2. Chondroitin Sulfate Proteoglycans in Neuronal Circuit Formation
Natural covalently bound hybrid molecules of core protein and linear polysaccharide chains, glycosaminoglycans, are proteoglycans [
2]. Depending on differences in the glycosaminoglycan chains, the molecules are classified into heparan sulfate proteoglycans (HSGP) with heparan sulfate (HS), chondroitin sulfate proteoglycans (CSPG) with chondroitin sulfate (CS), or keratan sulfate proteoglycans (KSPG) with keratan sulfate (KS). Each is involved in essential processes of nervous system development [
3,
4,
5,
6]. By interacting with various growth factors through their HS chains, HSPGs are involved in differentiation and circuit formation. It has been found that the glycosaminoglycan chains of CSPGs (CS) interact with other proteins such as growth factors to regulate their function indirectly; however, more recent findings indicate that CS is recognized directly through its receptors, exerting its effects as guidance cues [
7,
8,
9,
10]. Here, we review novel methods to examine CS structure-function relationship and its role in neuronal circuit formation.
Considering the structural components of CSPG, it is understandable that the function of CSPG is derived from its core protein and/or CS chains. The core proteins, depending on their structure, participate in forming extracellular matrices through interactions with other extracellular matrix molecules. The CS chains consist of repeating disaccharide units of glucuronic acid (GlcA) and N-acetylgalactosamine (GalNAc), forming straight chains of 20–200 units, with sulfate groups at various positions on the sugar chains. In many cases, effects of CS inhibit axonal outgrowth [
11,
12,
13,
14,
15,
16,
17,
18,
19,
20,
21]. For example, if the central nervous system is damaged in higher vertebrates, CS is accumulated in glial scar tissue, inhibiting axonal growth to prevent the axons from rewiring their pathway. Thus, CS is now becoming a major therapeutic target since animal experimental models of spinal cord injury showed that rewiring of the neuronal circuits is expected by clearance of pathologically accumulated CS [
22,
23,
24,
25]. In contrast, it has also been reported that CS exerts various effects on different neurons. For example, CS promotes neurites’ outgrowth of hippocampal neurons [
26,
27,
28,
29] and exerts trophic effects on rat retinal ganglion cells [
30]. Recently, it was reported that CS is involved in ocular dominance plasticity in the primary visual cortex [
31,
32,
33,
34]. In sum, the literature suggests that CS, although one molecular entity, exerts diverse effects by interacting with various substances having different functional specificity. To understand axonal guidance and neuronal circuit formation, it is important to elucidate the mechanisms by which CS exerts a variety of functions.
Sulfation at different positions of the backbone disaccharide units generates various units with different structures; the backbone disaccharide unit (GlcA-GalNAc) is designated as O-unit. Depending on the positions of the sulfate residues, the disaccharide units are designated as A-unit (GlcA-GalNAc4S), B-unit (IdoA-GalNAc4S, IdoA is C-5 epimer of GlcA), C-unit (GlcA-GalNAc6S), D-unit (GlcA2S-GalNAc6S), and E-unit (GlcA-GalNAc4S6S) (
Figure 2). Various combinations of these units generate tremendous structural diversity, even in 1 CS chain, which carries information like “unit clustering” or “unit sequences.” Recent reports indicate that cells may recognize specific unit sequences of CS, which has led to investigations into the molecular mechanism of CS recognition [
7,
35,
36,
37,
38,
39,
40,
41].
Figure 2.
Schematic representation of chondroitin sulfate (CS) units’ structures. CS is made up of unbranched polysaccharides composed of repeating units of disaccharide (glucuronic acid and N-acetylgalactosamine) that are classified into O-, A-, B-, C-, D-, and E-units depending on sulfation sites; the structural diversity of CS results from variable combinations of these units. (A)-unit of CS [GlcAβ1-3GalNAc(4-O-sulfate)];(C)-unit of CS [GlcAβ1-3GalNAc(6-O-sulfate)];(D)-unit of CS [GlcA(2-O-sulfate)β1-3GalNAc(6-O-sulfate)];(E)-unit of CS [GlcAβ1-3GalNAc(4,6-O-disulfate)];(B)-unit of CS is an epimer of C5 position on GlcAs [IdoAα1-3GalNAc(4-O-sulfate)].
Figure 2.
Schematic representation of chondroitin sulfate (CS) units’ structures. CS is made up of unbranched polysaccharides composed of repeating units of disaccharide (glucuronic acid and N-acetylgalactosamine) that are classified into O-, A-, B-, C-, D-, and E-units depending on sulfation sites; the structural diversity of CS results from variable combinations of these units. (A)-unit of CS [GlcAβ1-3GalNAc(4-O-sulfate)];(C)-unit of CS [GlcAβ1-3GalNAc(6-O-sulfate)];(D)-unit of CS [GlcA(2-O-sulfate)β1-3GalNAc(6-O-sulfate)];(E)-unit of CS [GlcAβ1-3GalNAc(4,6-O-disulfate)];(B)-unit of CS is an epimer of C5 position on GlcAs [IdoAα1-3GalNAc(4-O-sulfate)].
3. Obstacles in Studying CS Function
It has not necessarily been easy to investigate the relationship between the structure and function of CS. CS is biologically synthesized through multiple steps, for example, biosynthesis of (donor substrates of) individual monosaccharides, expression of saccharide carrier proteins, regulation of the carrier proteins’ expression, expression of enzymes involving CS chain elongation, activity regulation of the elongation enzymes, expression of CS sulfotransferases, and activity regulation of the sulfotransferases [
42]. Considering the formation of CS chains through these multiple steps, it is clear that the structural diversity of CS such as “percentage contents of the units” “the clustering” or “the unit sequences” is not primarily encoded in genetic information. Until recently, it has been difficult to generate chemically defined CS chains with a standard structure; generally one has been using CS chains purified from biological samples to investigate their function in the context of cellular environment. In the last few years, it is developed the synthetic method that positions sulfate groups at precise locations on the CS carbohydrate chain [
43]. This figures out one of the methodological problems and makes it possible to investigate precisely structure-function relationship of CS.
On the other hand, to study the function of CS, one still faces another methodological problem in investigating function of CS. CS in vivo covalently bound to its core protein, but CS samples commonly used are cut from the core proteins during the purification. Thus, to be precise, the effect of a CS chain without its core protein is not thought to be physiologically accurate. However, this has not been taken well into consideration, primarily due to the difficulty in assembling an appropriate experimental system.
Because CSPG is a natural hybrid molecule, whose information is not primarily within its genome, it is difficult to prepare standardized CSPG or CS products in vitro as recombinant proteins. One needs to produce CS and core proteins and combine them to build up the hybrid molecule. Again, one has to maneuver the different structural diversity of CS on the same core protein. Thus, in investigating the structure–function relationship of CS, it is ideal that the disaccharide unit contents are shown in purified CSPG with a specific core protein from a biological sample, producing a catalogue library of the same core proteins carrying different kinds of structurally diverse CS, and comparing their effects. It is easy to imagine the difficulty in making such a library.
These obstacles stem from, at least, two problems. The first problem is difficulty on obtaining CS with defined structure. The second is how to apply CS to cells physiologically as if it is on the core protein. To figure out the second problem, more physiological experiments are needed. In this review, we spotlight CS derivatives that were developed long ago [
44,
45] but have not been used enough and adequately in neuroscience. With using derivatives of CS, one is able to investigate effects of CS chains on proteins, which mimics CSPG artificially. One is also able to investigate CS on beads, which mimics dense distribution of CS localized in living tissue. To understand the function of CS in neuronal circuit formation, we introduce two kinds of CS derivatives. Both derivations here we review are on the reducing ends of the CS chains, leaving the non-reducing ends for cell interaction because CS chains are bound to their core protein at their reducing ends on native proteoglycans, which is to let the CS chains mimic the native features. Therefore, one expects that this effects the interaction of the glycans and cells like native CS chains on proteoglycans. Likewise, one expects that the free non-reducing ends of CS chains might affect cell behaviors differently like CS chains on the native proteoglycans. The CS derivatives open a way to investigate relationship between structural diversity of CS and its functional specificity. New ideas create novel values in the old techniques.
4. Biotinylated Chondroitin Sulfate and Artificial Chondroitin Sulfate Proteoglycan
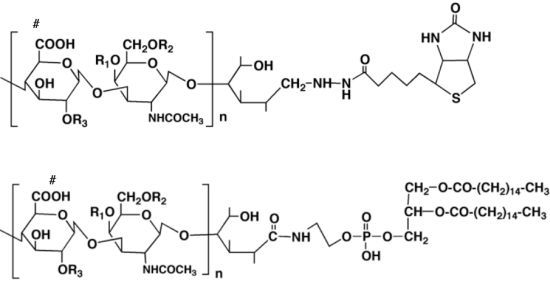
The first derivative is “biotinylated chondroitin sulfate (CS-biotin)” [
44], in which the reducing end of the CS purified from biological samples was chemically coupled with biotin (
Figure 3).
Figure 3.
Schematic drawings of biotinylated chondroitin sulfate (CS-biotin) (
A) and the CS-biotin-streptavidin complex (
B). The reducing end of CS is coupled with biotin and the CS-biotin-streptavidin complex is formed with high affinity between biotin and streptavidin; it is thought that the complex consists of a tetramer. (
C) Complex of CS-biotin and streptavidin in non-denaturing polyacrylamide gel double-stained with SYPRO Ruby (1-3) or Alcian blue (1’-3’). Samples loaded are shown below each lane: 1,1’ streptavidin; 2,2’ biotin-CS; 3,3’ mixed solution with CS-biotin and streptavidin. Open arrowheads: complex of CS-biotin and streptavidin; closed arrowhead: streptavidin; asterisk: unconjugated CS.(
D) Streptavidin-biotin-CSD complex under the influence of retinal axons
in vitro. (
a–
c) Retinal culture on streptavidin-biotin-CSD complex substrate, Staining: (
a) MO-225, (
b) axons,
c merged image.
Arrows indicate absence of MO-225 staining under the axons,
arrowheads indicate absence of MO-225 staining with no axons; the MO-225-negative areas are aligned like axon paths. (
d–
f) After mechanical removal of axons. Staining: (
d) MO-225 staining;(
e) anti-streptavidin staining;(
f) merged image.
Open arrowheads indicate the absence of MO-225 staining and presence of anti-streptavidin staining along the axon paths.
Bars 20 µm. (
C) and (
D) are adapted from Ando
et al. [
46].
Figure 3.
Schematic drawings of biotinylated chondroitin sulfate (CS-biotin) (
A) and the CS-biotin-streptavidin complex (
B). The reducing end of CS is coupled with biotin and the CS-biotin-streptavidin complex is formed with high affinity between biotin and streptavidin; it is thought that the complex consists of a tetramer. (
C) Complex of CS-biotin and streptavidin in non-denaturing polyacrylamide gel double-stained with SYPRO Ruby (1-3) or Alcian blue (1’-3’). Samples loaded are shown below each lane: 1,1’ streptavidin; 2,2’ biotin-CS; 3,3’ mixed solution with CS-biotin and streptavidin. Open arrowheads: complex of CS-biotin and streptavidin; closed arrowhead: streptavidin; asterisk: unconjugated CS.(
D) Streptavidin-biotin-CSD complex under the influence of retinal axons
in vitro. (
a–
c) Retinal culture on streptavidin-biotin-CSD complex substrate, Staining: (
a) MO-225, (
b) axons,
c merged image.
Arrows indicate absence of MO-225 staining under the axons,
arrowheads indicate absence of MO-225 staining with no axons; the MO-225-negative areas are aligned like axon paths. (
d–
f) After mechanical removal of axons. Staining: (
d) MO-225 staining;(
e) anti-streptavidin staining;(
f) merged image.
Open arrowheads indicate the absence of MO-225 staining and presence of anti-streptavidin staining along the axon paths.
Bars 20 µm. (
C) and (
D) are adapted from Ando
et al. [
46].
![Polymers 05 00254 g003]()
CS is able to be biotinylated with the method of Sinohara
et al. [
44]. CS (50 nmol) solution in 0.1 M MES buffer, pH 5.0 (100 µL), and EZ-Link biotin hydrazide (5 µmol, Pierce) solution in dimethylsulfoxide (100 µL) were mixed and incubated at 70 °C for 2 h, followed by the addition of sodium cyanoborohydrate (10 µmol), and further incubation at 70 °C for 16 h. The reaction mixture was subjected to a PD-10 desalting column and then to a Superdex 30 HR 16/600 column using 0.2 M ammonium acetate as an eluent. The elution was monitored by UV absorbance at 225 nm, and the biotin-conjugated CS was obtained by freeze-drying 3 times.
Ando
et al. [
46] generated complex of CS-biotin and streptavidin. The CS-biotin and streptavidin were mixed and electrophoresed on non-denaturing polyacrylamide gels. The protein portion of the complex was visualized fluorescently with SYPRO Ruby; subsequently, the carbohydrate portion of the complex was stained with Alcian blue in the same gels. A broad band stained with both SYPRO Ruby and Alcian blue was observed, indicating the formation of a complex between CS-biotin and streptavidin (
Figure 3C). Unlike the sharp band for streptavidin, the band for the complex was broad in the higher molecular-weight range because of the variation in the molecular weights and anionic charges of the CS with varying lengths [
46]. Because it is known that 1 biotin molecule binds to a streptavidin and streptavidin molecules form a tetramer, it is thought that the complex consists of a tetramer: 4 CS-biotins and 4 streptavidins [
47]. Because the binding between biotin and streptavidin is known to be one of the highest affinities in the natural world, it is thought that the complex is stable in physiological conditions; thus, the complex is expected to be utilized in functional experiments as “artificial CSPG” with 4 CS chains on the core. Since it was confirmed that streptavidin itself in the complex (as the “core protein”) has no effect on axonal elongation, this made it possible to examine CS function in neuronal circuit formation.
CS is inhibitory to axonal outgrowth of retinal ganglion cells. However, inspecting the distribution of CS around the pathway of retinal ganglion cells’ axons
in vivo, the retinal axons are seen running on a region with densely distributed CS [
48]. Indeed, a small but substantial population of retinal axons grows on a culture substrate containing CS. These results suggest that some, but not all, of the retinal axons are insensitive to the CS inhibitory effect. In the case of hippocampal neurons, it is reported that CS enhances neurite outgrowth [
26,
27,
28,
29]. These disparate findings indicate that the effects of CS differ between types of neurons, possibly because of varying sensitivity to CS.
Ando
et al. [
46] used CS-biotin in investigating axonal elongation
in vitro, which is a successful example of application of the CS derivative (
Figure 3D); thus, here we describe their approach. They used CS-biotin to generate the CS-biotin-streptavidin complex as an artificial CSPG, and further examined the interaction between the retinal axons and CS. The culture substrate was prepared using the CS-biotin-streptavidin complex and was double-stained with anti-CS (MO-225) and anti-streptavidin. Both the antibodies showed uniform staining throughout the substrate when there is no axon, indicating an even distribution of the complex in the substrate. A small population of retinal axons grew on the substrate with CS-biotin-streptavidin. Along the axonal trajectories, small regions were observed with decreasing signal of anti-CS antibody immunostaining, but those same regions had a uniform signal of anti-streptavidin staining. These results indicate that the density of epitopes against the carbohydrate portion of the complex, CS, was reduced or absent. However, the epitopes against the protein portion, streptavidin, were present along the axonal trajectory. The uniform staining of anti-streptavidin polyclonal antibody indicated that the complex was distributed uniformly on the substrate, and that antibodies readily accessed the complex on the substrate with no mechanical hindrance by the axons; thus, the decreasing signal of anti-CS antibody staining is likely an effect of the axons on the complex. Since the signal for carbohydrate portion (anti-CS staining) was merely reduced, it seems that the retinal axons have a selective influence on the CS carbohydrate chain in their local environment. It is a possibility that the retinal axons cut the carbohydrate chains of CS to reduce the length or modify the structure of CS, resulting in the observed epitope density reduction. However, further investigation is necessary to reveal its mechanism. Importantly, the reduction of the CS staining may have biological significance. For instance, axons may act upon their local environment to modify the carbohydrate chains of the CS and thus render the environment more suitable to their growth, which is thought to be an adaptation of the axons to their growth environment.
It has never been reported that neurons cut or modify carbohydrate chains of CS in their extracellular environment, nor has there been any investigation for such a mechanism. However, since CS is one of the important players in preventing axons from rewiring in central nervous system injury, for example, in spinal cord injury, it is meaningful to point out all possible intrinsic mechanisms to modify the carbohydrate chains of CS in terms of therapeutic application. The derivative of CS-biotin constitutes an essential contribution to this field of study.
As demonstrated above, the CS-biotin-streptavidin complex is useful in preparing artificial CSPG. Moreover, it can be used to prepare complexes of different types of CS chains on the same core protein. Since highly sulfated CS, for example D- or E-unit containing CS, is thought to be involved with neurite elongation or axonal guidance [
7,
28,
29,
36,
39], it would be intriguing to use the complex with D- or E-unit containing CS in future investigations. The CS-biotin-streptavidin complex enables us to examine the relationship between the structural diversity of CS and its functional specificity. Application of the CS-biotin derivative may be expected not only in the investigation of neuronal circuit formation, but also in other fields.
5. Lipid-Derivatized Chondroitin Sulfate and Polystyrene Beads Coated with Lipid-Derivatized Chondroitin Sulfate
Experiments using neuronal culture are important because one is able to examine the effect of CS on neuronal cells directly. “The spot assay” in which CS is spotted on the culture substrate, has been commonly used up to this point. Comparing axonal trajectories inside and outside the CS spot, one expects to study the effects of CS on axons. However, there seems to be methodological difficulties with this approach. The CS spot on the substrate surface is subsequently coated with extracellular matrix-like laminin; however, the CS prevents laminin from being coated in the spot, resulting in a laminin density difference between the inside (lower density) and outside (higher density) of the spot (Ichijo, unpublished observation). On the other hand, to form a boundary of the spot, the coating order is essential; in the reverse order of coating, the spot is not created because of its hydrophilic surface. Since growth cones prefer a higher density of laminin, it is difficult to distinguish the effects of CS from the general effects of laminin.
Additionally, “the neurite outgrowth assay” in which the CS is uniformly distributed on the culture substrate, the CS operates continuously on the whole neuronal cell body, making it difficult to analyze when and where the CS is causing an effect. Consequently, this technique might not be useful for examining CS effects on behaviors of neuronal growth cones. Thus, a two-dimensional substrate of CS is not necessarily suitable for investigating the effects of CS structural diversity on growth cones. Gilbert
et al. (2005) pointed out this difficulty; to understand the functional roles of CS diversity, they covalently coupled CS to three-dimensional hydrogels [
37].
To bypass the problems of the conventional assay and examine the effect of CS on axonal behavior directly, another derivative of CS is introduced. The amino group of phosphatidylethanolamine was chemically coupled on the reducing end of the CS chain to generate the “lipid-derivatized CS (CS-PE)”, which was originally developed by Sugiura
et al. (
Figure 4) [
45].
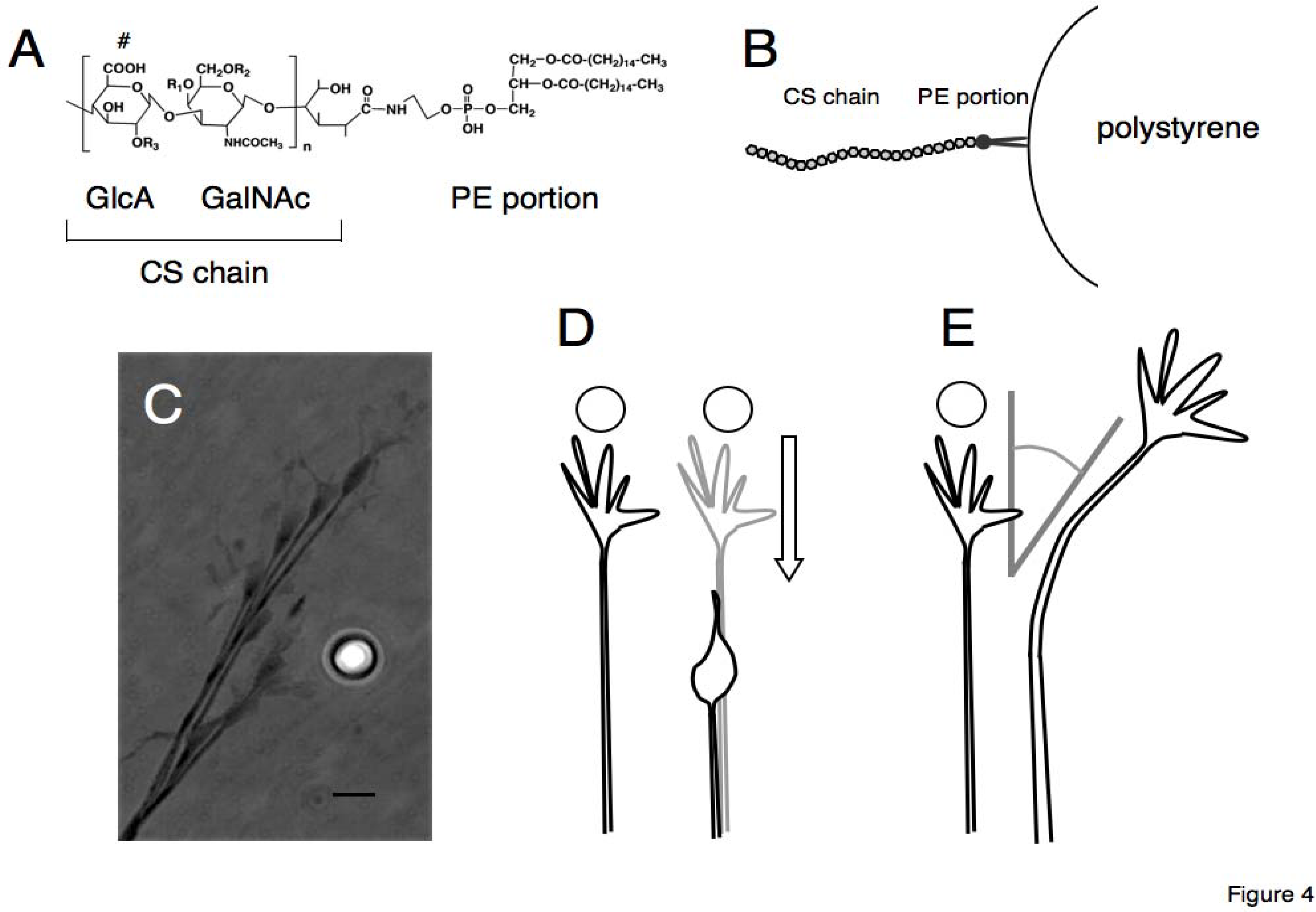
Figure 4.
Schematic drawings of lipid-derivatized CS (CS-PE) (A) and CS-PE bead (B). The reducing end of the CS chain is coupled with an amino group of phosphatidylethanolamine to generate CS-PE (A). The CS-PE hydrophobically adheres on the surface of polystyrene beads, forming the CS-PE beads with CS chains arranged outwardly (B). Behavior of a retinal growth cone encountering a CS-PE bead is shown in a photomontage of time-lapse images (C). Typical behaviors of the retinal growth cones encountering CSE-PE beads are shown in (D) retraction and (E) turning away from the beads.
Figure 4.
Schematic drawings of lipid-derivatized CS (CS-PE) (A) and CS-PE bead (B). The reducing end of the CS chain is coupled with an amino group of phosphatidylethanolamine to generate CS-PE (A). The CS-PE hydrophobically adheres on the surface of polystyrene beads, forming the CS-PE beads with CS chains arranged outwardly (B). Behavior of a retinal growth cone encountering a CS-PE bead is shown in a photomontage of time-lapse images (C). Typical behaviors of the retinal growth cones encountering CSE-PE beads are shown in (D) retraction and (E) turning away from the beads.
CS (5.0 µmol) was oxidized with iodine (50 µmol) in 5 mL of methanol-water (1:10, v/v) at room temperature for 6 h. The oxidized CS was precipitated with 3 volumes of 95% (v/v) ethanol containing 1.3% (w/v) potassium acetate. The precipitate was dissolved in water and passed through a Dowex 50 × 8 (H+) resin. The solution was concentrated at 40 °C in vacuo, and the remaining water was replaced by the addition of dimethylformamide (DMF) followed by evaporation several times. The DMF solution (5 mL) was allowed to stand at 4 °C for 72 h, to which dipalmitoyl-phosphatidyl ethanolamine (PE, 250 µmol) in 0.5 mL of chloroform was added. The mixture was stirred at 60 °C for 6 h, then concentrated in vacuo, and suspended with 0.5 mL of 0.2 M NaCl. The undissolved material (excess PE) was removed by centrifugation. The supernatant was subjected to a TSK-gel Phenyl 5PW column using 0.2 M NaCl as a wash and 30% (v/v) methanol-water as an eluent. The PE-conjugated CS was obtained via concentration and freeze-drying.
The CS-PE is amphipathic, consisting of a hydrophilic CS chain and a hydrophobic phosphatidylethanolamine. The PE portion of the CS-PE hydrophobically adheres on the surface of polystyrene beads, forming the CS-PE beads with CS chains arranged outwardly. Spreading the CS-PE beads on the culture dish, one is able to closely observe behaviors of axonal growth cones approaching and encountering the CS-PE beads in time-lapse movies. Moreover, by using each unit-enriched CS-PE (CSA-, CSC-, CSD-, or CSE-PE), one is able to prepare the CS-PE beads with various CS structural diversities (CSA-PE beads and so on).
This method, the “CS-PE beads assay” enables Simbo
et al. to compare effects of CS with different structural diversity on the behavior of retinal growth cones directly in time-lapse movies (
Figure 4) [
49]. Growth cones of the retinal ganglion cells avoided E-unit containing CS-PE beads (CSE-PE beads), exhibiting significant repulsion or turning away from the CSE-PE beads. The effects of CSE-PE on the retina growth cones were quenched by treatment of the beads with chondroitinase ABC. The beads with other types of CS (CSA-, CSC-, and CSD-PE) or the beads without CS had no significant effect. Therefore, the retinal growth cones differentiated the CS on the beads and selectively moved away from the E-unit-containing CS. Moreover, the repulsive effects of CSE-PE beads are different from the well-known proteinaceous repulsive guidance factors like ephrins or semaphorins [
50,
51], because CSE-PE beads induce various continuous degrees of moving away behaviors, much like volume control. Thus far, the mechanisms moderating the repulsive effects of CSE are not known; therefore, further investigation is needed. There are several possible hypotheses addressing the differential effects between heterogeneous CSE chains or differing sensitivity between retinal growth cones. Since CS consists of a linear carbohydrate chain with 20–200 disaccharide units, it is able to carry large amounts of information in the heterogeneous structure of CS, such as sequences of the units or clustering of the unit inside the chain, which might be suitable for fine regulation of axonal behaviors in contrast to the proteinaceous guidance factors. With polystyrene beads coated with lipid-derivatized CS, one is able to examine the structural diversity and functional specificity of CS. Lipid-derivatized CS is applicable not only in the investigation of axonal guidance but also other experimental approaches for biological phenomena.
6. Further Problems in CS Research
Here we have reviewed methodology using CS derivatives, CS-biotin and CS-PE, and their application in axonal guidance research. These methods have opened new possibilities although there is a limitation. There are still problems in CS research. The problem on how to elucidate structural diversity of CS is generally recognized, which might be also a bottleneck of applying the CS derivatives in neuroscience and biomedical sciences. One is not being able to determine the “sequence of the units in CS chain” practically. Until recently, there had been no useful method synthesizing particular CS chains. Therefore, one had to obtain biological samples of CS from many organisms, chemically analyze them with percentages of the unit contents, and classify them in terms of the dominant unit contents, into CSA, CSC, and so on. However, these CS samples are not regarded as CS with defined structures. To overcome the problem, Gama
et al. [
43] developed the chemically synthetic method that positions sulfate groups at precise locations on the CS carbohydrate chain and obtained CS carbohydrate chains with the standard structure, such as tetrasaccharides of defined sulfation pattern with A-, C-, E-, and so on. Sugiura
et al. [
52] have also recently developed a novel method of synthesizing CS species chemo-enzymatically with defined lengths and defined sulfate compositions. These open the way to generate a CS library with defined structure.
Investigating effects of defined CSs is a promising way to understand structure-function relationship of CS although it is not thought to be perfectly enough; one should consider effects of the defined CSs on their core protein. Combining CS-biotin or CS-PE with the defined CS structure would enable us to investigate the precise linkage between structure and function of CS in more physiological conditions. After completing this step of investigations, one is able to consider the problems of structural diversity and functional specificity of CS why the structure of CS is so diverse or what sequences of CS code.
Recently, the function of CS has been successfully investigated by analysis of genetic network between extracellular factors and their receptors with using genetically modified mice; thus, these genetic approaches seem promising in overcoming the obstacles in CS function research [
7,
8,
9,
10]. In fact, the existence of receptors specific to CS carbohydrate chains has been proposed. In contrast, methods using CS derivatives are thought to directly access the effect of CS on neurons, which is complementary to the genetic approaches. Our understandings of CS function and its structural diversity are expected to be promoted by the combined use of these direct
in vitro and indirect genetic approaches.
Novel ideas regarding other CS derivatives should lead to further breakthroughs and progress in this field of research. In addition to the derivatives described here, there appear to be other promising derivatives; for example, fluorescently labeled CS derivatives [
53]. By combining both existing and new ideas for the use of CS derivatives with genetic approaches, the linkage between the structure and function of CS can be elucidated.