Differential Expression of Six Rnase2 and Three Rnase3 Paralogs Identified in Blunt Snout Bream in Response to Aeromonas hydrophila Infection
Abstract
:1. Introduction
2. Materials and Methods
2.1. Collection of Samples
2.2. Identification of Rnase2 and Rnase3 Genes
2.3. Protein Alignment and Physicochemical Properties
2.4. Structural Predictions
2.5. Phylogenetic Analysis
2.6. Quantitative Analysis of Rnase2 and Rnase3 Messenger RNA Expression
2.7. Bacterial Challenge Experiment
2.8. Statistical Analyses
3. Results
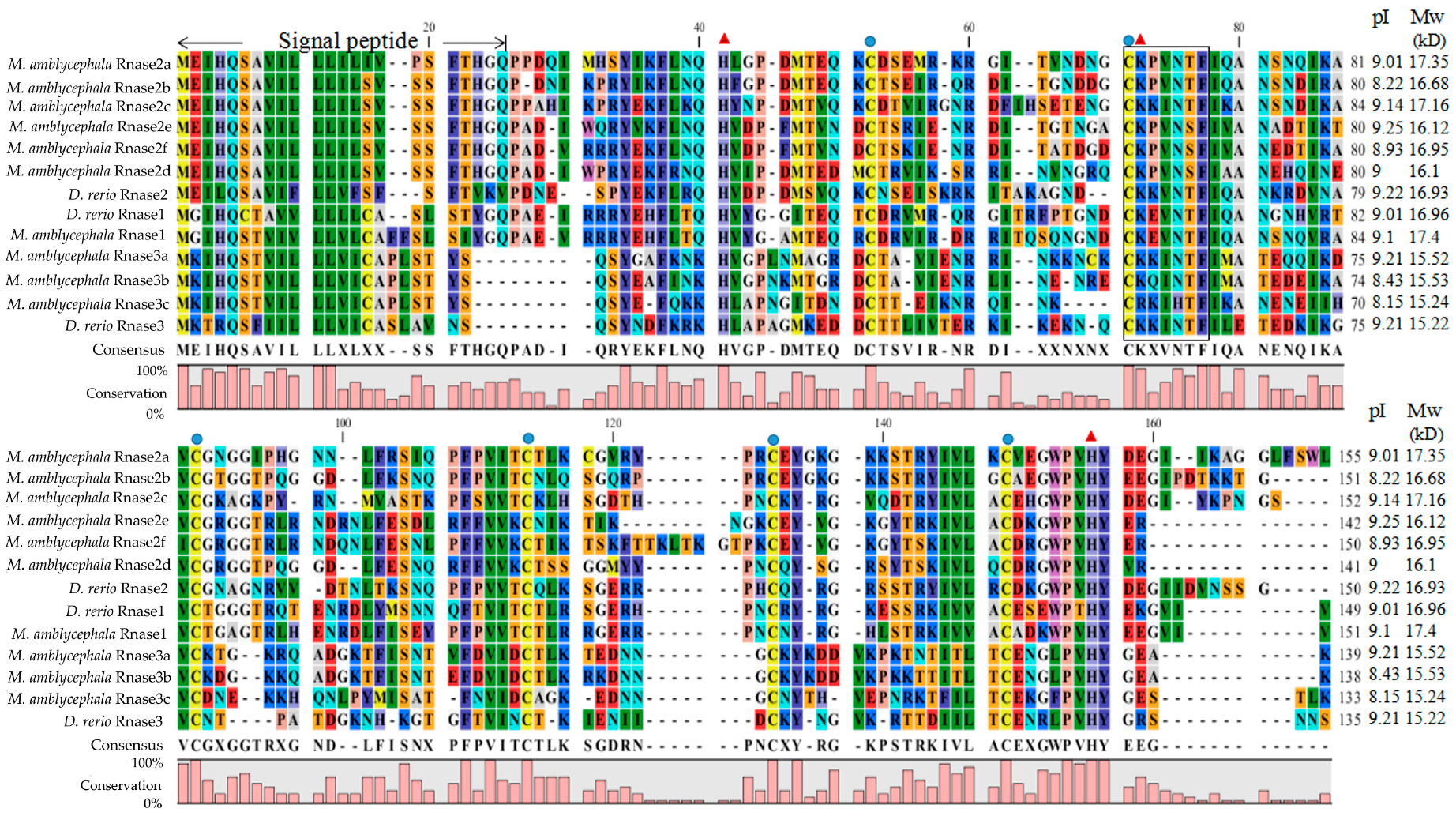
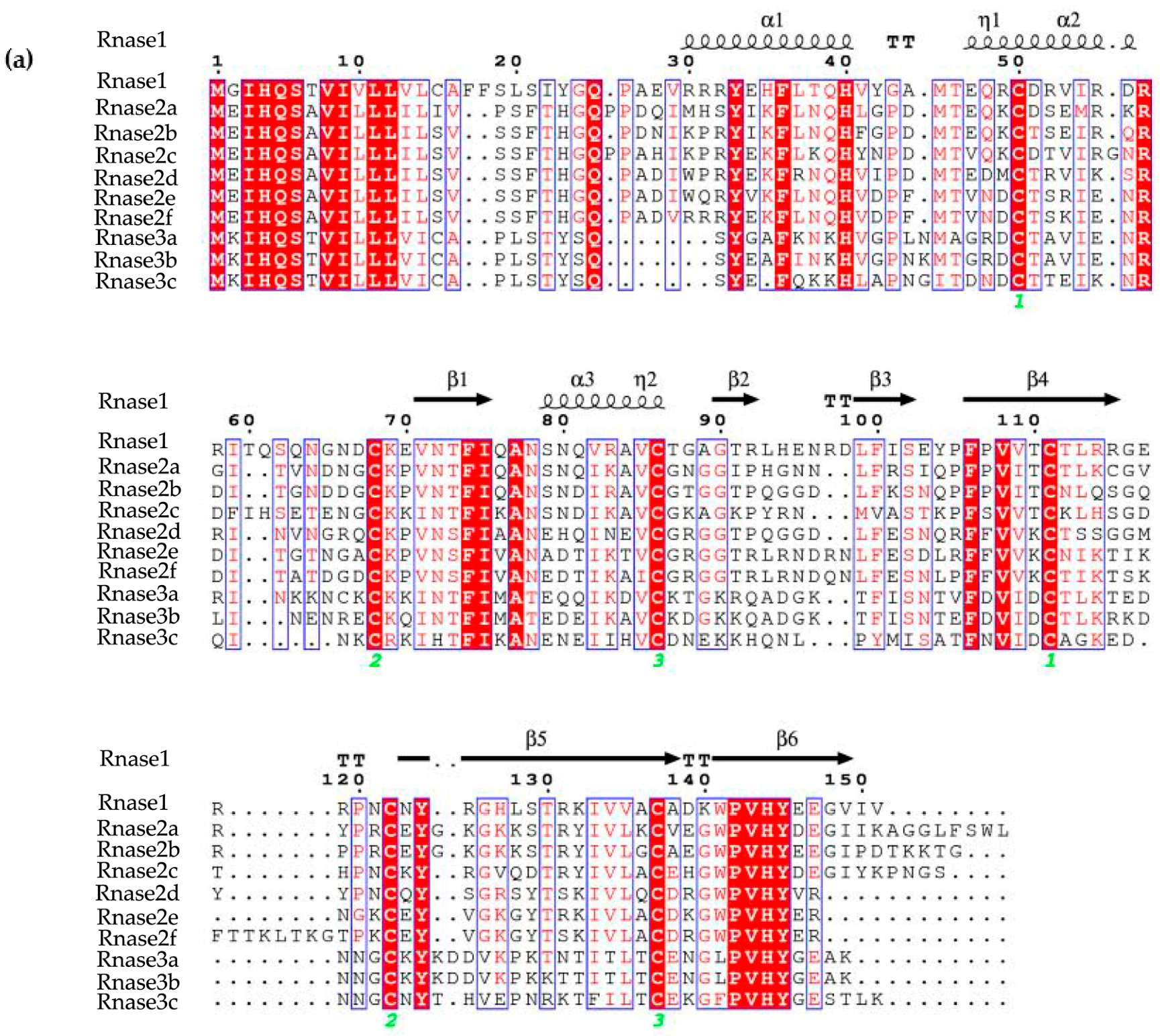
3.1. Protein Alignment
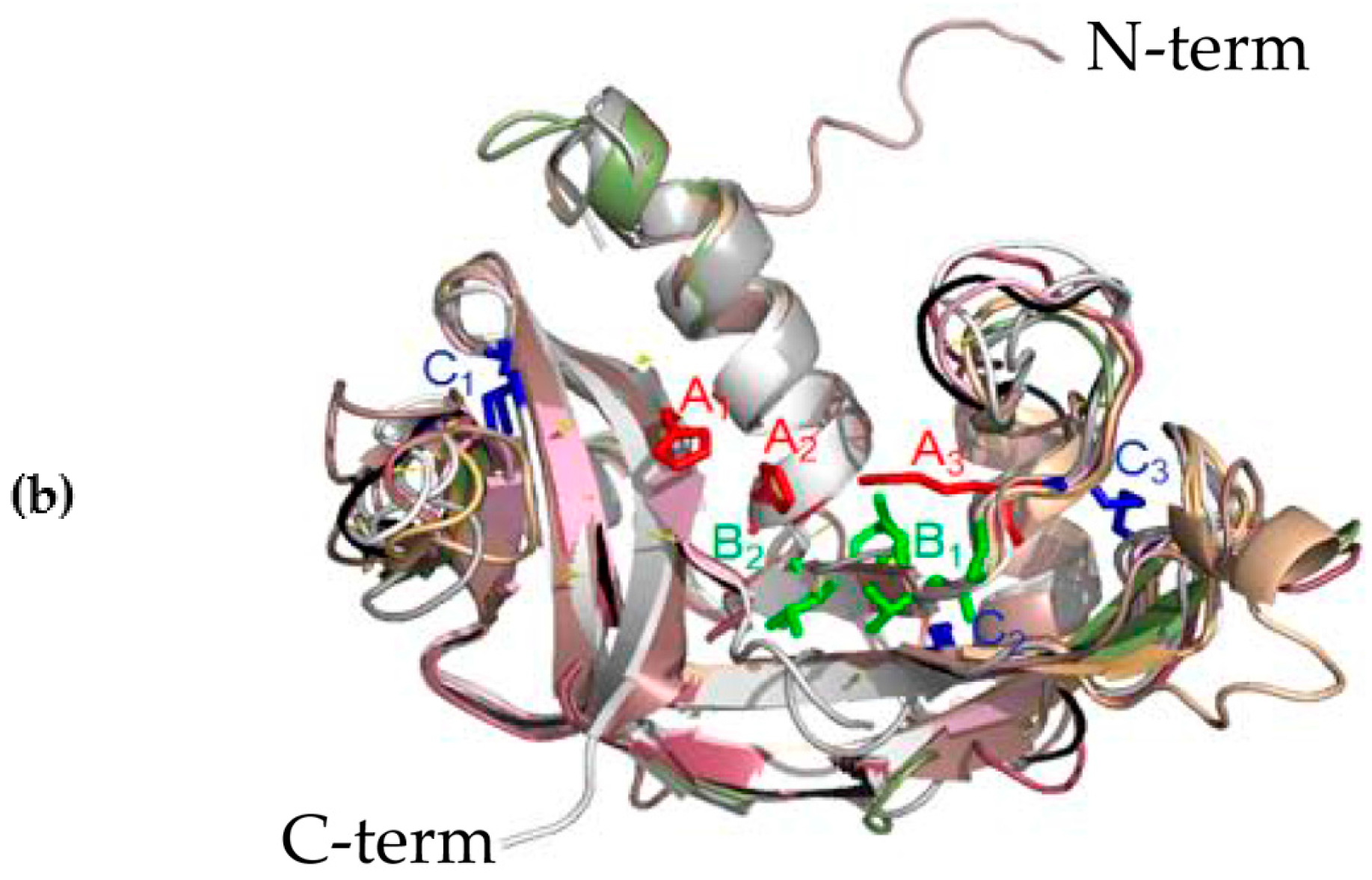
3.2. Structural Predictions
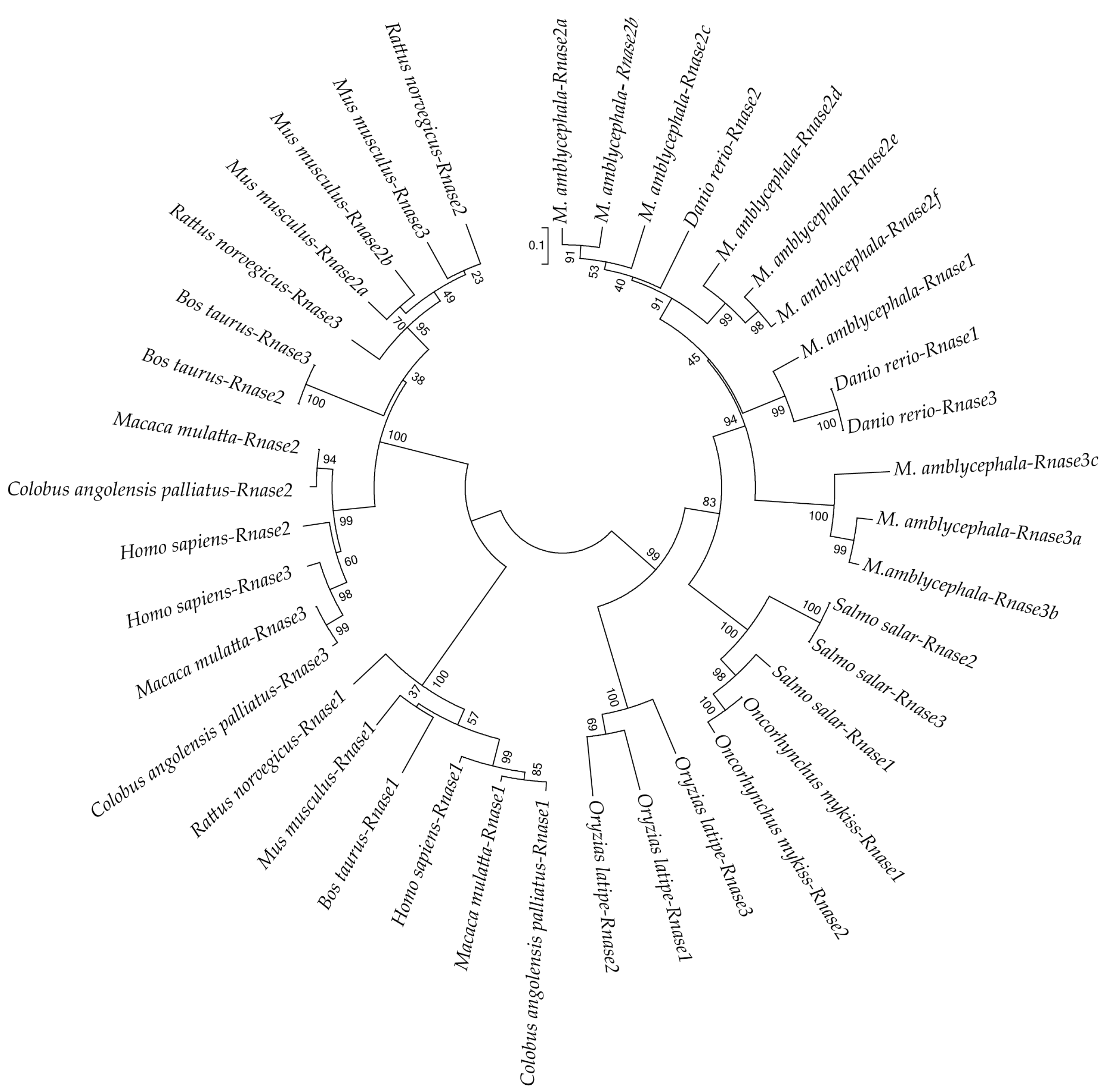
3.3. Phylogenetic Analysis
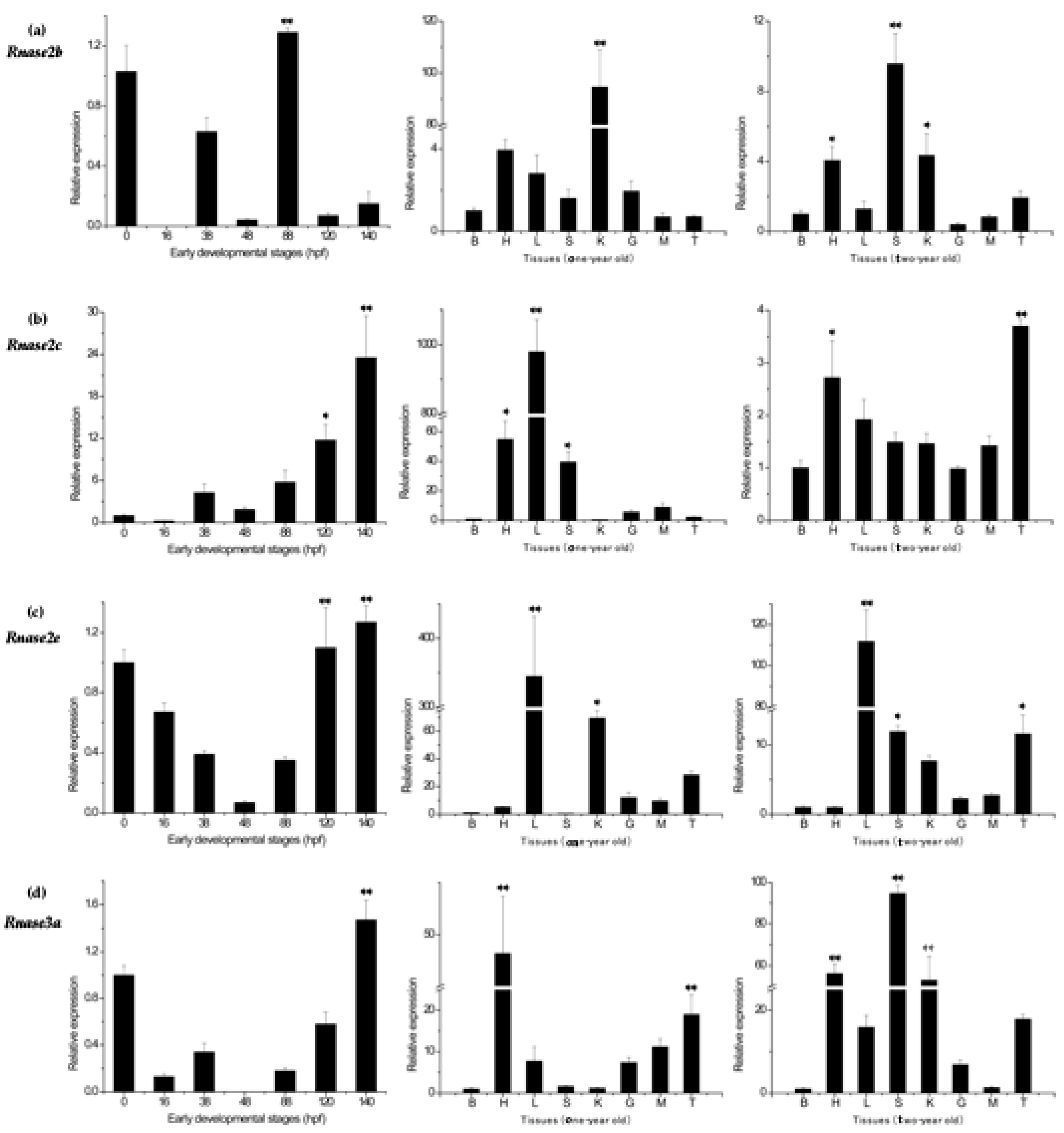
3.4. Rnase2 and Rnase3 Messenger RNA Expression Patterns
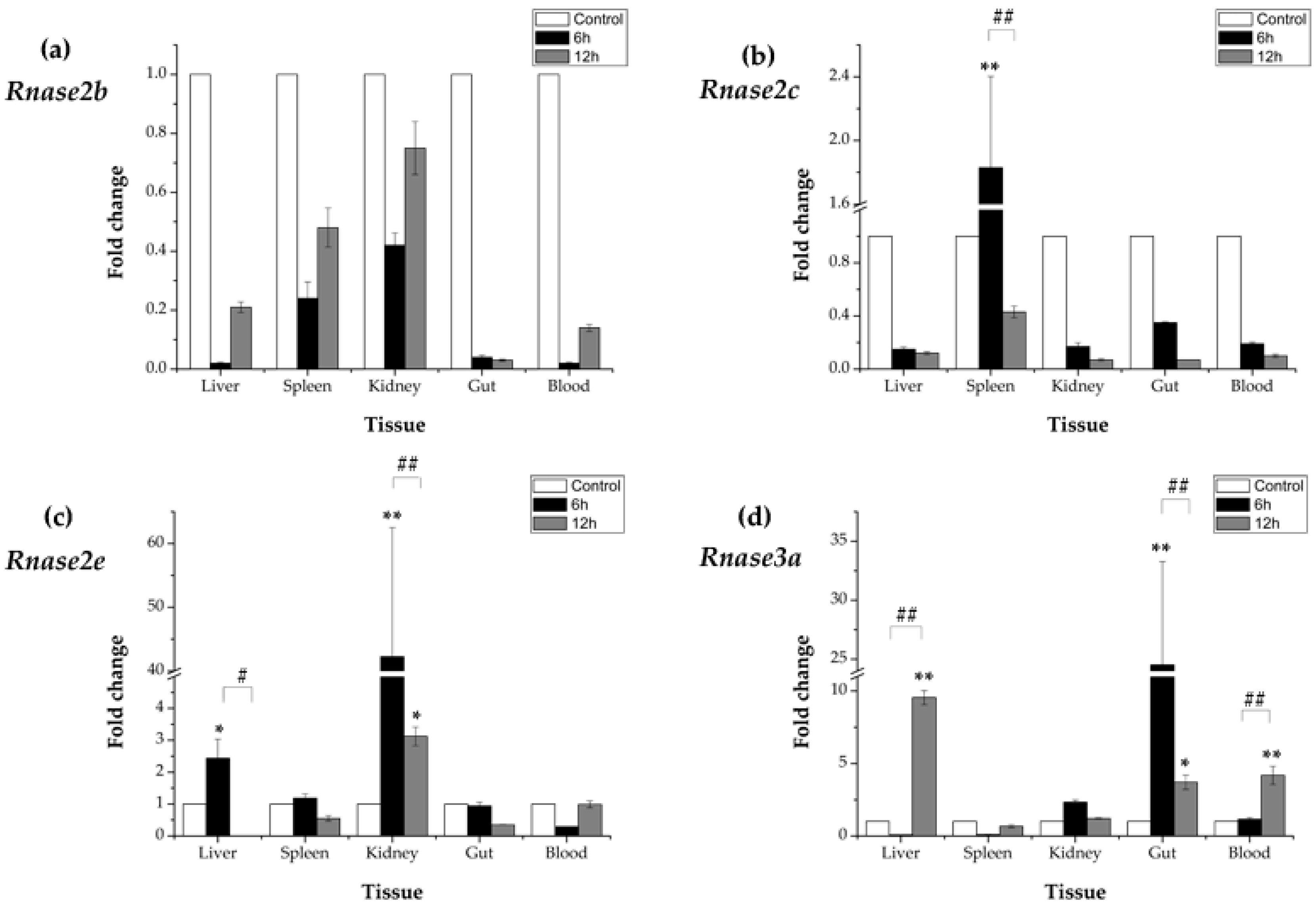
3.5. Expression Profiles of Rnase2 and Rnase3 Messenger RNA after Aeromonas hydrophila Challenge
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Barnard, E.A. Biological function of pancreatic ribonuclease. Nature 1969, 221, 340–344. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.; Beintema, J.J.; Zhang, J. The ribonuclease A superfamily of mammals and birds: Identifying new members and tracing evolutionary histories. Genomics 2005, 85, 208–220. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.; Zhang, J. Zebrafish ribonucleases are bactericidal: Implications for the origin of the vertebrate RnaseA superfamily. Mol. Biol. Evol. 2007, 24, 1259–1268. [Google Scholar] [CrossRef] [PubMed]
- Kazakou, K.; Holloway, D.E.; Prior, S.H.; Subramanian, V.; Acharya, K.R. Ribonuclease A homologues of the zebrafish: Polymorphism, crystal structures of two representatives and their evolutionary implications. J. Mol. Biol. 2008, 380, 206–222. [Google Scholar] [CrossRef] [PubMed]
- Pizzo, E.; Buonanno, P.; Di, M.A.; Ponticelli, S.; De, F.S.; Quarto, N.; Cubellis, M.V.; D’Alessio, G. Ribonucleases and angiogenins from fish. J. Biol. Chem. 2006, 281, 27454–27460. [Google Scholar] [CrossRef] [PubMed]
- Pizzo, E.; Merlino, A.; Turano, M.; Russo, K.I.; Coscia, F.; Zanfardino, A.; Varcamonti, M.; Furia, A.; Giancola, C.; Mazzarella, L.; et al. A new Rnase sheds light on the Rnase/angiogenin subfamily from zebrafish. Biochem. J. 2011, 433, 345–355. [Google Scholar] [CrossRef] [PubMed]
- Pizzo, E.; Varcamonti, M.; Di, M.A.; Zanfardino, A.; Giancola, C.; D’Alessio, G. Ribonucleases with angiogenic and bactericidal activities from the Atlantic salmon. FFBS J. 2008, 275, 1283–1295. [Google Scholar] [CrossRef] [PubMed]
- Lang, D.; Zhang, Y.; Yu, L. Molecular evolution of the ribonuclease A superfamily. Hereditas 2014, 36, 165–192. [Google Scholar]
- Wheeler, T.T.; Maqbool, N.J.; Gupta, S.K. Mapping, phylogenetic and expression analysis of the Rnase (RnaseA) locus in cattle. J. Mol. Evol. 2012, 74, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Tao, F.Y.; Wei, Z.; Ma, R.Y. Progress in bactericidal activity of ribonuclease A superfamily members. Chin. J. Antibiot. 2009, 34, 325–379. [Google Scholar]
- Xu, H.; Li, D. Research progress of human Ribonuclease A superfamily. Chem. Life 2012, 32, 174–179. [Google Scholar]
- Gupta, S.K.; Haigh, B.J.; Griffin, F.J.; Wheeler, T.T. The mammalian secreted Rnases: Mechanisms of action in host defence. Innate Immun. 2013, 19, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Sorrentino, S. The eight human “canonical” ribonucleases: Molecular diversity, catalytic properties, and special biological actions of the enzyme proteins. FEBS Lett. 2010, 584, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Attery, A.; Batra, J.K. Mouse eosinophil associated ribonucleases: Mechanism of cytotoxic, antibacterial and antiparasitic activities. Int. J. Biol. Macromol. 2017, 94, 445–450. [Google Scholar] [CrossRef] [PubMed]
- Acharya, K.R.; Ackerman, S.J. Eosinophil granule proteins: form and function. J. Biol. Chem. 2014, 289, 17406–17415. [Google Scholar] [CrossRef] [PubMed]
- Futami, J. Tissue-specific expression of pancreatic-type RNases and RNase inhibitor in humans. DNA Cell. Biol. 1997, 16, 413–419. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, H.F. Eosinophil-derived Neurotoxin/RNase 2: Connecting the past, the present and the future. Curr. Pharm. Biotechnol. 2008, 9, 135–140. [Google Scholar] [CrossRef] [PubMed]
- Bedoya, V.I.; Boasso, A.; Hardy, A.W.; Rybak, S.; Shearer, G.M.; Rugeles, M.T. Ribonucleases in HIV type 1 inhibition: effect of recombinant RNases on infection of primary T cells and immune activation-induced RNase gene and protein expression. Aids Res. Hum. Retrov. 2006, 22, 897–907. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, H.F.; Dyer, K.D.; Domachowske, J.B. Respiratory viruses and eosinophils: Exploring the connections. Antivir. Res. 2009, 83, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Koczera, P.; Martin, L.; Marx, G.; Tobias, S. The ribonuclease A superfamily in humans: Canonical Rnases as the buttress of innate immunity. Int. J. Mol. Sci. 2016, 17, 1278. [Google Scholar] [CrossRef] [PubMed]
- Gullberg, U.; Widegren, B.; Arnason, U.; Egesten, A.; Olsson, I. The cytotoxic eosinophil cationic protein (ECP) has ribonuclease activity. Biochem. Bioph. Res. Co. 1986, 139, 1239–1242. [Google Scholar] [CrossRef]
- Rosenberg, H.F.; Tenen, D.G.; Ackerman, S.J. Molecular cloning of the human eosinophil-derived neurotoxin: a member of the ribonuclease gene family. Proc. Natl. Acad. Sci. USA 1989, 86, 4460–4464. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, H.F.; Dyer, K.D. Eosinophil cationic protein and eosinophil-derived neurotoxin, evolution of novel function in a primate ribonuclease gene family. J. Biol. Chem. 1995, 270, 21539–21542. [Google Scholar] [CrossRef] [PubMed]
- Carreras, E.; Boix, E.; Rosenberg, H.F.; Cuchillo, C.M.; Nogués, M.V. Both aromatic and cationic residues contribute to the membrane-lytic and bactericidal activity of eosinophil cationic protein. Biochemistry 2003, 42, 6636–6644. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Batra, J.K. Role of unique basic residues in cytotoxic, antibacterial and antiparasitic activities of human eosinophil cationic protein. Biol. Chem. 2011, 392, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Pulido, D.; Torrent, M.; Andreu, D.; Nogués, M.V.; Boix, E. Two human host defense ribonucleases against mycobacteria, the eosinophil cationic protein (RNase3) and RNase7. Antimicrob. Agents. Chemother. 2013, 57, 3797–3805. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Rosenberg, H.F.; Nei, M. Positive Darwinian selection after gene duplication in primate ribonuclease genes. Proc. Natl. Acad. Sci. USA 1998, 95, 3708–3713. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J. Evolution by gene duplication: An update. Trends Ecol. Evol. 2003, 18, 292–298. [Google Scholar] [CrossRef]
- Liu, H.; Wang, W. Expression patterns and functional novelty of ribonuclease 1 in herbivorous Megalobrama amblycephala. Int. J. Mol. Sci. 2016, 17, 786. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Luo, W.; Liu, H.; Zeng, C.; Liu, X.; Yi, S.; Wang, W. Transcriptome analysis and SSR/SNP markers information of the blunt snout bream (Megalobrama amblycephala). PLoS ONE 2012, 7, e42637. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Zhao, Z.; Brown, P.B.; Cui, H.; Xie, J.; Tsion, H.M.; Ge, X. Dietary vitamin A requirement of juvenile Wuchang bream (Megalobrama amblycephala) determined by growth and disease resistance. Aquaculture 2016, 450, 23–30. [Google Scholar] [CrossRef]
- Liu, H.; Chen, C.; Gao, Z.; Min, J.; Gu, Y.; Jian, J.; Jiang, X.; Cai, H.; Ebersberger, I.; Xu, M.; et al. The draft genome of blunt snout bream (Megalobrama amblycephala) reveals the development of intermuscular bone and adaptation to herbivorous diet. Gigascience 2017, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, 597–603. [Google Scholar] [CrossRef] [PubMed]
- SignalP4.1 Server. Available online: http://www.cbs.dtu.dk/services/SignalP/ (accessed on 10 August 2017).
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Gallo Cassarino, T.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic. Acids Res. 2014, 42, W252. [Google Scholar] [CrossRef] [PubMed]
- Gouet, P.; Robert, X.; Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic. Acids. Res. 2003, 31, 3320–3323. [Google Scholar] [CrossRef] [PubMed]
- De Lano, W.L. The Py MOL Molecular Graphics System; De Lano Scientific: San Carlos, CA, USA, 2002. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- GenBank. Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 10 August 2017).
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Li, Z.; Yang, L.; Wang, J.; Shi, W.; Pawar, R.A.; Liu, Y.; Xu, C.; Cong, W.; Hu, Q.; Lu, T.; et al. β-Actin is a useful internal control for tissue-specific gene expression studies using quantitative real-time PCR in the half-smooth tongue sole Cynoglossus semilaevis challenged with LPS or Vibrio anguillarum. Fish Shellfish Immun. 2010, 29, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Giusti, D.; Gatouillat, G.; Jan, S.L.; Plée, J.; Bernard, P.; Antonicelli, F.; Pham, B.N. Eosinophil Cationic Protein (ECP), a predictive marker of bullous pemphigoid severity and outcome. Sci. Rep. 2017, 7, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Arnold, U. Stability and folding of amphibian ribonuclease A superfamily members in comparison with mammalian homologues. FEBS J. 2014, 281, 3559–3575. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Rosenberg, H.F. Complementary advantageous substitutions in the evolution of an antiviral RNase of higher primates. Proc. Natl. Acad. Sci. USA 2002, 99, 5486–5491. [Google Scholar] [CrossRef] [PubMed]
- Harder, J.; Schröder, J.M. RNase7, a novel innate immune defense antimicrobial protein of healthy human skin. J. Biol. Chem. 2002, 277, 46779–46784. [Google Scholar] [CrossRef] [PubMed]
- Nitto, T.; Dyer, K.D.; Czapiga, M.; Rosenberg, H.F. Evolution and function of leukocyte RNaseA ribonucleases of the avian species, Gallus gallus. J. Biol. Chem. 2006, 281, 25622–25634. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Stappenbeck, T.S.; Hong, C.V.; Gordon, J.I. Angiogenins: A new class of microbicidal proteins involved in innate immunity. Nat. Immun. 2003, 4, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Boix, E.; Salazar, V.A.; Torrent, M.; Pulido, D.; Nogués, M.V.; Moussaoui, M. Structural determinants of the eosinophil cationic protein antimicrobial activity. Biol. Chem. 2012, 393, 801–815. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J. Parallel adaptive origins of digestive RNases in Asian and African leaf monkeys. Nat. Genet. 2006, 38, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, H.F. Eosinophil-derived neurotoxin (EDN/RNase2) and the mouse eosinophil-associated RNases (mEars): Expanding roles in promoting host defense. Int. J. Mol. Sci. 2015, 16, 15442–15455. [Google Scholar] [CrossRef] [PubMed]
- Goo, S.M.; Cho, S. The expansion and functional diversification of the mammalian ribonuclease a superfamily epitomizes the efficiency of multigene families at generating biological novelty. Genome Biol. Evol. 2013, 5, 2124. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef] [PubMed]
- Rugeles, M.T.; Trubey, C.M.; Bedoya, V.I.; Pinto, L.A.; Oppenheim, J.J.; Rybak, S.M.; Shearer, G.M. Ribonuclease is partly responsible for the HIV-1 inhibitory effect activated by HLA alloantigen recognition. Aids. 2003, 17, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Gagne, D.; Narayanan, C.; Bafna, K.; Charest, L.A.; Agarwal, P.K.; Doucet, N. Sequence-specific backbone resonance assignments and microsecond timescale molecular dynamics simulation of human eosinophil-derived neurotoxin. Biomol. NMR Assignm. 2017, 11, 143–149. [Google Scholar] [CrossRef] [PubMed]
- Pulido, D.; Garcia-Mayoral, M.F.; Moussaoui, M.; Velázquez, D.; Torrent, M.; Bruix, M.; Boix, E. Structural basis for endotoxin neutralization by the Eosinophil Cationic Protein. FEBS J. 2016, 283, 4176–4191. [Google Scholar] [CrossRef] [PubMed]
| Primer Name | Primer Sequence (5′-3′) | Fragment Length |
|---|---|---|
| β-actin | F:ACCCACACCGTGCCCATCTA R:CGGACAATTTCTCTTTCGGCTG | 204 |
| Rnase2a | F:GTCAACCACCAGACCAAAT R:AAGGCTGAATGCTCCTAAAC | 237 |
| Rnase2b | F:GATGGCTGCAAACCTGTCA R:CGAGTGGACTTCTTCCCTTT | 194 |
| Rnase2c | F:GATATGACCGTGCAGAAGTG R:TCTGTATTTACAGTTTGGGTGT | 240 |
| Rnase2d | F:CCTTCACTCACGGACAACC R:GCACTGACGACCATTTACAT | 140 |
| Rnase2e | F:AGAGGAGGAACTCGACTGAG R:AGCCTTTATCACAAGCCAAC | 157 |
| Rnase2f | F:AAGCAATTTGTGGCAGAG R:GGAGTTCCCTTAGTTAGTTTAG | 133 |
| Rnase3a | F:AGGCAAGCGGATGGAAAG R:CACCATAATGTACTGGGAGACC | 166 |
| Rnase3b | F:AATTAAAGCCGTTTGTAAGG R:ACCATAATGTACTGGGAGACC | 193 |
| Rnase3c | F:TTGTCCACTTACAGCCAGAG R:AACCGTTGTTATCTTCTTTGC | 256 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Geng, R.; Liu, H.; Wang, W. Differential Expression of Six Rnase2 and Three Rnase3 Paralogs Identified in Blunt Snout Bream in Response to Aeromonas hydrophila Infection. Genes 2018, 9, 95. https://doi.org/10.3390/genes9020095
Geng R, Liu H, Wang W. Differential Expression of Six Rnase2 and Three Rnase3 Paralogs Identified in Blunt Snout Bream in Response to Aeromonas hydrophila Infection. Genes. 2018; 9(2):95. https://doi.org/10.3390/genes9020095
Chicago/Turabian StyleGeng, Ruijing, Han Liu, and Weimin Wang. 2018. "Differential Expression of Six Rnase2 and Three Rnase3 Paralogs Identified in Blunt Snout Bream in Response to Aeromonas hydrophila Infection" Genes 9, no. 2: 95. https://doi.org/10.3390/genes9020095




