Molecular Markers Improve Breeding Efficiency in Apomictic Poa Pratensis L.
Abstract
:1. Introduction
2. Materials and Methods
3. Results and Discussion
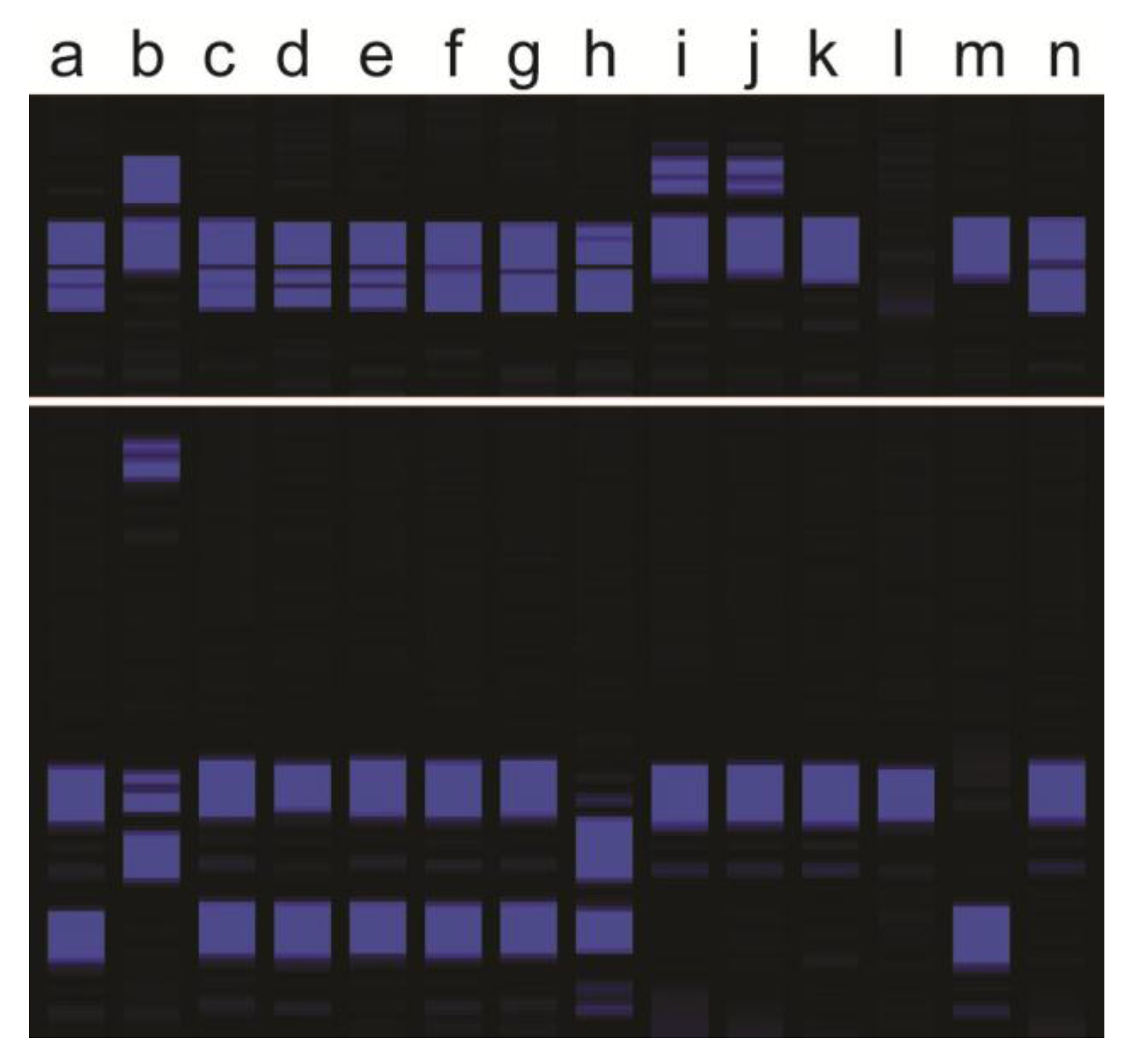
3.1. Hybrid Detection
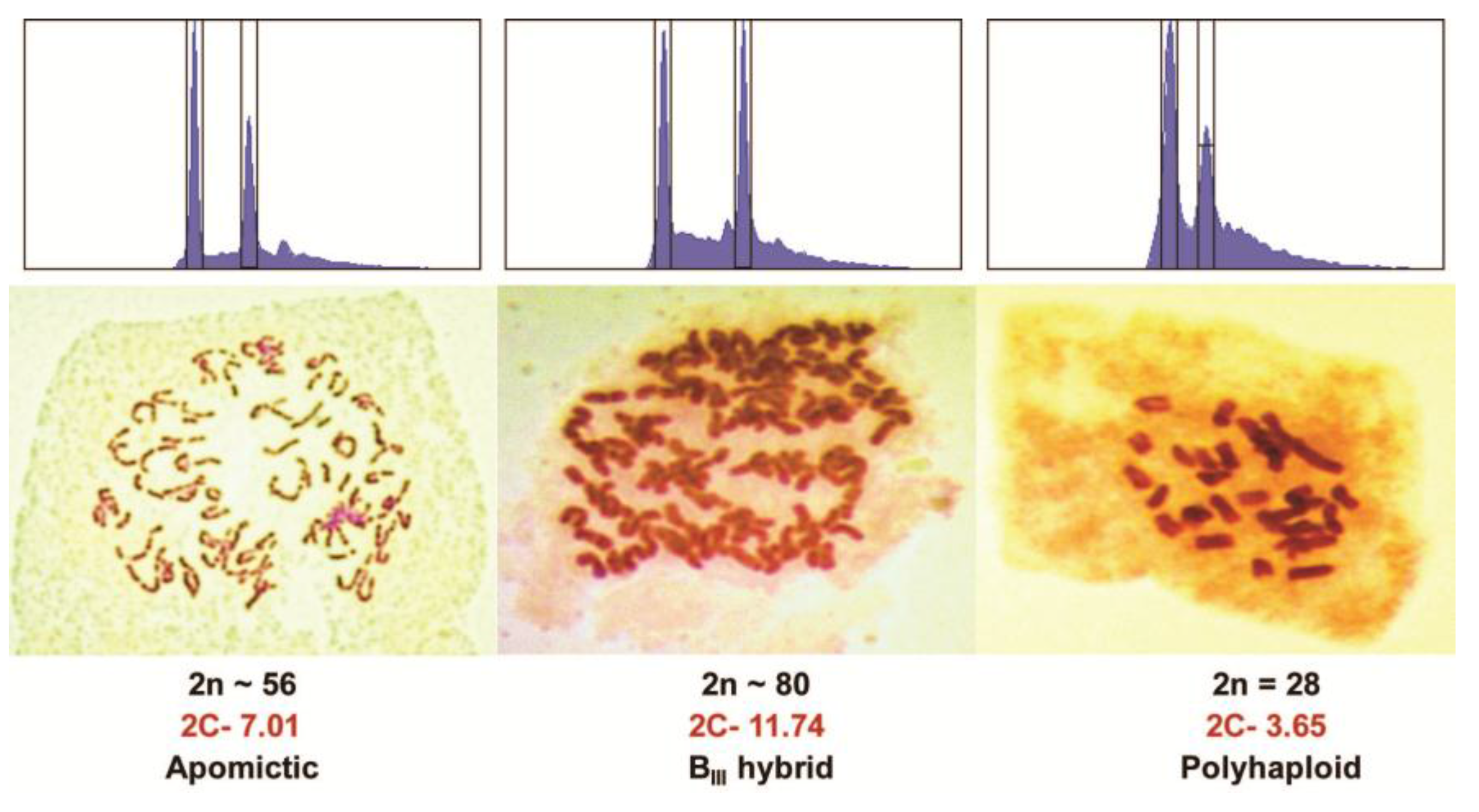
3.2. Apomictic Progeny Detection
3.3. Cultivar Differentiation
3.4. Emerging Efforts
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gillespie, L.J.; Soreng, R.J. A phylogenetic analysis of the bluegrass genus Poa based on cpDNA restriction site data. Syst. Bot. 2005, 30, 84–105. [Google Scholar] [CrossRef]
- Evans, R.A.; Young, J.A.; Roundy, B.A. Seedbed Requirements for Germination of Sandberg Bluegrass. Agron. J. 1977, 69, 817–820. [Google Scholar] [CrossRef]
- Cabi, E.; Soreng, R.J.; Gillespie, L.; Amiri, N. Poa densa (Poaceae), an overlooked Turkish steppe grass, and the evolution of bulbs in Poa. Willdenowia 2016, 46, 201–211. [Google Scholar] [CrossRef]
- Huff, D.R. Annual Bluegrass. In Turfgrass Biology, Genetics, and Breeding; Casler, M.D., Duncan, R.R., Eds.; John Wiley and Sons, Inc.: Hoboken, NJ, USA, 2003; pp. 39–52. [Google Scholar]
- Hurley, R. Rough Bluegrass. In Turfgrass Biology, Genetics, and Breeding; Casler, M.D., Duncan, R.R., Eds.; John Wiley and Sons, Inc.: Hoboken, NJ, USA, 2003; pp. 67–74. [Google Scholar]
- Read, J.C.; Anderson, S.J. Texas Bluegrass. In Turfgrass Biology, Genetics, and Breeding; Casler, M.D., Duncan, R.R., Eds.; John Wiley and Sons, Inc.: Hoboken, NJ, USA, 2003; pp. 61–66. [Google Scholar]
- Speckmann, G.J.; Van Dijk, G.E. Chromosome number and plant morphology in some ecotypes of Poa pratensis L. Euphytica 1972, 21, 171–180. [Google Scholar] [CrossRef]
- Eaton, T.; Curley, J.C.; Williamson, R.; Jung, G. Determination of the Level of Variation in Polyploidy among Kentucky Bluegrass Cultivars by Means of Flow Cytometry. Crop Sci. 2004, 44, 2168–2174. [Google Scholar] [CrossRef]
- Huff, D.R.; Bara, J.M. Determining Genetic Origins of Aberrant Progeny from Facultative Apomictic Kentucky Bluegrass Using a Combination of Flow-Cytometry and Silver-Stained Rapd Markers. Theor. Appl. Genet. 1993, 87, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Barcaccia, G.; Mazzucato, A.; Belardinelli, A.; Pezzotti, M.; Lucretti, S.; Falcinelli, M. Inheritance of parental genomes in progenies of Poa pratensis L. from sexual and apomictic genotypes as assessed by RAPD markers and flow cytometry. Theor. Appl. Genet. 1997, 95, 516–524. [Google Scholar] [CrossRef]
- Murovec, J.; Kastelec, D.; Vilhar, B.; Cop, J.; Bohanec, B. High variability of nuclear DNA content in cultivars and natural populations of Poa pratensis L. in relation to morphological characters. Acta Biol. Crac. 2009, 5, 45–52. [Google Scholar]
- Barcaccia, G.; Albertini, E. Apomixis in plant reproduction: A novel perspective on an old dilemma. Plant Reprod. 2013, 26, 159–179. [Google Scholar] [CrossRef] [PubMed]
- Huff, D.R. Kentucky bluegrass. In Turfgrass Biology, Genetics, and Breeding; Casler, M.D., Duncan, R.R., Eds.; John Wiley and Sons, Inc.: Hoboken, NJ, USA, 2003; pp. 27–38. [Google Scholar]
- Meyer, W.A.; Rose, B.L.; Johnson-Cicalese, J.M.; Funk, C.R. Registration of Midnight Kentucky Bluegrass. Crop Sci. 1984, 24, 822–823. [Google Scholar] [CrossRef]
- Kindiger, B. Frequency of androgenesis in Poa arachnifera × P. ligularis and P. poiformis hybridizations. Grassl. Sci. 2015, 61, 177–180. [Google Scholar] [CrossRef]
- Lickfeldt, D.W.; Voigt, T.B.; Hamblin, A.M. Composition and characteristics of blended Kentucky bluegrass stands. Hortscience 2002, 37, 1124–1126. [Google Scholar]
- Joshi, A.; Bushman, B.S.; Pickett, B.; Robbins, M.D.; Staub, J.E.; Johnson, P.G. Phylogenetic relationships among low-ploidy species of Poa using chloroplast sequences. Genome 2016, 60, 384–392. [Google Scholar] [CrossRef] [PubMed]
- Bushman, B.S.; Warnke, S.E.; Amundsen, K.L.; Combs, K.M.; Johnson, P.G. Molecular Markers Highlight Variation within and among Kentucky Bluegrass Varieties and Accessions. Crop Sci. 2013, 53, 2245–2254. [Google Scholar]
- Benham, J.J. Genographer Version 1.6.0; Montana State University: Bozeman, MT, USA, 2001. [Google Scholar]
- Warnke, S.E.; Thammina, C.S.; Amundsen, K.; Miljanic, P.; Hershman, H. High-Resolution Melt Analysis of Simple Sequence Repeats for Bentgrass Species Differentiation. Int. Turfgrass Soc. Res. J. 2017, 13, 466–470. [Google Scholar] [CrossRef]
- Griffin, P.C.; Robin, C.; Hoffmann, A.A. A next-generation sequencing method for overcoming the multiple gene copy problem in polyploid phylogenetics, applied to Poa grasses. BMC Biol. 2011, 9, 19. [Google Scholar] [CrossRef] [PubMed]
- Niemann, J.; Wojciechowski, A.; Janowicz, J. Identification of apomixis in the Kentucky bluegrass (Poa pratensis L.) using auxin test. Acta Soc. Bot. Pol. 2012, 81, 217–221. [Google Scholar] [CrossRef]
- Wieners, R.R.; Fei, S.-Z.; Johnson, R.C. Characterization of a USDA Kentucky Bluegrass (Poa pratensis L.) Core Collection for Reproductive Mode and DNA Content by Flow Cytometry. Genet. Resour. Crop Evol. 2006, 53, 1531–1541. [Google Scholar] [CrossRef]
- Matzk, F.; Meister, A.; Schubert, I. An efficient screen for reproductive pathways using mature seeds of monocots and dicots. Plant J. 2000, 21, 97–108. [Google Scholar] [CrossRef] [PubMed]
- Kelley, A.M.; Johnson, P.G.; Waldron, B.S.; Peel, M.D. A survey of apomixis and ploidy levels among Poa L. (Poaceae) using flow cytometry. Crop Sci. 2009, 45, 1395–1402. [Google Scholar] [CrossRef]
- Lickfeldt, D.W.; Voigt, T.B.; Hamblin, A.M. Cultivar composition and spatial patterns in Kentucky bluegrass blends. Crop Sci. 2002, 42, 842–847. [Google Scholar] [CrossRef]
- Huff, D.R. Characterization of Kentucky bluegrass cultivars using RAPD makers. Int. Turfgrass Soc. Res. J. 2001, 9, 169–175. [Google Scholar]
- Curley, J.; Jung, G. RAPD-based genetic relationships in Kentucky bluegrass: Comparison of cultivars, interspecific hybrids, and plant introductions. Crop Sci. 2004, 44, 1299–1306. [Google Scholar] [CrossRef]
- Honig, J.A.; Bonos, S.A.; Meyer, W.A. Isolation and Characterization of 88 Polymorphic Microsatellite Markers in Kentucky Bluegrass (Poa pratensis L.). HortScience 2010, 45, 1759–1763. [Google Scholar]
- Raggi, L.; Bitocchi, E.; Russi, L.; Marconi, G.; Sharbel, T.F.; Veronesi, F.; Albertini, E. Understanding Genetic Diversity and Population Structure of a Poa pratensis Worldwide Collection through Morphological, Nuclear and Chloroplast Diversity Analysis. PLoS ONE 2015, 10, e0124709. [Google Scholar] [CrossRef] [PubMed]
- Honig, J.A.; Averello, V.; Bonos, S.A.; Meyer, W.A. Classification of Kentucky Bluegrass (Poa pratensis L.) Cultivars and Accessions Based on Microsatellite (Simple Sequence Repeat) Markers. HortScience 2012, 47, 1356–1366. [Google Scholar]
| Progeny Type | Maternal Gamete 1 | Paternal Gamete | Fusion Method | Embryo Ploidy 2 | Endosperm Ploidy 3 |
|---|---|---|---|---|---|
| Apomict | unreduced | -- | Parthenogenesis | 2C | 5C (6C) |
| Polyhaploid | reduced | -- | Parthenogenesis | C | 3C (4C) |
| BII hybrid | reduced | reduced | Fertilization | 2C | 3C |
| BIII hybrid (M) 4 | unreduced | reduced | Fertilization | 3C | 5C |
| BIII hybrid (P) | reduced | unreduced | Fertilization | 3C | 4C |
| BIV hybrid | unreduced | unreduced | Fertilization | 4C | 6C |
| Progeny Type | NorthStar × PI371768 | PI371768 × NorthStar | Washington × PI440603 | PI440603 × Washington | PI499557 × PI440603 | PI440603 × PI499557 |
|---|---|---|---|---|---|---|
| Apomict | 78 | 91 | 67 | 88 | 83 | 86 |
| Polyhaploid | 3 | 0 | 10 | 3 | 1 | 2 |
| BII Hybrid | 1 | 0 | 0 | 1 | 0 | 0 |
| BIII Hybrid | 1 | 0 | 0 | 3 | 3 | 0 |
| Total Tested | 83 | 91 | 77 | 95 | 87 | 88 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bushman, B.S.; Joshi, A.; Johnson, P.G. Molecular Markers Improve Breeding Efficiency in Apomictic Poa Pratensis L. Agronomy 2018, 8, 17. https://doi.org/10.3390/agronomy8020017
Bushman BS, Joshi A, Johnson PG. Molecular Markers Improve Breeding Efficiency in Apomictic Poa Pratensis L. Agronomy. 2018; 8(2):17. https://doi.org/10.3390/agronomy8020017
Chicago/Turabian StyleBushman, B. Shaun, Alpana Joshi, and Paul G. Johnson. 2018. "Molecular Markers Improve Breeding Efficiency in Apomictic Poa Pratensis L." Agronomy 8, no. 2: 17. https://doi.org/10.3390/agronomy8020017





