomarker discovery and identification of potential controls and consequences of miRNA dysregulation. J. Urol. 2011, 186, 1077–1083. [Google Scholar] [CrossRef] [PubMed]
- Caramuta, S.; Egyhazi, S.; Rodolfo, M.; Witten, D.; Hansson, J.; Larsson, C.; Lui, W.O. MicroRNA expression profiles associated with mutational status and survival in malignant melanoma. J. Investig. Dermatol. 2010, 130, 2062–2070. [Google Scholar] [CrossRef]
- Nikiforova, M.N.; Tseng, G.C.; Steward, D.; Diorio, D.; Nikiforov, Y.E. MicroRNA expression profiling of thyroid tumors: biological significance and diagnostic utility. J. Clin. Endocrinol. Metabol. 2008, 93, 1600–1608. [Google Scholar] [CrossRef]
- Lulla, R.R.; Costa, F.F.; Bischof, J.M.; Chou, P.M.; de, F.B.M.; Vanin, E.F.; Soares, M.B. Identification of differentially expressed microRNAs in osteosarcoma. Sarcoma 2011, 2011. [Google Scholar] [CrossRef]
- Thayanithy, V.; Sarver, A.L.; Kartha, R.V.; Li, L.; Angstadt, A.Y.; Breen, M.; Steer, C.J.; Modiano, J.F.; Subramanian, S. Perturbation of 14q32 miRNAs-cMYC gene network in osteosarcoma. Bone 2012, 50, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Zavadil, J.; Ye, H.; Liu, Z.; Wu, J.; Lee, P.; Hernando, E.; Soteropoulos, P.; Toruner, G.A.; Wei, J.J. Profiling and functional analyses of microRNAs and their target gene products in human uterine leiomyomas. PloS ONE 2010, 5. [Google Scholar] [CrossRef]
- Miyachi, M.; Tsuchiya, K.; Yoshida, H.; Yagyu, S.; Kikuchi, K.; Misawa, A.; Iehara, T.; Hosoi, H. Circulating muscle-specific microRNA, miR-206, as a potential diagnostic marker for rhabdomyosarcoma. Biochem. Biophys. Res. Commun. 2010, 400, 89–93. [Google Scholar] [CrossRef]
- Koelz, M.; Lense, J.; Wrba, F.; Scheffler, M.; Dienes, H.P.; Odenthal, M. Down-regulation of miR-221 and miR-222 correlates with pronounced Kit expression in gastrointestinal stromal tumors. Int. J. Oncology 2011, 38, 503–511. [Google Scholar]
- Ban, J.; Jug, G.; Mestdagh, P.; Schwentner, R.; Kauer, M.; Aryee, D.N.; Schaefer, K.L.; Nakatani, F.; Scotlandi, K.; Reiter, M.; Strunk, D.; Speleman, F.; Vandesompele, J.; Kovar, H. Hsa-mir-145 is the top EWS-FLI1-repressed microRNA involved in a positive feedback loop in Ewing's sarcoma. Oncogene 2011, 30, 2173–2180. [Google Scholar] [CrossRef]
- Saydam, O.; Senol, O.; Wurdinger, T.; Mizrak, A.; Ozdener, G.B.; Stemmer-Rachamimov, A.O.; Yi, M.; Stephens, R.M.; Krichevsky, A.M.; Saydam, N.; Brenner, G.J.; Breakefield, X.O. miRNA-7 attenuation in Schwannoma tumors stimulates growth by upregulating three oncogenic signaling pathways. Cancer Res. 2011, 71, 852–861. [Google Scholar] [CrossRef] [PubMed]
- Sarver, A.L.; Li, L.; Subramanian, S. MicroRNA miR-183 functions as an oncogene by targeting the transcription factor EGR1 and promoting tumor cell migration. Cancer Res. 2010, 70, 9570–9580. [Google Scholar] [CrossRef]
- Subramanian, S.; Thayanithy, V.; West, R.B.; Lee, C.H.; Beck, A.H.; Zhu, S.; Downs-Kelly, E.; Montgomery, K.; Goldblum, J.R.; Hogendoorn, P.C.; et al. Genome-wide transcriptome analyses reveal p53 inactivation mediated loss of miR-34a expression in malignant peripheral nerve sheath tumours. J. Pathol. 2010, 220, 58–70. [Google Scholar] [CrossRef]
- Yamagishi, M.; Nakano, K.; Miyake, A.; Yamochi, T.; Kagami, Y.; Tsutsumi, A.; Matsuda, Y.; Sato-Otsubo, A.; Muto, S.; Utsunomiya, A.; et al. Polycomb-mediated loss of miR-31 activates NIK-dependent NF-kappaB pathway in adult T cell leukemia and other cancers. Cancer Cell. 2012, 21, 121–135. [Google Scholar] [CrossRef]
- Careccia, S.; Mainardi, S.; Pelosi, A.; Gurtner, A.; Diverio, D.; Riccioni, R.; Testa, U.; Pelosi, E.; Piaggio, G.; Sacchi, A.; et al. A restricted signature of miRNAs distinguishes APL blasts from normal promyelocytes. Oncogene 2009, 28, 4034–4040. [Google Scholar] [CrossRef] [PubMed]
- Pigazzi, M.; Manara, E.; Baron, E.; Basso, G. miR-34b targets cyclic AMP-responsive element binding protein in acute myeloid leukemia. Cancer Res. 2009, 69, 2471–2478. [Google Scholar] [CrossRef]
- Ghosh, A.K.; Shanafelt, T.D.; Cimmino, A.; Taccioli, C.; Volinia, S.; Liu, C.G.; Calin, G.A.; Croce, C.M.; Chan, D.A.; Giaccia, A.J.; et al. Aberrant regulation of pVHL levels by microRNA promotes the HIF/VEGF axis in CLL B cells. Blood 2009, 113, 5568–5574. [Google Scholar] [CrossRef] [PubMed]
- Stamatopoulos, B.; Meuleman, N.; Haibe-Kains, B.; Saussoy, P.; Van Den Neste, E.; Michaux, L.; Heimann, P.; Martiat, P.; Bron, D.; Lagneaux, L. microRNA-29c and microRNA-223 down-regulation has in vivo significance in chronic lymphocytic leukemia and improves disease risk stratification. Blood 2009, 113, 5237–5245. [Google Scholar] [CrossRef]
- Fang, C.; Zhu, D.X.; Dong, H.J.; Zhou, Z.J.; Wang, Y.H.; Liu, L.; Fan, L.; Miao, K.R.; Liu, P.; Xu, W.; Li, J.Y. Serum microRNAs are promising novel biomarkers for diffuse large B cell lymphoma. Ann. Hematol. 2012, 91, 553–559. [Google Scholar] [CrossRef]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The microRNA spectrum in 12 body fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef]
- Mahn, R.; Heukamp, L.C.; Rogenhofer, S.; von Ruecker, A.; Muller, S.C.; Ellinger, J. Circulating microRNAs (miRNA) in serum of patients with prostate cancer. Urology 2011, 77, 1265.e9–1265.e16. [Google Scholar] [CrossRef]
- Yamada, Y.; Enokida, H.; Kojima, S.; Kawakami, K.; Chiyomaru, T.; Tatarano, S.; Yoshino, H.; Kawahara, K.; Nishiyama, K.; Seki, N.; Nakagawa, M. MiR-96 and miR-183 detection in urine serve as potential tumor markers of urothelial carcinoma: Correlation with stage and grade, and comparison with urinary cytology. Cancer Sci. 2011, 102, 522–529. [Google Scholar] [CrossRef]
- Baraniskin, A.; Kuhnhenn, J.; Schlegel, U.; Maghnouj, A.; Zollner, H.; Schmiegel, W.; Hahn, S.; Schroers, R. Identification of microRNAs in the cerebrospinal fluid as biomarker for the diagnosis of glioma. J. Neuro-Oncology 2012, 14, 29–33. [Google Scholar] [CrossRef]
- Wu, L.; Zhou, H.; Lin, H.; Qi, J.; Zhu, C.; Gao, Z.; Wang, H. Circulating microRNAs are elevated in plasma from severe preeclamptic pregnancies. Reproduction 2012, 143, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Todd, N.W.; Liu, Z.; Zhan, M.; Fang, H.; Peng, H.; Alattar, M.; Deepak, J.; Stass, S.A.; Jiang, F. Altered miRNA expression in sputum for diagnosis of non-small cell lung cancer. Lung Cancer 2010, 67, 170–176. [Google Scholar] [CrossRef]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Hunter, M.P.; Ismail, N.; Zhang, X.; Aguda, B.D.; Lee, E.J.; Yu, L.; Xiao, T.; Schafer, J.; Lee, M.L.; Schmittgen, T.D.; Nana-Sinkam, S.P.; Jarjoura, D.; Marsh, C.B. Detection of microRNA expression in human peripheral blood microvesicles. PLoS ONE 2008, 3. [Google Scholar] [CrossRef]
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Fichtlscherer, S.; Zeiher, A.M.; Dimmeler, S. Circulating microRNAs: biomarkers or mediators of cardiovascular diseases? Arterioscler. Thromb. Vasc. Biol. 2011, 31, 2383–2390. [Google Scholar] [CrossRef]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–433. [Google Scholar] [CrossRef]
- Gilad, S.; Meiri, E.; Yogev, Y.; Benjamin, S.; Lebanony, D.; Yerushalmi, N.; Benjamin, H.; Kushnir, M.; Cholakh, H.; Melamed, N.; Bentwich, Z.; Hod, M.; Goren, Y.; Chajut, A. Serum microRNAs are promising novel biomarkers. PloS ONE 2008, 3. [Google Scholar] [CrossRef]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef]
- Hu, Z.; Chen, X.; Zhao, Y.; Tian, T.; Jin, G.; Shu, Y.; Chen, Y.; Xu, L.; Zen, K.; Zhang, C.; Shen, H. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J. Clin. Oncol. 2010, 28, 1721–1726. [Google Scholar] [CrossRef]
- Cummins, J.M.; He, Y.; Leary, R.J.; Pagliarini, R.; Diaz, L.A., Jr.; Sjoblom, T.; Barad, O.; Bentwich, Z.; Szafranska, A.E.; Labourier, E.; et al. The colorectal microRNAome. Proc. Natl. Acad. Sci. USA 2006, 103, 3687–3692. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.T.; Baldwin, D.A.; Scearce, L.M.; Oberholtzer, J.C.; Tobias, J.W.; Mourelatos, Z. Microarray-based, high-throughput gene expression profiling of microRNAs. Nat. Methods 2004, 1, 155–161. [Google Scholar] [CrossRef]
- Ajit, S.K. Circulating microRNAs as biomarkers, therapeutic targets, and signaling molecules. Sensors 2012, 12, 3359–3369. [Google Scholar] [CrossRef]
- Steer, C.J.; Subramanian, S. Circulating microRNAs as biomarkers: a new frontier in diagnostics. Liver Transplant. 2012, 18, 265–269. [Google Scholar] [CrossRef]
- LaConti, J.J.; Shivapurkar, N.; Preet, A.; Deslattes Mays, A.; Peran, I.; Kim, S.E.; Marshall, J.L.; Riegel, A.T.; Wellstein, A. Tissue and serum microRNAs in the Kras(G12D) transgenic animal model and in patients with pancreatic cancer. PloS ONE 2011, 6. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sundarbose, K.; Kartha, R.V.; Subramanian, S. MicroRNAs as Biomarkers in Cancer. Diagnostics 2013, 3, 84-104. https://doi.org/10.3390/diagnostics3010084
Sundarbose K, Kartha RV, Subramanian S. MicroRNAs as Biomarkers in Cancer. Diagnostics. 2013; 3(1):84-104. https://doi.org/10.3390/diagnostics3010084
Chicago/Turabian StyleSundarbose, Kamini, Reena V. Kartha, and Subbaya Subramanian. 2013. "MicroRNAs as Biomarkers in Cancer" Diagnostics 3, no. 1: 84-104. https://doi.org/10.3390/diagnostics3010084
APA StyleSundarbose, K., Kartha, R. V., & Subramanian, S. (2013). MicroRNAs as Biomarkers in Cancer. Diagnostics, 3(1), 84-104. https://doi.org/10.3390/diagnostics3010084






