Hydrogen-Bonding in Two Pyridinium Salts of [Mo2O4Cl4(μ2-dmsH)]3−Complex (dmsH− = a Half-Neutralized Form of 2,2-Dimethylsuccinic Acid)
Abstract
:1. Introduction
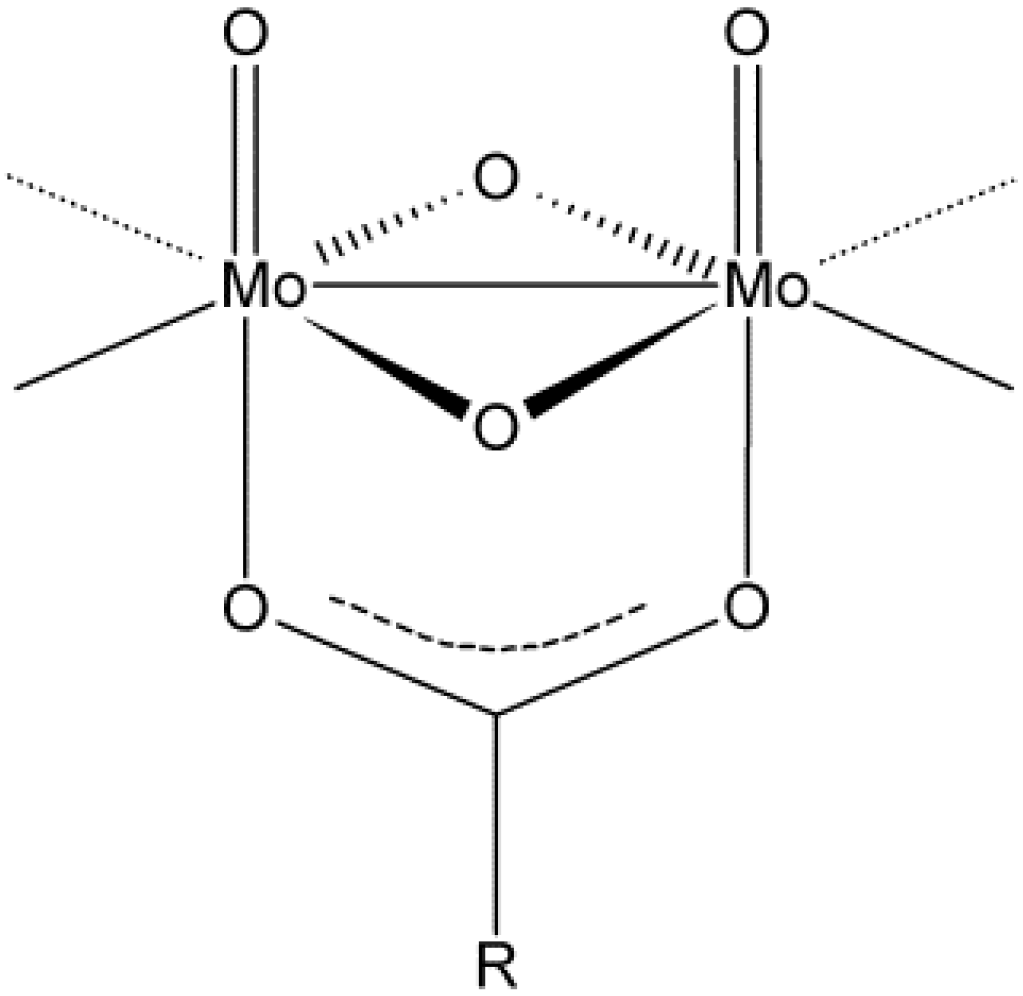

2. Results and Discussion
2.1. Solid State Structures
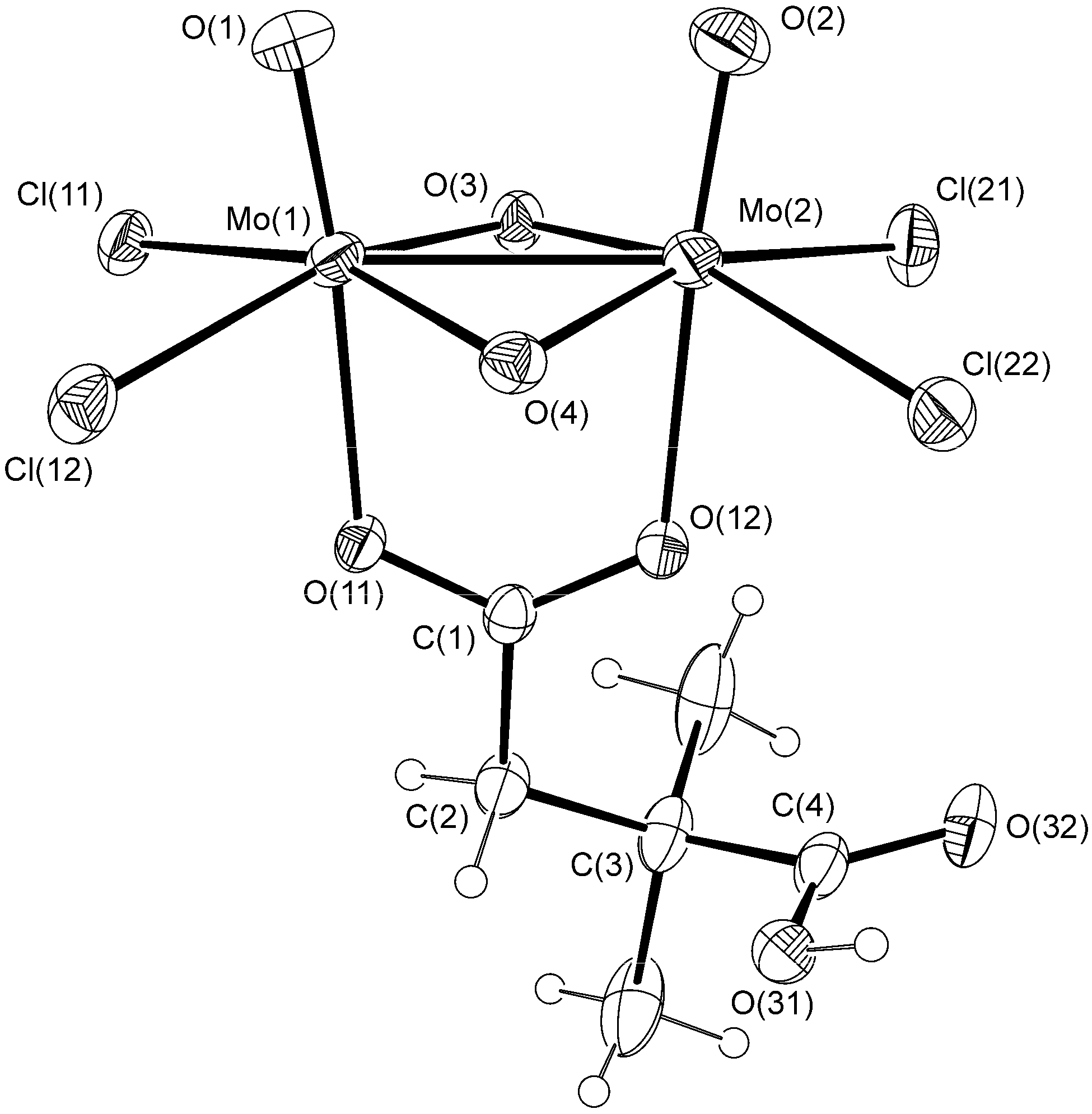
| Bond | 1a | 2 | |
|---|---|---|---|
| Mo–Mo | 2.5827(4) | 2.5713(4) | 2.5929(7) |
| Fold angleb | 157.9(1) | 156.6(1) | 160.7(3) |
| Mo–Cl | 2.4407(9)−2.4718(9) | 2.4522(9)−2.4722(9) | 2.438(2)−2.472(2) |
| Mo–O(carboxylate) | 2.286(2), 2.344(2) | 2.324(2), 2.366(2) | 2.236(4), 2.287(4) |
| C–O(carboxylate) | 1.258(4), 1.288(4) | 1.255(4), 1.275(4) | 1.265(6), 1.271(6) |
| C–O(COOH) | 1.254(4), 1.293(4) | 1.245(4), 1.291(4) | 1.223(7), 1.312(8) |
| Species | La | Mo–O(L) | Mo–Mo | Fold angle | Ref. |
|---|---|---|---|---|---|
| 1 | dmsH− | 2.286(2), 2.344(2)2.324(2), 2.366(2) | 2.5827(4)2.5713(4) | 157.9(1)156.6(1) | - |
| 2 | dmsH− | 2.236(4), 2.287(4) | 2.5929(7) | 160.7(3) | - |
| [Mo2O4Cl4(μ2-OOCCH3)]3− | acetate | 2.354(1), 2.366(1) | 2.5727(2) | 157.01(9) | [14] |
| [Mo2O4Cl4(μ2-Hmal)]3− | Hmal− | 2.330(2), 2.357(2) | 2.5859(3) | 160.08(5) | [15] |
| [Mo2O4Cl4(μ2-Hmale)]3−b | Hmale− | 2.380(2), 2.438(2)2.371(2), 2.377(2) | 2.5916(3)2.5951(3) | 156.49(9)158.3(1) | [16] |
| [Mo2O4Cl4(μ2-btcH2)]3− | btcH2− | 2.269(2), 2.281(2) | 2.5962(4) | 159.85(5) | [9] |

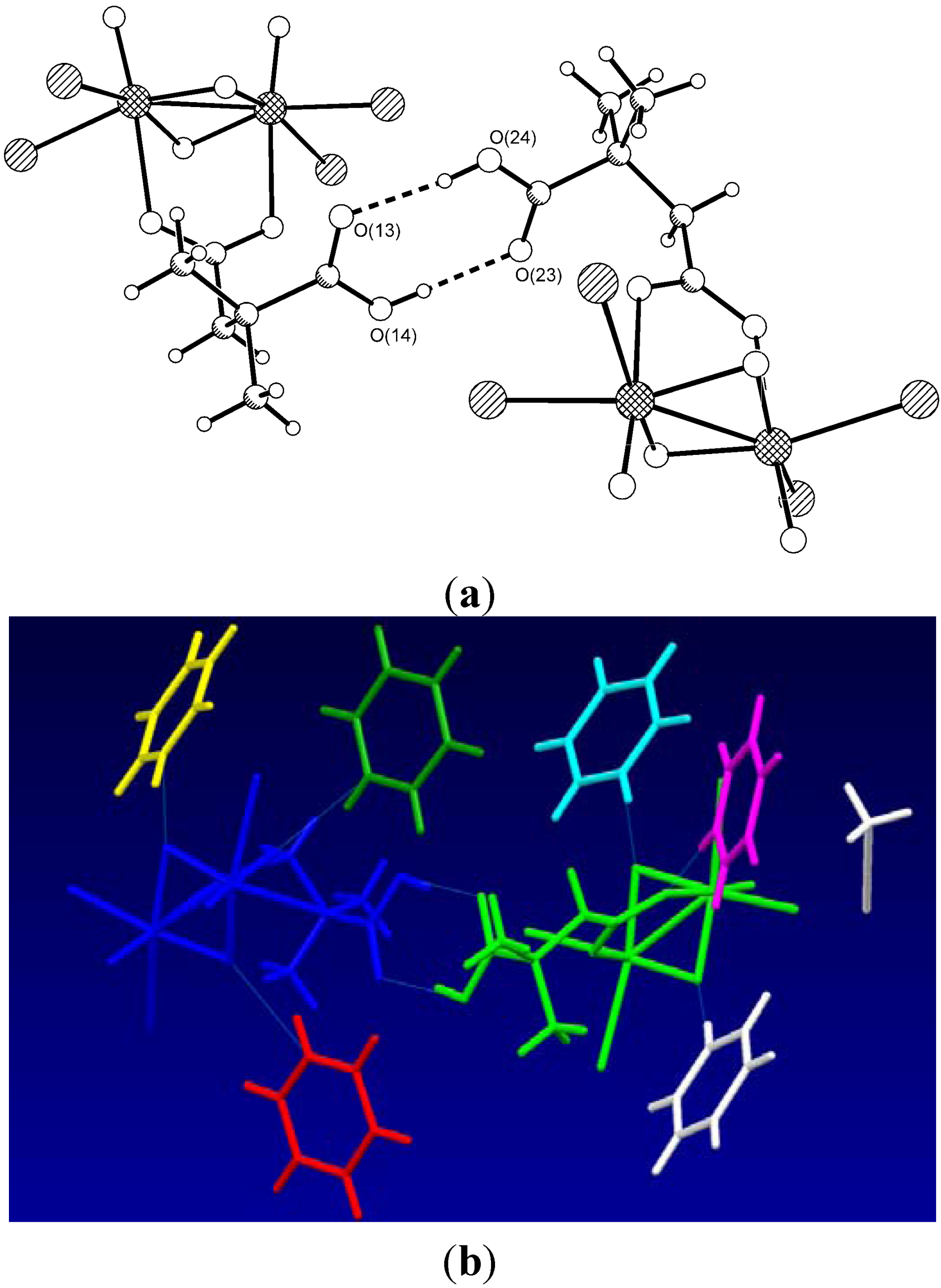
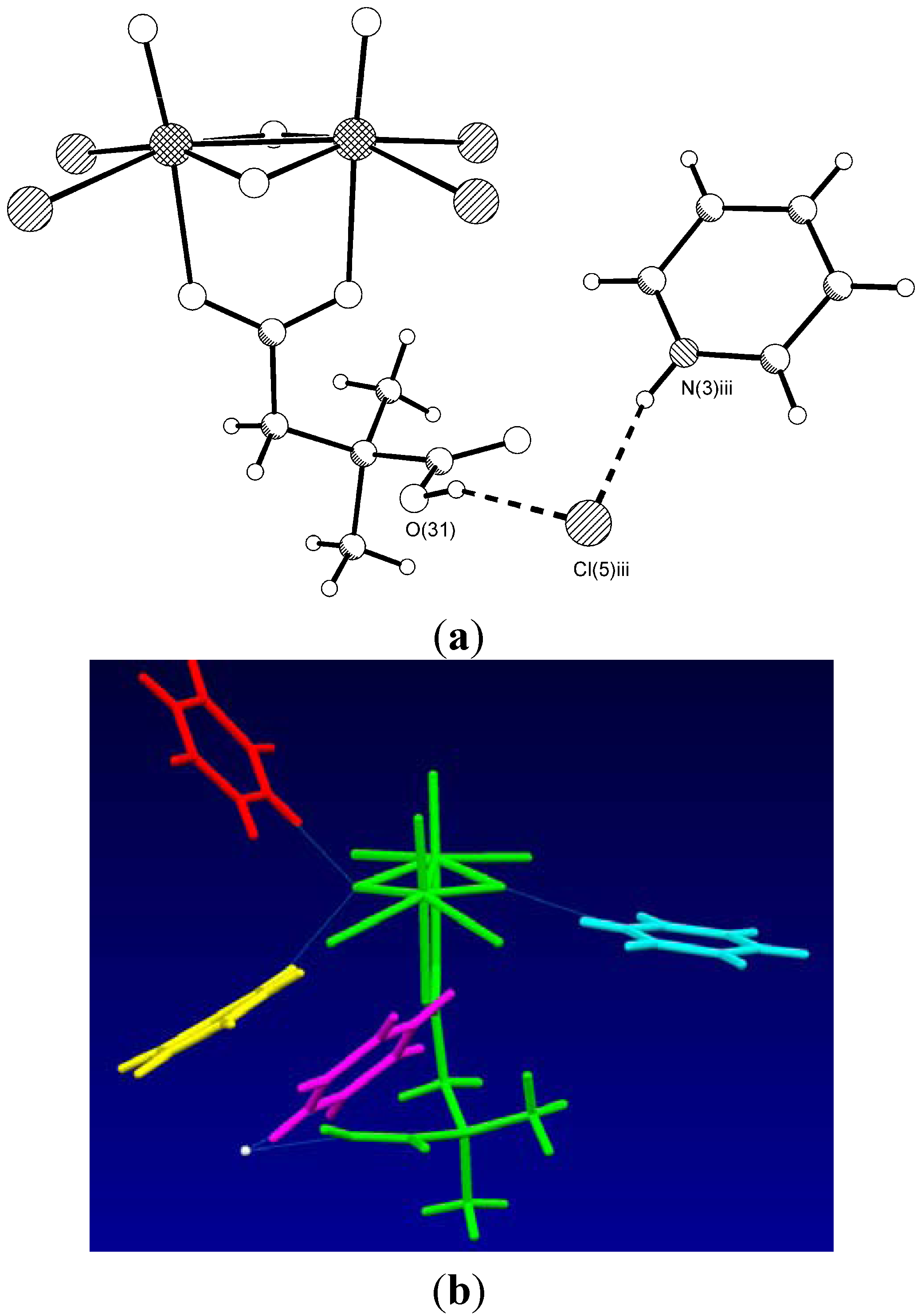


| Type | Donor atom···acceptor atom a | Length (Å) b |
|---|---|---|
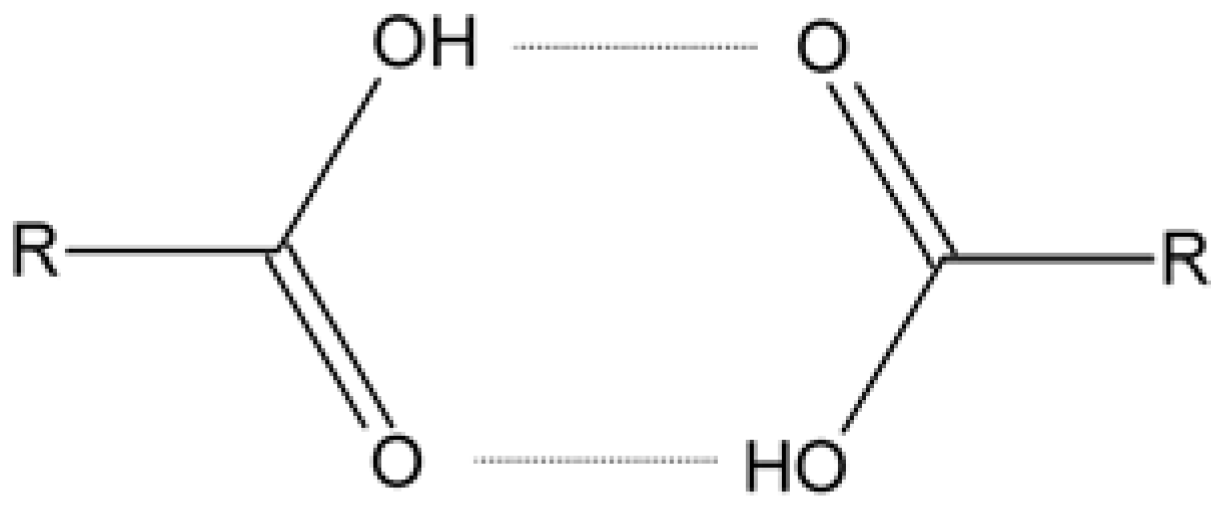
| COOH···COOH | O(14)···O(23) | 2.630(3) |
| COOH···COOH | O(24)···O(13) | 2.609(3) |
| PyH+··· μ2-O | N(1)···O(7) | 2.767(5) |
| PyH+··· μ2-O | N(2)···O(8) | 2.716(4) |
| PyH+··· μ2-O | N(4)···O(5) i | 2.620(4) |
| PyH+··· μ2-O | N(5)···O(6) ii | 2.722(4) |
| PyH+···COO− c | N(3)···O(11) ii | 2.737(4) |
| PyH+···COO− c | N(6)···O(22) i | 2.833(4) |
| Type | Donor atom···acceptor atom a | Length (Å) b |
|---|---|---|
| COOH···Cl− | O(31)···Cl(5) iii | 3.044(4) |
| PyH+···Cl− | N(3) iii··· Cl(5) iii | 3.047(5) |
| PyH+··· μ2-O | N(1)···O(4) iv | 2.620(8) |
| PyH+··· μ2-O | N(2)···O(4) | 2.807(7) |
| PyH+··· μ2-O | N(4)···O(3) | 2.666(6) |
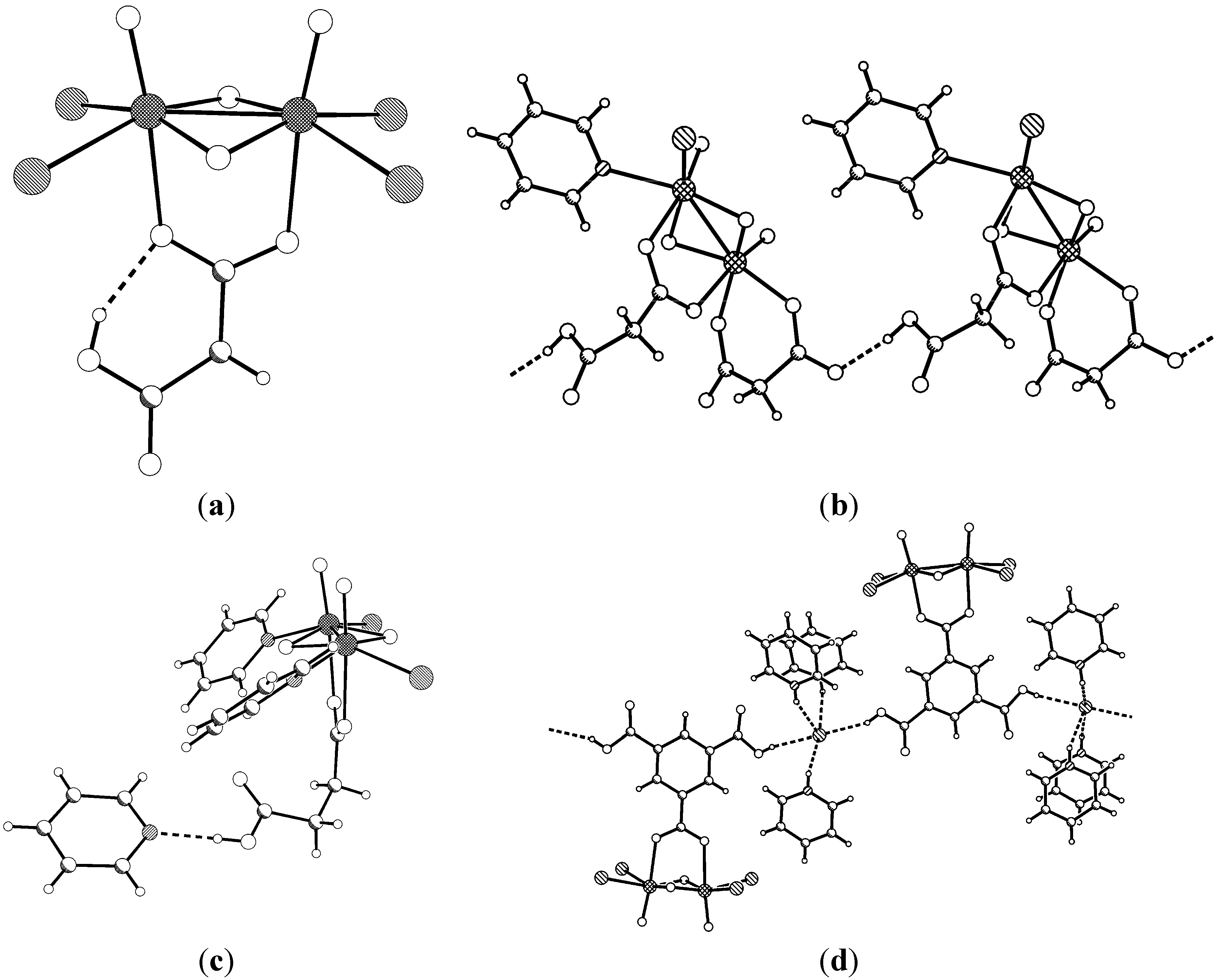
2.2. Infrared Spectroscopy
3. Experimental Section
3.1. General
3.2. Preparation of (PyH)3[Mo2O4Cl4(μ2-dmsH)]·1/2CH3CN (1)
3.3. Preparation of (PyH)4[Mo2O4Cl4(μ2-dmsH)]Cl (2)
3.4. X-ray Crystallography
| Compound | 1 | 2 |
|---|---|---|
| Empirical formula | C22H28.5Cl4Mo2N3.5O8 | C26H33Cl5Mo2N4O8 |
| Formula weight | 803.7 | 898.7 |
| Crystal system | monoclinic | monoclinic |
| Space group | P 21/n | P 21/n |
| T (K) | 150(2) | 150(2) |
| a (Å) | 9.9123(1) | 9.0254(1) |
| b (Å) | 17.0078(1) | 25.7214(4) |
| c (Å) | 36.1401(3) | 16.2035(2) |
| α (deg) | 90 | 90 |
| β (deg) | 97.4302(4) | 97.0941(7) |
| γ (deg) | 90 | 90 |
| V (Å3) | 6041.57(9) | 3732.78(8) |
| Dcalcd (g/cm3) | 1.767 | 1.599 |
| Z | 8 | 4 |
| λ (Å) | 0.71073 | 0.71073 |
| μ (mm−1) | 1.233 | 1.077 |
| Collected reflections | 19959 | 12861 |
| Unique reflections, Rint | 10966, 0.028 | 6785, 0.0281 |
| Observed reflections | 8997 | 5449 |
| R1 a (I > 2σ(I) | 0.0291 | 0.0526 |
| wR2 b (all data) | 0.0658 | 0.1283 |
4. Conclusions
Acknowledgments
Conflict of Interest
References and Notes
- Cotton, F.A. Structure and bonding in molecular oxo-molybdenum compounds. J. Less Common. Met. 1974, 36, 13–22. [Google Scholar] [CrossRef]
- Chae, H.K.; Klemperer, W.G.; Marquart, T.A. High-nuclearity oxomolybdenum(V) complexes. Coord. Chem. Rev. 1993, 128, 209–224. [Google Scholar]
- Khan, M.I.; Müller, A.; Dillinger, S.; Bögge, H.; Chen, Q.; Zubieta, J. Cation inclusion within the mixed-valence polyanion cluster [(MoVIO3)4MoV12O28(OH)12]8−: Syntheses and structures of (NH4)7[NaMo16(OH)12O40]·4H2O and (Me2NH2)6[H2Mo16(OH)12O40]. Angew. Chem. Int. Ed. Engl. 1993, 32, 1780–1782. [Google Scholar] [CrossRef]
- Dolbecq, A.; Lisnard, L.; Mialane, P.; Marrot, J.; Benard, M.; Rohmer, M.-M.; Secheresse, F. Synthesis and characterization of octa- and hexanuclear polyoxomolybdate wheels: Role of the inorganic template and the counterion. Inorg. Chem. 2006, 45, 5898–5910. [Google Scholar]
- James, S.L. Metal-organic frameworks. Chem. Soc. Rev. 2003, 32, 276–288. [Google Scholar] [CrossRef]
- Cotton, F.A.; Donahue, J.P.; Lin, C.; Murillo, C.A. The simplest supramolecular complexes containing pairs of Mo2(formamidinate)3 units linked with various dicarboxylates: Preparative methods, structures, and electrochemistry. Inorg. Chem. 2001, 40, 1234–1244. [Google Scholar] [CrossRef]
- Cotton, F.A.; Daniels, L.M.; Lin, C.; Murillo, C.A. Square and triangular arrays based on Mo24+ and Rh24+ units. J. Am. Chem. Soc. 1999, 121, 4538–4539. [Google Scholar]
- Cotton, F.A.; Lin, C.; Murillo, C.A. Supramolecular squares with Mo24+ corners. Inorg. Chem. 2001, 40, 478–484. [Google Scholar] [CrossRef]
- Modec, B.; Brenčič, J.V. Anions of 1,3,5-benzenetricarboxylic and heptanedioic acids serving as bridges between dimolybdenum(V) metal−metal bonded units: Preparation and structural characterization of dinuclear and tetranuclear complexes. Eur. J. Inorg. Chem. 2005, 2005, 4325–4334. [Google Scholar] [CrossRef]
- Modec, B.; Brenčič, J.V. Novel methanol-containing oxomolybdate(V) complexes: Synthesis and structural characterisation of intermediates in the formation of {Mo2O4}2+ clusters from [MoOCl4(H2O)]− and [MoOBr4]− precursors. Eur. J. Inorg. Chem. 2005, 2005, 1698–1709. [Google Scholar] [CrossRef]
- Cotton, F.A.; Morehouse, S.M. The molecular structure of a diamagnetic, doubly oxygen-bridged, binuclear complex of molybdenum(V) containing a metal–metal bond. Inorg. Chem. 1965, 4, 1377–1381. [Google Scholar] [CrossRef]
- Stiefel, E.I. The coordination and bioinorganic chemistry of molybdenum. In Progress in Inorganic Chemistry; John Wiley & Sons Inc.: Hoboken, NJ, USA, 1977; pp. 1–223. [Google Scholar]
- A relative orientation of the dmsH− ligand may be given by the dihedral angle between the planes of its COO− and COOH functions. For the two [Mo2O4Cl4(μ2-dmsH)]3− anions of the asymmetric unit in 1 the angles are 78.2(3) and 66.8(3)°. A value of 83.8(5)° was calculated for the dmsH− ligand in 2.
- Modec, B. Acetato complexes of molybdenum(V): A novel tetranuclear core based on the metal−metal bonded {Mo2O4}2+ units. Inorg. Chim. Acta 2007, 361, 2863–2870. [Google Scholar] [CrossRef]
- Modec, B.; Dolenc, D.; Brenčič, J.V. New Molybdenum(V) complexes based on the {Mo2O4}2+ structural core with esters or anions of malonic and succinic acid. Inorg. Chim. Acta 2007, 360, 663–678. [Google Scholar] [CrossRef]
- Modec, B.; Brenčič, J.V. Novel hydrogenmaleato molybdenum(V) complexes based on a dinuclear metal−metal bonded unit: Syntheses and structural characterization of (PyH)3[Mo2O4Cl4(OOCCH=CHCOOH)] and (PyH)3[Mo2O4Br4(OOCCH=CHCOOH)]·CH3CN. Inorg. Chem. Commun. 2004, 7, 516–520. [Google Scholar] [CrossRef]
- Douglas, B.; McDaniel, D.; Alexander, J. Concepts and Models of Inorganic Chemistry, 3rd ed; John Wiley & Sons Inc.: New York, NY, USA, 1994; p. 102. [Google Scholar]
- Leiserowitz, L. Molecular packing modes. Carboxylic acids. Acta Crystallogr. Sect. B 1976, 32, 775–802. [Google Scholar] [CrossRef]
- Allen, F.H.; Kennard, O.; Taylor, R. Systematic analysis of structural data as a research technique in organic chemistry. Acc. Chem. Res. 1983, 16, 146–153. [Google Scholar] [CrossRef]
- Nakamoto, K. Infrared and Raman Spectra of Inorganic and Coordination Compounds. Part B: Applications, in: Coordination, Organometallic, and Bioinorganic Chemistry, 5th ed; John Wiley & Sons, Inc.: New York, NY, USA, 1997; pp. 59–62. [Google Scholar]
- Deacon, G.B.; Phillips, R.J. Relationships between the carbon–oxygen stretching frequencies of carboxylato complexes and the type of carboxylate coordination. Coord. Chem. Rev. 1980, 33, 227–250. [Google Scholar] [CrossRef]
- Colthup, N.B.; Daly, L.H.; Wiberley, S.E. Introduction to Infrared and Raman Spectroscopy; Academic Press: New York, NY, USA, 1964; pp. 257–260. [Google Scholar]
- Otwinowski, Z.; Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997, 276, 307–326. [Google Scholar] [CrossRef]
- Sheldrick, G.M. SHELXS-97; University of Göttingen: Göttingen, Germany, 1997; Programs for Crystal Structure Solution and Refinement. [Google Scholar]
- Farrugia, L.J. WinGX suite for small-molecule single-crystal crystallography. J. Appl. Cryst. 1999, 32, 837–838. [Google Scholar] [CrossRef]
- Farrugia, L.J. ORTEP-3 for Windows–A version of ORTEP-III with a graphical user interface (GUI). J. Appl. Cryst. 1997, 30, 565. [Google Scholar]
- SHELXTL, version 5.03 ed; Siemens Analytical X-ray Instrument Division: Madison, WI, USA, 1994.
- Macrae, C.F.; Edgington, P.R.; McCabe, P.; Pidcock, E.; Shields, G.P.; Taylor, R.; Towler, M.; van de Streek, J. Mercury: Visualization and analysis of crystal structures. J. Appl. Cryst. 2006, 39, 453–457. [Google Scholar] [CrossRef]
- CrystalMaker for Windows, version 2.6.1 ed; CrystalMaker Software, Ltd.: Oxfordshire, UK, 2007.
- CCDC. Available online: www.ccdc.cam.ac.uk/data_request/cif (accessed on 18 March).
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Modec, B. Hydrogen-Bonding in Two Pyridinium Salts of [Mo2O4Cl4(μ2-dmsH)]3−Complex (dmsH− = a Half-Neutralized Form of 2,2-Dimethylsuccinic Acid). Crystals 2013, 3, 275-288. https://doi.org/10.3390/cryst3020275
Modec B. Hydrogen-Bonding in Two Pyridinium Salts of [Mo2O4Cl4(μ2-dmsH)]3−Complex (dmsH− = a Half-Neutralized Form of 2,2-Dimethylsuccinic Acid). Crystals. 2013; 3(2):275-288. https://doi.org/10.3390/cryst3020275
Chicago/Turabian StyleModec, Barbara. 2013. "Hydrogen-Bonding in Two Pyridinium Salts of [Mo2O4Cl4(μ2-dmsH)]3−Complex (dmsH− = a Half-Neutralized Form of 2,2-Dimethylsuccinic Acid)" Crystals 3, no. 2: 275-288. https://doi.org/10.3390/cryst3020275






