MicroRNAs in the Stressed Heart: Sorting the Signal from the Noise
Abstract
:1. Introduction
2. A ‘Systems’ Approach is Key to Understanding MicroRNA Signaling
3. Assessing MicroRNAs Altered in the Stressed Heart
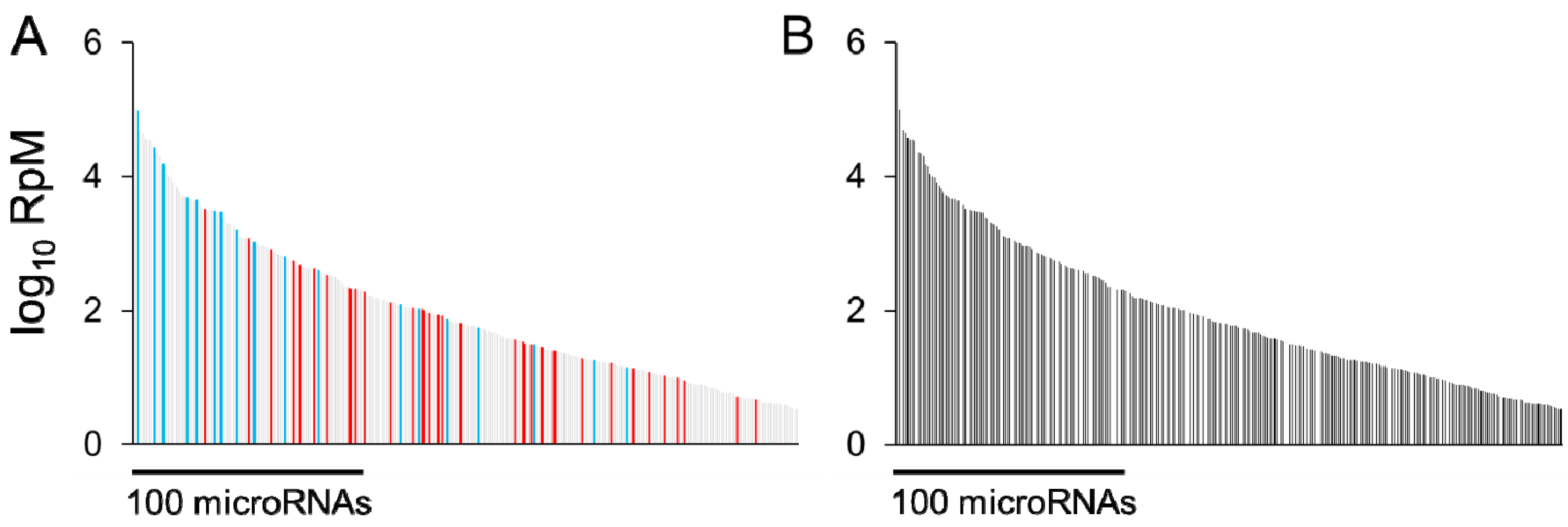
3.1. Which Altered MicroRNAs may be of the Most Biological Significance?
3.2. Challenges Faced by MicroRNA Target Prediction Algorithms
3.3. Quantifying MicroRNA-Dependent Regulation of mRNAs

3.3.1. Assays for mRNA Presence in, and Targeting by, the RISC
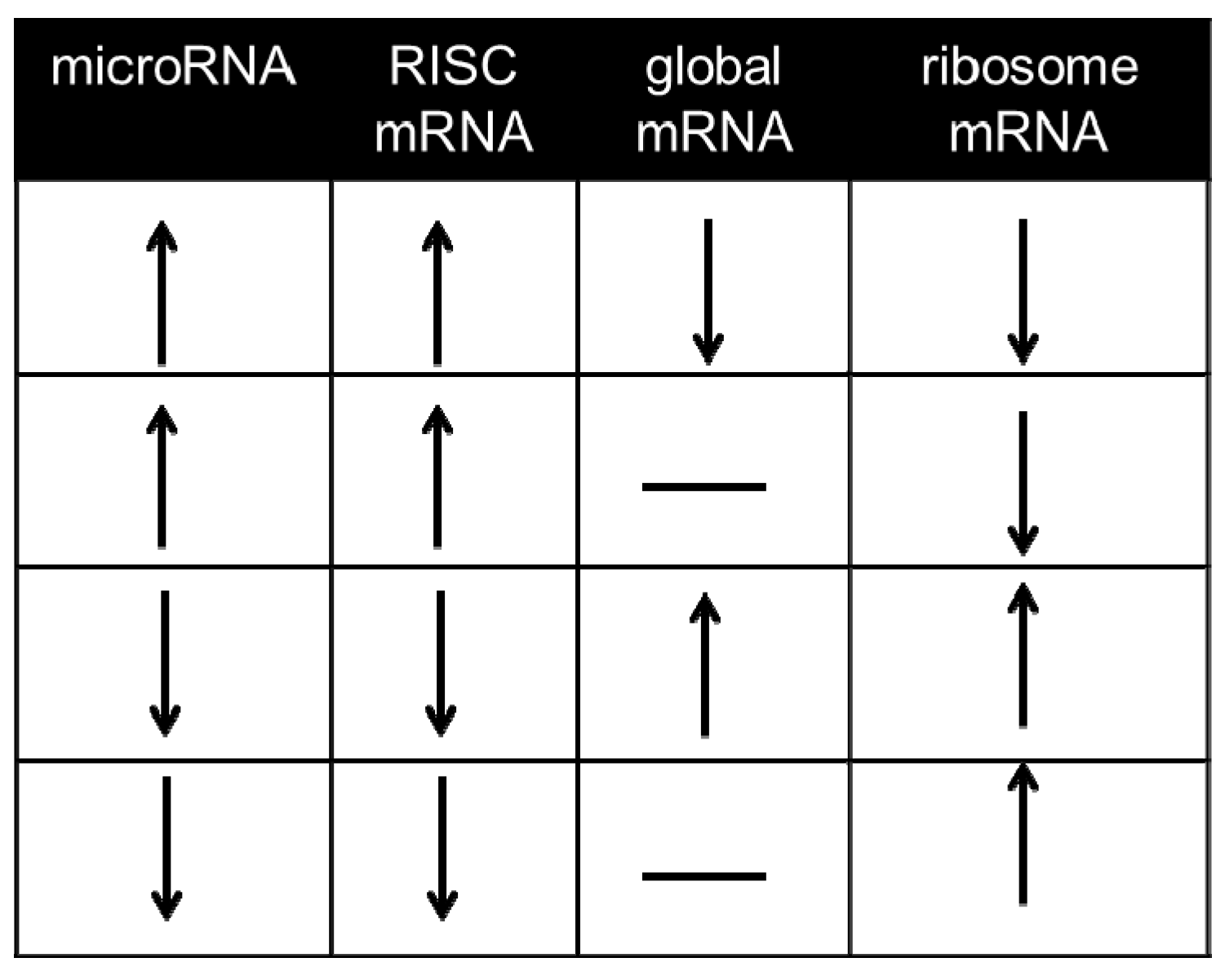
3.3.2. Pinning the MicroRNA to the Target—Approaches in the Absence of CLIP Experiments
3.3.3. Assays for Altered Translation of mRNAs
4. Assessing microRNAs not Altered in the Stressed Heart
5. Contextual Interpretation of Direct MicroRNA-mRNA Interaction Data
- mRNA sequence and structural considerations that permit microRNA/RISC-binding. While these parameters are likely to be quite stable among different contexts, what might be the effects of alternate 3′UTRs, other splice variants, or polymorphisms on microRNA binding sites? (A noteworthy example is the human angiotensin II type I receptor polymorphism that disrupts binding of miR-155 [81], a microRNA that also has cardiac functions [82].)
- Abundances of microRNAs and mRNAs and their possible stoichiometries are likely to differ between cell and tissue contexts.
- Competition may take place between mRNAs of different abundances but similar binding sites for access to the same microRNA. Furthermore, microRNAs have been reported to bind to long noncoding RNAs (lncRNAs), e.g., [83], amongst other competing endogenous RNAs (ceRNAs) such as circular RNAs [84]; at least one study has cast doubts on the significance of ceRNAs in titrating microRNA availability [85]. Nonetheless, the binding of microRNAs to mRNAs is a process that depends on multiple RNA interaction equilibria and will be altered by abundances of these other participants.
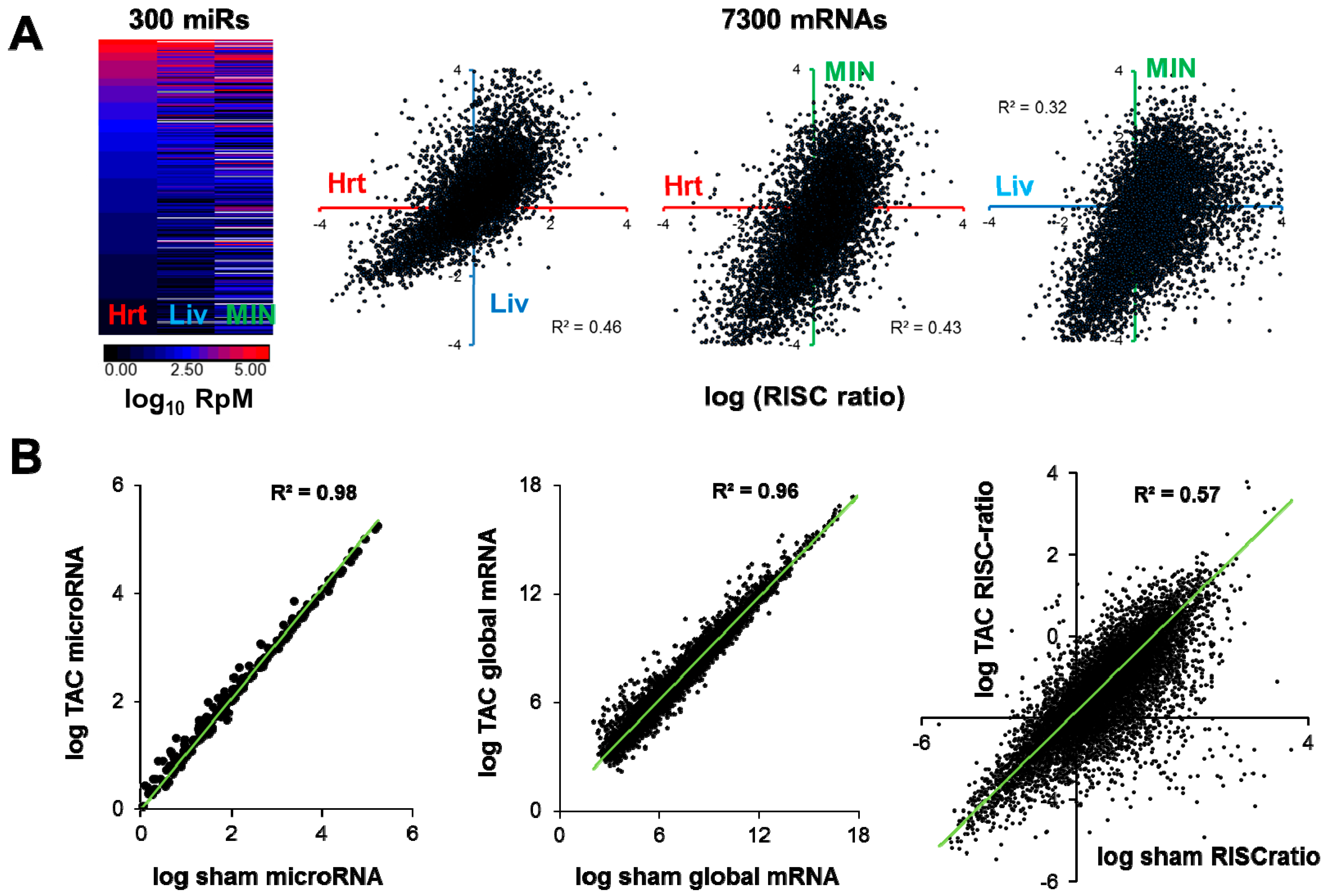
6. Validation, Perturbation and Integration
7. MicroRNA Therapeutics for Stress Relief?
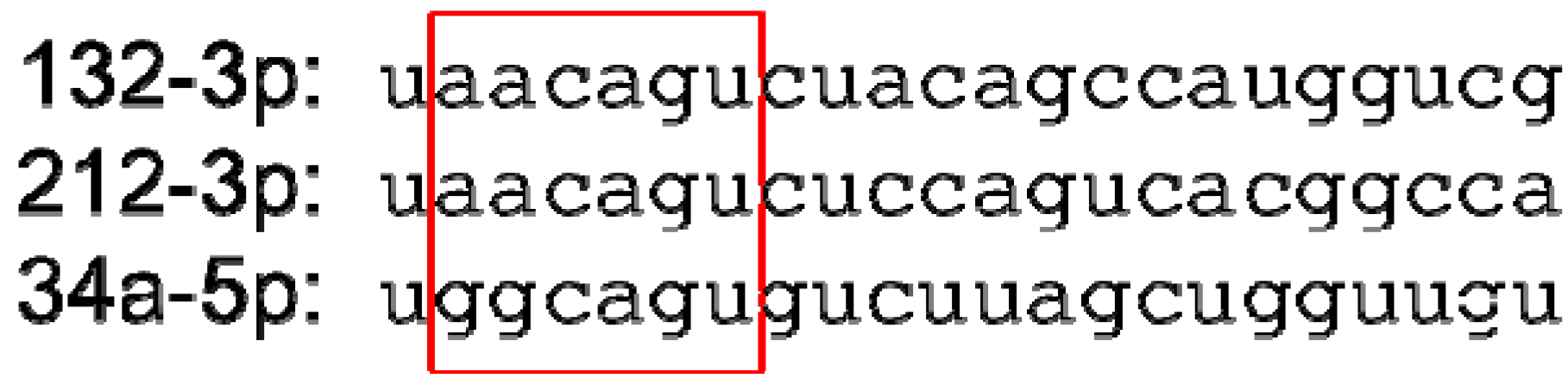
8. Conclusions
Acknowledgments
Conflicts of Interest
References
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: MicroRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.S.; Phillips, M.D.; Betel, D.; Mu, P.; Ventura, A.; Siepel, A.C.; Chen, K.C.; Lai, E.C. Widespread regulatory activity of vertebrate microRNA* species. RNA 2011, 17, 312–326. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Ransom, J.F.; Li, A.; Vedantham, V.; Von Drehle, M.; Muth, A.N.; Tsuchihashi, T.; McManus, M.T.; Schwartz, R.J.; Srivastava, D. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell 2007, 129, 303–317. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Bezprozvannaya, S.; Williams, A.H.; Qi, X.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N. microRNA-133a regulates cardiomyocyte proliferation and suppresses smooth muscle gene expression in the heart. Genes Dev. 2008, 22, 3242–3254. [Google Scholar] [CrossRef] [PubMed]
- Da Costa Martins, P.A.; Salic, K.; Gladka, M.M.; Armand, A.S.; Leptidis, S.; El Azzouzi, H.; Hansen, A.; Coenen-de Roo, C.J.; Bierhuizen, M.F.; Van der Nagel, R.; et al. MicroRNA-199b targets the nuclear kinase Dyrk1a in an auto-amplification loop promoting calcineurin/NFAT signalling. Nat. Cell Biol. 2010, 12, 1220–1227. [Google Scholar] [CrossRef] [PubMed]
- Matkovich, S.J.; Hu, Y.; Eschenbacher, W.H.; Dorn, L.E.; Dorn, G.W., II. Direct and indirect involvement of microRNA-499 in clinical and experimental cardiomyopathy. Circ. Res. 2012, 111, 521–531. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.P.; Chen, J.; Seok, H.Y.; Zhang, Z.; Kataoka, M.; Hu, X.; Wang, D.Z. MicroRNA-22 regulates cardiac hypertrophy and remodeling in response to stress. Circ. Res. 2013, 112, 1234–1243. [Google Scholar] [CrossRef] [PubMed]
- Gurha, P.; Abreu-Goodger, C.; Wang, T.; Ramirez, M.O.; Drumond, A.L.; van Dongen, S.; Chen, Y.; Bartonicek, N.; Enright, A.J.; Lee, B.; et al. Targeted deletion of microRNA-22 promotes stress-induced cardiac dilation and contractile dysfunction. Circulation 2012, 125, 2751–2761. [Google Scholar] [CrossRef] [PubMed]
- Gladka, M.M.; da Costa Martins, P.A.; De Windt, L.J. Small changes can make a big difference—MicroRNA regulation of cardiac hypertrophy. J. Mol. Cell. Cardiol. 2012, 52, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Kuppusamy, K.T.; Sperber, H.; Ruohola-Baker, H. MicroRNA regulation and role in stem cell maintenance, cardiac differentiation and hypertrophy. Curr. Mol. Med. 2013, 13, 757–764. [Google Scholar] [CrossRef] [PubMed]
- Ooi, J.Y.; Bernardo, B.C.; McMullen, J.R. The therapeutic potential of miRNAs regulated in settings of physiological cardiac hypertrophy. Future Med. Chem. 2014, 6, 205–222. [Google Scholar] [CrossRef] [PubMed]
- Fiedler, J.; Thum, T. MicroRNAs in myocardial infarction. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 201–205. [Google Scholar] [CrossRef] [PubMed]
- Greco, S.; Gaetano, C.; Martelli, F. HypoxamiR regulation and function in ischemic cardiovascular diseases. Antioxid. Redox Signal. 2013. [Google Scholar] [CrossRef]
- Topkara, V.K.; Mann, D.L. Role of microRNAs in cardiac remodeling and heart failure. Cardiovasc. Drugs Ther. 2011, 25, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Dorn, I.I.; Gerald, W. MicroRNAs in cardiac disease. Transl. Res. 2011, 157, 226–235. [Google Scholar] [CrossRef] [PubMed]
- Melman, Y.F.; Shah, R.; Das, S. MicroRNAs in Heart Failure: Is the Picture Becoming Less miRky? Circ. Heart Fail. 2014, 7, 203–214. [Google Scholar] [CrossRef]
- Lusis, A.J.; Weiss, J.N. Cardiovascular networks: Systems-based approaches to cardiovascular disease. Circulation 2010, 121, 157–170. [Google Scholar] [CrossRef] [PubMed]
- Dewey, F.E.; Wheeler, M.T.; Ashley, E.A. Systems biology of heart failure, challenges and hopes. Curr. Opin. Cardiol. 2011, 26, 314–321. [Google Scholar] [CrossRef] [PubMed]
- Parikh, V.N.; Jin, R.C.; Rabello, S.; Gulbahce, N.; White, K.; Hale, A.; Cottrill, K.A.; Shaik, R.S.; Waxman, A.B.; Zhang, Y.Y.; et al. MicroRNA-21 integrates pathogenic signaling to control pulmonary hypertension: Results of a network bioinformatics approach. Circulation 2012, 125, 1520–1532. [Google Scholar] [CrossRef] [PubMed]
- Gerstein, M.B.; Kundaje, A.; Hariharan, M.; Landt, S.G.; Yan, K.K.; Cheng, C.; Mu, X.J.; Khurana, E.; Rozowsky, J.; Alexander, R.; et al. Architecture of the human regulatory network derived from ENCODE data. Nature 2012, 489, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Judson, R.L.; Greve, T.S.; Parchem, R.J.; Blelloch, R. MicroRNA-based discovery of barriers to dedifferentiation of fibroblasts to pluripotent stem cells. Nat. Struct. Mol. Biol. 2013, 20, 1227–1235. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2011, 39, D152–157. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.J.; Zhang, C.; Gantman, E.C.; Mele, A.; Darnell, J.C.; Darnell, R.B. Mapping Argonaute and conventional RNA-binding protein interactions with RNA at single-nucleotide resolution using HITS-CLIP and CIMS analysis. Nat. Protoc. 2014, 9, 263–293. [Google Scholar] [CrossRef] [PubMed]
- Hafner, M.; Lianoglou, S.; Tuschl, T.; Betel, D. Genome-wide identification of miRNA targets by PAR-CLIP. Methods 2012, 58, 94–105. [Google Scholar] [CrossRef] [PubMed]
- Erhard, F.; Dolken, L.; Jaskiewicz, L.; Zimmer, R. PARma: Identification of microRNA target sites in AGO-PAR-CLIP data. Genome Biol. 2013, 14, R79. [Google Scholar] [CrossRef] [PubMed]
- Khorshid, M.; Hausser, J.; Zavolan, M.; van Nimwegen, E. A biophysical miRNA-mRNA interaction model infers canonical and noncanonical targets. Nat. Methods 2013, 10, 253–255. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Mallick, B.; Long, D.; Rennie, W.A.; Wolenc, A.; Carmack, C.S.; Ding, Y. CLIP-based prediction of mammalian microRNA binding sites. Nucleic Acids Res. 2013, 41, e138. [Google Scholar] [CrossRef] [PubMed]
- Rennie, W.; Liu, C.; Carmack, C.S.; Wolenc, A.; Kanoria, S.; Lu, J.; Long, D.; Ding, Y. STarMir: A web server for prediction of microRNA binding sites. Nucleic Acids Res. W1 2014, W114–W118. [Google Scholar] [CrossRef]
- Care, A.; Catalucci, D.; Felicetti, F.; Bonci, D.; Addario, A.; Gallo, P.; Bang, M.L.; Segnalini, P.; Gu, Y.; Dalton, N.D.; et al. MicroRNA-133 controls cardiac hypertrophy. Nat. Med. 2007, 13, 613–618. [Google Scholar] [CrossRef] [PubMed]
- Van Rooij, E.; Quiat, D.; Johnson, B.A.; Sutherland, L.B.; Qi, X.; Richardson, J.A.; Kelm, R.J., Jr.; Olson, E.N. A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance. Dev. Cell 2009, 17, 662–673. [Google Scholar] [CrossRef] [PubMed]
- Matkovich, S.J.; Wang, W.; Tu, Y.; Eschenbacher, W.H.; Dorn, L.E.; Condorelli, G.; Diwan, A.; Nerbonne, J.M.; Dorn, G.W., 2nd. MicroRNA-133a protects against myocardial fibrosis and modulates electrical repolarization without affecting hypertrophy in pressure-overloaded adult hearts. Circ. Res. 2010, 106, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Patrick, D.M.; Montgomery, R.L.; Qi, X.; Obad, S.; Kauppinen, S.; Hill, J.A.; Van Rooij, E.; Olson, E.N. Stress-dependent cardiac remodeling occurs in the absence of microRNA-21 in mice. J. Clin. Invest. 2010, 120, 3912–3916. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Matkovich, S.J.; Hecker, P.A.; Zhang, Y.; Dorn, G.W., II. Epitranscriptional orchestration of genetic reprogramming is an emergent property of stress-regulated cardiac microRNAs. Proc. Natl. Acad. Sci. USA 2012, 109, 19864–19869. [Google Scholar] [CrossRef] [PubMed]
- Matkovich, S.J.; Hu, Y.; Dorn, G.W. Regulation of cardiac microRNAs. Circ. Res. 2013, 113, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Mencia, A.; Modamio-Hoybjor, S.; Redshaw, N.; Morin, M.; Mayo-Merino, F.; Olavarrieta, L.; Aguirre, L.A.; del Castillo, I.; Steel, K.P.; Dalmay, T.; et al. Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss. Nat. Genet. 2009, 41, 609–613. [Google Scholar] [CrossRef] [PubMed]
- Dorn, G.W., 2nd; Matkovich, S.J.; Eschenbacher, W.H.; Zhang, Y. A human 3′ miR-499 mutation alters cardiac mRNA targeting and function. Circ. Res. 2012, 110, 958–967. [Google Scholar]
- Humphreys, D.T.; Hynes, C.J.; Patel, H.R.; Wei, G.H.; Cannon, L.; Fatkin, D.; Suter, C.M.; Clancy, J.L.; Preiss, T. Complexity of murine cardiomyocyte miRNA biogenesis, sequence variant expression and function. PLoS One 2012, 7, e30933. [Google Scholar] [CrossRef] [PubMed]
- Garcia, D.M.; Baek, D.; Shin, C.; Bell, G.W.; Grimson, A.; Bartel, D.P. Weak seed-pairing stability and high target-site abundance decrease the proficiency of lsy-6 and other microRNAs. Nat. Struct. Mol. Biol. 2011, 18, 1139–1146. [Google Scholar] [CrossRef] [PubMed]
- Grimson, A.; Farh, K.K.; Johnston, W.K.; Garrett-Engele, P.; Lim, L.P.; Bartel, D.P. MicroRNA targeting specificity in mammals: Determinants beyond seed pairing. Mol. Cell 2007, 27, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Krek, A.; Grun, D.; Poy, M.N.; Wolf, R.; Rosenberg, L.; Epstein, E.J.; MacMenamin, P.; Da Piedade, I.; Gunsalus, K.C.; Stoffel, M.; et al. Combinatorial microRNA target predictions. Nat. Genet. 2005, 37, 495–500. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; El Naqa, I.M. Prediction of both conserved and nonconserved microRNA targets in animals. Bioinformatics 2008, 24, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Wang, X. miRDB: A microRNA target prediction and functional annotation database with a wiki interface. RNA 2008, 14, 1012–1017. [Google Scholar] [CrossRef] [PubMed]
- Chi, S.W.; Hannon, G.J.; Darnell, R.B. An alternative mode of microRNA target recognition. Nat. Struct. Mol. Biol. 2012, 19, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Hausser, J.; Syed, A.P.; Bilen, B.; Zavolan, M. Analysis of CDS-located miRNA target sites suggests that they can effectively inhibit translation. Genome Res. 2013, 23, 604–615. [Google Scholar] [CrossRef] [PubMed]
- Shin, C.; Nam, J.W.; Farh, K.K.; Chiang, H.R.; Shkumatava, A.; Bartel, D.P. Expanding the microRNA targeting code: Functional sites with centered pairing. Mol. Cell 2010, 38, 789–802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsang, J.S.; Ebert, M.S.; van Oudenaarden, A. Genome-wide dissection of microRNA functions and cotargeting networks using gene set signatures. Mol. Cell 2010, 38, 140–153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matkovich, S.J.; Zhang, Y.; Van Booven, D.; Dorn, G.W., 2nd. Deep mRNA sequencing for in vivo functional analysis of cardiac transcriptional regulators. Application to Gαq. Circ. Res. 2010, 106, 1459–1467. [Google Scholar]
- Baillat, D.; Shiekhattar, R. Functional dissection of the human TNRC6 (GW182-related) family of proteins. Mol. Cell. Biol. 2009, 29, 4144–4155. [Google Scholar] [CrossRef] [PubMed]
- Cambronne, X.A.; Shen, R.; Auer, P.L.; Goodman, R.H. Capturing microRNA targets using an RNA-induced silencing complex (RISC)-trap approach. Proc. Natl. Acad. Sci. USA 2012, 109, 20473–20478. [Google Scholar] [CrossRef] [PubMed]
- Hafner, M.; Landthaler, M.; Burger, L.; Khorshid, M.; Hausser, J.; Berninger, P.; Rothballer, A.; Ascano, M., Jr.; Jungkamp, A.C.; Munschauer, M.; et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 2010, 141, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Chi, S.W.; Zang, J.B.; Mele, A.; Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 2009, 460, 479–486. [Google Scholar] [PubMed]
- Wang, W.X.; Wilfred, B.R.; Hu, Y.; Stromberg, A.J.; Nelson, P.T. Anti-Argonaute RIP-Chip shows that miRNA transfections alter global patterns of mRNA recruitment to microribonucleoprotein complexes. RNA 2010, 16, 394–404. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.X.; Wilfred, B.R.; Xie, K.; Jennings, M.H.; Hu, Y.H.; Stromberg, A.J.; Nelson, P.T. Individual microRNAs (miRNAs) display distinct mRNA targeting “rules”. RNA Biol. 2010, 7, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Matkovich, S.J.; Van Booven, D.J.; Eschenbacher, W.H.; Dorn, G.W., 2nd. RISC RNA sequencing for context-specific identification of in vivo microRNA targets. Circ. Res. 2011, 108, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Boudreau, R.L.; Jiang, P.; Gilmore, B.L.; Spengler, R.M.; Tirabassi, R.; Nelson, J.A.; Ross, C.A.; Xing, Y.; Davidson, B.L. Transcriptome-wide discovery of microRNA binding sites in human brain. Neuron 2014, 81, 294–305. [Google Scholar] [CrossRef] [PubMed]
- Tattikota, S.G.; Rathjen, T.; McAnulty, S.J.; Wessels, H.H.; Akerman, I.; van de Bunt, M.; Hausser, J.; Esguerra, J.L.; Musahl, A.; Pandey, A.K.; et al. Argonaute2 Mediates Compensatory Expansion of the Pancreatic beta Cell. Cell MeTable 2014, 19, 122–134. [Google Scholar] [CrossRef]
- Karginov, F.V.; Conaco, C.; Xuan, Z.; Schmidt, B.H.; Parker, J.S.; Mandel, G.; Hannon, G.J. A biochemical approach to identifying microRNA targets. Proc. Natl. Acad. Sci. USA 2007, 104, 19291–19296. [Google Scholar] [CrossRef] [PubMed]
- Helwak, A.; Kudla, G.; Dudnakova, T.; Tollervey, D. Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 2013, 153, 654–665. [Google Scholar] [CrossRef] [PubMed]
- Fabian, M.R.; Sonenberg, N. The mechanics of miRNA-mediated gene silencing: A look under the hood of miRISC. Nat. Struct. Mol. Biol. 2012, 19, 586–593. [Google Scholar] [PubMed]
- Djuranovic, S.; Nahvi, A.; Green, R. miRNA-mediated gene silencing by translational repression followed by mRNA deadenylation and decay. Science 2012, 336, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Kameswaran, V.; Bramswig, N.C.; McKenna, L.B.; Penn, M.; Schug, J.; Hand, N.J.; Chen, Y.; Choi, I.; Vourekas, A.; Won, K.J.; et al. Epigenetic regulation of the DLK1-MEG3 microRNA cluster in human type 2 diabetic islets. Cell MeTable 2014, 19, 135–145. [Google Scholar] [CrossRef]
- Burger, K.; Muhl, B.; Kellner, M.; Rohrmoser, M.; Gruber-Eber, A.; Windhager, L.; Friedel, C.C.; Dolken, L.; Eick, D. 4-thiouridine inhibits rRNA synthesis and causes a nucleolar stress response. RNA Biol. 2013, 10. [Google Scholar]
- Androsavich, J.R.; Chau, B.N. Non-inhibited miRNAs shape the cellular response to anti-miR. Nucleic Acids Res. 2014. [Google Scholar] [CrossRef]
- Kishore, S.; Jaskiewicz, L.; Burger, L.; Hausser, J.; Khorshid, M.; Zavolan, M. A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins. Nat. Methods 2011, 8, 559–564. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Long, D.; Lee, R.; Williams, P.; Chan, C.Y.; Ambros, V.; Ding, Y. Potent effect of target structure on microRNA function. Nat. Struct. Mol. Biol. 2007, 14, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Rehmsmeier, M.; Steffen, P.; Hochsmann, M.; Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 2004, 10, 1507–1517. [Google Scholar] [CrossRef] [PubMed]
- Meijer, H.A.; Kong, Y.W.; Lu, W.T.; Wilczynska, A.; Spriggs, R.V.; Robinson, S.W.; Godfrey, J.D.; Willis, A.E.; Bushell, M. Translational repression and eIF4A2 activity are critical for microRNA-mediated gene regulation. Science 2013, 340, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Ingolia, N.T.; Brar, G.A.; Rouskin, S.; McGeachy, A.M.; Weissman, J.S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 2012, 7, 1534–1550. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, M.J.; Vernal, R.; Dolznig, H.; Mullner, E.W.; Garcia-Sanz, J.A. Isolation of polysome-bound mRNA from solid tissues amenable for RT-PCR and profiling experiments. RNA 2007, 13, 414–421. [Google Scholar] [CrossRef] [PubMed]
- Drawnel, F.M.; Wachten, D.; Molkentin, J.D.; Maillet, M.; Aronsen, J.M.; Swift, F.; Sjaastad, I.; Liu, N.; Catalucci, D.; Mikoshiba, K.; et al. Mutual antagonism between IP(3)RII and miRNA-133a regulates calcium signals and cardiac hypertrophy. J. Cell Biol. 2012, 199, 783–798. [Google Scholar] [CrossRef] [PubMed]
- Baek, D.; Villen, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Selbach, M.; Schwanhausser, B.; Thierfelder, N.; Fang, Z.; Khanin, R.; Rajewsky, N. Widespread changes in protein synthesis induced by microRNAs. Nature 2008, 455, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Mallat, Y.; Tritsch, E.; Ladouce, R.; Winter, D.L.; Friguet, B.; Li, Z.; Mericskay, M. Proteome modulation in H9c2 cardiac cells by microRNAs miR-378 and miR-378*. Mol. Cell. Proteomics 2014, 13, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Raddatz, K.; Albrecht, D.; Hochgrafe, F.; Hecker, M.; Gotthardt, M. A proteome map of murine heart and skeletal muscle. Proteomics 2008, 8, 1885–1897. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.F.; Murchison, E.P.; Tang, R.; Callis, T.E.; Tatsuguchi, M.; Deng, Z.; Rojas, M.; Hammond, S.M.; Schneider, M.D.; Selzman, C.H.; et al. Targeted deletion of Dicer in the heart leads to dilated cardiomyopathy and heart failure. Proc. Natl. Acad. Sci. USA 2008, 105, 2111–2116. [Google Scholar] [CrossRef] [PubMed]
- Carrer, M.; Liu, N.; Grueter, C.E.; Williams, A.H.; Frisard, M.I.; Hulver, M.W.; Bassel-Duby, R.; Olson, E.N. Control of mitochondrial metabolism and systemic energy homeostasis by microRNAs 378 and 378*. Proc. Natl. Acad. Sci. USA 2012, 109, 15330–15335. [Google Scholar] [CrossRef] [PubMed]
- Ganesan, J.; Ramanujam, D.; Sassi, Y.; Ahles, A.; Jentzsch, C.; Werfel, S.; Leierseder, S.; Loyer, X.; Giacca, M.; Zentilin, L.; et al. MiR-378 controls cardiac hypertrophy by combined repression of mitogen-activated protein kinase pathway factors. Circulation 2013, 127, 2097–2106. [Google Scholar] [CrossRef] [PubMed]
- Mendell, J.T.; Olson, E.N. MicroRNAs in stress signaling and human disease. Cell 2012, 148, 1172–1187. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.M.; Buckenberger, J.A.; Jiang, J.; Malana, G.E.; Nuovo, G.J.; Chotani, M.; Feldman, D.S.; Schmittgen, T.D.; Elton, T.S. The human angiotensin II type 1 receptor +1166 A/C polymorphism attenuates microRNA-155 binding. J. Biol. Chem. 2007, 282, 24262–24269. [Google Scholar] [CrossRef] [PubMed]
- Seok, H.Y.; Chen, J.; Kataoka, M.; Huang, Z.P.; Ding, J.; Yan, J.; Hu, X.; Wang, D.Z. Loss of microRNA-155 protects the heart from pathological cardiac hypertrophy. Circ. Res. 2014, 114, 1585–1595. [Google Scholar] [CrossRef] [PubMed]
- Kallen, A.N.; Zhou, X.B.; Xu, J.; Qiao, C.; Ma, J.; Yan, L.; Lu, L.; Liu, C.; Yi, J.S.; Zhang, H.; et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol. Cell 2013, 52, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Rinn, J.; Pandolfi, P.P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014, 505, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Denzler, R.; Agarwal, V.; Stefano, J.; Bartel, D.P.; Stoffel, M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol. Cell 2014, 54, 766–776. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Broad_Institute GSEA. Available online: http://www.broadinstitute.org/gsea/index.jsp (accessed on 18 July 2014).
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Thomson_Reuters MetaCore. Available online: http://www.genego.com (accessed on 18 July 2014).
- Gurha, P.; Wang, T.; Larimore, A.H.; Sassi, Y.; Abreu-Goodger, C.; Ramirez, M.O.; Reddy, A.K.; Engelhardt, S.; Taffet, G.E.; Wehrens, X.H.; et al. microRNA-22 promotes heart failure through coordinate suppression of PPAR/ERR-nuclear hormone receptor transcription. PLoS One 2013, 8, e75882. [Google Scholar] [CrossRef] [PubMed]
- Terentyev, D.; Belevych, A.E.; Terentyeva, R.; Martin, M.M.; Malana, G.E.; Kuhn, D.E.; Abdellatif, M.; Feldman, D.S.; Elton, T.S.; Gyorke, S. miR-1 overexpression enhances Ca(2+) release and promotes cardiac arrhythmogenesis by targeting PP2A regulatory subunit B56alpha and causing CaMKII-dependent hyperphosphorylation of RyR2. Circ. Res. 2009, 104, 514–521. [Google Scholar] [CrossRef] [PubMed]
- Abdi, A.; Tahoori, M.B.; Emamian, E.S. Fault diagnosis engineering of digital circuits can identify vulnerable molecules in complex cellular pathways. Sci. Signal 2008, 1, ra10. [Google Scholar] [PubMed]
- Ryall, K.A.; Holland, D.O.; Delaney, K.A.; Kraeutler, M.J.; Parker, A.J.; Saucerman, J.J. Network reconstruction and systems analysis of cardiac myocyte hypertrophy signaling. J. Biol. Chem. 2012, 287, 42259–42268. [Google Scholar] [CrossRef] [PubMed]
- Van Rooij, E.; Purcell, A.L.; Levin, A.A. Developing microRNA therapeutics. Circ. Res. 2012, 110, 496–507. [Google Scholar] [CrossRef] [PubMed]
- Krutzfeldt, J.; Rajewsky, N.; Braich, R.; Rajeev, K.G.; Tuschl, T.; Manoharan, M.; Stoffel, M. Silencing of microRNAs in vivo with “antagomirs”. Nature 2005, 438, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Thum, T.; Gross, C.; Fiedler, J.; Fischer, T.; Kissler, S.; Bussen, M.; Galuppo, P.; Just, S.; Rottbauer, W.; Frantz, S.; et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 2008, 456, 980–984. [Google Scholar] [CrossRef] [PubMed]
- Montgomery, R.L.; Hullinger, T.G.; Semus, H.M.; Dickinson, B.A.; Seto, A.G.; Lynch, J.M.; Stack, C.; Latimer, P.A.; Olson, E.N.; van Rooij, E. Therapeutic inhibition of miR-208a improves cardiac function and survival during heart failure. Circulation 2011, 124, 1537–1547. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, B.C.; Gao, X.M.; Winbanks, C.E.; Boey, E.J.; Tham, Y.K.; Kiriazis, H.; Gregorevic, P.; Obad, S.; Kauppinen, S.; Du, X.J.; et al. Therapeutic inhibition of the miR-34 family attenuates pathological cardiac remodeling and improves heart function. Proc. Natl. Acad. Sci. USA 2012, 109, 17615–17620. [Google Scholar] [CrossRef] [PubMed]
- Ucar, A.; Gupta, S.K.; Fiedler, J.; Erikci, E.; Kardasinski, M.; Batkai, S.; Dangwal, S.; Kumarswamy, R.; Bang, C.; Holzmann, A.; et al. The miRNA-212/132 family regulates both cardiac hypertrophy and cardiomyocyte autophagy. Nat. Commun. 2012, 3, 1078. [Google Scholar] [CrossRef] [PubMed]
- Boon, R.A.; Iekushi, K.; Lechner, S.; Seeger, T.; Fischer, A.; Heydt, S.; Kaluza, D.; Treguer, K.; Carmona, G.; Bonauer, A.; et al. MicroRNA-34a regulates cardiac ageing and function. Nature 2013, 495, 107–110. [Google Scholar] [CrossRef] [PubMed]
- Li, R.C.; Tao, J.; Guo, Y.B.; Wu, H.D.; Liu, R.F.; Bai, Y.; Lv, Z.Z.; Luo, G.Z.; Li, L.L.; Wang, M.; et al. In vivo suppression of microRNA-24 prevents the transition toward decompensated hypertrophy in aortic-constricted mice. Circ. Res. 2013, 112, 601–605. [Google Scholar] [CrossRef] [PubMed]
- Thum, T.; Chau, N.; Bhat, B.; Gupta, S.K.; Linsley, P.S.; Bauersachs, J.; Engelhardt, S. Comparison of different miR-21 inhibitor chemistries in a cardiac disease model. J. Clin. Invest. 2011, 121, 461–462. [Google Scholar] [CrossRef] [PubMed]
- Patrick, D.M.; Kauppinen, S.; van Rooij, E.; Olson, E.N. Response to Thum; et al. J. Clin. Invest. 2011, 121, 462–463. [Google Scholar] [CrossRef]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; Van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Bouchie, A. First microRNA mimic enters clinic. Nat. Biotechnol. 2013, 31, 577. [Google Scholar] [CrossRef] [PubMed]
- Grueter, C.E.; Van Rooij, E.; Johnson, B.A.; DeLeon, S.M.; Sutherland, L.B.; Qi, X.; Gautron, L.; Elmquist, J.K.; Bassel-Duby, R.; Olson, E.N. A cardiac microRNA governs systemic energy homeostasis by regulation of MED13. Cell 2012, 149, 671–683. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Matkovich, S.J. MicroRNAs in the Stressed Heart: Sorting the Signal from the Noise. Cells 2014, 3, 778-801. https://doi.org/10.3390/cells3030778
Matkovich SJ. MicroRNAs in the Stressed Heart: Sorting the Signal from the Noise. Cells. 2014; 3(3):778-801. https://doi.org/10.3390/cells3030778
Chicago/Turabian StyleMatkovich, Scot J. 2014. "MicroRNAs in the Stressed Heart: Sorting the Signal from the Noise" Cells 3, no. 3: 778-801. https://doi.org/10.3390/cells3030778



