Microbial Endophytes that Live within the Seeds of Two Tomato Hybrids Cultivated in Argentina
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Community: Structure and Diversity
2.2. Isolation of Bacteria from Tomato Seeds
2.3. Extraction of Total DNA, PCR Amplification and Sequencing of the 16S rDNA Partial Gene
2.4. In Vitro Antagonism of Endophytic Bacteria towards Tomato Pathogens
2.4.1. In Vivo Bioassays of the Pathogen Inhibition Effects of Bacteria
2.4.2. Inhibitory Activity of the Cell-Free Supernatant of Endophytic Bacteria against Fungal Pathogens
2.4.3. Effect of Volatiles from Endophytic Bacteria against Fungal Pathogens
2.5. Bacterial Effect on Tomato Growth
2.6. Siderophore and Phytohormone Production and Phosphate Solubilization
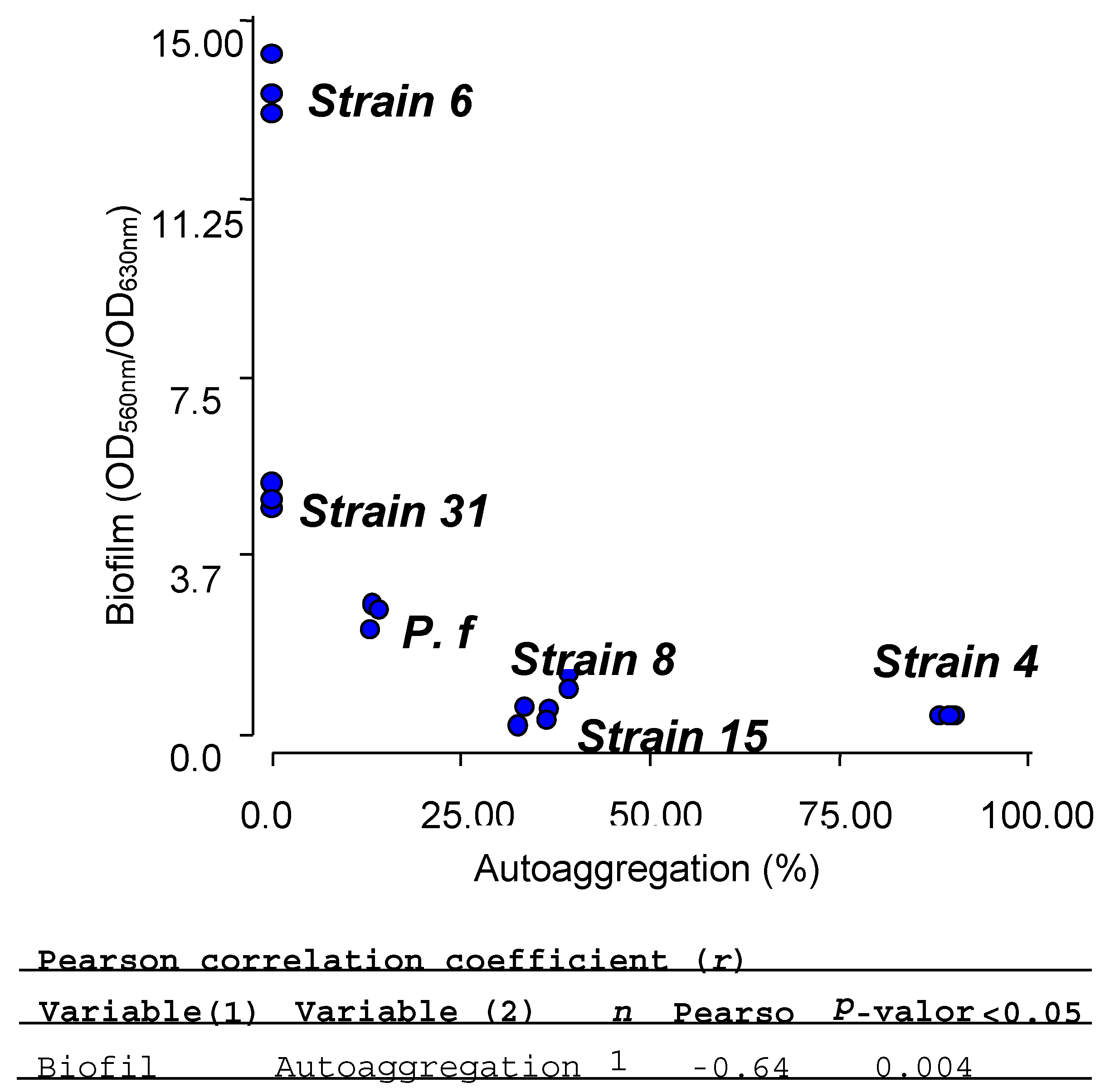
2.7. Biofilm and Autoaggregation Assays
3. Results
3.1. Bacterial Community
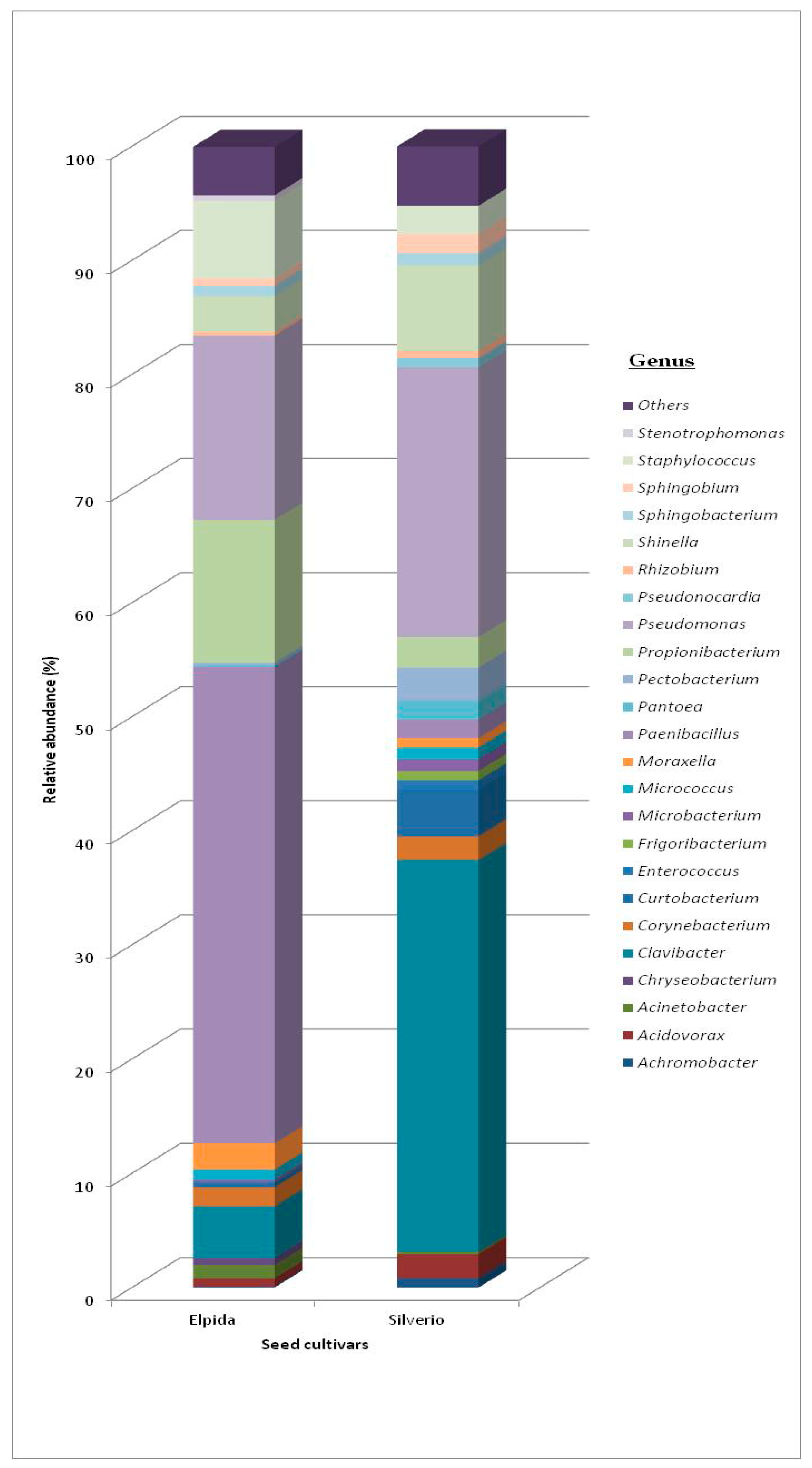
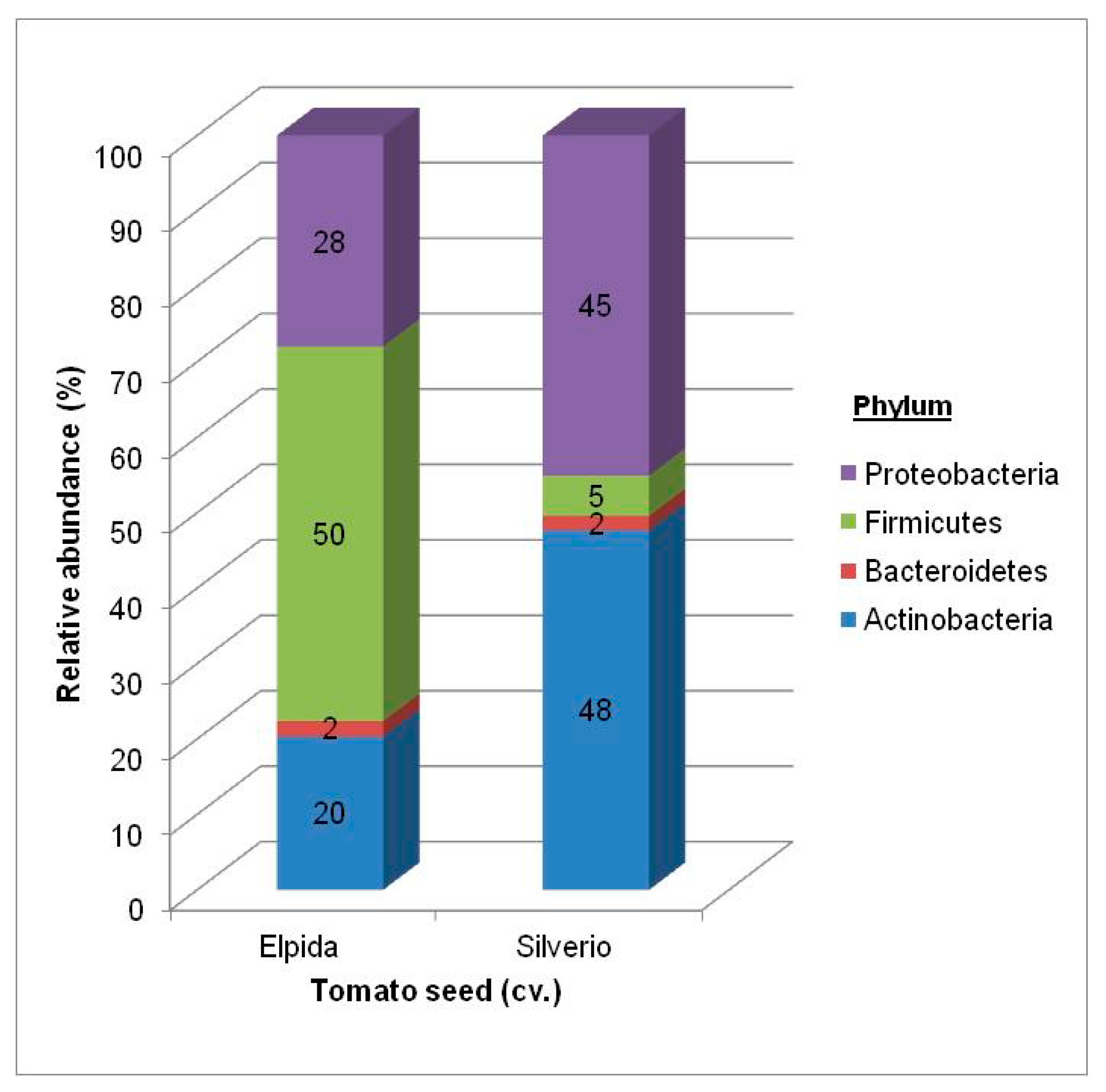
3.1.1. Total Bacterial Community Structure and Diversity
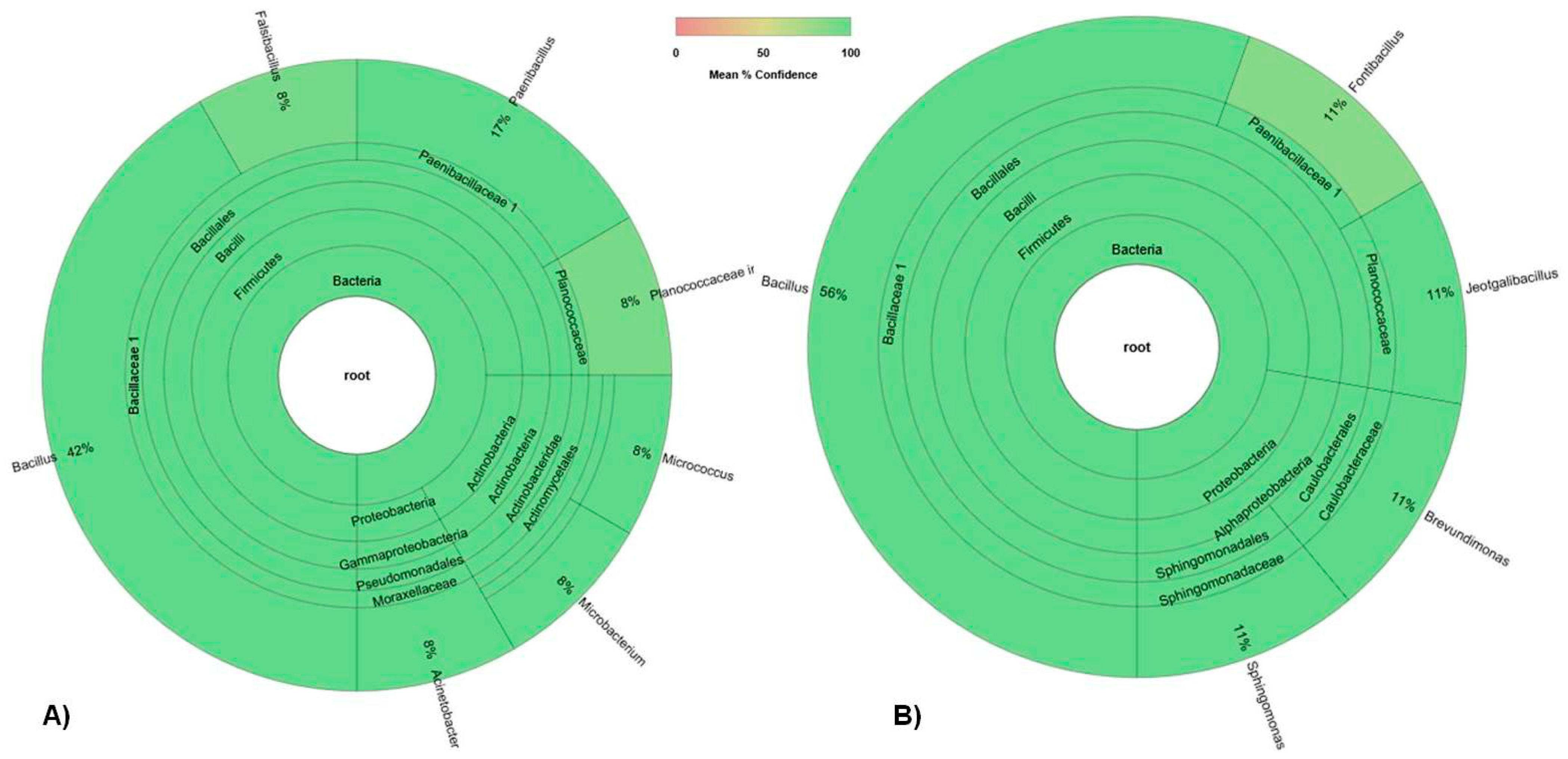
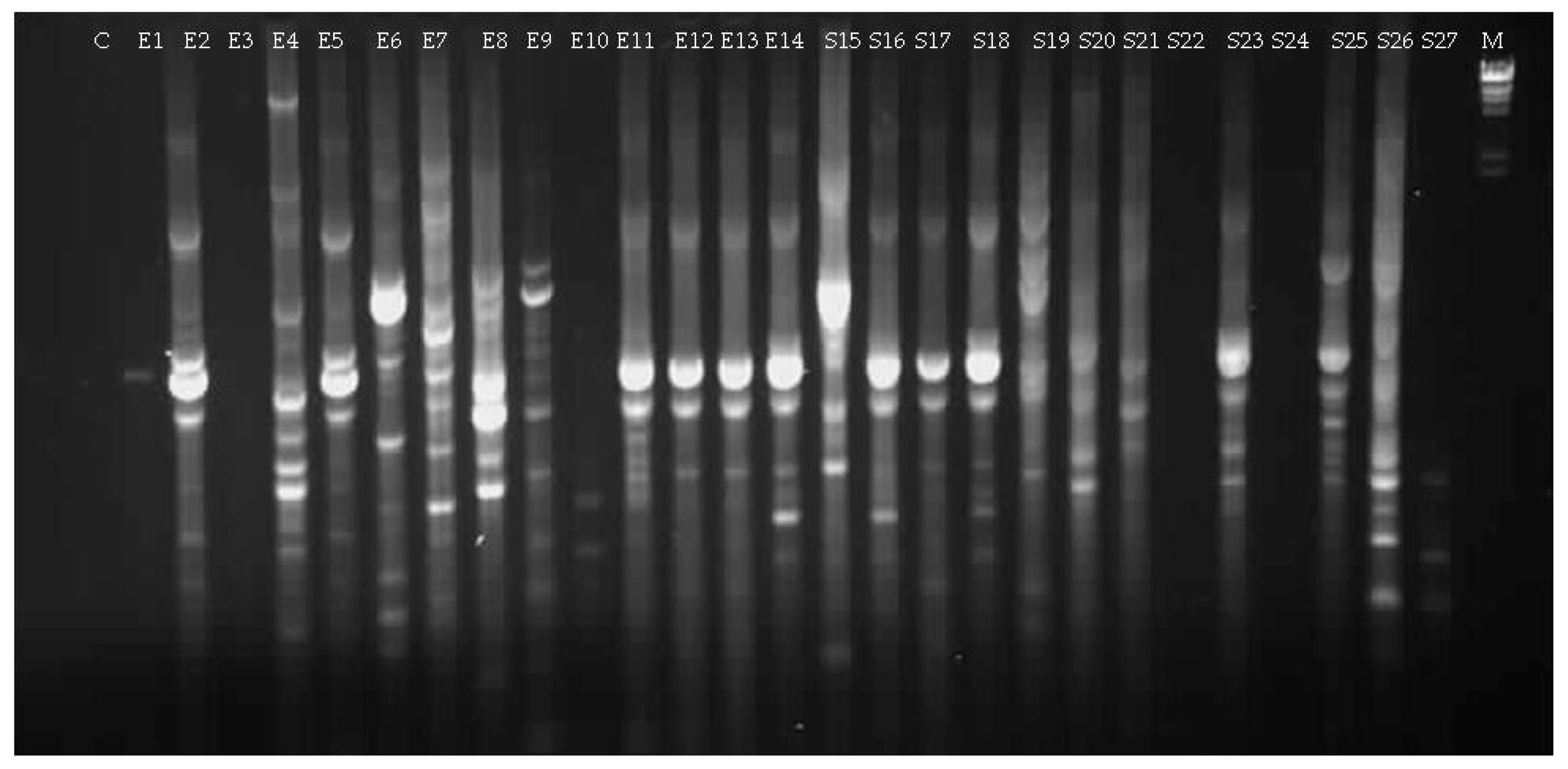
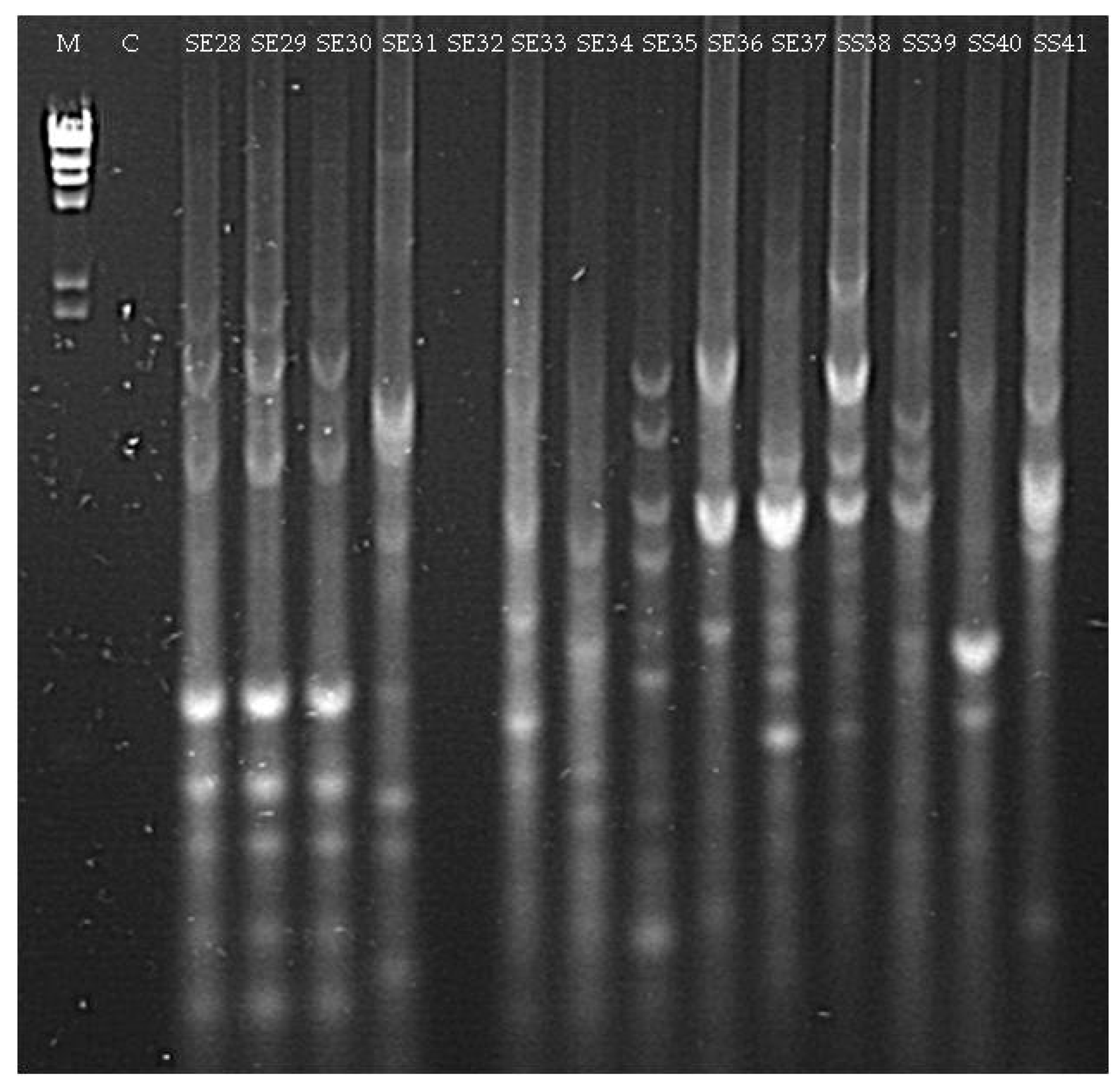
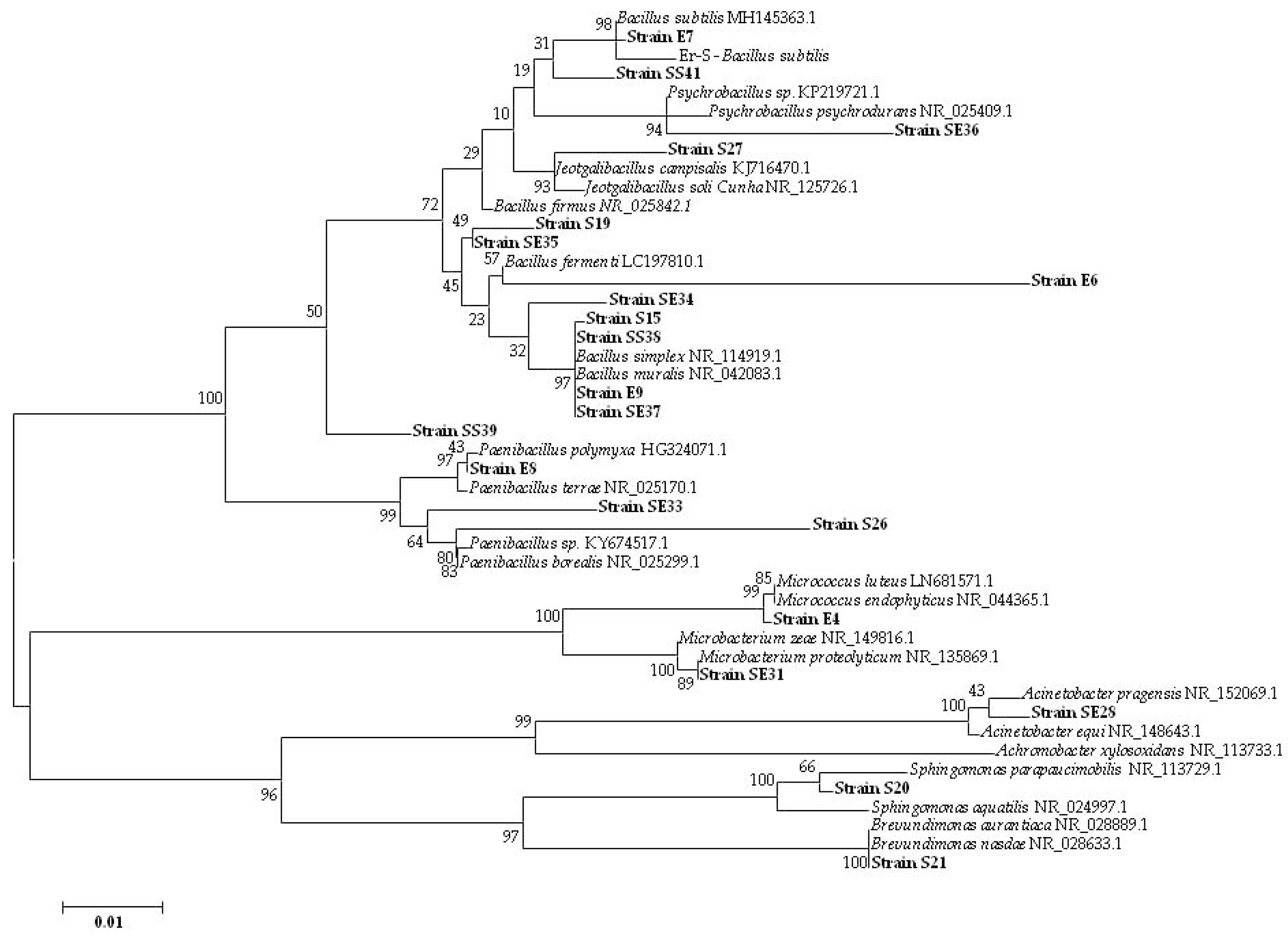
3.1.2. Culturable Bacterial Community
3.2. In Vitro Antagonism of Endophytic Bacteria towards Tomato Pathogens
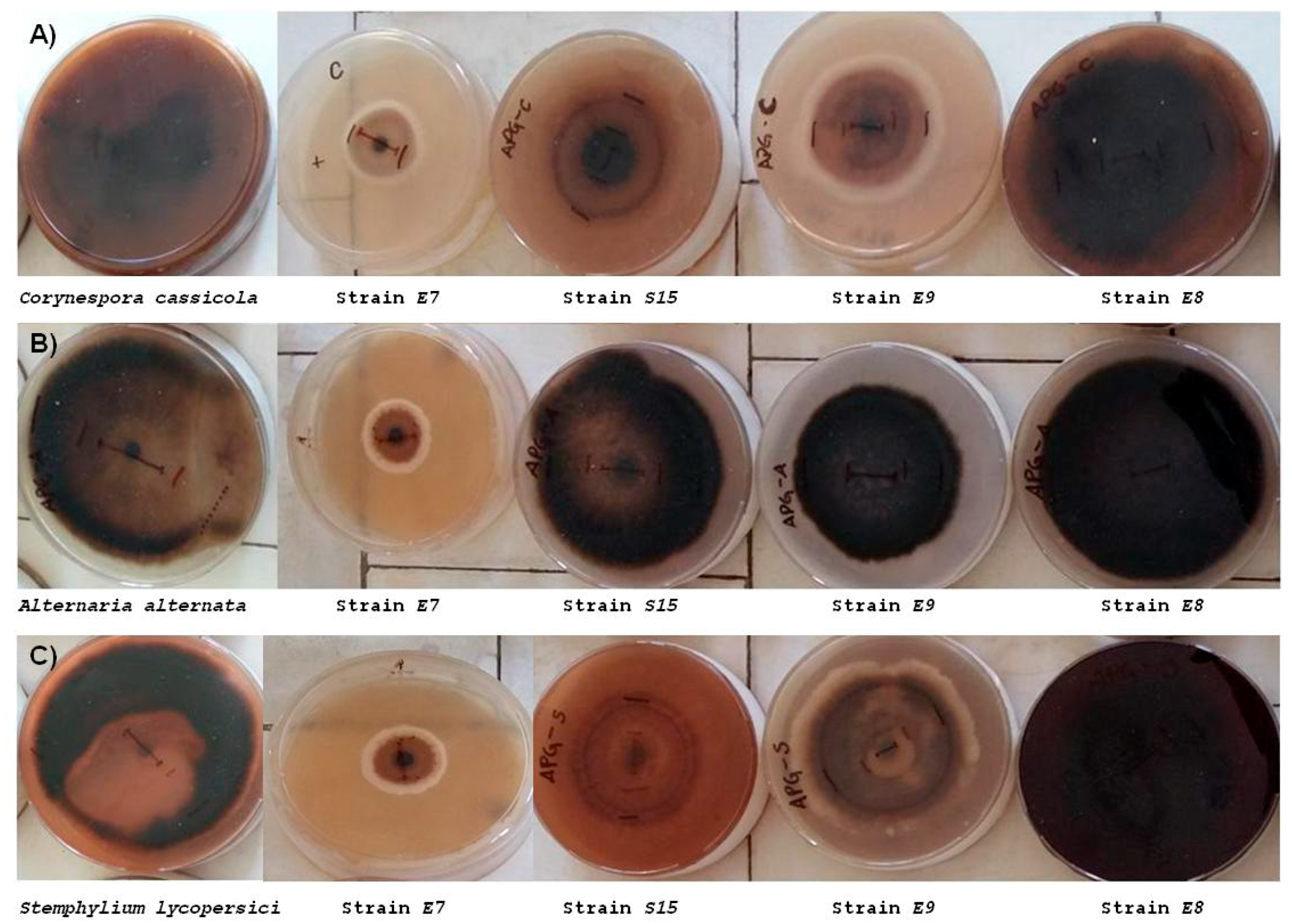
3.2.1. In Vivo Antagonism of Endophytic Bacteria towards Tomato Fungal Pathogens
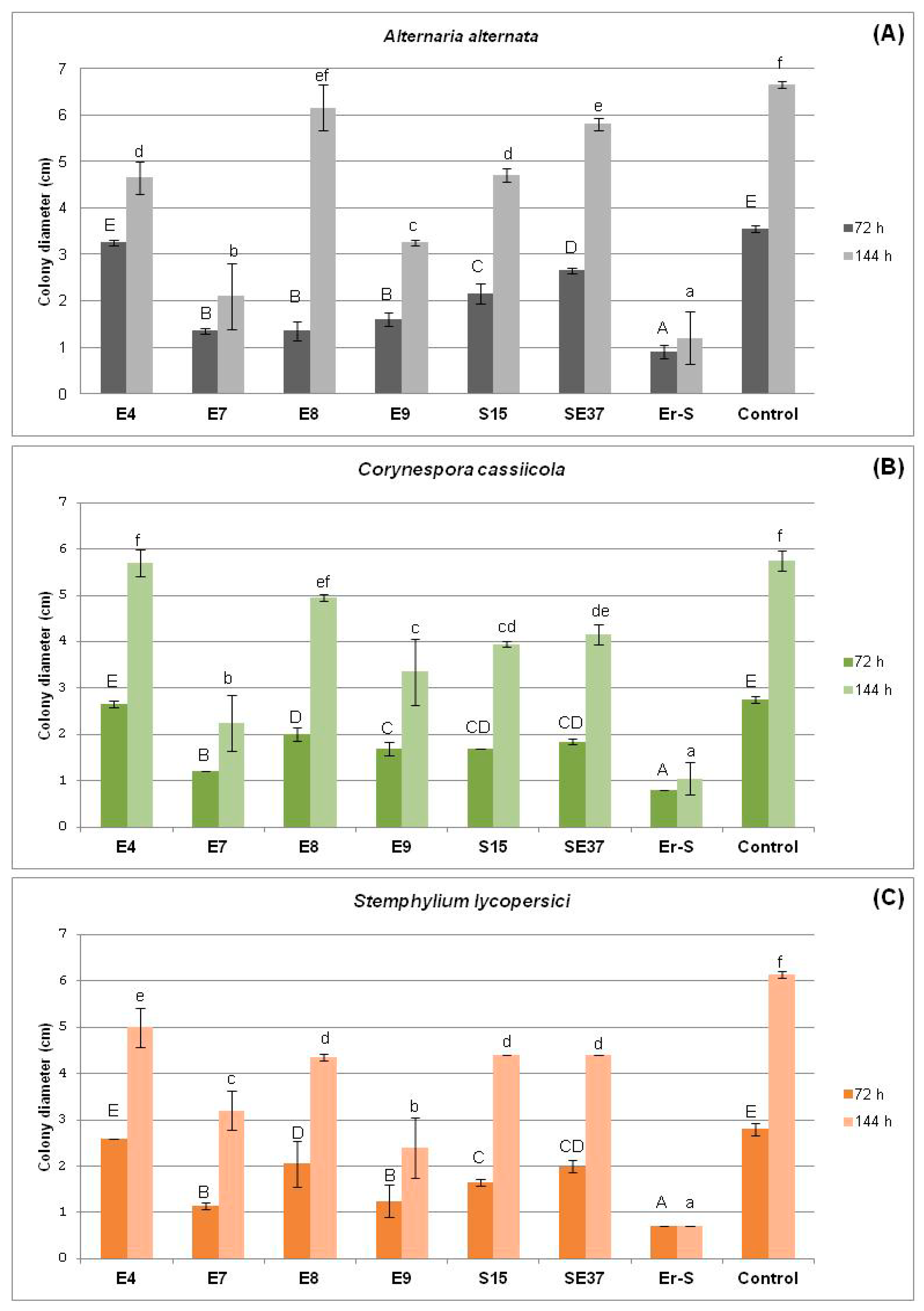
3.2.2. Effect of the Cell-Free Supernatant of Endophytic Bacteria against Fungal Pathogens
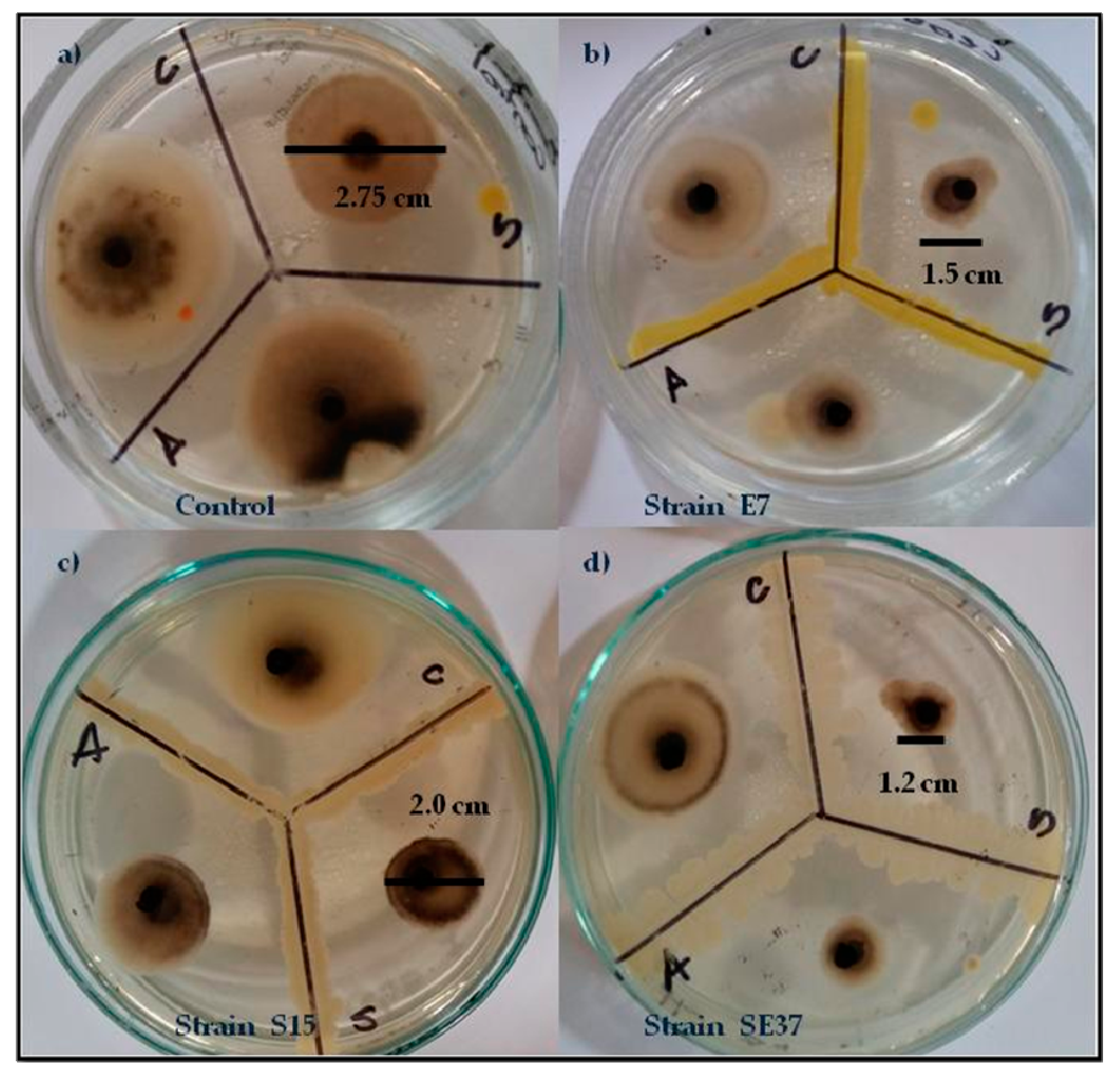
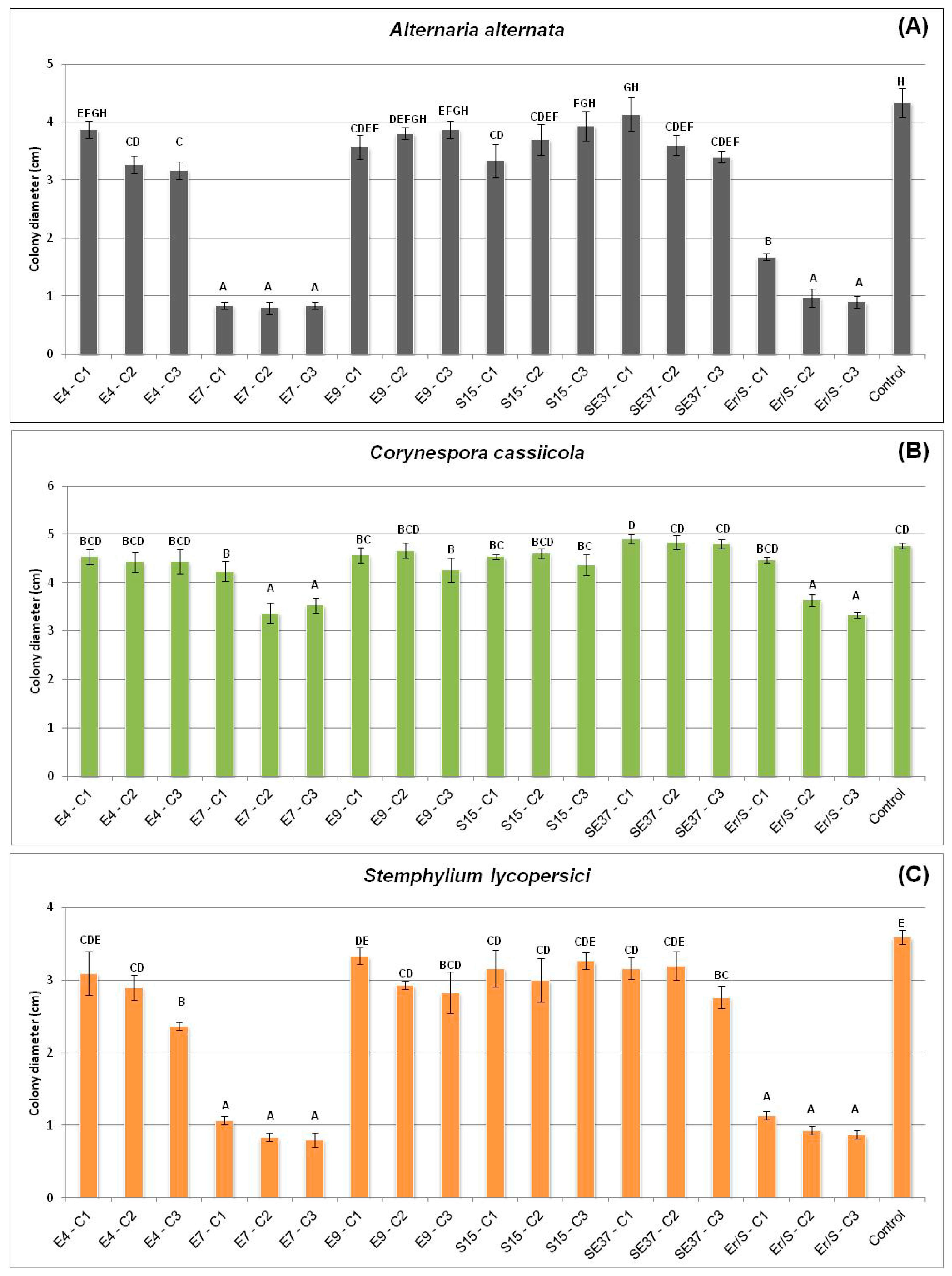
3.2.3. Effect of Volatiles from Endophytic Bacteria on Fungal Pathogen Growth
3.3. Plant Growth Promotion Assays
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
References
- Food and Agriculture Organization of the United Nations, FAOSTAT. Available online: http://faostat.fao.org (accessed on 1 June 2018).
- Hallmann, J.; Quadt-Hallmann, A.; Mahaffee, W.; Kloepper, J. Bacterial endophytes in agricultural crops. Can. J. Microbiol. 1997, 4, 895–914. [Google Scholar] [CrossRef]
- Hardoim, P.; Van Overbeek, L.; Berg, G.; Pirttilä, A.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The hidden world within plants: Ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol. Mol. Biol. Rev. 2015, 79, 293–320. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.; Saldierna-Guzmán, J.P.; Shay, J. Transmission of Bacterial Endophytes. Microorganisms 2017, 5, 70. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, R.; Pieterse, C.; Bakker, P. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, D.; Ansari, M.; Sahoo, R.; Tuteja, N. Biofertilizers function as key player in sustainable agriculture by improving soil fertility, plant tolerance and crop productivity. Microb. Cell Fact. 2014, 13, 66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vejan, P.; Abdullah, R.; Khadiran, T.; Ismail, S.; Nasrulhaq-Boyce, A. Role of plant growth promoting rhizobacteria in agricultural sustainability—A review. Molecules 2016, 21, 573. [Google Scholar] [CrossRef] [PubMed]
- Malfanova, N.; Lugtenberg, B.; Berg, G. Endophytic Bacteria with Plant Growth Promoting and Biocontrol Abilities. Ph.D. Thesis, Leiden University, Leiden, The Netherlands, 2013. [Google Scholar]
- Turner, T.; James, E.; Poole, P. The plant microbiome. Genome Biol. 2013, 14, 209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rosenblueth, M.; Martínez-Romero, E. Bacterial endophytes and their interactions with hosts. Mol. Plant-Microbe Interact. 2006, 19, 827–837. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, P.; van Overbeek, L.; van Elsas, J. Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol. 2008, 16, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, F.; Ahmad, I.; Khan, M. Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol. Res. 2008, 163, 173–181. [Google Scholar] [CrossRef] [PubMed]
- Compant, S.; Clément, C.; Sessitsch, A. Plant growth-promoting bacteria in the rhizo-and endosphere of plants: Their role, colonization, mechanisms involved and prospects for utilization. Soil Biol. Biochem. 2010, 42, 669–678. [Google Scholar] [CrossRef] [Green Version]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; van Themaat, E.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Ann. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef] [PubMed]
- Gaiero, J.; McCall, C.; Thompson, K.; Day, N.; Best, A.; Dunfield, K. Inside the root microbiome: Bacterial root endophytes and plant growth promotion. Am. J. Bot. 2013, 100, 1738–1750. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nelson, E. Microbial dynamics and interactions in the spermosphere. Annu. Rev. Phytopathol. 2004, 42, 271–309. [Google Scholar] [CrossRef] [PubMed]
- Ewald, P. Transmission modes and evolution of the parasitism-mutualism continuum. Ann. N. Y. Acad. Sci. 1987, 503, 295–306. [Google Scholar] [CrossRef] [PubMed]
- Rudgers, J.; Afkhami, M.; Rúa, M.; Davitt, A.; Hammer, S.; Huguet, V. A fungus among us: Broad patterns of endophyte distribution in the grasses. Ecology 2009, 90, 1531–1539. [Google Scholar] [CrossRef] [PubMed]
- Cankar, K.; Kraigher, H.; Ravnikar, M.; Rupnik, M. Bacterial endophytes from seeds of Norway spruce (Picea abies L. Karst). FEMS Microbiol. Lett. 2005, 244, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Sheng, J.; Chen, L.; Men, Y.; Gan, L.; Guo, S.; Shen, L. Bacterial community compositions of tomato (Lycopersicum esculentum Mill.) seeds and plant growth promoting activity of ACC deaminase producing Bacillus subtilis (HYT-12-1) on tomato seedlings. World. J. Microbiol. Biotech. 2014, 30, 835–845. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, W.; Munakata, N.; Horneck, G.; Melosh, H.; Setlow, P. Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments. Microbiol. Mol. Boil. Rev. 2000, 64, 548–572. [Google Scholar] [CrossRef]
- Truyens, S.; Weyens, N.; Cuypers, A.; Vangronsveld, J. Bacterial seed endophytes: Genera, vertical transmission and interaction with plants. Environ. Microbiol. Rep. 2015, 7, 40–50. [Google Scholar] [CrossRef]
- Tian, B.; Zhang, C.; Ye, Y.; Wen, J.; Wu, Y.; Wang, H.; Li, L.; Cai, S.; Cai, W.; Cheng, Z.; et al. Beneficial traits of bacterial endophytes belonging to the core communities of the tomato root microbiome. Agric. Ecosyst. Environ. 2017, 247, 149–156. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Rott, M.; Schlaeppi, K.; van Themaat, E.; Ahmadinejad, N.; Assenza, F.; Rauf, P.; Huettel, B.; Reinhardt, R.; Schmelzer, E.; et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 2012, 488, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Sessitsch, A.; Hardoim, P.; Döring, J.; Weilharter, A.; Krause, A.; Woyke, T.; Mitter, B.; Hauberg-Lotte, L.; Friedrich, F.; Rahalkar, M.; et al. Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. Mol. Plant-Microbe Interact. 2012, 25, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Müller, D.; Srinivas, G.; Garrido-Oter, R.; Potthoff, E.; Rott, M.; Dombrowski, N.; Münch, P.; Spaepen, S.; Remus-Emsermann, M.; et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 2015, 528, 364–369. [Google Scholar] [CrossRef] [PubMed]
- Romero, F.; Marina, M.; Pieckenstain, F. The communities of tomato (Solanum lycopersicum L.) leaf endophytic bacteria, analyzed by 16S-ribosomal RNA gene pyrosequencing. FEMS Microbiol. Lett. 2014, 351, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Weisburg, W.; Barns, S.; Pelletier, D.; Lane, D. 16S ribosomal DNA amplification for phylogenetic study. J. Bacterial. 1991, 173, 697–703. [Google Scholar] [CrossRef]
- Lane, D. 16S/23S rRNA sequencing. In Nucleic Acid Techniques in Bacterial Systematic; Stackebrandt, E., Goodfellow, M., Eds.; John Wiley and Sons: New York, NY, USA, 1991; pp. 115–175. [Google Scholar]
- Schloss, P.; Westcott, S.; Ryabin, T.; Hall, J.; Hartmann, M.; Hollister, E.; Lesniewski, R.; Oakley, B.; Parks, D.; Robinson, C.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2012, 41, D590–D596. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.; Bayles, D.; Looft, T.; Trachsel, J.; Bass, B.; Alt, D.; Bearson, S.; Nicholson, T.; Casey, T. Pipeline for amplifying and analyzing amplicons of the V1–V3 region of the 16S rRNA gene. BMC Res. Notes 2016, 9, 380. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.; Haas, B.; Clemente, J.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hill, M. Diversity and evenness: A unifying notation and its consequences. Ecology 1973, 54, 427–432. [Google Scholar] [CrossRef]
- Medina, R.; Gara, P.; Fernández-González, A.; Rosso, J.; Del Panno, M. Remediation of a soil chronically contaminated with hydrocarbons through persulfate oxidation and bioremediation. Sci. Total Environ. 2018, 618, 518–530. [Google Scholar] [CrossRef] [PubMed]
- Surette, M.; Sturz, A.; Lada, R.; Nowak, J. Bacterial endophytes in processing carrots (Daucus carota L. var. sativus): Their localization, population density, biodiversity and their effects on plant growth. Plant Soil 2003, 253, 381–390. [Google Scholar] [CrossRef]
- López, S.; Balatti, P. Closely related strains of Bradyrhizobium contained in commercial inoculates of soybean are identified by a set of PCR reactions. Genet. Eng. Biotech. J. 2012, 2011, 1–8. [Google Scholar]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baysal, Ö.; Lai, D.; Xu, H.; Siragusa, M.; Çalışkan, M.; Carimi, F.; Teixeira, J.; Tör, M. A proteomic approach provides new insights into the control of soil-borne plant pathogens by Bacillus species. PLoS ONE 2013, 8, e53182. [Google Scholar] [CrossRef] [PubMed]
- Medina, R.; López, S.; Franco, M.; Rollan, C.; Ronco, B.; Saparrat, M.; De Wit, P.; Balatti, P. A Survey on Occurrence of Cladosporium fulvum Identifies Race 0 and Race 2 in Tomato-Growing Areas of Argentina. Plant Dis. 2015, 99, 1732–1737. [Google Scholar] [CrossRef]
- Alexander, D.; Zuberer, D. Use of chrome azurol S reagents to evaluate siderophore production by rhizosphere bacteria. Biol. Fert. Soils 1991, 12, 39–45. [Google Scholar] [CrossRef]
- Castagno, L.; Estrella, M.; Sannazzaro, A.; Grassano, A.; Ruiz, O. Phosphate-solubilization mechanism and in vitro plant growth promotion activity mediated by Pantoea eucalypti isolated from Lotus tenuis rhizosphere in the Salado River Basin (Argentina). J. Appl. Microbiol. 2011, 110, 1151–1165. [Google Scholar] [CrossRef] [PubMed]
- Bric, J.; Bostock, R.; Silverstone, S. Rapid in situ assay for indoleacetic acid production by bacteria immobilized on a nitrocellulose membrane. Appl. Environ. Microbiol. 1991, 57, 535–538. [Google Scholar] [PubMed]
- Sorroche, F.; Spesia, M.; Zorreguieta, A.; Giordano, W. A positive correlation between bacterial autoaggregation and biofilm formation in native Sinorhizobium meliloti isolates from Argentina. Appl. Environ. Microbiol. 2012, 78, 4092–4101. [Google Scholar] [CrossRef] [PubMed]
- Kloepper, J.; Beauchamp, C. A review of issues related to measuring colonization of plant roots by bacteria. Can. J. Microbiol. 1992, 38, 1219–1232. [Google Scholar] [CrossRef]
- Guan, B.; Zhou, D.; Zhang, H.; Tian, Y.; Japhet, W.; Wang, P. Germination responses of Medicago ruthenica seeds to salinity, alkalinity, and temperature. J. Arid Environ. 2009, 73, 135–138. [Google Scholar] [CrossRef]
- Bacilio-Jiménez, M.; Aguilar-Flores, S.; Ventura-Zapata, E.; Pérez-Campos, E.; Bouquelet, S.; Zenteno, E. Chemical characterization of root exudates from rice (Oryza sativa) and their effects on the chemotactic response of endophytic bacteria. Plant Soil 2003, 249, 271–277. [Google Scholar] [CrossRef]
- Cottyn, B.; Regalado, E.; Lanoot, B.; De Cleene, M.; Mew, T.; Swings, J. Bacterial populations associated with rice seed in the tropical environment. Phytopathology 2001, 91, 282–292. [Google Scholar] [CrossRef] [PubMed]
- Mano, H.; Tanaka, F.; Watanabe, A.; Kaga, H.; Okunishi, S.; Morisaki, H. Culturable surface and endophytic bacterial flora of the maturing seeds of rice plants (Oryza sativa) cultivated in a paddy field. Microbes Environ. 2006, 21, 86–100. [Google Scholar] [CrossRef]
- Compant, S.; Mitter, B.; Colli-Mull, J.; Gangl, H.; Sessitsch, A. Endophytes of grapevine flowers, berries, and seeds: Identification of cultivable bacteria, comparison with other plant parts, and visualization of niches of colonization. Microb. Ecol. 2011, 62, 188–197. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, P.; Hardoim, C.; Van Overbeek, L.; Van Elsas, J. Dynamics of seed-borne rice endophytes on early plant growth stages. PLoS ONE 2012, 7, e30438. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Carvalhais, L.; Crawford, M.; Singh, E.; Dennis, P.; Pieterse, C.; Schenk, P. Inner plant values: Diversity, colonization and benefits from endophytic bacteria. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Simon, H.; Smith, K.; Dodsworth, J.; Guenthner, B.; Handelsman, J.; Goodman, R. Influence of tomato genotype on growth of inoculated and indigenous bacteria in the spermosphere. Appl. Environ. Microbiol. 2001, 67, 514–520. [Google Scholar] [CrossRef] [PubMed]
- Adams, P.; Kloepper, J. Effect of host genotype on indigenous bacterial endophytes of cotton (Gossypium hirsutum L.). Plant Soil 2002, 240, 181–189. [Google Scholar] [CrossRef]
- Franco-Correa, M.; Chavarro-Anzola, V. Actinobacteria as Plant Growth-Promoting Rhizobacteria. In Actinobacteria-Basics and Biotechnological Applications; InTech: Rijeka, Croatia, 2016. [Google Scholar] [Green Version]
- Sangthong, C.; Setkit, K.; Prapagdee, B. Improvement of cadmium phytoremediation after soil inoculation with a cadmium-resistant Micrococcus sp. Environ. Sci. Pollut. Res. 2016, 23, 756–764. [Google Scholar] [CrossRef] [PubMed]
- Prapagdee, B.; Chanprasert, M.; Mongkolsuk, S. Bioaugmentation with cadmium-resistant plant growth-promoting rhizobacteria to assist cadmium phytoextraction by Helianthus annuus. Chemosphere 2013, 92, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Vílchez, J.I.; Niehaus, K.; Dowling, D.; González-López, J.; Manzanera, M. Protection of pepper plants from drought by Microbacterium sp. 3J1 by modulation of the plant’s glutamine and α-ketoglutarate content: A comparative metabolomics approach. Front. Microbiol. 2018, 9, 284. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, K.; Stauber, T.; Ewald, D. Paenibacillus—A predominant endophytic bacterium colonising tissue cultures of woody plants. Plant Cell Tissue Organ Cult. 2008, 93, 347–351. [Google Scholar] [CrossRef]
- Cheba, B.A. Diversity, Phyto Beneficial Activities and Agrobiotechnology of Plant Growth Promoting Bacillus and Paenibacillus. In Proceedings of the ICBB 2016: 18th International Conference on Biotechnology and Bioengineering, Stockholm, Sweden, 11–12 July 2016. [Google Scholar]
- Grady, E.; MacDonald, J.; Liu, L.; Richman, A.; Yuan, Z. Current knowledge and perspectives of Paenibacillus: A review. Microb. Cell Fact. 2016, 15, 203. [Google Scholar] [CrossRef] [PubMed]
- Timmusk, S.; Grantcharova, N.; Wagner, E. Paenibacillus polymyxa invades plant roots and forms biofilms. Appl. Environ. Microb. 2005, 71, 7292–7300. [Google Scholar] [CrossRef] [PubMed]
| Sample | Total Sequences | 0H | 1H | 2H |
|---|---|---|---|---|
| Elpida seed | 11,495 | 35,147 | 3.6 | 1.37 |
| Silverio seed | 12,167 | 62,867 | 3.7 | 1.39 |
| Phyla | Class | Order Elpida Seed | Genus Elpida Seed | Order Silverio Seed | Genus Silverio Seed |
|---|---|---|---|---|---|
| Actinobacteria | Actinobacteria | Actinomycetales 14.3% | Clavibacter (61%) Corynebacterium (20%) Micrococcus (11%) Curtobacterium (6%) Microbacterium (2%) | Actinomycetales 27.3% | Clavibacter (81%) Corynebacterium (6%) Micrococcus (3%) Curtobacterium (7%) Microbacterium (3%) |
| Bacteroidetes | Flavobacteria | Flavobacteriales 0.7% | Flavobacterium (30%) | Flavobacteriales 1.3% | Flavobacterium (54%) |
| Sphingobacteria | Sphingobacteriales 0.5% | Sphingobacterium (100%) | Sphingobacteriales 1.7% | Sphingobacterium (100%) | |
| Firmicutes | Bacilli | Bacillales 63.3% | Paenibacillus (92%) Staphylococcus (8%) | Bacillales 2.7% | Paenibacillus (26%) Staphylococcus (74%) |
| Lactobacillales 0.5% | Lactobacillales 0.7% | ||||
| Proteobacteria | Alpha | Rhizobiales 2.7% | Shinella (70%) Sphingobium (15%) Rhizobium, Ensifer, Sinorhizobium (15%) | Rhizobiales 16.0% | Shinella (70%) Sphingobium (15%) Rhizobium, Ensifer, Sinorhizobium (15%) |
| Sphingomonadales 0.7% | Sphingomonadales 3.3% | ||||
| Beta | Burkholderiales 0.5% | Achromobacter (20%) Acidovorax (80%) | Burkholderiales 5.0% | Achromobacter (29%) Acidovorax (71%) | |
| Gamma | Enterobacteriales 0.6% | Pantoea, Pectobacterium, Serratia (3%) | Enterobacteriales 4.0% | Pantoea, Pectobacterium, Serratia (10%) | |
| Pseudomonadales 14.6% | Pseudomonas (75%) Moraxella (14%) Acinetobacter (8%) | Pseudomonadales 37.3% | Pseudomonas (89%) Moraxella (0.5%) Acinetobacter (0.5%) |
| Isolate (Origin) | Closest Match in NCBI Database (Accession Number) | Identity (%) |
|---|---|---|
| E4 (seed Elpida) | Micrococcus sp. (MG963203) | 99 |
| E6 (seed Elpida) | Bacillus sp. (MG963204) | 92 |
| E7 (seed Elpida) | Bacillus sp. (MG963205) | 99 |
| E8 (seed Elpida) | P. polymyxa (MG963206) | 99 |
| E9 (seed Elpida) | Bacillus sp. (MG963207) | 98 |
| S15 (seed Silverio) | Bacillus sp. (MG963209) | 99 |
| S19 (seed Silverio) | Bacillus sp. (MG963210) | 99 |
| S20 (seed Silverio) | Sphingomonas sp. (MG963211) | 96 |
| S21 (seed Silverio) | Brevundimonas sp. (MG963212) | 99 |
| S26 (seed Silverio) | Paenibacillus sp. (MG963213) | 91 |
| S27 (seed Silverio) | Jeotgalibacillus sp. (MG963214) | 99 |
| SE28 (seedling Elpida) | Acinetobacter sp. (MG963215) | 98 |
| SE31 (seedling Elpida) | Microbacterium sp. (MG963216) | 99 |
| SE33 (seedling Elpida) | Paenibacillus sp. (MG963217) | 99 |
| SE34 (seedling Elpida) | Bacillus sp. (MG963218) | 99 |
| SE35 (seedling Elpida) | Bacillus sp. (MG963219) | 99 |
| SE36 (seedling Elpida) | Psychrobacillus sp. (MG963220) | 97 |
| SE37 (seedling Elpida) | Bacillus sp. (MG963221) | 98 |
| SS38 (seedling Silverio) | Bacillus sp. (MG963222) | 99 |
| SS39 (seedling Silverio) | Bacillus sp. (MG963223) | 99 |
| SS41 (seedling Silverio) | Bacillus sp. (MG963224) | 96 |
| Er-S | B. subtilis (MG963208) | 99 |
| Strain | A. alternata | C. cassiicola | S. lycopersici |
|---|---|---|---|
| SE37 | 1.65 ± 0.289 a | 2.95 ± 0.06 bc | 1.4 ± 0.231 a |
| E4 | 2.05 ± 0.289 ab | 2.55 ± 0.289 a | 1.4 ± 0.231 a |
| E8 | 2 ± 0.115 ab | 2.8 ± 0.115 ab | 1.55 ± 0.06 ab |
| E7 | 2.35 ± 0.173 bc | 2.85 ± 0.06 ab | 1.85 ± 0.06 bc |
| Er-S | 2.45 ± 0.404 bc | 3 ± 0.08 bc | 1.9 ± 0.115 bc |
| S15 | 2.55 ± 0.173 bc | 3.25 ± 0.289 c | 2 ± 0.115 cd |
| E9 | 2.75 ± 0.289 cd | 2.7 ± 0.115 ab | 2.25 ± 0.06 cde |
| E6 | 2.9 ± 0.115 cde | 3.6 ± 0.115 d | 2.35 ± 0.404 def |
| S19 | 2.9 ± 0.08 cde | 3.7 ± 0.115 d | 2.6 ± 0.115 efg |
| SE31 | 3.15 ± 0.289 def | 3.7 ± 0.115 d | 2.6 ± 0.115 efg |
| Control | 3.6 ± 0.08 f | 4.05 ± 0.06 e | 2.75 ± 0.06 fg |
| SE33 | 3.35 ± 0.06 ef | 4.05 ± 0.06 e | 2.9 ± 0.115 g |
| SE36 | 3.25 ± 0.289 def | 4.3 ± 0.08 e | 2.95 ± 0.06 g |
| Isolate | Fresh Weight Root (g) | Root Volume (mL) | Dry Weight Root (g) | Dry Air Weight (g) |
|---|---|---|---|---|
| Strain E4 | 5.09 ± 0.7 a | 4.57 ± 0.5 ab | 0.31 ± 0.1 bc | 0.59 ± 0.1 cde |
| Strain E6 | 5.43 ± 0.8 a | 5.44 ± 0.6 a | 0.38 ± 0.1 ab | 0.55 ± 0.1 def |
| Strain E8 | 5.10 ± 0.6 a | 5.06 ± 0.8 a | 0.35 ± 0.1 ab | 0.56 ± 0.1 de |
| Strain S15 | 5.59 ± 0.3 a | 5.56 ± 0.4 a | 0.44 ± 0.0 a | 0.57 ± 0.1 cde |
| Strain S21 | 3.64 ± 0.3 bc | 3.63 ± 0.5 bcd | 0.25 ± 0.0 cd | 0.41 ± 0.1 ef |
| Strain SE28 | 2.36 ± 0.4 d | 2.07 ± 0.6 e | 0.19 ± 0.0 d | 0.52 ± 0.1 ef |
| Strain SE31 | 3.93 ± 0.7 b | 3.44 ± 0.5 cd | 0.38 ± 0.1 ab | 0.76 ± 0.1 abc |
| Strain SE36 | 3.29 ± 0.2 bcd | 3.38 ± 0.5 cd | 0.25 ± 0.0 cd | 0.74 ± 0.1 bcd |
| Strain SS38 | 3.26 ± 0.5 bcd | 3.50 ± 0.5 cd | 0.28 ± 0.1 bcd | 0.86 ± 0.1 ab |
| Strain SS39 | 3.71 ± 0.7 bc | 3.50 ± 0.7 cd | 0.26 ± 0.1 cd | 0.94 ± 0.2 a |
| P. fluorescens | 4.01 ± 0.7 b | 4.00 ± 0.9 bc | 0.29 ± 0.1 bcd | 0.84 ± 0.2 ab |
| Control | 2.96 ± 0.3 cd | 2.84 ± 0.2 de | 0.19 ± 0.0 d | 0.36 ± 0.0 f |
| Isolate 1 | Source | IAA Production | Siderophore Production | Phosphate Solubilization |
|---|---|---|---|---|
| E7 | Seeds Elpida | + | + | + |
| E8 | + | |||
| S15 | Seeds Silverio | + | ||
| S19 | + | |||
| S27 | + | |||
| SE28 | Seedling Elpida | + | + | |
| SE35 | + | |||
| SE36 | + | |||
| SE37 | + | |||
| SS38 | Seedling Silverio | + |
| Isolate | Biofilm (OD560nm/OD630nm) | Autoaggregation (%) |
|---|---|---|
| E4 | 0.38 ± 0.02 | 89.41 ± 1.08 |
| E6 | 13.58 ± 0.62 | 0 |
| E8 | 0.44 ± 0.23 | 34.16 ± 2.33 |
| S15 | 0.86 ± 0.52 | 38.14 ± 1.55 |
| SE31 | 5.00 ± 0.26 | 0 |
| PF | 2.51 ± 0.26 | 13.54 ± 0.55 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
López, S.M.Y.; Pastorino, G.N.; Franco, M.E.E.; Medina, R.; Lucentini, C.G.; Saparrat, M.C.N.; Balatti, P.A. Microbial Endophytes that Live within the Seeds of Two Tomato Hybrids Cultivated in Argentina. Agronomy 2018, 8, 136. https://doi.org/10.3390/agronomy8080136
López SMY, Pastorino GN, Franco MEE, Medina R, Lucentini CG, Saparrat MCN, Balatti PA. Microbial Endophytes that Live within the Seeds of Two Tomato Hybrids Cultivated in Argentina. Agronomy. 2018; 8(8):136. https://doi.org/10.3390/agronomy8080136
Chicago/Turabian StyleLópez, Silvina Marianela Yanil, Graciela Noemi Pastorino, Mario Emilio Ernesto Franco, Rocio Medina, César Gustavo Lucentini, Mario Carlos Nazareno Saparrat, and Pedro Alberto Balatti. 2018. "Microbial Endophytes that Live within the Seeds of Two Tomato Hybrids Cultivated in Argentina" Agronomy 8, no. 8: 136. https://doi.org/10.3390/agronomy8080136





