Molecular Expression of the Scribble Complex Genes, Dlg, Scrib and Lgl, in Silkworm, Bombyx mori
Abstract
:1. Introduction
2. Materials and Methods
2.1. Insects
2.2. Gene Identification

2.3. RNA Isolation, cDNA Synthesis
2.4. qRT-RCR
| Gene | Primers | Sequences | GenBank |
|---|---|---|---|
| Scrib | Scrib F | 5' TAGTTCAGCAACTGGAACGC 3' | BGIBMGA005373 |
| Scrib R | 5' TTCTAGCCATGCGAATTGAG 3' | ||
| Dlg | Dlg F | 5' AACGCTGACGGAGAAATCTT 3' | BGIBMGA010382 |
| Dlg R | 5' GAGTGTACGCGATCGTCAAT 3' | ||
| Lgl | Lgl F | 5' TCGTCTTCCGAATTACAACG 3' | BGIBMGA005570 |
| Lgl R | 5' GGCACCTCTTCCTTATGCTC 3' | ||
| Rp49 | Rp49 F | 5' CAGGCGGTTCAAGGGTCAATAC 3' | AY769302 |
| Rp49 R | 5' TGCTGGGCTCTTTCCACGA 3' |
2.5. Tissue-Specific Distribution and Developmental Expression of Scrib, Dlg and Lgl
3. Results
3.1. Tissue-Specific Distribution and Developmental Expression of the Scrib Gene

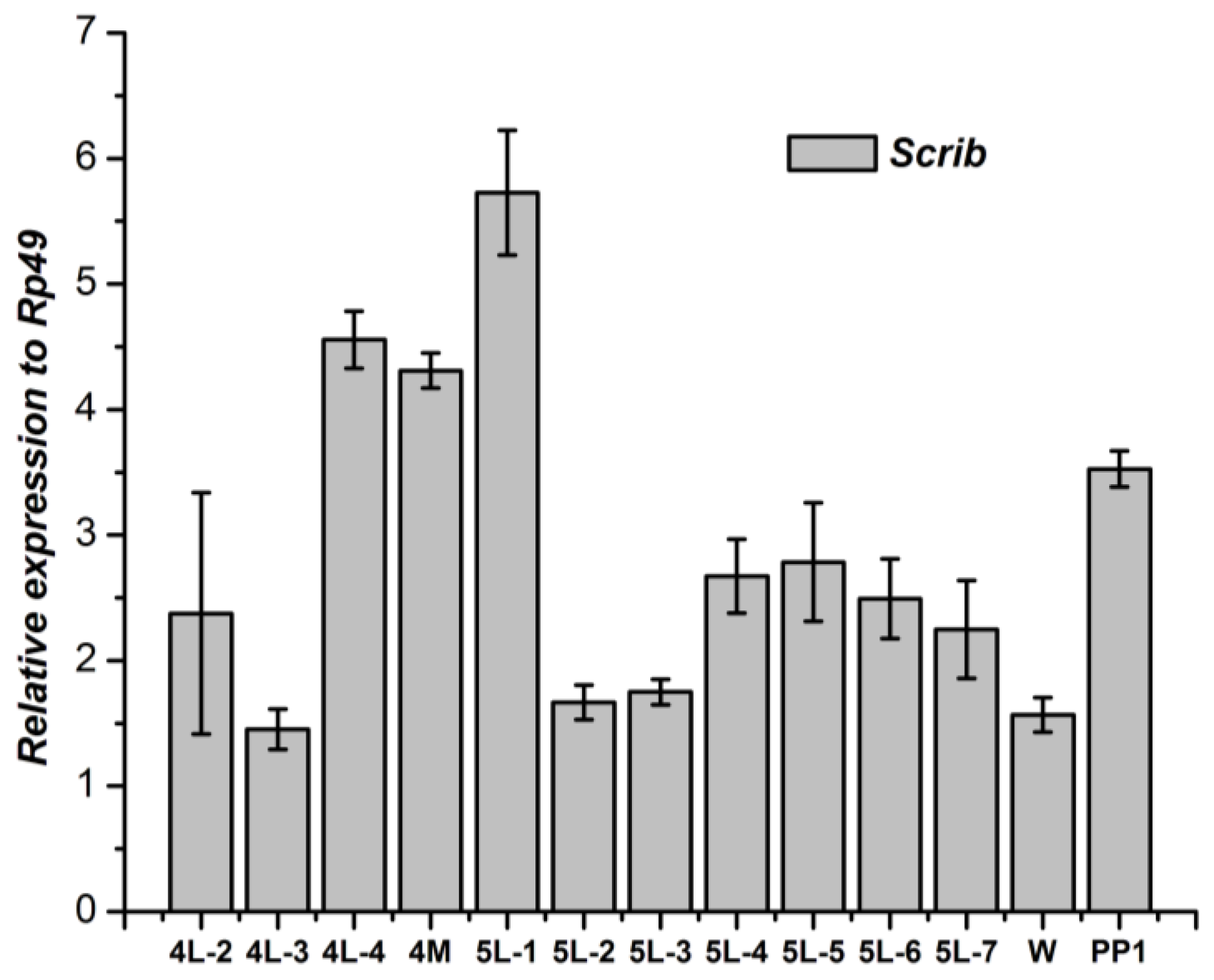
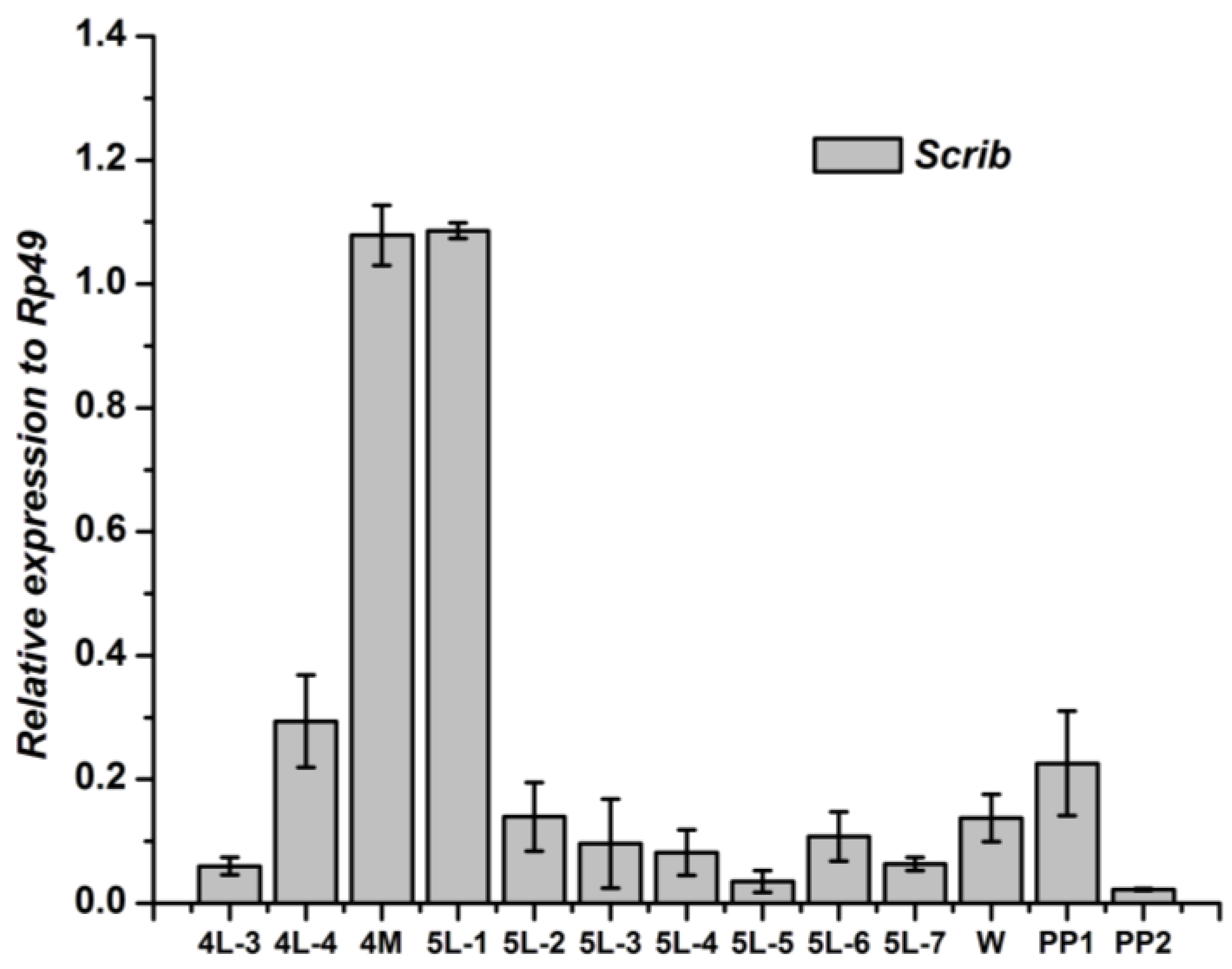
3.2. Tissue-Specific Distribution and Developmental Expression of Dlg Gene
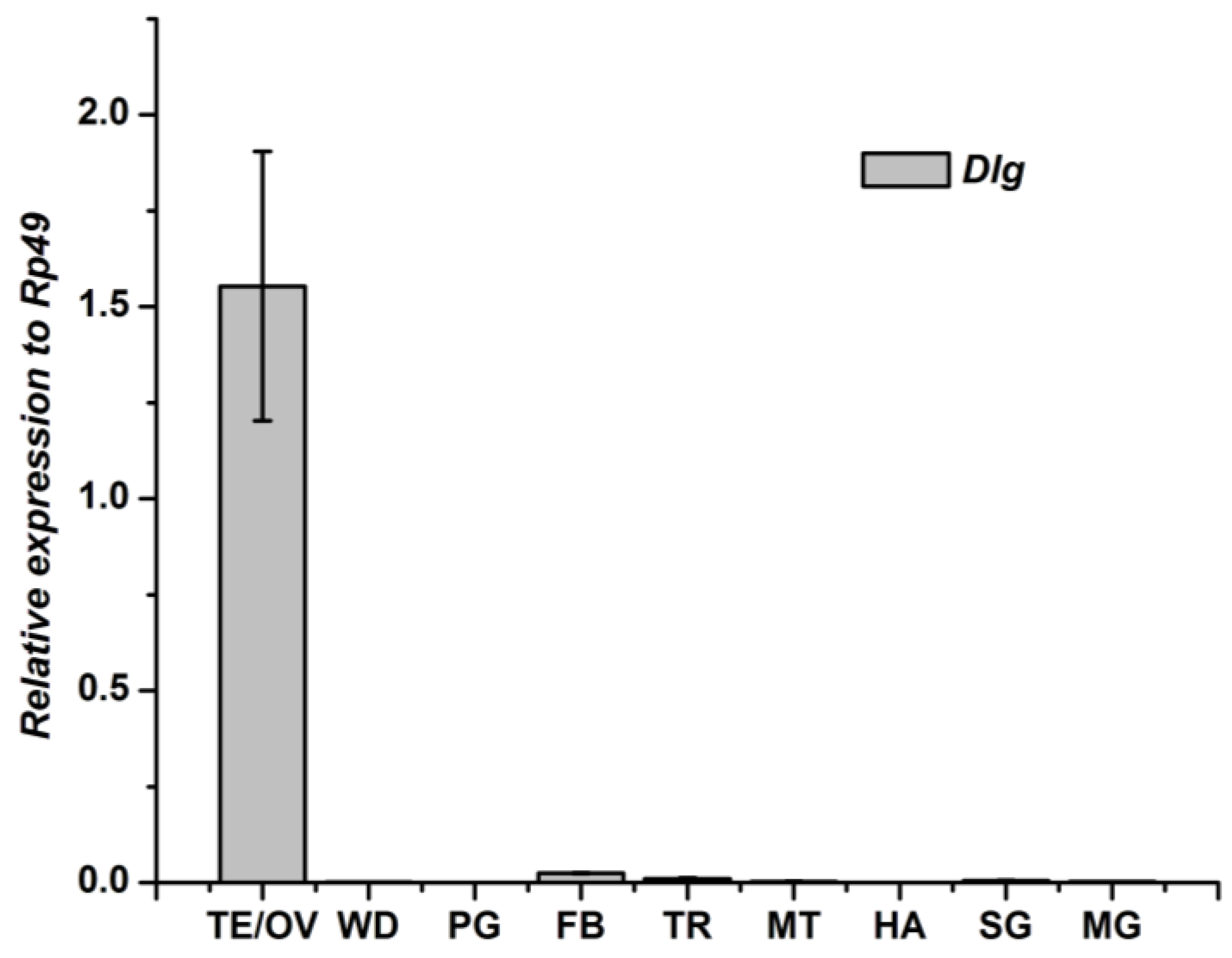
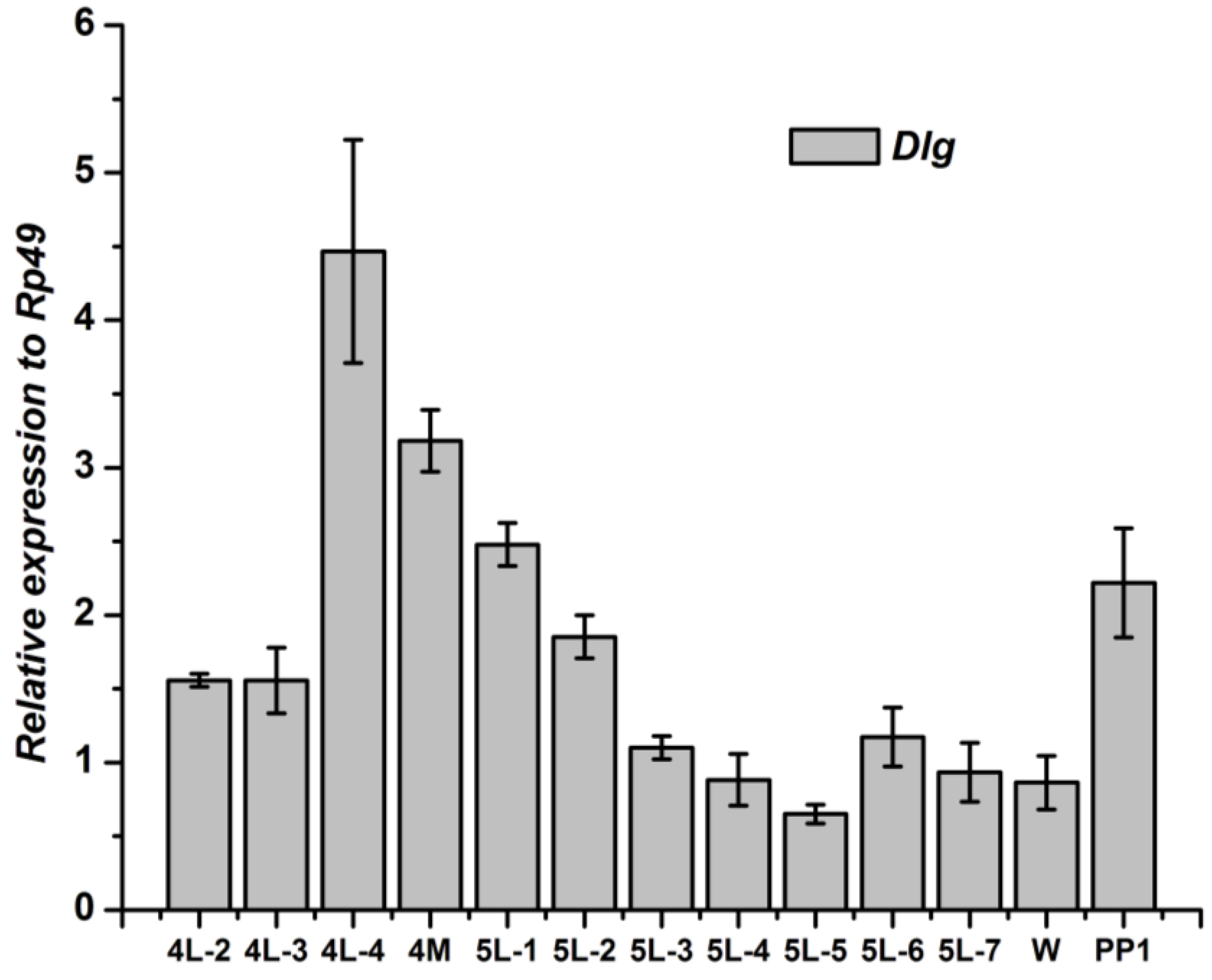

3.3. Tissue-Specific Distribution and Developmental Expression of Lgl Gene
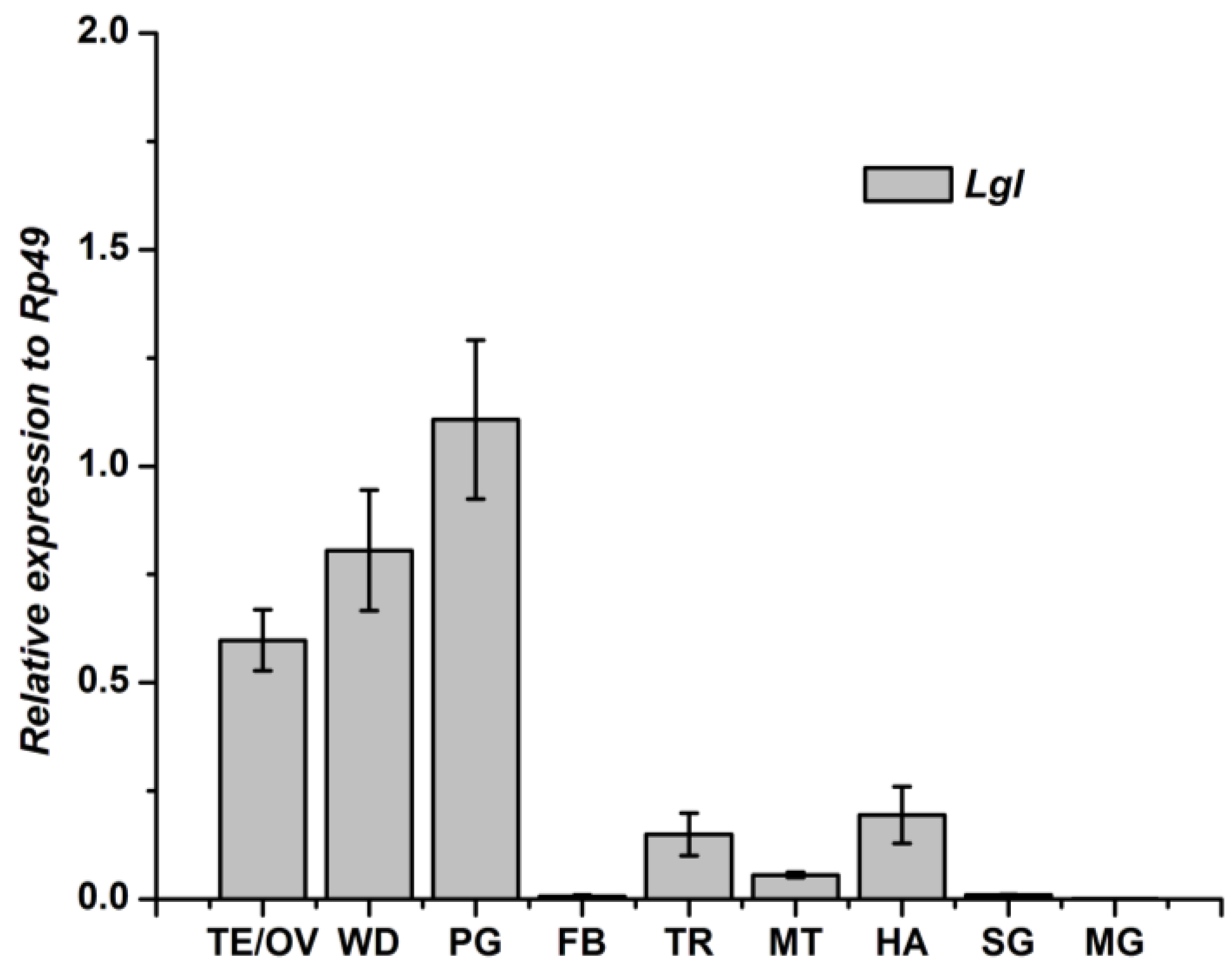
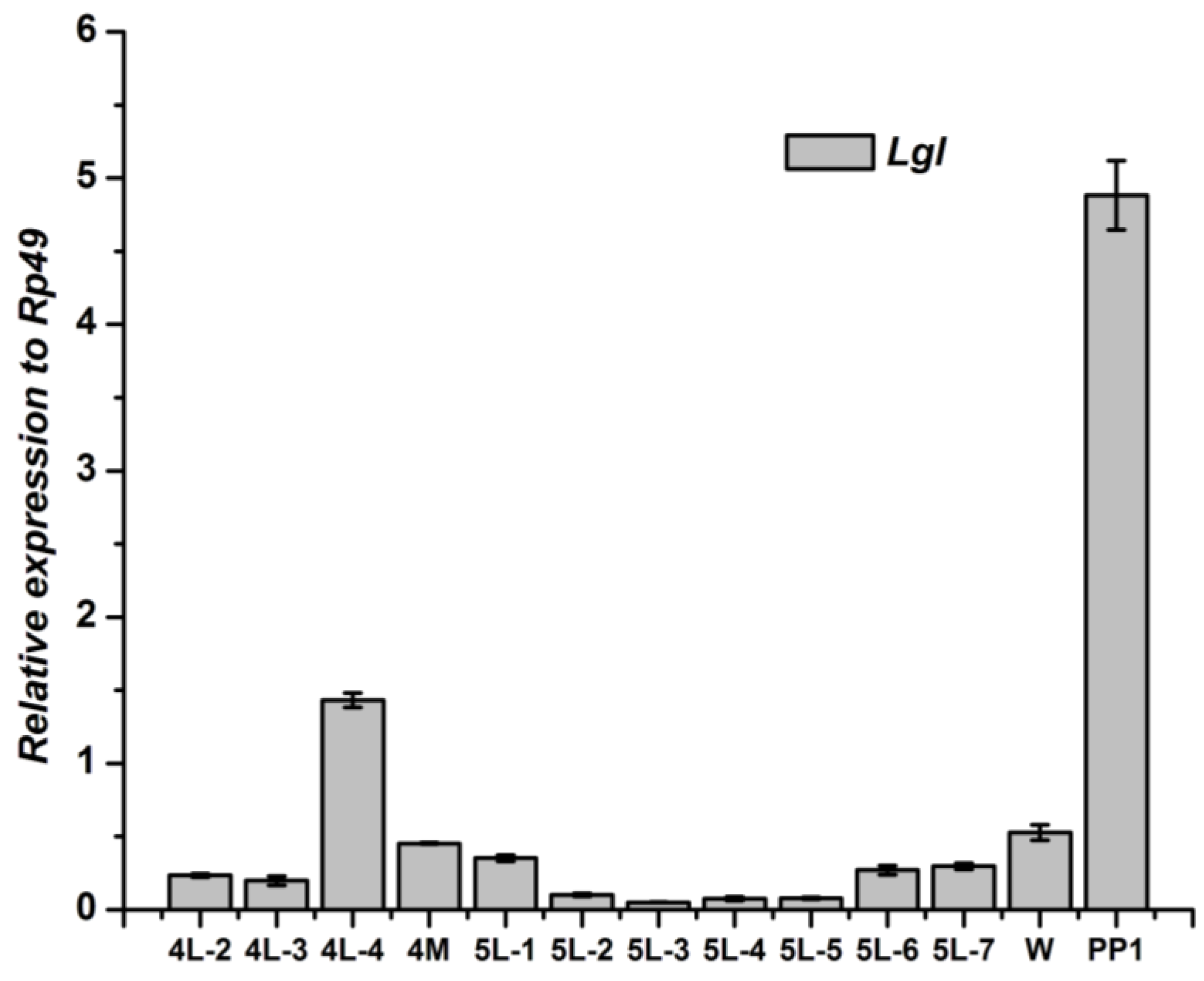

4. Discussion
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Elsum, I.; Yates, L.; Humbert, P.O.; Richardson, H.E. The Scribble-Dlg-Lgl polarity module in development and cancer: From flies to man. Essays Biochem. 2012, 53, 141–168. [Google Scholar] [CrossRef]
- Assémat, E.; Bazellières, E.; Pallesi-Pocachard, E.; Le Bivic, A.; Massey-Harroche, D. Polarity complex proteins. BBA Biomembranes 2008, 1778, 614–630. [Google Scholar] [CrossRef]
- Bissell, M.J.; Radisky, D. Putting tumours in context. Nat. Rev. Cancer 2001, 1, 46–54. [Google Scholar] [CrossRef]
- Humbert, P.; Russell, S.; Richardson, H. Dlg, Scribble and Lgl in cell polarity, cell proliferation and cancer. Bioessays 2003, 25, 542–553. [Google Scholar] [CrossRef]
- Bilder, D. Epithelial polarity and proliferation control: Links from the Drosophila neoplastic tumor suppressors. Gene Dev. 2004, 18, 1909–1925. [Google Scholar] [CrossRef]
- Brumby, A.M.; Richardson, H.E. Scribble mutants cooperate with oncogenic Ras or Notch to cause neoplastic overgrowth in Drosophila. EMBO J. 2003, 22, 5769–5779. [Google Scholar] [CrossRef]
- Bilder, D.; Perrimon, N. Localization of apical epithelial determinants by the basolateral PDZ protein Scribble. Nature 2000, 403, 676–680. [Google Scholar] [CrossRef]
- Woods, D.F.; Bryant, P.J. Molecular cloning of the lethal (1) discs large-1 oncogene of Drosophila. Dev. Biol. 1989, 134, 222–235. [Google Scholar] [CrossRef]
- Enomoto, M.; Igaki, T. Deciphering tumor-suppressor signaling in flies: Genetic link between Scribble/Dlg/Lgl and the Hippo pathway. J. Genet. Genomics 2011, 38, 461–470. [Google Scholar] [CrossRef]
- Jacob, L.; Opper, M.; Metzroth, B.; Phannavong, B.; Mechler, B.M. Structure of the l (2) gl gene of Drosophila and delimitation of its tumor suppressor domain. Cell 1987, 50, 215–225. [Google Scholar] [CrossRef]
- Thomas, U.; Phannavong, B.; Müller, B.; Garner, C.C.; Gundelfinger, E.D. Functional expression of rat synapse-associated proteins SAP97 and SAP102 in Drosophila dlg-1 mutants: Effects on tumor suppression and synaptic bouton structure. Mech. Dev. 1997, 62, 161–174. [Google Scholar] [CrossRef]
- International Silkworm Genome Consortium. The genome of a lepidopteran model insect, the silkworm Bombyx mori. Insect Biochem. Mol. Biol. 2008, 38, 1036–1045.
- Wei, Z.J.; Yu, M.; Tang, S.M.; Yi, Y.Z.; Hong, G.Y.; Jiang, S.T. Transcriptional regulation of the gene for prothoracicotropic hormone in the silkworm, Bombyx mori. Mol. Biol. Rep. 2011, 38, 1121–1127. [Google Scholar] [CrossRef]
- Xia, H.; Zhang, C.; Feng, F.; Yuan, Y.; Zhou, Y.; Liu, X.; Chen, K. Molecular cloning, expression and characterization of a novel vacuolar protein sorting 4 gene in silkworm, Bombyx mori. Mol. Biol. Rep. 2012, 39, 10339–10346. [Google Scholar] [CrossRef]
- Tang, S.; Wu, J.; Zhao, X.; Wang, H.; Qiu, Z.; Shen, X.; Guo, X. Molecular cloning and characterization of hatching enzyme-like geneII (BmHEL II) in the silkworm Bombyx mori. Biochem. Biophys. Res. Commun. 2012, 419, 194–199. [Google Scholar] [CrossRef]
- Tian, L.; Guo, E.; Diao, Y.; Zhou, S.; Peng, Q.; Cao, Y.; Li, S. Genome-wide regulation of innate immunity by juvenile hormone and 20-hydroxyecdysone in the Bombyx fat body. BMC Genomics 2010, 11, 549. [Google Scholar] [CrossRef]
- Duan, J.; Li, R.; Cheng, D.; Fan, W.; Zha, X.; Cheng, T.; Xia, Q. SilkDB v2. 0: A platform for silkworm (Bombyx mori) genome biology. Nucleic Acids Res. 2010, 38, D453–D456. [Google Scholar] [CrossRef]
- Klämbt, C.; Schmidt, O. Developmental expression and tissue distribution of the lethal (2) giant larvae protein of Drosophila melanogaster. EMBO J. 1986, 5, 2955–2961. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Qi, H.-S.; Liu, S.-M.; Li, S.; Wei, Z.-J. Molecular Expression of the Scribble Complex Genes, Dlg, Scrib and Lgl, in Silkworm, Bombyx mori. Genes 2013, 4, 264-274. https://doi.org/10.3390/genes4020264
Qi H-S, Liu S-M, Li S, Wei Z-J. Molecular Expression of the Scribble Complex Genes, Dlg, Scrib and Lgl, in Silkworm, Bombyx mori. Genes. 2013; 4(2):264-274. https://doi.org/10.3390/genes4020264
Chicago/Turabian StyleQi, Hai-Sheng, Shu-Min Liu, Sheng Li, and Zhao-Jun Wei. 2013. "Molecular Expression of the Scribble Complex Genes, Dlg, Scrib and Lgl, in Silkworm, Bombyx mori" Genes 4, no. 2: 264-274. https://doi.org/10.3390/genes4020264






