Aberrant Promoter Hypermethylation of RASSF Family Members in Merkel Cell Carcinoma
Abstract
:1. Introduction
2. Experimental
2.1. CpG Island Prediction, PCR Product Size and Digestion Products
2.2. Merkel Cell Carcinoma and Controls
2.3. DNA Isolation
2.4. Methylation Analysis by COBRA
3. Results
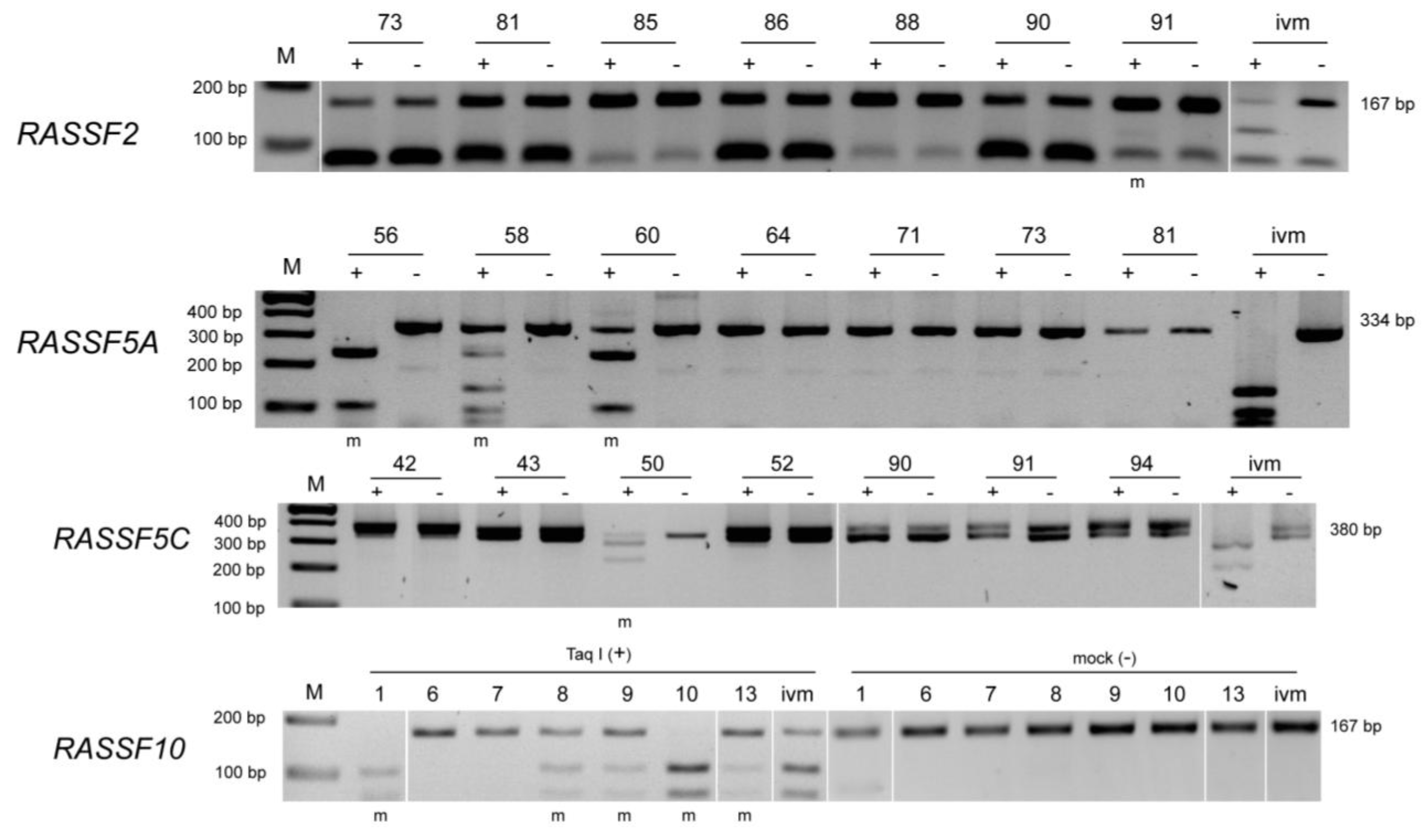
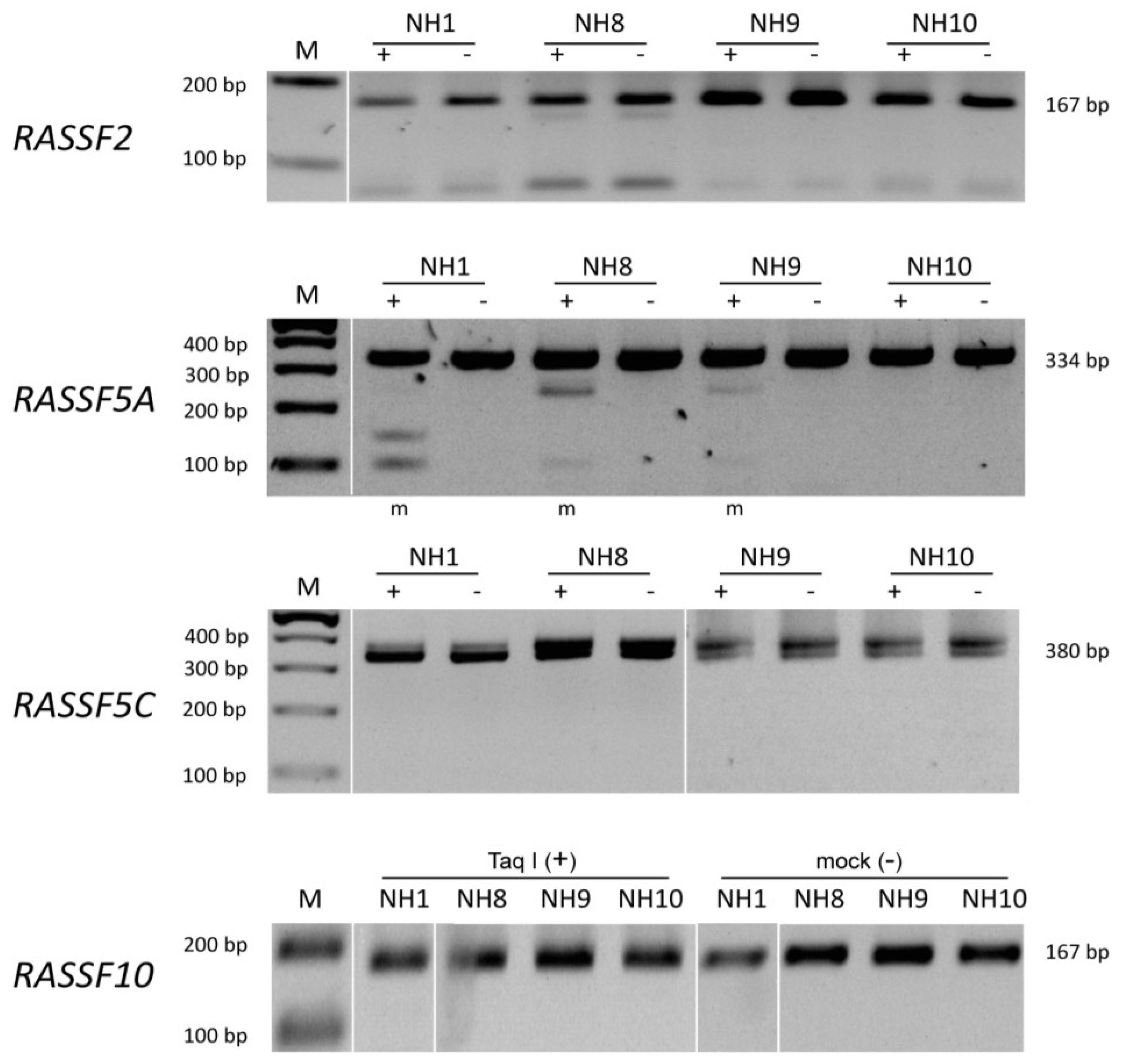
| Gene | Tumor samples | Control samples |
|---|---|---|
| RASSF2 | 7% (3/43) | 0% (0/9) |
| RASSF5A | 44% (17/39) | 43% (3/7) |
| RASSF5C | 8% (2/26) | 0% (0/11) |
| RASSF10 | 23% (19/84) | 0% (0/14) |
4. Discussion
5. Conclusions
Abbreviations
| MCC | Merkel cell carcinoma |
| RASSF | Ras-Association Domain Family |
| TSG | tumor suppressor gene |
| COBRA | combined bisulfite restriction analysis |
Acknowledgements
Conflicts of Interest
References
- Hodgson, N.C. Merkel cell carcinoma: Changing incidence trends. J. Surg. Oncol. 2005, 89, 1–4. [Google Scholar] [CrossRef]
- Grabowski, J.; Saltzstein, S.L.; Sadler, G.R.; Tahir, Z.; Blair, S. A comparison of Merkel cell carcinoma and melanoma: Results from the california cancer registry. Clin. Med. Oncol. 2008, 2, 327–333. [Google Scholar]
- Spurgeon, M.E.; Lambert, P.F. Merkel cell polyomavirus: A newly discovered human virus with oncogenic potential. Virology 2013, 435, 118–130. [Google Scholar] [CrossRef]
- Kassem, A.; Schopflin, A.; Diaz, C.; Weyers, W.; Stickeler, E.; Werner, M.; Zur Hausen, A. Frequent detection of Merkel cell polyomavirus in human Merkel cell carcinomas and identification of a unique deletion in the VP1 gene. Cancer Res. 2008, 68, 5009–5013. [Google Scholar] [CrossRef]
- Feng, H.; Shuda, M.; Chang, Y.; Moore, P.S. Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science 2008, 319, 1096–1100. [Google Scholar] [CrossRef]
- Becker, J.C.; Houben, R.; Ugurel, S.; Trefzer, U.; Pfohler, C.; Schrama, D. Mc polyomavirus is frequently present in Merkel cell carcinoma of european patients. J. Invest. Dermatol. 2009, 129, 248–250. [Google Scholar] [CrossRef]
- Helmbold, P.; Lahtz, C.; Enk, A.; Herrmann-Trost, P.; Marsch, W.; Kutzner, H.; Dammann, R.H. Frequent occurrence of RASSF1A promoter hypermethylation and Merkel cell polyomavirus in Merkel cell carcinoma. Mol. Carcinog. 2009, 48, 903–909. [Google Scholar] [CrossRef]
- Martel-Jantin, C.; Filippone, C.; Cassar, O.; Peter, M.; Tomasic, G.; Vielh, P.; Briere, J.; Petrella, T.; Aubriot-Lorton, M.H.; Mortier, L.; et al. Genetic variability and integration of Merkel cell polyomavirus in Merkel cell carcinoma. Virology 2012, 426, 134–142. [Google Scholar] [CrossRef]
- Martel-Jantin, C.; Pedergnana, V.; Nicol, J.T.; Leblond, V.; Tregouet, D.A.; Tortevoye, P.; Plancoulaine, S.; Coursaget, P.; Touze, A.; Abel, L.; et al. Merkel cell polyomavirus infection occurs during early childhood and is transmitted between siblings. J. Clin. Virol. 2013, 58, 288–291. [Google Scholar] [CrossRef]
- Helmbold, P.; Lahtz, C.; Herpel, E.; Schnabel, P.A.; Dammann, R.H. Frequent hypermethylation of RASSF1A tumour suppressor gene promoter and presence of Merkel cell polyomavirus in small cell lung cancer. Eur. J. Cancer 2009, 45, 2207–2211. [Google Scholar] [CrossRef]
- Lasithiotaki, I.; Antoniou, K.M.; Derdas, S.P.; Sarchianaki, E.; Symvoulakis, E.K.; Psaraki, A.; Spandidos, D.A.; Stathopoulos, E.N.; Siafakas, N.M.; Sourvinos, G. The presence of Merkel cell polyomavirus is associated with deregulated expression of Braf and Bcl-2 genes in non-small cell lung cancer. Int. J. Cancer 2013, 133, 604–611. [Google Scholar] [CrossRef]
- Houben, R.; Michel, B.; Vetter-Kauczok, C.S.; Pfohler, C.; Laetsch, B.; Wolter, M.D.; Leonard, J.H.; Trefzer, U.; Ugurel, S.; Schrama, D.; et al. Absence of classical map kinase pathway signalling in Merkel cell carcinoma. J. Invest. Dermatol. 2006, 126, 1135–1142. [Google Scholar] [CrossRef]
- Van Gele, M.; Kaghad, M.; Leonard, J.H.; van Roy, N.; Naeyaert, J.M.; Geerts, M.L.; van Belle, S.; Cocquyt, V.; Bridge, J.; Sciot, R.; et al. Mutation analysis of P73 and TP53 in Merkel cell carcinoma. Br. J. Cancer 2000, 82, 823–826. [Google Scholar] [CrossRef]
- Lassacher, A.; Heitzer, E.; Kerl, H.; Wolf, P. p14ARF hypermethylation is common but INK4a-ARF locus or p53 mutations are rare in Merkel cell carcinoma. J. Invest. Dermatol. 2008, 128, 1788–1796. [Google Scholar] [CrossRef]
- Taberlay, P.C.; Jones, P.A. DNA methylation and cancer. Fortschr. Arzneimittelforsch. 2011, 67, 1–23. [Google Scholar]
- Bergman, Y.; Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 2013, 20, 274–281. [Google Scholar] [CrossRef]
- Richter, A.M.; Pfeifer, G.P.; Dammann, R.H. The RASSF proteins in cancer; from epigenetic silencing to functional characterization. Biochim. Biophys. Acta 2009, 1796, 114–128. [Google Scholar]
- Endoh, M.; Tamura, G.; Honda, T.; Homma, N.; Terashima, M.; Nishizuka, S.; Motoyama, T. RASSF2, a potential tumour suppressor, is silenced by CpG island hypermethylation in gastric cancer. Br. J. Cancer 2005, 93, 1395–1399. [Google Scholar] [CrossRef]
- Schagdarsurengin, U.; Richter, A.M.; Hornung, J.; Lange, C.; Steinmann, K.; Dammann, R.H. Frequent epigenetic inactivation of RASSF2 in thyroid cancer and functional consequences. Mol. Cancer 2010, 9, e264. [Google Scholar] [CrossRef]
- Zhang, Z.; Sun, D.; van do, N.; Tang, A.; Hu, L.; Huang, G. Inactivation of RASSF2A by promoter methylation correlates with lymph node metastasis in nasopharyngeal carcinoma. Int. J. Cancer 2007, 120, 32–38. [Google Scholar] [CrossRef]
- Hesson, L.; Dallol, A.; Minna, J.D.; Maher, E.R.; Latif, F. NORE1A, a homologue of RASSF1A tumour suppressor gene is inactivated in human cancers. Oncogene 2003, 22, 947–954. [Google Scholar] [CrossRef]
- Macheiner, D.; Heller, G.; Kappel, S.; Bichler, C.; Stattner, S.; Ziegler, B.; Kandioler, D.; Wrba, F.; Schulte-Hermann, R.; Zochbauer-Muller, S.; et al. NORE1B, a candidate tumor suppressor, is epigenetically silenced in human hepatocellular carcinoma. J. Hepatol. 2006, 45, 81–89. [Google Scholar] [CrossRef]
- Schagdarsurengin, U.; Richter, A.M.; Wohler, C.; Dammann, R.H. Frequent epigenetic inactivation of RASSF10 in thyroid cancer. Epigenetics 2009, 4, 571–576. [Google Scholar] [CrossRef]
- Richter, A.M.; Walesch, S.K.; Würl, P.; Taubert, H.; Dammann, R.H. The tumor suppressor RASSF10 is upregulated upon contact inhibition and frequently epigenetically silenced in cancer. Oncogenesis 2012, 1, e18. [Google Scholar] [CrossRef]
- Helmbold, P.; Richter, A.M.; Walesch, S.K.; Skorokhod, A.; Marsch, W.C.; Enk, A.; Dammann, R.H. RASSF10 promoter hypermethylation is frequent in malignant melanoma of the skin but uncommon in nevus cell nevi. J. Invest. Dermatol. 2012, 132, 687–694. [Google Scholar] [CrossRef]
- Hill, V.K.; Underhill-Day, N.; Krex, D.; Robel, K.; Sangan, C.B.; Summersgill, H.R.; Morris, M.; Gentle, D.; Chalmers, A.D.; Maher, E.R.; et al. Epigenetic inactivation of the RASSF10 candidate tumor suppressor gene is a frequent and an early event in gliomagenesis. Oncogene 2010, 30, 978–989. [Google Scholar]
- Hesson, L.B.; Dunwell, T.L.; Cooper, W.N.; Catchpoole, D.; Brini, A.T.; Chiaramonte, R.; Griffiths, M.; Chalmers, A.D.; Maher, E.R.; Latif, F. The novel RASSF6 and RASSF10 candidate tumour suppressor genes are frequently epigenetically inactivated in childhood leukaemias. Mol. Cancer 2009, 8, e42. [Google Scholar] [CrossRef] [Green Version]
- Shinawi, T.; Hill, V.; Dagklis, A.; Baliakas, P.; Stamatopoulos, K.; Agathanggelou, A.; Stankovic, T.; Maher, E.R.; Ghia, P.; Latif, F. KIBRA gene methylation is associated with unfavorable biological prognostic parameters in chronic lymphocytic leukemia. Epigenetics 2012, 7, 211–215. [Google Scholar] [CrossRef]
- Rice, P.; Longden, I.; Bleasby, A. EMBOSS: The European Molecular Biology Open Software Suite. Available online: http://www.ebi.ac.uk/Tools/seqstats/emboss_cpgplot/ (accessed on 30 April 2013).
- Underhill-Day, N.; Hill, V.; Latif, F. N-terminal RASSF family: RASSF7–RASSF10. Epigenetics 2011, 6, 284–292. [Google Scholar] [CrossRef]
- Vos, M.D.; Ellis, C.A.; Elam, C.; Ulku, A.S.; Taylor, B.J.; Clark, G.J. RASSF2 is a novel K-Ras-specific effector and potential tumor suppressor. J. Biol. Chem. 2003, 278, 28045–28051. [Google Scholar]
- Hesson, L.B.; Wilson, R.; Morton, D.; Adams, C.; Walker, M.; Maher, E.R.; Latif, F. CpG island promoter hypermethylation of a novel Ras-effector gene RASSF2A is an early event in colon carcinogenesis and correlates inversely with K-ras mutations. Oncogene 2005, 24, 3987–3994. [Google Scholar] [CrossRef]
- Akino, K.; Toyota, M.; Suzuki, H.; Mita, H.; Sasaki, Y.; Ohe-Toyota, M.; Issa, J.P.; Hinoda, Y.; Imai, K.; Tokino, T. The Ras effector RASSF2 is a novel tumor-suppressor gene in human colorectal cancer. Gastroenterology 2005, 129, 156–169. [Google Scholar] [CrossRef]
- Park, H.W.; Kang, H.C.; Kim, I.J.; Jang, S.G.; Kim, K.; Yoon, H.J.; Jeong, S.Y.; Park, J.G. Correlation between hypermethylation of the RASSF2A promoter and K-ras/BRAF mutations in microsatellite-stable colorectal cancers. Int. J. Cancer 2007, 120, 7–12. [Google Scholar] [CrossRef]
- Tommasi, S.; Dammann, R.; Jin, S.G.; Zhang, X.F.; Avruch, J.; Pfeifer, G.P. RASSF3 and NORE1: Identification and cloning of two human homologues of the putative tumor suppressor gene RASSF1. Oncogene 2002, 21, 2713–2720. [Google Scholar] [CrossRef]
- Chen, J.; Lui, W.O.; Vos, M.D.; Clark, G.J.; Takahashi, M.; Schoumans, J.; Khoo, S.K.; Petillo, D.; Lavery, T.; Sugimura, J.; et al. The t(1;3) breakpoint-spanning genes LSAMP and NORE1 are involved in clear cell renal cell carcinomas. Cancer Cell 2003, 4, 405–413. [Google Scholar] [CrossRef]
- Aoyama, Y.; Avruch, J.; Zhang, X.F. Nore1 inhibits tumor cell growth independent of Ras or the MST1/2 kinases. Oncogene 2004, 23, 3426–3433. [Google Scholar] [CrossRef]
- Vos, M.D.; Martinez, A.; Ellis, C.A.; Vallecorsa, T.; Clark, G.J. The pro-apoptotic Ras effector Nore1 may serve as a Ras-regulated tumor suppressor in the lung. J. Biol. Chem. 2003, 278, 21938–21943. [Google Scholar]
- Song, H.; Kim, H.; Lee, K.; Lee, D.H.; Kim, T.S.; Song, J.Y.; Lee, D.; Choi, D.; Ko, C.Y.; Kim, H.S.; et al. Ablation of RASSF2 induces bone defects and subsequent haematopoietic anomalies in mice. EMBO J. 2012, 31, 1147–1159. [Google Scholar] [CrossRef]
- Moshnikova, A.; Frye, J.; Shay, J.W.; Minna, J.D.; Khokhlatchev, A.V. The growth and tumor suppressor NORE1A is a cytoskeletal protein that suppresses growth by inhibition of the ERK pathway. J. Biol. Chem. 2006, 281, 8143–8152. [Google Scholar]
- Katagiri, K.; Ohnishi, N.; Kabashima, K.; Iyoda, T.; Takeda, N.; Shinkai, Y.; Inaba, K.; Kinashi, T. Crucial functions of the Rap1 effector molecule RAPL in lymphocyte and dendritic cell trafficking. Nat. Immunol. 2004, 5, 1045–1051. [Google Scholar] [CrossRef]
- Wei, Z.; Chen, X.; Chen, J.; Wang, W.; Xu, X.; Cai, Q. RASSF10 is epigenetically silenced and functions as a tumor suppressor in gastric cancer. Biochem. Biophys. Res. Commun. 2013, 432, 632–637. [Google Scholar] [CrossRef]
- Dansranjavin, T.; Wagenlehner, F.; Gattenloehner, S.; Steger, K.; Weidner, W.; Dammann, R.; Schagdarsurengin, U. Epigenetic down regulation of RASSF10 and its possible clinical implication in prostate carcinoma. Prostate 2012. [Google Scholar] [CrossRef]
- Dumitrescu, R.G. Epigenetic markers of early tumor development. Methods Mol. Biol. 2012, 863, 3–14. [Google Scholar] [CrossRef]
Supplementary Files
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Richter, A.M.; Haag, T.; Walesch, S.; Herrmann-Trost, P.; Marsch, W.C.; Kutzner, H.; Helmbold, P.; Dammann, R.H. Aberrant Promoter Hypermethylation of RASSF Family Members in Merkel Cell Carcinoma. Cancers 2013, 5, 1566-1576. https://doi.org/10.3390/cancers5041566
Richter AM, Haag T, Walesch S, Herrmann-Trost P, Marsch WC, Kutzner H, Helmbold P, Dammann RH. Aberrant Promoter Hypermethylation of RASSF Family Members in Merkel Cell Carcinoma. Cancers. 2013; 5(4):1566-1576. https://doi.org/10.3390/cancers5041566
Chicago/Turabian StyleRichter, Antje M., Tanja Haag, Sara Walesch, Peter Herrmann-Trost, Wolfgang C. Marsch, Heinz Kutzner, Peter Helmbold, and Reinhard H. Dammann. 2013. "Aberrant Promoter Hypermethylation of RASSF Family Members in Merkel Cell Carcinoma" Cancers 5, no. 4: 1566-1576. https://doi.org/10.3390/cancers5041566





