Genome-Wide Analysis of the Biosynthesis and Deactivation of Gibberellin-Dioxygenases Gene Family in Camellia sinensis (L.) O. Kuntze
Abstract
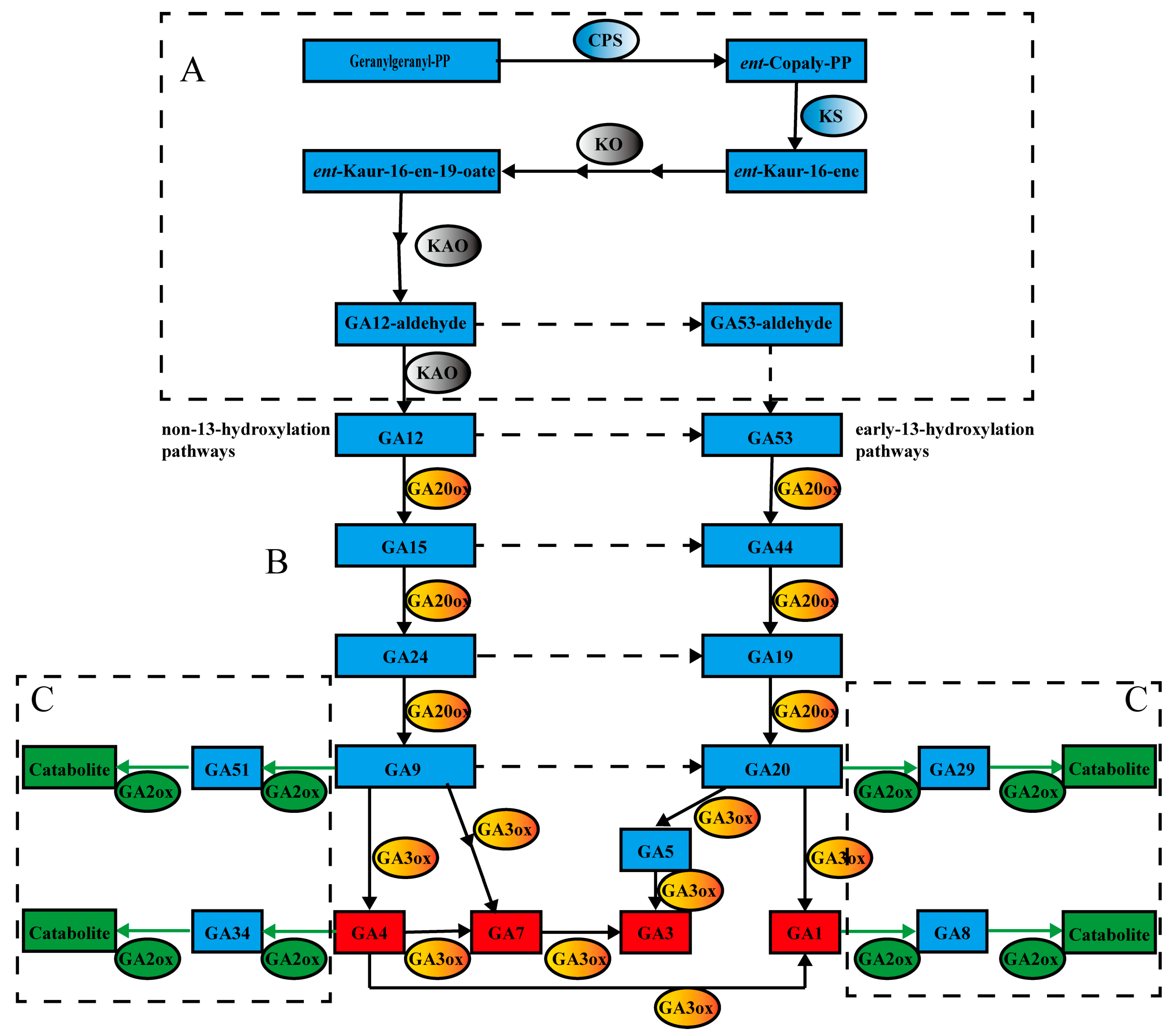
:1. Introduction
2. Materials and Methods
2.1. Plant Materials and Abiotic Treatments
2.2. RNA Isolation and cDNA Synthesis
2.3. Data Mining for GAox Protein Genes
2.4. Cloning the Full-Length of cDNA of GAox Protein Genes
2.5. Analysis of Sequences
2.6. Gene Expression Analysis by qRT-PCR
3. Results
3.1. Isolation, Identification and Annotation Information of the GAox Family Genes in Tea Plants
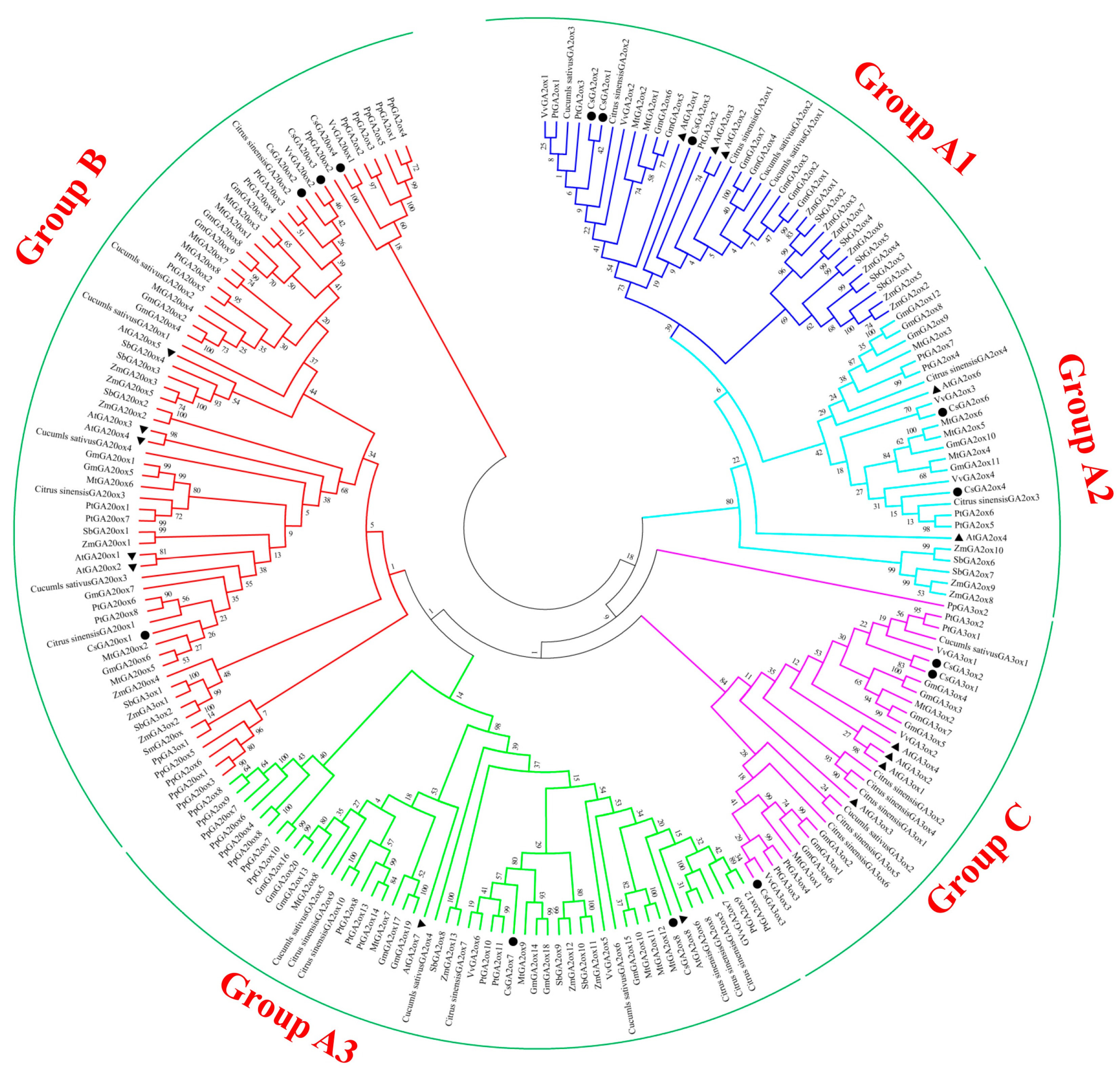
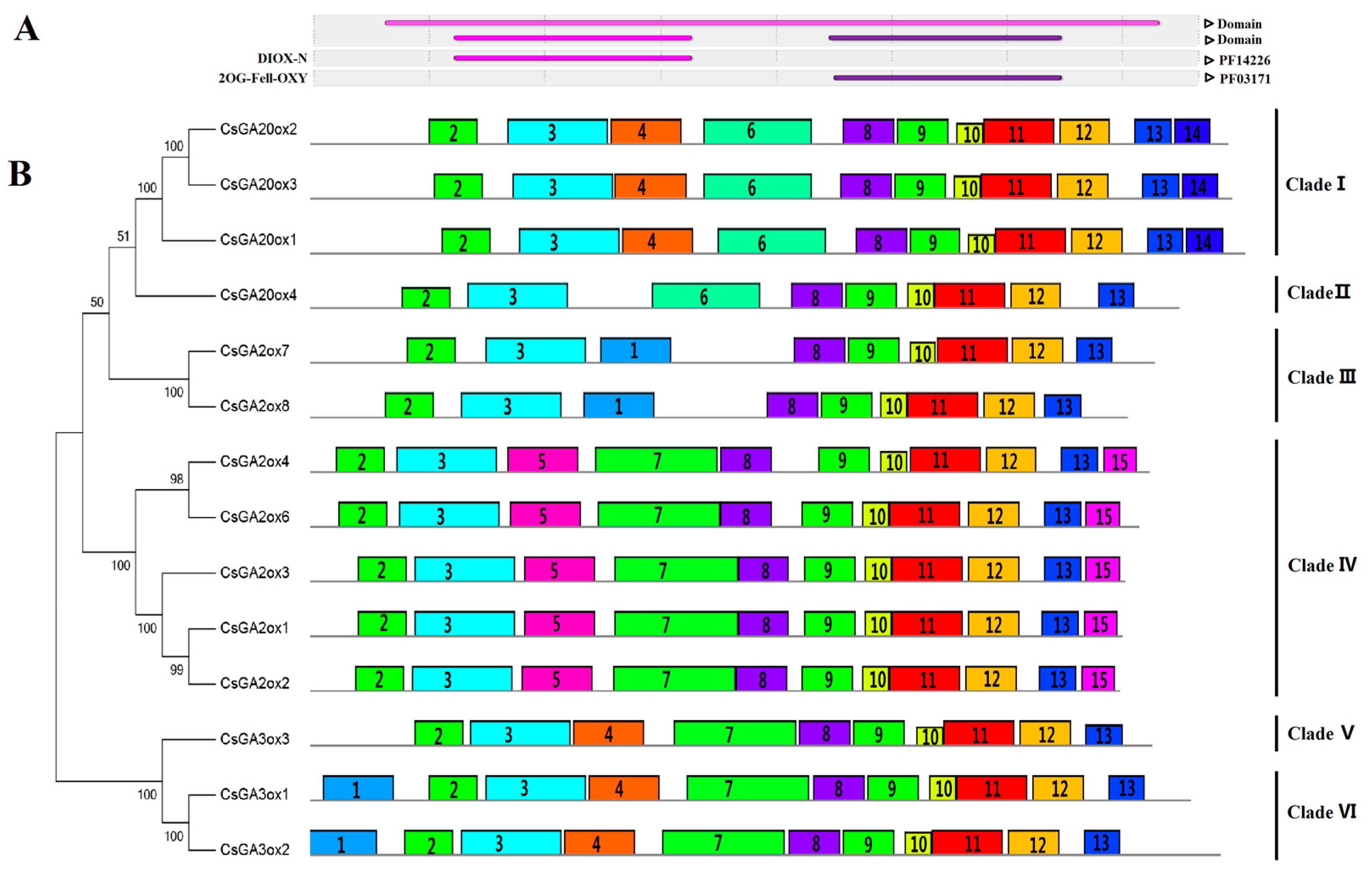
3.2. Similarity and Phylogenetic Gene Structure Analysis of the GAox Genes
3.3. Tissue-Specific Expression of CsGAoxes
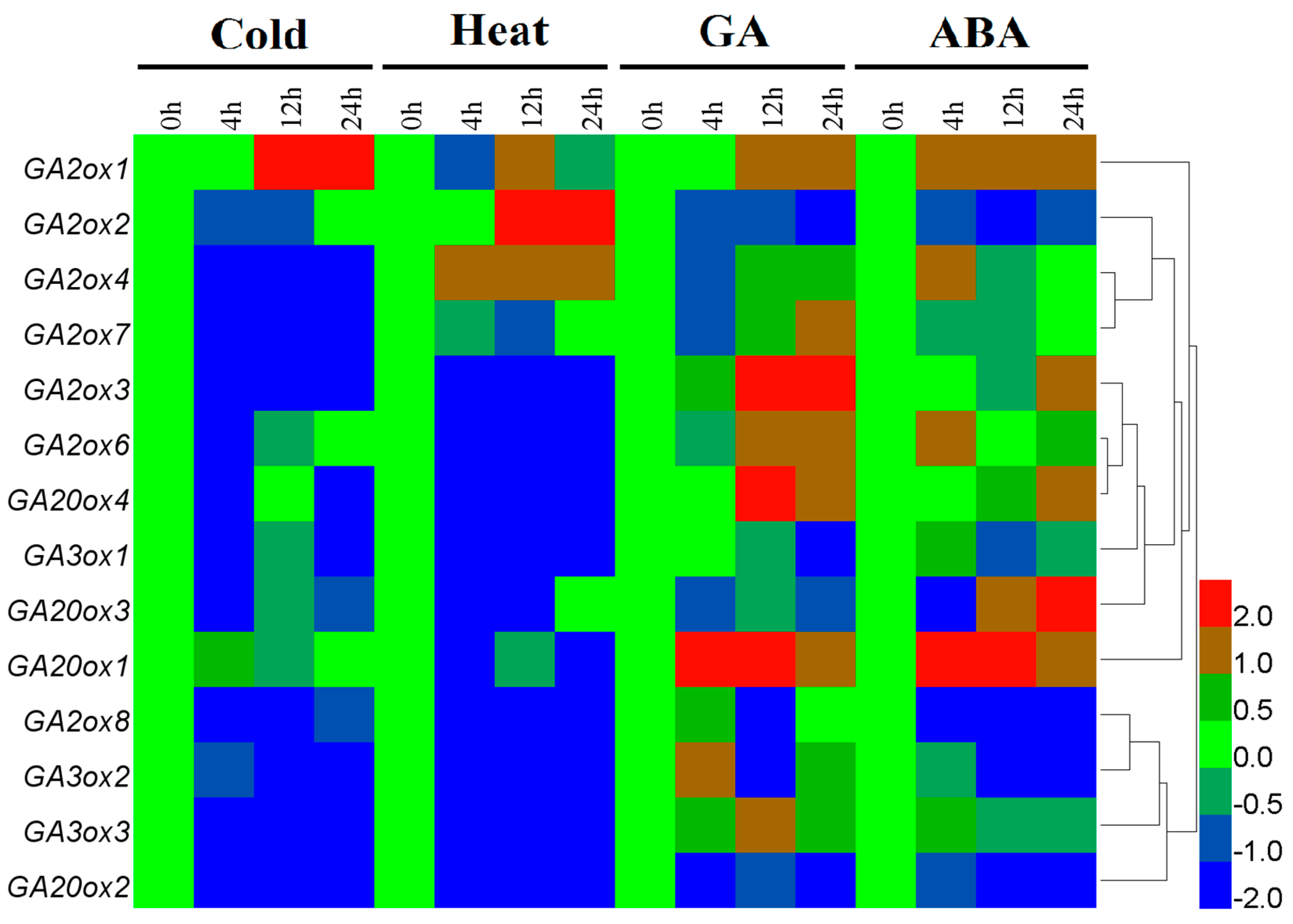
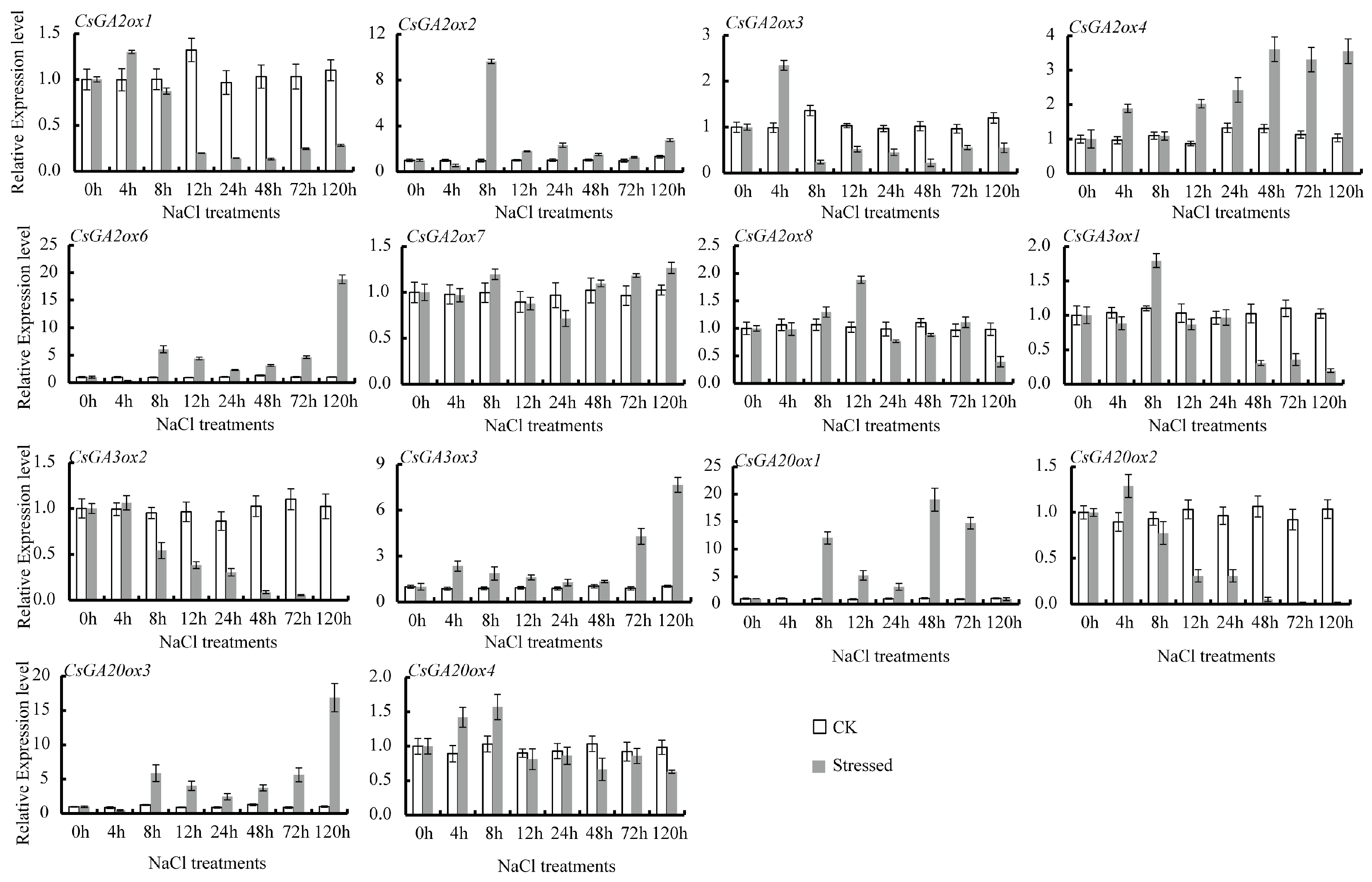
3.4. Differential Expression of CsGAox Genes under Abiotic Stresses
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gao, X.; Zhang, Y.; He, Z.; Fu, X. Hormone Metabolism & Signaling in Plants; Li, J.Y., Li, C.Y., Smith, S.M., Eds.; Academic Press: New York, NY, USA, 2017; pp. 107–160. [Google Scholar] [CrossRef]
- Hedden, P.; Kamiya, Y. Gibberellin biosynthesis: enzymes, genes and their regulation. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1997, 48, 431–460. [Google Scholar] [CrossRef] [PubMed]
- Olszewski, N.; Gubler, F. Gibberellin signaling: biosynthesis, catabolism, and response pathways. Plant Cell 2002, 14, S61–S80. [Google Scholar] [PubMed]
- Vera-Sirera, F.; Gomez, M.D.; Perez-Amador, M.A. DELLA proteins, a group of GRAS transcription regulators that mediate gibberellin signaling. Plant Transcr. Factors. 2016, 313–328. [Google Scholar]
- Hedden, P.; Thomas, S.G. Gibberellin biosynthesis and its regulation. Biochem. J. 2012, 444, 11–25. [Google Scholar] [CrossRef] [PubMed]
- Hedden, P.; Phillips, A.L. Gibberellin metabolism: new insights revealed by the genes. Trends Plant Sci. 2000, 5, 523–530. [Google Scholar] [CrossRef]
- Huerta, L.; Garcialor, A.; Garciamartinez, J.L. Characterization of gibberellin 20-oxidases in the citrus hybrid carrizo citrange. Tree Physiol. 2009, 29, 569–577. [Google Scholar] [CrossRef] [PubMed]
- Sponsel, V.M.; Hedden, P. Plant Hormones. Biosynthesis, Signal Transduction, Action; Davies, P.J., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2004; pp. 63–94. [Google Scholar]
- De, C.E.; De, L.V. 2-oxoglutarate-dependent dioxygenase and related enzymes: biochemical characterization. Phytochemistry 1994, 36, 1093–1107. [Google Scholar]
- Phillips, A.L.; Ward, D.A.; Uknes, S.; Appleford, N.E.; Lange, T.; Huttly, A.K.; Gaskin, P.; Graebe, J.E.; Hedden, P. Isolation and expression of three gibberellin 20-oxidase cDNA clones from Arabidopsis. Plant Physiol. 1995, 108, 1049–1057. [Google Scholar] [CrossRef] [PubMed]
- Schomburg, F.M.; Bizzell, C.M.; Lee, D.J.; Zeevaart, J.A.; Amasino, R.M. Overexpression of a novel class of gibberellin 2-oxidases decreases gibberellin levels and creates dwarf plants. Plant Cell 2003, 15, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Vidal, A.M.; Gisbert, C.; Talón, M.; Primomillo, E.; Lópezdíaz, I.; Garcíamartínez, J.L. The ectopic overexpression of a citrus gibberellin 20-oxidase enhances the non-13-hydroxylation pathway of gibberellin biosynthesis and induces an extremely elongated phenotype in tobacco. Physiol. Plant 2001, 112, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.L.; Li, L.; Wu, K.; Peeters, A.J.; Gage, D.A.; Zeevaart, J.A. The GA5 locus of Arabidopsis thaliana encodes a multifunctional gibberellin 20-oxidase: molecular cloning and functional expression. Proc. Natl. Acad. Sci. USA 1995, 92, 6640–6644. [Google Scholar] [CrossRef] [PubMed]
- Lange, T.; Hedden, P.; Graebe, J.E. Expression cloning of a gibberellin 20-oxidase, a multifunctional enzyme involved in gibberellin biosynthesis. Proc. Natl. Acad. Sci. USA 1994, 91, 8552–8556. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.; Phillips, A.; Hedden, P. Molecular cloning and functional expression of gibberellin 2-oxidases, multifunctional enzymes involved in gibberellin deactivation. Proc. Natl. Acad. Sci. USA 1999, 96, 4698–4703. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.C.; Ross, J.J.; Smith, J.J.; Lester, D.R.; Reid, J.B. Feed-forward regulation of gibberellin deactivation in pea. J. Plant Growth Regul. 2001, 20, 87–94. [Google Scholar] [CrossRef]
- Barua, D.N. Seasonal Dormancy in Tea (Camellia sinensis L.). Nature 1969, 224, 514. [Google Scholar] [CrossRef]
- Nagar, P.K.; Kumar, A. Changes in endogenous gibberellin activity during winter dormancy in tea (Camellia sinensis (L.) O. Kuntze). Acta Physiol. Plant 2000, 22, 439–443. [Google Scholar] [CrossRef]
- Yue, C.; Zeng, J.M.; Zhang, Z.F.; Wang, X.C.; Cao, H.L. Research Progress in the Phytohormone of Tea Plant (Camellia sinensis). J. Tea Sci. 2012, 32, 382–392. [Google Scholar]
- Xia, E.H.; Zhang, H.B.; Sheng, J.; Li, K.; Zhang, Q.J.; Kim, C.; Zhang, Y.; Liu, Y.; Zhu, T.; Li, W.; et al. The tea tree genome provides insights into tea flavor and independent evolution of caffeine biosynthesis. Mol. Plant 2017, 10, 866–877. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.N.; Tang, L.; Hou, Y.; Wang, P.; Yang, H.; Wei, C.L. Differential transcriptome analysis of leaves of tea plant (Camellia sinensis) provides comprehensive insights into the defense responses to Ectropis oblique attack using RNA-Seq. Funct. Integr. Genom. 2016, 16, 383–398. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.P. Textbook of Tea Cultivation; China Agriculture Press: Beijing, China, 2008; pp. 176–177. [Google Scholar]
- Li, H.; Huang, W.; Wang, G.L.; Wu, Z.J.; Zhuang, J. Expression profile analysis of ascorbic acid-related genes in response to temperature stress in the tea plant, Camellia sinensis (L.) O. Kuntze. Genet. Mol. Res. 2016, 15. [Google Scholar] [CrossRef] [PubMed]
- Li, X.W.; Feng, Z.G.; Yang, H.M.; Zhu, X.P.; Liu, J.; Yuan, H.Y. A novel cold-regulated gene from Camellia sinensis, CsCOR1, enhances salt- and dehydration-tolerance in tobacco. Biochem. Biophys. Res. Commun. 2010, 394, 354–359. [Google Scholar] [CrossRef] [PubMed]
- Yue, C.; Cao, H.; Wang, L.; Zhou, Y.; Hao, X.; Zeng, J.; Wang, X.; Yang, Y. Molecular cloning and expression analysis of tea plant aquaporin (AQP) gene family. Plant Physiol. Biochem. 2014, 83, 65–76. [Google Scholar] [CrossRef] [PubMed]
- The University of California. Phytozome. Available online: https://phytozome.jgi.doe.gov/pz/portal.html (accessed on 9 May 2016).
- The Arabidopsis Biological Resource Center. TAIR. Available online: http://www.arabidopsis.org/ (accessed on 9 July 2017).
- Wang, D.; Pei, K.; Fu, Y.; Sun, Z.; Li, S.; Liu, H.; Tang, K.; Han, B.; Tao, Y. Genome-wide analysis of the auxin response factors (ARF) gene family in rice (Oryza sativa). Gene 2007, 394, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Wang, M.; Li, X.; Jiu, S.; Wang, C.; Fang, J. Genome-wide analysis of the sucrose synthase gene family in grape (Vitis vinifera): structure, evolution, and expression profiles. Genes 2017, 8, 111. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schafferi, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Altschul, S.F.; Wootton, J.C.; Gertz, E.M.; Agarwala, R.; Morgulis, A.; Schäffer, A.A.; Yu, Y.K. Protein database searches using compositionally adjusted substitution matrices. FEBS J. 2005, 272, 5101–5109. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 2012, 40, 302–305. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhard, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 2016, 44, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.; Zaretskaya, I.; Raytselis, Y.; Merezhuk, Y.; McGinnis, S.; Madden, T.L. NCBI BLAST: a better web interface. Nucleic Acids Res. 2008, 36, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; William, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–532. [Google Scholar] [PubMed]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, k. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 2007, 35, 585–587. [Google Scholar] [CrossRef] [PubMed]
- Emanuelsson, O.; Brunak, S.; von Heijne, G.; Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2007, 2, 953–971. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Hunter, S.; Apweiler, R.; Attwood, T.K.; Bairoch, A.; Bataman, A.; Binns, D.; Bork, P.; Das, U.; Daugherty, L.; Duquenne, L.; et al. InterPro: the integrative protein signature database. Nucleic Acids Res. 2009, 37, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Erik, V.B.; Asad, M.; Marc, D.L. Validation of reference genes for gene expression analysis in chicory (Cichorium intybus) using quantitative real-time PCR. BMC Mol. Biol. 2010, 11, 15. [Google Scholar] [CrossRef] [Green Version]
- Gohain, B.; Bandyopadhyay, T.; Borchetia, S.; Bharalee, R.; Gupta, S.; Bhorali, P.; Agarwala, N.; Das, S. Identification and validation of stable reference genes in Camellia Species. J. Bioteachnol. Pharm. Res. 2011, 2, 9–18. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Tang, D.; Shen, Y.; Qin, B.; Hong, L.; You, A.; Li, M.; Wang, X.; Yu, H.; Gu, M.; et al. Activation of gibberellin 2-oxidase 6 decreases active gibberellin levels and creates a dominant semi-dwarf phenotype in rice (Oryza sativa L.). J. Genet. Genom. 2010, 37, 23–36. [Google Scholar] [CrossRef]
- Lo, S.F.; Yang, S.Y.; Chen, K.T.; Hsing, Y.I.; Zeevaart, J.A.; Chen, L.J.; Yu, S.M. A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 2008, 20, 2603–2618. [Google Scholar] [CrossRef] [PubMed]
- Shan, C.; Mei, Z.; Duan, J.; Chen, H.; Feng, H.; Cai, W. OsGA2ox5, a gibberellin metabolism enzyme, is involved in plant growth, the root gravity response and salt stress. PLoS One 2014, 9, e87110. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S.; Kamiya, Y. Gibberellin biosynthesis: its regulation by endogenous and environmental signals. Plant Cell Physiol. 2000, 41, 251–257. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Lange, T.; Altpeter, F. Cloning and characterization of a cDNA encoding a multifunctional gibberellin 20-oxidase from perennial ryegrass (Lolium perenne L.). Plant Sci. 2002, 163, 147–155. [Google Scholar] [CrossRef]
- Lee, D.J.; Zeevaart, J.A. Molecular cloning of GA 2-oxidase3 from spinach and its ectopic expression in Nicotiana sylvestris. Plant Physiol. 2005, 138, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Deng, W.; Wang, Y.; Liu, Z.; Cheng, H.; Xue, Y. HemI: a toolkit for illustrating heatmaps. PLoS ONE 2014, 9, e111988. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.L.; Li, Q.; Deng, Y.W.; Lou, Y.G.; Wang, M.Y.; Zhou, G.X.; Zhang, Y.Y.; He, Z.H. Altered disease development in the eui mutants and Eui overexpressors indicates that gibberellins negatively regulate rice basal disease resistance. Mol. Plant 2008, 1, 528–537. [Google Scholar] [CrossRef] [PubMed]
- Razem, F.A.; Baron, K.; Hill, R.D. Turning on gibberellin and abscisic acid signaling. Curr. Opin. Plant Biol. 2006, 9, 454–459. [Google Scholar] [CrossRef] [PubMed]
- Weiss, D.; Ori, N. Mechanisms of cross talk between gibberellin and other hormones. Plant Physiol. 2007, 144, 1240–1246. [Google Scholar] [CrossRef] [PubMed]
- Bethke, P.C.; Jones, R.L. Gibberellin signaling. Curr. Opin. Plant Biol. 1998, 1, 440–446. [Google Scholar] [CrossRef]
- Phillips, A. Gibberellins in Arabidopsis. Plant Physiol. Biochem. 1998, 36, 115–124. [Google Scholar] [CrossRef]
- Venkatesh, J.; Park, S.W. Genome-wide analysis and expression profiling of DNA-binding with one zinc finger (Dof) transcription factor family in potato. Plant Physiol. Biochem. 2015, 94, 73–85. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Liu, J.; Dong, C.; Cheng, Z.M. The CBL and CIPK gene family in grapevine (Vitis vinifera): genome-wide analysis and expression profiles in response to various abiotic stresses. Front. Plant Sci. 2017, 8, 978. [Google Scholar] [CrossRef] [PubMed]
- Chiang, H.; Hwang, l.; Goodma, H. Isolation of the Arabidopsis GA4 locus. Plant Cell 1995, 7, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Plackett, A.; Powers, S.; Fernandez-Garcia, N.; Urbanova, T.; Takebayashi, Y.; Seo, M.; Jikumaru, Y.; Benlloch, R.; Nilsson, O.; Ruiz-Rivero, O.; et al. Analysis of the developmental roles of the Arabidopsis gibberellin 20-oxidases demonstrates that GA20ox1, -2, and -3 are the dominant paralogs. Plant Cell 2012, 24, 941–960. [Google Scholar] [CrossRef] [PubMed]
- Reinecke, D.M.; Wickramarathna, A.D.; Ozga, J.A.; Kurepin, L.V.; Jin, A.L.; Good, A.G.; Pharis, R.P. Gibberellin 3-oxidase gene expression patterns influence gibberellin biosynthesis, growth, and development in pea. Plant Physiol. 2013, 163, 929–945. [Google Scholar] [CrossRef] [PubMed]
- Sponsel, V.; Schmidt, F.; Porter, S.; Nakayama, M.; Kohlstruk, S.; Estelle, M. Characterization of new gibberellin-responsive semidwarf mutants of Arabidopsis. Plant Physiol. 1997, 115, 1009–1020. [Google Scholar] [CrossRef] [PubMed]
- Coles, J.P.; Phillips, A.L.; Croker, S.J.; Garcia-Lepe, R.; Lewis, M.J.; Hedden, P. Modification of gibberellin production and plant development in Arabidopsis by sense and antisense expression of gibberellin 20-oxidase genes. Plant J. 1999, 17, 547–556. [Google Scholar] [PubMed]
- Yamaguchi, S. Gibberellin metabolism and its regulation. Annu. Rev. Plant Biol. 2008, 59, 225–251. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Ueguchi-Tanaka, M.; Sentoku, N.; Kitano, H.; Matsuoka, M.; Kobayashi, M. Cloning and functional analysis of two gibberellin 3β-hydroxylase genes that are differently expressed during the growth of rice. Proc. Natl. Acad. Sci. USA 2001, 98, 8909–8914. [Google Scholar] [CrossRef] [PubMed]
- Plackett, A.; Thomas, S.; Wilson, Z.; Hedden, P. Gibberellin control of stamen development: a fertile field. Trends Plant Sci. 2011, 16, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S.; Smith, M.W.; Brown, R.G.S.; Kamiya, Y.; Sun, T. Phytochrome regulation and differential expression of gibberellin 3β-hydroxylase genes in germinating Arabidopsis seeds. Plant Cell 1998, 10, 2115–2126. [Google Scholar] [PubMed]
- Huang, S.; Raman, A.S.; Ream, J.E.; Fujiwara, H.; Cerny, R.E.; Brown, S.M. Overexpression of 20-oxidase confers a gibberellin overproduction phenotype in Arabidopsis. Plant Physiol. 1998, 118, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Israelsson, M.; Mellerowicz, E.; Chono, M.; Gullberg, J.; Mortz, T. Cloning and overproduction of gibberellin 3-oxidase in hybrid aspen trees. Effects on gibberellin homeostasis and development. Plant Physiol. 2004, 135, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, T.; Kobayashi, M.; Itoh, H.; Tagiri, A.; Kayaka, H.; Lwahori, S.; Matsuoka, M. Expression of a gibberellin 2-oxidase gene around the shoot apex is related to phase transition in rice. Plant Physiol. 2001, 125, 1508–1516. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.J.; Zeevaart, J.A. Differential regulation of RNA levels of gibberellin dioxygenases by photoperiod in spinach. Plant Physiol. 2002, 130, 2085–2094. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Gong, F.; Cheminant, S.; Alioua, M.; Hedden, P.; Genschik, P. The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism. Plant Cell 2008, 20, 2117–2129. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, Y.; Ogawa, M.; Kuwahara, A.; Hanada, A.; Kamiya, Y.; Yamaguchi, S. Activation of gibberellin biosynthesis and response pathways by low temperature during imbibition of Arabidopsis thaliana seeds. Plant Cell 2004, 16, 367–378. [Google Scholar] [CrossRef] [PubMed]
- Toh, S.; Imamura, A.; Watanabe, A.; Nakabayashi, K.; Okamoto, M.; Jikumaru, Y.; Hanada, A.; Aso, Y.; Ishiyama, K.; Tamura, N.; et al. High temperature-induced abscisic acid biosynthesis and its role in the inhibition of gibberellin action in Arabidopsis seeds. Plant Physiol. 2008, 146, 1368–1385. [Google Scholar] [CrossRef] [PubMed]
- Oh, E.; Yamaguchi, S.; Hu, J.; Yusuke, J.; Jung, B.; Paik, L.; Lee, H.S.; Sun, T.P.; Kamiya, Y.; Choi, G. PIL5, a phytochrome-interacting bHLH protein, regulates gibberellin responsiveness by binding directly to the GAI and RGA promoters in Arabidopsis seeds. Plant Cell 2007, 19, 1192–1208. [Google Scholar] [CrossRef] [PubMed]
- Seo, M.; Hanada, A.; Kuwahara, A.; Endo, A.; Okamoto, M.; Yamauchi, Y.; North, H.; Marion-Poll, A.; Sun, T.P.; Koshiba, T.; et al. Regulation of hormone metabolism in Arabidopsis seeds: phytochrome regulation of abscisic acid metabolism and abscisic acid regulation of gibberellin metabolism. Plant J. 2006, 48, 354–366. [Google Scholar] [CrossRef] [PubMed]
- Zentella, R.; Zhang, Z.L.; Park, M.; Thomas, S.G.; Endo, A.; Murase, K.; Fleet, C.M.; Jikumary, Y.; Nambara, E.; Kamiya, Y.; Sun, T.P. Global analysis of DELLA direct targets in early gibberellin signaling in Arabidopsis. Plant Cell 2007, 19, 3037–3057. [Google Scholar] [CrossRef] [PubMed]
- Yaish, M.W.; El-Kereamy, A.; Zhu, T.; Beatty, P.H.; Good, A.G.; Bi, Y.M.; Rothstein, S.J. The APETALA-2-like transcription factor OsAP2-39 controls key interactions between abscisic acid and gibberellin in rice. PLoS Genet. 2010, 6, e1001098. [Google Scholar] [CrossRef] [PubMed]
- Colebrook, E.H.; Thomas, S.G.; Phillips, A.L.; Hedden, P. The role of gibberellin signalling in plant responses to abiotic stress. J. Exp. Biol. 2014, 217, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Magome, H.; Yamaguchi, S.; Hanada, A.; Kamiya, Y.; Oda, K. The DDF1 transcriptional activator upregulates expression of a gibberellin-deactivating gene, GA2ox7, under high-salinity stress in Arabidopsis. Plant J. 2008, 56, 613–626. [Google Scholar] [CrossRef] [PubMed]



| Gene names | Accession Numbers | Gene ID a | Number of Deduced Amino Acid | Molecular Weight (kDa) | Subcellular Location (WoLF PSORT/TargetP) | Groups | Theoretical pI |
|---|---|---|---|---|---|---|---|
| CsGA20ox1 | KC193604 | CSA007392 | 383 | 42.94 | nucl | B | 6.06 |
| CsGA20ox2 | KY296366 | NF | 376 | 43.02 | nucl/cyto | B | 6.98 |
| CsGA20ox3 | MF370231 | CSA000105 | 378 | 43.03 | nucl/cyto | B | 6.94 |
| CsGA20ox4 | MF370232 | CSA002490 | 356 | 40.31 | cyto | B | 5.30 |
| CsGA2ox1 | KY296367 | CSA032124 | 333 | 37.18 | nucl/cyto | A1 | 5.93 |
| CsGA2ox2 | KY296368 | CSA026961 | 332 | 37.31 | cyto/nucl | A1 | 7.61 |
| CsGA2ox3 | KY296369 | CSA004444 | 334 | 37.56 | nucl/cyto/chlo | A1 | 5.47 |
| CsGA2ox4 | MF370234 | NF | 344 | 38.67 | nucl | A2 | 5.73 |
| CsGA2ox6 | MF370235 | CSA014596 | 340 | 38.02 | nucl/chol/mito | A2 | 7.94 |
| CsGA2ox7 | MF370236 | CSA013052 | 346 | 39.24 | cyto/nucl | A3 | 6.79 |
| CsGA2ox8 | MF370237 | CSA034015 | 335 | 38.53 | cyto/nucl | A3 | 5.33 |
| CsGA3ox1 | KF703743 | CSA002905 | 361 | 40.29 | chlo/mito/nucl | C | 6.67 |
| CsGA3ox2 | KF703744 | CSA020111 | 373 | 41.08 | nucl/cyto | C | 8.22 |
| CsGA3ox3 | MF370233 | CSA034282 | 345 | 38.72 | cyto/nucl | C | 5.66 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pan, C.; Tian, K.; Ban, Q.; Wang, L.; Sun, Q.; He, Y.; Yang, Y.; Pan, Y.; Li, Y.; Jiang, J.; et al. Genome-Wide Analysis of the Biosynthesis and Deactivation of Gibberellin-Dioxygenases Gene Family in Camellia sinensis (L.) O. Kuntze. Genes 2017, 8, 235. https://doi.org/10.3390/genes8090235
Pan C, Tian K, Ban Q, Wang L, Sun Q, He Y, Yang Y, Pan Y, Li Y, Jiang J, et al. Genome-Wide Analysis of the Biosynthesis and Deactivation of Gibberellin-Dioxygenases Gene Family in Camellia sinensis (L.) O. Kuntze. Genes. 2017; 8(9):235. https://doi.org/10.3390/genes8090235
Chicago/Turabian StylePan, Cheng, Kunhong Tian, Qiuyan Ban, Leigang Wang, Qilu Sun, Yan He, Yuanfei Yang, Yuting Pan, Yeyun Li, Jiayue Jiang, and et al. 2017. "Genome-Wide Analysis of the Biosynthesis and Deactivation of Gibberellin-Dioxygenases Gene Family in Camellia sinensis (L.) O. Kuntze" Genes 8, no. 9: 235. https://doi.org/10.3390/genes8090235





