Nanomaterials and Autophagy: New Insights in Cancer Treatment
Abstract
:1. Introduction
2. Nanomedicine, Nanomaterials and Application in Cancer
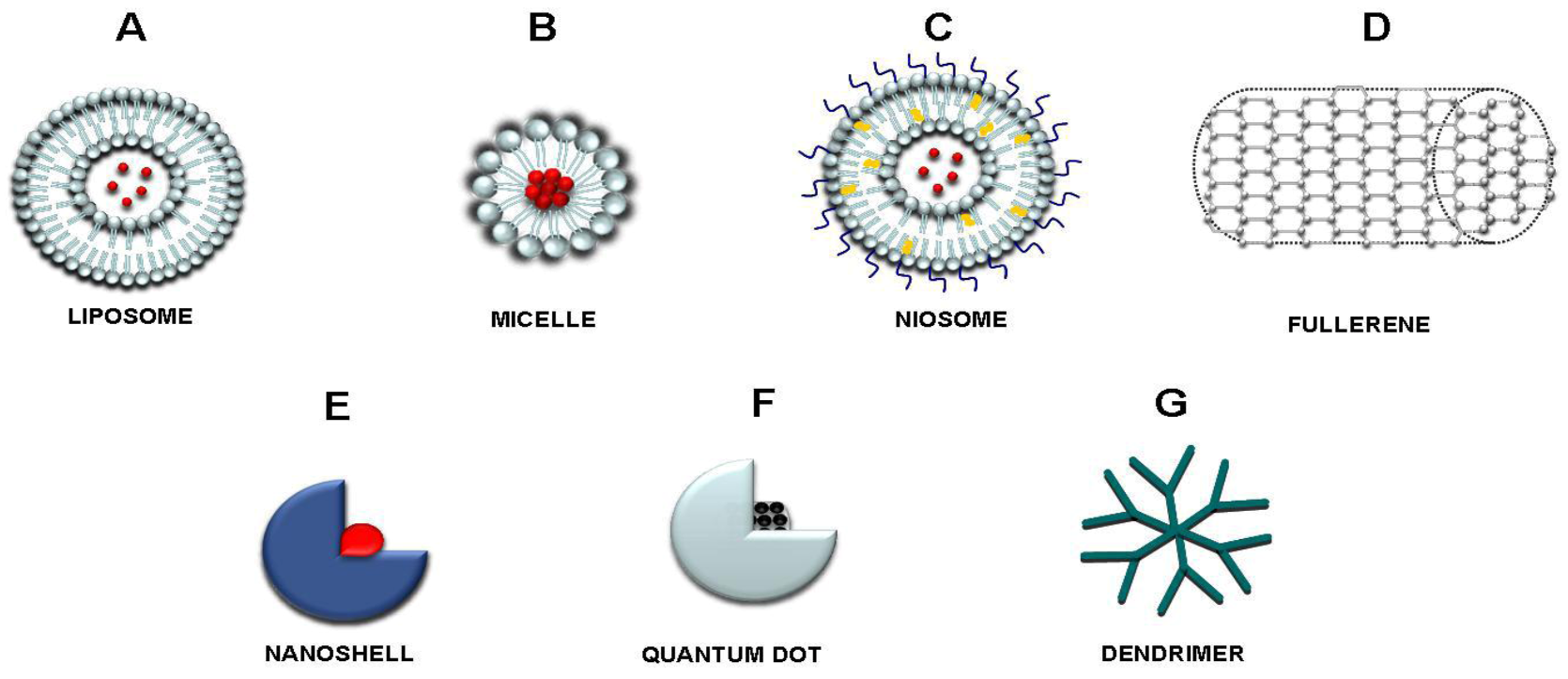
| Delivered drugs | Nanomaterials | Reference |
|---|---|---|
| Camptotechin | Si-NPs | [15] |
| Kahalalide F | Au-NPs | [16] |
| Docetaxel | Zn-NPs | [17] |
| Gemcitabine | immunoliposomes | [18] |
| Paclitaxel | cationic liposomes | [19] |
| Paclitaxel | tetrahexyloxy-tetra-paminocalix[4]arene (A4C6) | [20] |
| Doxorubicin | polymalic acid backbone | [21] |
| Doxorubucin | PEG liposomes | [22] |
| Doxorubucin | PEG-LPD(liposomes polycation DNA) | [23] |
| Myocet (doxorubicin) | non PEG liposomes | [23] |
| DaunoXome (daunorubicin) | unilamellar liposomes | [24] |
| Cisplatin | MWCNTs | [25] |
| Gemcitabine | SWCNTs | [26] |
| Doxorubicin | MWCNTs@poly(ethylene glycol-b-propilene sulphide) | [27] |
| Cisplatin-EGF | SWNTs | [28] |
| Gemcitabine | Fe3O4@poly(ethylene glycol)-NPs | [29] |
| Doxorubicin | Fe3O4@gelatin-NPs | [30] |
| 5-Fluorouracil | Fe3O4@ethylcellulose-NPs | [31] |
| Daunorubicin | Fe3O4-NPs | [32] |
| Cisplatin | Fe3O4@poly ε-caprolactone-NPs | [33] |
| Paclitaxel | Fe3O4@poly[aniline-co-sodium N-(1-butyric acid)]-NPs | [34] |
| 1,3-Bis(2-chloroethyl)-1-nitrosourea (BCNU) | Fe3O4@poly[aniline-co-N-(1-butyric acid) aniline] | [35] |
2.1. Nanosafety
3. Autophagy-Assisted Cancer Therapy: Old and New Insights
3.1. Autophagy and Cancer
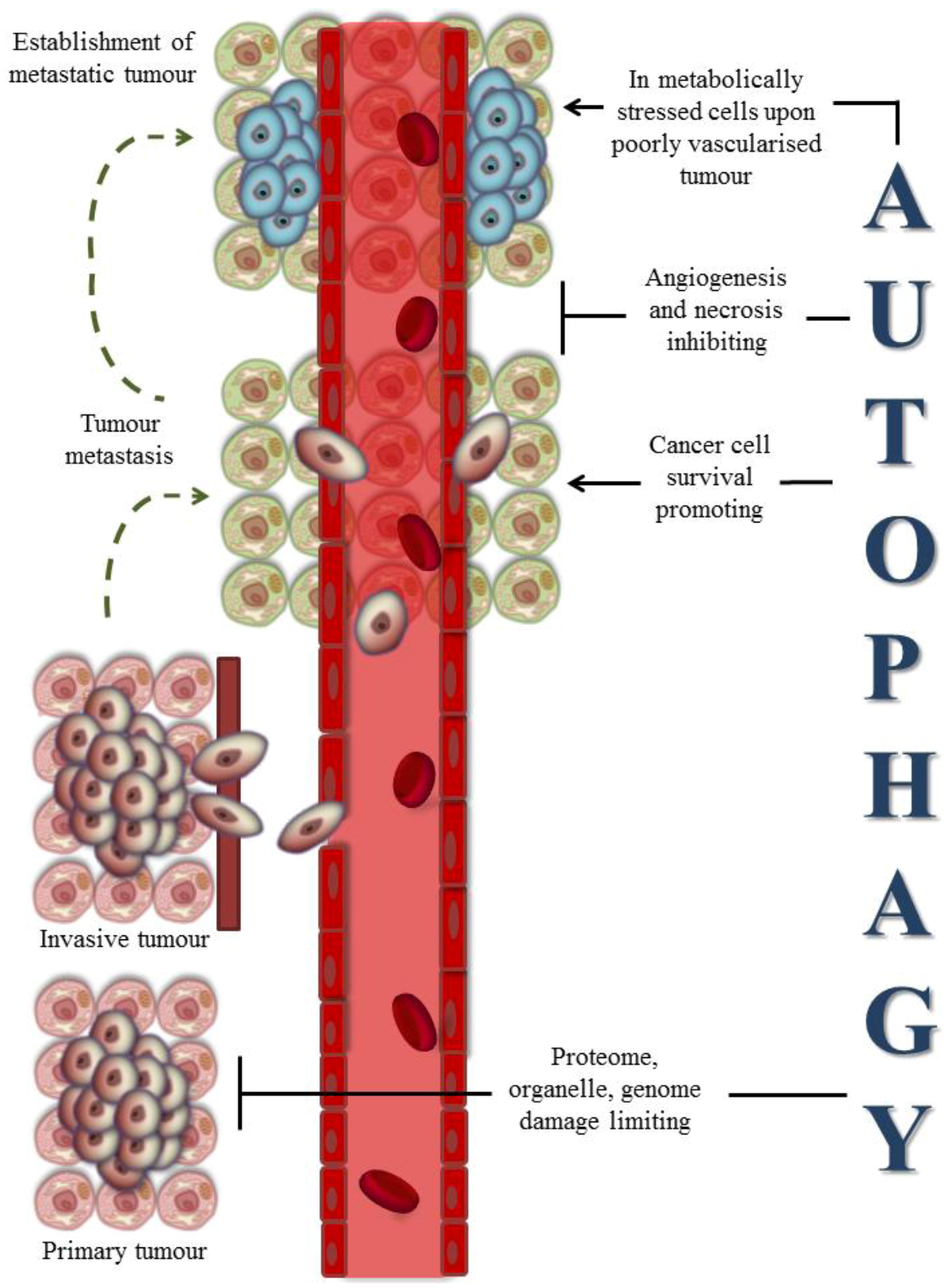
3.2. Autophagy and Cancer Therapies
3.3. Nanomaterials-Induced Autophagy: A Tool in Cancer Therapy?
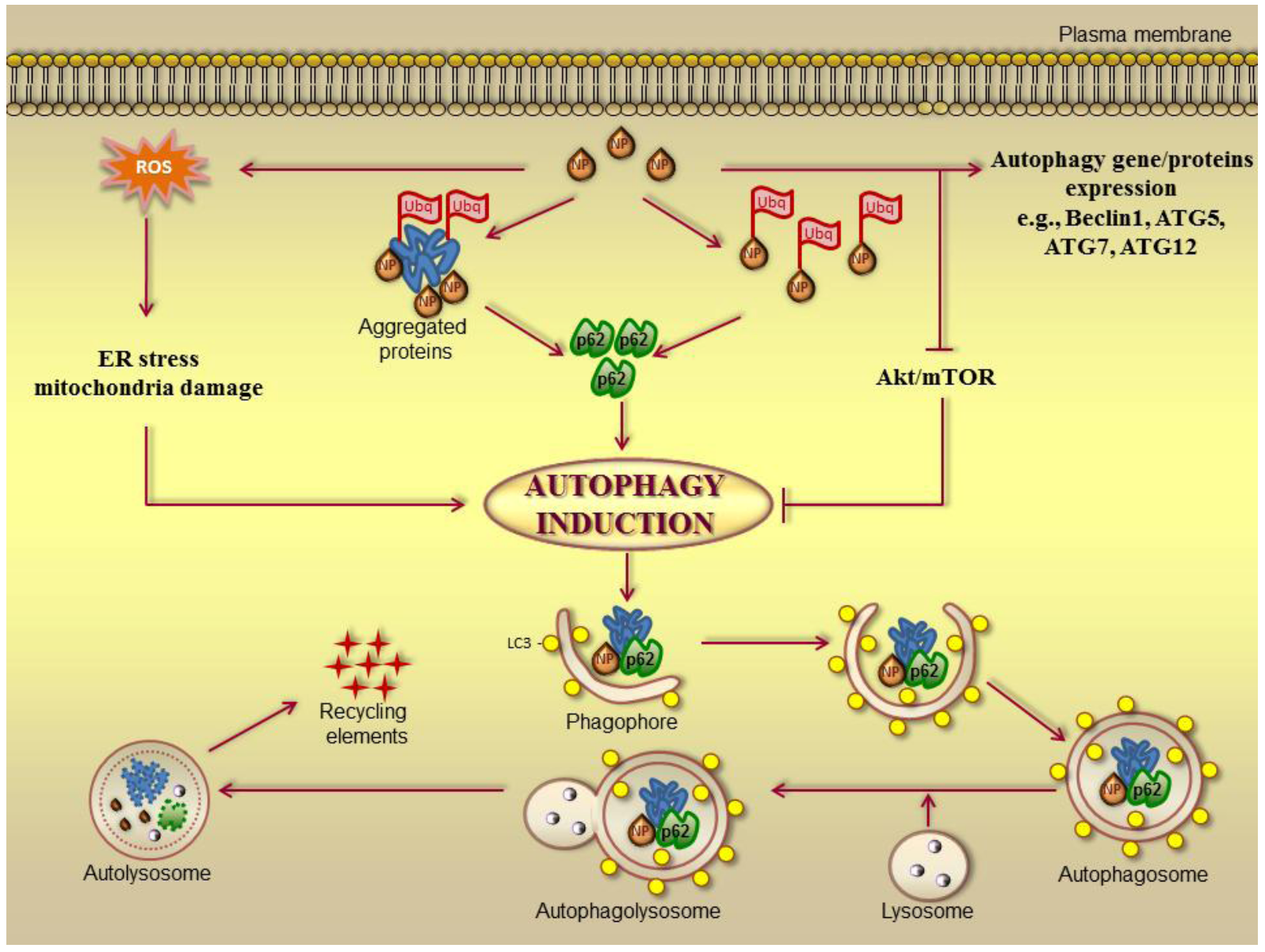
| Nanomaterials | Model | Delivered molecule | Reference |
|---|---|---|---|
| GNP-Chl | MCF-7 human breast cancer cells | Chloroquine | [155] |
| Fe@Au-NPs | OEMC1 human oral cancer cells | - | [146] |
| Magnetic NPs (C225-NPs) | NSCLC non-small cell lung cancer cells | Anti-EGFR antibody | [144] |
| C60(Nd)-NPs | HeLa cervix cancer cells MCF-7 cells | doxorubicin | [156] |
| nC60 | C6 rat glioma cells U251 human glioma cells | - | [145] |
| FeO-NPs | A549 human lung epithelial cancer cells | - | [147] |
| nC60 | HeLa cervix cancer cells MCF-7 cells | Doxorubicin | [151] |
| α-Al2O3-NPs | C57BL/6 with 3LL lung tumour | Antigen tumour derived | [141] |
4. Conclusions
References
- Jain, K.K. Nanotechnology in clinical laboratory diagnostics. Clin. Chim. Acta 2005, 358, 37–54. [Google Scholar] [CrossRef]
- ASTM E2456 - 06 Standard Terminology Relating to Nanotechnology. ASTM International, 2006.
- Nevozhay, D.; Kańska, U.; Budzyńska, R.; Boratyński, J. Current status of research on conjugates and related drug delivery systems in the treatment of cancer and other diseases. Postepy Hig. Med. Dosw. 2007, 61, 350–360. [Google Scholar]
- Ai, J.; Biazar, E.; Jafarpour, M.; Montazeri, M.; Majdi, A.; Aminifard, S.; Zafari, M.; Akbari, H.R.; Rad, H.G. Nanotoxicology and nanoparticle safety in biomedical designs. Int. J. Nanomedicine 2011, 6, 1117–1127. [Google Scholar]
- Duncan, R. The dawning era of polymer therapeutics. Nat. Rev. Drug Discov. 2003, 2, 347–360. [Google Scholar] [CrossRef]
- Torchilin, V.P. Recent advances with liposomes as pharmaceutical carriers. Nat. Rev. Drug Discov. 2005, 4, 145–160. [Google Scholar] [CrossRef]
- Blanco, E.; Kessinger, C.W.; Sumer, B.D.; Gao, J. Multifunctional micellar nanomedicine for cancer therapy. Exp. Biol. Med. 2009, 234, 123–131. [Google Scholar]
- Sankhyan, A.; Pawar, P. Recent trends in noisome as vesicular drug delivery system. J. Appl. Pharm. Sci. 2012, 2, 20–32. [Google Scholar]
- Sanvicens, N.; Marco, M.P. Multifunctional nanoparticles—Properties and prospects for their use in human medicine. Trends Biotechnol. 2008, 26, 425–433. [Google Scholar] [CrossRef]
- Svenson, S.; Tomalia, D.A. Dendrimers in biomedical applications-reflections on the field. Adv. Drug Deliv. Rev. 2005, 57, 2106–2129. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, B.; Qiao, W.; Yin, T. A novel controlled release drug delivery system for multiple drugs based on electrospun nanofibers containing nanoparticles. J. Pharm Sci. 2010, 99, 4805–4811. [Google Scholar] [CrossRef]
- Ferrari, M. Cancer nanotechnology: Opportunities and challenges. Nat. Rev. Cancer 2005, 5, 161–171. [Google Scholar] [CrossRef]
- Peer, D.; Karp, J.M.; Hong, S.; Farokhzad, O.C.; Margalit, R.; Langer, R. Nanocarriers as an emerging platform for cancer therapy. Nat. Nanotechnol. 2007, 2, 751–760. [Google Scholar]
- Heath, J.R.; Davis, M.E. Nanotechnology and cancer. Annu. Rev. Med. 2008, 59, 251–265. [Google Scholar] [CrossRef]
- Botella, P.; Abasolo, I.; Fernández, Y.; Muniesa, C.; Miranda, S.; Quesada, M.; Ruiz, J.; Schwartz, S., Jr.; Corma, A. Surface-modified silica nanoparticles for tumor-targeted delivery of camptothecin and its biological evaluation. J. Control. Release 2011, 156, 246–257. [Google Scholar] [CrossRef]
- Hosta, L.; Pla-Roca, M.; Arbiol, J.; López-Iglesias, C.; Samitier, J.; Cruz, L.J.; Kogan, M.J.; Albericio, F. Conjugation of Kahalalide F with gold nanoparticles to enhance in vitro antitumoral activity. Bioconjug. Chem. 2009, 20, 138–146. [Google Scholar] [CrossRef]
- Conte, C.; Ungaro, F.; Maglio, G.; Tirino, P.; Siracusano, G.; Sciortino, M.T.; Leone, N.; Palma, G.; Barbieri, A.; Arra, C.; et al. Biodegradable core-shell nanoassemblies for the delivery of docetaxel and Zn(II)-phthalocyanine inspired by combination therapy for cancer. J. Control. Release 2013, 10, 40–52. [Google Scholar]
- Kim, I.Y.; Kang, Y.S.; Lee, D.S.; Park, H.J.; Choi, E.K.; Oh, Y.K.; Son, H.J.; Kim, J.S. Antitumor activity of EGFR targeted pH-sensitive immunoliposomes encapsulating gemcitabine in A549 xenograft nude mice. J. Control. Release 2009, 140, 55–60. [Google Scholar] [CrossRef]
- Strieth, S.; Eichhorn, M.E.; Werner, A.; Sauer, B.; Teifel, M.; Michaelis, U.; Berghaus, A.; Dellian, M. Paclitaxel encapsulated in cationic liposomes increases tumor microvessel leakiness and improves therapeutic efficacy in combination with Cisplatin. Clin. Cancer Res. 2008, 14, 4603–4611. [Google Scholar] [CrossRef]
- Weeden, C.; Hartlieb, K.J.; Lim, L.Y. Preparation and physicochemical characterization of a novel paclitaxel-loaded amphiphilic aminocalixarene nanoparticle platform for anticancer chemotherapy. J. Pharm. Pharmacol. 2012, 64, 1403–1411. [Google Scholar] [CrossRef]
- Patil, R.; Portilla-Arias, J.; Ding, H.; Konda, B.; Rekechenetskiy, A.; Inoue, S.; Black, K.L.; Holler, E.; Ljubimova, J.Y. Cellular delivery of doxorubicin via pH-controlled hydrazone linkage using multifunctional nano vehicle based on poly(β-L-malic acid). Int. J. Mol. Sci. 2012, 13, 11681–11693. [Google Scholar] [CrossRef]
- Hatakeyama, H.; Akita, H.; Ishida, E.; Hashimoto, K.; Kobayashi, H.; Aoki, T.; Yasuda, J.; Obata, K.; Kikuchi, H.; Ishida, T.; et al. Tumor targeting of doxorubicin by anti-MT1-MMP antibody-modified PEG liposomes. Int. J. Pharm. 2007, 342, 194–200. [Google Scholar] [CrossRef]
- Chen, Y.; Wu, J.J.; Huang, L. Nanoparticles targeted with NGR motif deliver c-myc siRNA and doxorubicin for anticancer therapy. Mol. Ther. 2010, 18, 828–834. [Google Scholar] [CrossRef]
- Hofheinz, R.D.; Gnad-Vogt, S.U.; Beyer, U.; Hochhaus, A. Liposomal encapsulated anti-cancer drugs. Anticancer Drugs 2005, 16, 691–707. [Google Scholar] [CrossRef]
- Tripisciano, C.; Costa, S.; Kalenczuk, R.J.; Borowiak-Palen, E. Cisplatin filled multiwalled carbon nanotubes—A novel molecular hybrid of anticancer drug container. Eur. Phys. J. B 2010, 75, 141–146. [Google Scholar] [CrossRef]
- Arsawang, U.; Saengsawang, O.; Rungrotmongkol, T.; Sornmee, P.; Wittayanarakul, K.; Remsungnen, T.; Hannongbua, S. How do carbon nanotubes serve as carriers for gemcitabine transport in a drug delivery system? J. Mol. Graph. Model. 2011, 29, 591–596. [Google Scholar] [CrossRef]
- Di Crescenzo, A.; Velluto, D.; Hubbell, J.A.; Fontana, A. Biocompatible dispersions of carbon nanotubes: A potential tool for intracellular transport of anticancer drugs. Nanoscale 2011, 3, 925–928. [Google Scholar] [CrossRef]
- Bhirde, A.A.; Patel, V.; Gavard, J.; Zhang, G.; Sousa, A.A.; Masedunskas, A.; Leapman, R.D.; Weigert, R.; Gutkind, J.S.; Rusling, J.F. Targeted killing of cancer cells in vivo and in vitro with EGF-directed carbon nanotube-based drug delivery. ACS Nano 2009, 3, 307–316. [Google Scholar] [CrossRef]
- Tong, Q.; Li, H.; Li, W.; Chen, H.; Shu, X.; Lu, X.; Wang, G. In vitro and in vivo anti-tumor effects of gemcitabine loaded with a new drug delivery system. J. Nanosci. Nanotechnol. 2011, 11, 3651–3658. [Google Scholar] [CrossRef]
- Gaihre, B.; Khil, M.S.; Lee, D.R.; Kim, H.Y. Gelatin-coated magnetic iron oxide nanoparticles as carrier system: Drug loading and in vitro drug release study. Int. J. Pharm. 2009, 365, 180–189. [Google Scholar] [CrossRef]
- Arias, J.L.; López-Viota, M.; Delgado, A.V.; Ruiz, M.A. Iron/ethylcellulose (core/shell) nanoplatform loaded with 5-fluorouracil for cancer targeting. Colloids Surf. B Biointerfaces 2010, 77, 111–116. [Google Scholar] [CrossRef]
- Wu, W.; Chen, B.; Cheng, J.; Wang, J.; Xu, W.; Liu, L.; Xia, G.; Wei, H.; Wang, X.; Yang, M.; et al. Biocompatibility of Fe3O4/DNR magnetic nanoparticles in the treatment of hematologic malignancies. Int. J. Nanomedicine 2010, 5, 1079–1084. [Google Scholar]
- Yang, J.; Park, S.B.; Yoon, H.G.; Huh, Y.M.; Haam, S. Preparation of poly epsilon-caprolactone nanoparticles containing magnetite for magnetic drug carrier. Int. J. Pharm. 2006, 324, 185–190. [Google Scholar] [CrossRef]
- Hua, M.Y.; Yang, H.W.; Chuang, C.K.; Tsai, R.Y.; Chen, W.J.; Chuang, K.L.; Chang, Y.H.; Chuang, H.C.; Pang, S.T. Magnetic-nanoparticle-modified paclitaxel for targeted therapy for prostate cancer. Biomaterials 2010, 31, 7355–7363. [Google Scholar] [CrossRef]
- Hua, M.Y.; Liu, H.L.; Yang, H.W.; Chen, P.Y.; Tsai, R.Y.; Huang, C.Y.; Tseng, I.C.; Lyu, L.A.; Ma, C.C.; Tang, H.J.; et al. The effectiveness of a magnetic nanoparticle-based delivery system for BCNU in the treatment of gliomas. Biomaterials 2011, 32, 516–527. [Google Scholar] [CrossRef]
- Agostinis, P.; Berg, K.; Cengel, K.A.; Foster, T.H.; Girotti, A.W.; Gollnick, S.O.; Hahn, S.M.; Hamblin, M.R.; Juzeniene, A.; Kessel, D.; et al. Photodynamic therapy of cancer: An update. CA Cancer J. Clin. 2011, 61, 250–281. [Google Scholar] [CrossRef]
- Verma, S.; Watt, G.M.; Mai, Z.; Hasan, T. Strategies for enhanced photodynamic therapy effects. Photochem. Photobiol. 2007, 83, 996–1005. [Google Scholar] [CrossRef]
- Markovic, Z.M.; Ristic, B.Z.; Arsikin, K.M.; Klisic, D.G.; Harhaji-Trajkovic, L.M.; Todorovic-Markovic, B.M.; Kepic, D.P.; Kravic-Stevovic, T.K.; Jovanovic, S.P.; Milenkovic, M.M.; Milivojevic, D.D.; et al. Graphene quantum dots as autophagy-inducing photodynamic agents. Biomaterials 2012, 33, 7084–7092. [Google Scholar] [CrossRef]
- Lkhagvadulam, B.; Kim, J.H.; Yoon, I.; Shim, Y.K. Size-dependent photodynamic activity of gold nanoparticles conjugate of water soluble purpurin-18-N-methyl-d-glucamine. Int. J. Biomed. Biotech. 2013, Article ID 720579. [Google Scholar]
- Allard, E.; Hindre, F.; Passirani, C.; Lemaire, L.; Lepareur, N.; Noiret, N.; Menei, P.; Benoit, J.P. 188Re-loaded lipid nanocapsules as a promising radiopharmaceutical carrier for internal radiotherapy of malignant gliomas. Eur. J. Nucl. Med. Mol. Imaging 2008, 35, 1838–1846. [Google Scholar] [CrossRef] [Green Version]
- Vanpouille-Box, C.; Lacoeuille, F.; Roux, J.; Aubé, C.; Garcion, E.; Lepareur, N.; Oberti, F.; Bouchet, F.; Noiret, N.; Garin, E.; et al. Lipid nanocapsules loaded with rhenium-188 reduce tumor progression in a rat hepatocellular carcinoma model. PLoS One 2011, 6, e16926. [Google Scholar]
- Hamdy, S.; Haddadi, A.; Hung, R.W.; Lavasanifar, A. Targeting dendritic cells with nano-particulate PLGA cancer vaccine formulations. Adv. Drug Deliv. Rev. 2011, 63, 943–955. [Google Scholar] [CrossRef]
- Uto, T.; Wang, X.; Sato, K.; Haraguchi, M.; Akagi, T.; Akashi, M.; Baba, M. Targeting of antigen to dendritic cells with poly(gamma-glutamic acid) nanoparticles induces antigen-specific humoral and cellular immunity. J. Immunol. 2007, 178, 2979–2986. [Google Scholar]
- Uto, T.; Akagi, T.; Toyama, M.; Nishi, Y.; Shima, F.; Akashi, M.; Baba, M. Comparative activity of biodegradable nanoparticles with aluminum adjuvants: Antigen uptake by dendritic cells and induction of immune response in mice. Immunol. Lett. 2011, 140, 36–43. [Google Scholar] [CrossRef]
- Uto, T.; Akagi, T.; Yoshinaga, K.; Toyama, M.; Akashi, M.; Baba, M. The induction of innate and adaptive immunity by biodegradable poly(γ-glutamic acid) nanoparticles via a TLR4 and MyD88 signaling pathway. Biomaterials 2011, 32, 5206–5212. [Google Scholar] [CrossRef]
- Vanpouille-Box, C.; Lacoeuille, F.; Belloche, C.; Lepareur, N.; Lemaire, L.; LeJeune, J.J.; Benoît, J.P.; Menei, P.; Couturier, O.F.; Garcion, E.; et al. Tumor eradication in rat glioma and bypass of immunosuppressive barriers using internal radiation with (188)Re-lipid nanocapsules. Biomaterials 2011, 32, 6781–6790. [Google Scholar] [CrossRef] [Green Version]
- Somia, N.; Verma, I.M. Gene therapy: trials and tribulations. Nat. Rev. Genet. 2000, 1, 91–99. [Google Scholar] [CrossRef]
- Pichon, C.; Billiet, L.; Midoux, P. Chemical vectors for gene delivery: Uptake and intracellular trafficking. Curr. Opin. Biotechnol. 2010, 21, 640–645. [Google Scholar] [CrossRef]
- Li, C.; Li, L.; Keates, A.C. Targeting cancer gene therapy with magnetic nanoparticles. Oncotarget 2012, 3, 365–370. [Google Scholar]
- Podila, R.; Brown, J.M. Toxicity of Engineered Nanomaterials: A Physicochemical Perspective. J. Biochem. Mol. Toxicol. 2012. [Google Scholar] [CrossRef]
- Li, N.; Xia, T.; Nel, A.E. The role of oxidative stress in ambient particulate matter-induced lung diseases and its implications in the toxicity of engineered nanoparticles. Free Radic. Biol. Med. 2008, 44, 1689–1699. [Google Scholar] [CrossRef]
- Becker, H.; Herzberg, F.; Schulte, A.; Kolossa-Gehring, M. The carcinogenic potential of nanomaterials, their release from products and options for regulating them. Int. J. Hyg. Environ. Health. 2011, 214, 231–238. [Google Scholar] [CrossRef]
- Magaye, R.; Zhao, J.; Bowman, L.; Ding, M. Genotoxicity and carcinogenicity of cobalt-, nickel- and copper-based nanoparticles. Exp. Ther. Med. 2012, 4, 551–561. [Google Scholar]
- Meng, J.; Xing, J.; Wang, Y.; Lu, J.; Zhao, Y.; Gao, X.; Wang, P.C.; Jia, L.; Liang, X. Epigenetic modulation of human breast cancer by metallofullerenol nanoparticles: In vivo treatment and in vitro analysis. Nanoscale 2011, 3, 4713–4719. [Google Scholar] [CrossRef]
- Yang, X.; Liu, J.; He, H.; Zhou, L.; Gong, C.; Wang, X.; Yang, L.; Yuan, J.; Huang, H.; He, L.; et al. SiO2 nanoparticles induce cytotoxicity and protein expression alteration in HaCaT cells. Part. Fibre Toxicol. 2010. [Google Scholar] [CrossRef]
- Chen, M.; von Mikecz, A. Formation of nucleoplasmic protein aggregates impairs nuclear function in response to SiO2 nanoparticles. Exp. Cell Res. 2005, 305, 51–62. [Google Scholar] [CrossRef]
- Gong, C.; Tao, G.; Yang, L.; Liu, J.; Liu, Q.; Zhuang, Z. SiO2 nanoparticles induce global genomic hypomethylation in HaCaT cells. Biochem. Biophys. Res. Commun. 2010, 397, 397–400. [Google Scholar] [CrossRef]
- Stern, S.T.; Adiseshaiah, P.P.; Crist, R.M. Autophagy and lysosomal dysfunction as emerging mechanisms of nanomaterial toxicity. Part. Fibre Toxicol. 2012. [Google Scholar] [CrossRef]
- Stern, S.T.; Johnson, D.N. Role for nanomaterial-autophagy interaction in neurodegenerative disease. Autophagy 2008, 4, 1097–1100. [Google Scholar]
- Carew, J.S.; Nawrocki, S.T.; Cleveland, J.L. Modulating autophagy for therapeutic benefit. Autophagy 2007, 3, 464–467. [Google Scholar]
- Itakura, E.; Mizushima, N. Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy 2010, 6, 764–776. [Google Scholar] [CrossRef]
- Mijaljica, D.; Prescott, M.; Devenish, R.J. Microautophagy in mammalian cells: Revisiting a 40-year-old conundrum. Autophagy 2011, 7, 673–682. [Google Scholar] [CrossRef]
- Li, W.; Yang, Q.; Mao, Z. Chaperone-mediated autophagy: Machinery, regulation and biological consequences. Cell. Mol. Life Sci. 2011, 68, 749–763. [Google Scholar] [CrossRef]
- Rabinowitz, J.D.; White, E. Autophagy and metabolism. Science 2010, 330, 1344–1348. [Google Scholar] [CrossRef]
- Nakatogawa, H.; Suzuki, K.; Kamada, Y.; Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 2009, 10, 458–467. [Google Scholar] [CrossRef]
- Eskelinen, E.L.; Reggiori, F.; Baba, M.; Kovács, A.L.; Seglen, P.O. Seeing is believing: The impact of electron microscopy on autophagy research. Autophagy 2011, 7, 935–956. [Google Scholar] [CrossRef]
- Klionsky, D.J.; Abdalla, F.C.; Abeliovich, H.; Abraham, R.T.; Acevedo-Arozena, A.; Adeli, K.; Agholme, L.; Agnello, M.; Agostinis, P.; Aguirre-Ghiso, J.A.; et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012, 8, 445–544. [Google Scholar] [CrossRef]
- Kroemer, G.; Mariño, G.; Levine, B. Autophagy and the integrated stress response. Mol. Cell 2010, 40, 280–293. [Google Scholar] [CrossRef]
- Inui, M.; Martello, G.; Piccolo, S. MicroRNA control of signal transduction. Nat. Rev. Mol. Cell Biol. 2010, 11, 252–263. [Google Scholar]
- Frankel, L.B.; Lund, A.H. MicroRNA regulation of autophagy. Carcinogenesis 2012, 33, 2018–2025. [Google Scholar] [CrossRef]
- Levine, B.; Kroemer, G. Autophagy in the pathogenesis of disease. Cell 2008, 132, 27–42. [Google Scholar] [CrossRef]
- Mathew, R.; White, E. Autophagy in tumorigenesis and energy metabolism: Friend by day, foe by night. Curr. Opin. Genet. Dev. 2011, 21, 113–119. [Google Scholar] [CrossRef]
- Yang, Z.J.; Chee, C.E.; Huang, S.; Sinicrope, F. Autophagy modulation for cancer therapy. Cancer Biol. Ther. 2011, 11, 169–176. [Google Scholar] [CrossRef]
- Maiuri, M.C.; Tasdemir, E.; Criollo, A.; Morselli, E.; Vicencio, J.M.; Carnuccio, R.; Kroemer, G. Control of autophagy by oncogenes and tumor suppressor genes. Cell Death Differ. 2009, 16, 87–93. [Google Scholar] [CrossRef]
- Zhou, S.; Zhao, L.; Kuang, M.; Zhang, B.; Liang, Z.; Yi, T.; Wei, Y.; Zhao, X. Autophagy in tumorigenesis and cancer therapy: Dr. Jekyll or Mr. Hyde? Cancer Lett. 2012, 323, 115–127. [Google Scholar] [CrossRef]
- Hoshino, I.; Matsubara, H. MicroRNAs in cancer diagnosis and therapy: From bench to bedside. Surg. Today 2012. [Google Scholar] [CrossRef]
- Wild, L.; Flanagan, J.M. Genome-wide hypomethylation in cancer may be a passive consequence of transformation. Biochim. Biophys. Acta 2010, 1806, 50–57. [Google Scholar]
- Jiang, Y.; Liu, S.; Chen, X.; Cao, Y.; Tao, Y. Genome-wide distribution of DNA methylation and DNA demethylation and related chromatin regulators in cancer. Biochim. Biophys. Acta 2012, 1835, 155–163. [Google Scholar]
- Feng, W.; Marquez, R.T.; Lu, Z.; Liu, J.; Lu, K.H.; Issa, J.P.; Fishman, D.M.; Yu, Y.; Bast, R.C. Imprinted tumor suppressor genes ARHI and PEG3 are the most frequently downregulated in human ovarian cancers by loss of heterozygosity and promoter methylation. Cancer 2008, 112, 1489–1502. [Google Scholar] [CrossRef]
- Tschan, M.P.; Jost, M.; Batliner, J.; Fey, M.F. The autophagy gene ULK1 plays a role in AML differentiation and is negatively regulated by the oncogenic MicroRNA-106a. In 52nd Annual Meeting and Exposition, Lausanne, Switzerland, 11–13 May 2011.
- Miller, T.E.; Ghoshal, K.; Ramaswamy, B.; Roy, S.; Datta, J.; Shapiro, C.L.; Jacob, S.; Majumder, S. MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J. Biol. Chem. 2008, 283, 29897–29903. [Google Scholar]
- Korkmaz, G.; le Sage, C.; Tekirdag, K.A.; Agami, R.; Gozuacik, D. miR-376b controls starvation and mTOR inhibition-related autophagy by targeting ATG4C and BECN1. Autophagy 2012, 8, 65–76. [Google Scholar]
- Abraham, D.; Jackson, N.; Gundara, J.S.; Zhao, J.; Gill, A.J.; Delbridge, L.; Robinson, B.G.; Sidhu, S.B. MicroRNA profiling of sporadic and hereditary medullary thyroid cancer identifies predictors of nodal metastasis, prognosis, and potential therapeutic targets. Clin. Cancer Res. 2011, 17, 4772–4781. [Google Scholar] [CrossRef]
- Zhu, H.; Wu, H.; Liu, X.; Li, B.; Chen, Y.; Ren, X.; Liu, C.G.; Yang, J.M. Regulation of autophagy by a beclin 1-targeted microRNA, miR-30a, in cancer cells. Autophagy 2009, 5, 816–823. [Google Scholar]
- Frankel, L.B.; Wen, J.; Lees, M.; Høyer-Hansen, M.; Farkas, T.; Krogh, A.; Jäättelä, M.; Lund, A.H. microRNA-101 is a potent inhibitor of autophagy. EMBO J. 2011, 30, 4628–4641. [Google Scholar] [CrossRef]
- Lisanti, M.P.; Martinez-Outschoorn, U.E.; Chiavarina, B.; Pavlides, S.; Whitaker-Menezes, D.; Tsirigos, A.; Witkiewicz, A.; Lin, Z.; Balliet, R.; Howell, A.; et al. Understanding the “lethal” drivers of tumor-stroma co-evolution: emerging role(s) for hypoxia, oxidative stress and autophagy/mitophagy in the tumor micro-environment. Cancer Biol. Ther. 2010, 10, 537–542. [Google Scholar] [CrossRef]
- Morselli, E.; Galluzzi, L.; Kepp, O.; Vicencio, J.M.; Criollo, A.; Maiuri, M.C.; Kroemer, G. Anti- and pro-tumor functions of autophagy. Biochim. Biophys. Acta 2009, 1793, 1524–1532. [Google Scholar] [CrossRef]
- Levine, B.; Abrams, J. p53: The Janus of autophagy? Nat. Cell Biol. 2008, 10, 637–639. [Google Scholar] [CrossRef]
- Nicholson, K.M.; Anderson, N.G. The protein kinase B/Akt signalling pathway in human malignancy. Cell Signal. 2002, 14, 381–395. [Google Scholar] [CrossRef]
- Mills, K.R.; Reginato, M.; Debnath, J.; Queenan, B.; Brugge, J.S. Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) is required for induction of autophagy during lumen formation in vitro. Proc. Natl. Acad. Sci. USA 2004, 101, 3438–3443. [Google Scholar]
- Prins, J.; Ledgerwood, E.; Ameloot, P.; Vandenabeele, P.; Faraco, P.; Bright, N.; O’Rahilly, S.; Bradley, J. Tumour necrosis factor induced autophagy and mitochondrial morphological abnormalities are mediated by TNFR-I and/or TNFR-II and do not invariably lead to cell death. Biochem. Soc. Trans. 1998, 26, S314. [Google Scholar]
- Maiuri, M.C.; Criollo, A.; Kroemer, G. Crosstalk between apoptosis and autophagy within the Beclin 1 interactome. EMBO J. 2010, 29, 515–516. [Google Scholar] [CrossRef]
- Han, J.; Hou, W.; Goldstein, L.A.; Lu, C.; Stolz, D.B.; Yin, X.M.; Rabinowich, H. Involvement of protective autophagy in TRAIL resistance of apoptosis-defective tumor cells. J. Biol. Chem. 2008, 283, 19665–19677. [Google Scholar]
- Thorburn, J.; Moore, F.; Rao, A.; Barclay, W.W.; Thomas, L.R.; Grant, K.W.; Cramer, S.D.; Thorburn, A. Selective inactivation of a Fas-associated death domain protein (FADD)-dependent apoptosis and autophagy pathway in immortal epithelial cells. Mol. Biol. Cell. 2005, 16, 1189–1199. [Google Scholar] [CrossRef]
- Rikiishi, H. Novel insights into the interplay between apoptosis and autophagy. Int. J. Cell Biol. 2012. [Google Scholar] [CrossRef]
- Mathew, R.; Karantza-Wadsworth, V.; White, E. Role of autophagy in cancer. Nat. Rev. Cancer 2007, 7, 961–967. [Google Scholar] [CrossRef]
- Amaravadi, R.K.; Lippincott-Schwartz, J.; Yin, X.M.; Weiss, W.A.; Takebe, N.; Timmer, W.; DiPaola, R.S.; Lotze, M.T.; White, E. Principles and current strategies for targeting autophagy for cancer treatment. Clin. Cancer Res. 2011, 17, 654–666. [Google Scholar] [CrossRef]
- Schoenlein, P.V.; Periyasamy-Thandavan, S.; Samaddar, J.S.; Jackson, W.H.; Barrett, J.T. Autophagy facilitates the progression of ERalpha-positive breast cancer cells to antiestrogen resistance. Autophagy 2009, 5, 400–403. [Google Scholar] [CrossRef]
- Vazquez-Martin, A.; Oliveras-Ferraros, C.; Menendez, J.A. Autophagy facilitates the development of breast cancer resistance to the anti-HER2 monoclonal antibody trastuzumab. PLoS One 2009, 4, e6251. [Google Scholar]
- Milani, M.; Rzymski, T.; Mellor, H.R.; Pike, L.; Bottini, A.; Generali, D.; Harris, A.L. The role of ATF4 stabilization and autophagy in resistance of breast cancer cells treated with Bortezomib. Cancer Res. 2009, 69, 4415–4423. [Google Scholar]
- Bauvy, C.; Gane, P.; Arico, S.; Codogno, P.; Ogier-Denis, E. Autophagy delays sulindac sulfide-induced apoptosis in the human intestinal colon cancer cell line HT-29. Exp. Cell Res. 2001, 268, 139–149. [Google Scholar] [CrossRef]
- Bellodi, C.; Lidonnici, M.R.; Hamilton, A.; Helgason, G.V.; Soliera, A.R.; Ronchetti, M.; Galavotti, S.; Young, K.W.; Selmi, T.; Yacobi, R.; et al. Targeting autophagy potentiates tyrosine kinase inhibitor-induced cell death in Philadelphia chromosome-positive cells, including primary CML stem cells. J. Clin. Invest. 2009, 119, 1109–1123. [Google Scholar] [CrossRef]
- Gupta, A.; Roy, S.; Lazar, A.J.; Wang, W.L.; McAuliffe, J.C.; Reynoso, D.; McMahon, J.; Taguchi, T.; Floris, G.; Debiec-Rychter, M.; et al. Autophagy inhibition and antimalarials promote cell death in gastrointestinal stromal tumor (GIST). Proc. Natl. Acad. Sci. USA 2010, 107, 14333–14338. [Google Scholar] [CrossRef]
- Shingu, T.; Fujiwara, K.; Bögler, O.; Akiyama, Y.; Moritake, K.; Shinojima, N.; Tamada, Y.; Yokoyama, T.; Kondo, S. Inhibition of autophagy at a late stage enhances imatinib-induced cytotoxicity in human malignant glioma cells. Int. J. Cancer 2009, 124, 1060–1071. [Google Scholar] [CrossRef]
- Wu, Z.; Chang, P.C.; Yang, J.C.; Chu, C.Y.; Wang, L.Y.; Chen, N.T.; Ma, A.H.; Desai, S.J.; Lo, S.H.; Evans, C.P.; et al. Autophagy blockade sensitizes prostate cancer cells towards src family kinase inhibitors. Genes Cancer 2010, 1, 40–49. [Google Scholar] [CrossRef]
- Carew, J.S.; Nawrocki, S.T.; Kahue, C.N.; Zhang, H.; Yang, C.; Chung, L.; Houghton, J.A.; Huang, P.; Giles, F.J.; Cleveland, J.L. Targeting autophagy augments the anticancer activity of the histone deacetylase inhibitor SAHA to overcome Bcr-Abl-mediated drug resistance. Blood 2007, 110, 313–322. [Google Scholar] [CrossRef]
- Li, J.; Hou, N.; Faried, A.; Tsutsumi, S.; Takeuchi, T.; Kuwano, H. Inhibition of autophagy by 3-MA enhances the effect of 5-FU-induced apoptosis in colon cancer cells. Ann. Surg. Oncol. 2009, 16, 761–771. [Google Scholar] [CrossRef]
- Li, J.; Hou, N.; Faried, A.; Tsutsumi, S.; Kuwano, H. Inhibition of autophagy augments 5-fluorouracil chemotherapy in human colon cancer in vitro and in vivo model. Eur. J. Cancer 2010, 46, 1900–1909. [Google Scholar] [CrossRef]
- Qian, W.; Liu, J.; Jin, J.; Ni, W.; Xu, W. Arsenic trioxide induces not only apoptosis but also autophagic cell death in leukemia cell lines via up-regulation of Beclin-1. Leuk. Res. 2007, 31, 329–339. [Google Scholar] [CrossRef]
- Maiuri, M.C.; Zalckvar, E.; Kimchi, A.; Kroemer, G. Self-eating and self-killing: Crosstalk between autophagy and apoptosis. Nat. Rev. Mol. Cell. Biol. 2007, 8, 741–752. [Google Scholar] [CrossRef]
- Kondo, Y.; Kanzawa, T.; Sawaya, R.; Kondo, S. The role of autophagy in cancer development and response to therapy. Nat. Rev. Cancer 2005, 5, 726–734. [Google Scholar] [CrossRef]
- Voss, V.; Senft, C.; Lang, V.; Ronellenfitsch, M.W.; Steinbach, J.P.; Seifert, V.; Kögel, D. The pan-Bcl-2 inhibitor (−)-gossypol triggers autophagic cell death in malignant glioma. Mol. Cancer Res. 2010, 8, 1002–1016. [Google Scholar] [CrossRef]
- Lian, J.; Wu, X.; He, F.; Karnak, D.; Tang, W.; Meng, Y.; Xiang, D.; Ji, M.; Lawrence, T.S.; Xu, L. A natural BH3 mimetic induces autophagy in apoptosis-resistant prostate cancer via modulating Bcl-2-Beclin1 interaction at endoplasmic reticulum. Cell Death Differ. 2011, 18, 60–71. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: the next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Townsend, K.N.; Hughson, L.R.; Schlie, K.; Poon, V.I.; Westerback, A.; Lum, J.J. Autophagy inhibition in cancer therapy: Metabolic considerations for antitumor immunity. Immunol. Rev. 2012, 249, 176–194. [Google Scholar] [CrossRef]
- Wildenberg, M.E.; Vos, A.C.; Wolfkamp, S.C.; Duijvestein, M.; Verhaar, A.P.; Te Velde, A.A.; van den Brink, G.R.; Hommes, D.W. Autophagy attenuates the adaptive immune response by destabilizing the immunologic synapse. Gastroenterology 2012, 142, 1493–1503. [Google Scholar] [CrossRef]
- Chemali, M.; Radtke, K.; Desjardins, M.; English, L. Alternative pathways for MHC class I presentation: a new function for autophagy. Cell. Mol. Life Sci. 2011, 68, 1533–1541. [Google Scholar] [CrossRef]
- Schmid, D.; Pypaert, M.; Münz, C. Antigen-loading compartments for major histocompatibility complex class II molecules continuously receive input from autophagosomes. Immunity 2007, 26, 79–92. [Google Scholar] [CrossRef]
- Gerriets, V.A.; Rathmell, J.C. Metabolic pathways in T cell fate and function. Trends Immunol. 2012, 33, 168–173. [Google Scholar] [CrossRef]
- Kovacs, J.R.; Li, C.; Yang, Q.; Li, G.; Garcia, I.G.; Ju, S.; Roodman, D.G.; Windle, J.J.; Zhang, X.; Lu, B. Autophagy promotes T-cell survival through degradation of proteins of the cell death machinery. Cell Death Differ. 2012, 19, 144–152. [Google Scholar] [CrossRef]
- Pua, H.H.; Guo, J.; Komatsu, M.; He, Y.W. Autophagy is essential for mitochondrial clearance in mature T lymphocytes. J. Immunol. 2009, 182, 4046–4055. [Google Scholar] [CrossRef]
- Jia, W.; He, Y.W. Temporal regulation of intracellular organelle homeostasis in T lymphocytes by autophagy. J. Immunol. 2011, 186, 5313–5322. [Google Scholar] [CrossRef]
- Hubbard, V.M.; Valdor, R.; Patel, B.; Singh, R.; Cuervo, A.M.; Macian, F. Macroautophagy regulates energy metabolism during effector T cell activation. J. Immunol. 2010, 185, 7349–7357. [Google Scholar] [CrossRef]
- Li, Y.; Wang, L.X.; Yang, G.; Hao, F.; Urba, W.J.; Hu, H.M. Efficient cross-presentation depends on autophagy in tumor cells. Cancer Res. 2008, 68, 6889–6895. [Google Scholar] [CrossRef]
- Li, Y.; Wang, L.X.; Pang, P.; Cui, Z.; Aung, S.; Haley, D.; Fox, B.A.; Urba, W.J.; Hu, H.M. Tumor-derived autophagosome vaccine: Mechanism of cross-presentation and therapeutic efficacy. Clin. Cancer Res. 2011, 17, 7047–7057. [Google Scholar]
- Yi, Y.; Zhou, Z.; Shu, S.; Fang, Y.; Twitty, C.; Hilton, T.L.; Aung, S.; Urba, W.J.; Fox, B.A.; Hu, H.M.; et al. Autophagy-assisted antigen cross-presentation: Autophagosome as the argo of shared tumor-specific antigens and DAMPs. Oncoimmunology 2012, 1, 976–978. [Google Scholar] [CrossRef]
- Kepp, O.; Galluzzi, L.; Martins, I.; Schlemmer, F.; Adjemian, S.; Michaud, M.; Sukkurwala, A.Q.; Menger, L.; Zitvogel, L.; Kroemer, G. Molecular determinants of immunogenic cell death elicited by anticancer chemotherapy. Cancer Metastasis Rev. 2011, 30, 61–69. [Google Scholar] [CrossRef]
- Twitty, C.G.; Jensen, S.M.; Hu, H.M.; Fox, B.A. Tumor-derived autophagosome vaccine: induction of cross-protective immune responses against short-lived proteins through a p62-dependent mechanism. Clin. Cancer Res. 2011, 17, 6467–6481. [Google Scholar] [CrossRef]
- Inguscio, V.; Panzarini, E.; Dini, L. Autophagy contributes to the death/survival balance in cancer photodynamic therapy. Cells 2012, 1, 464–491. [Google Scholar] [CrossRef]
- Reiners, J.J., Jr.; Agostinis, P.; Berg, K.; Oleinick, N.L.; Kessel, D. Assessing autophagy in the context of photodynamic therapy. Autophagy 2010, 6, 7–18. [Google Scholar] [CrossRef]
- Dewaele, M.; Maes, H.; Agostinis, P. ROS-mediated mechanisms of autophagy stimulation and their relevance in cancer therapy. Autophagy 2010, 6, 838–854. [Google Scholar] [CrossRef]
- Kessel, D.; Reiners, J.J., Jr. Apoptosis and autophagy after mitochondrial or endoplasmic reticulum photodamage. Photochem. Photobiol. 2007, 83, 1024–1028. [Google Scholar] [CrossRef]
- Kessel, D.; Arroyo, A.S. Apoptotic and autophagic responses to Bcl-2 inhibition and photodamage. Photochem. Photobiol. Sci. 2007, 6, 1290–1295. [Google Scholar] [CrossRef]
- Dini, L.; Inguscio, V.; Tenuzzo, B.; Panzarini, E. Rose bengal acetate photodynamic therapy-induced autophagy. Cancer Biol. Ther. 2010, 10, 1048–1055. [Google Scholar] [CrossRef]
- Panzarini, E.; Inguscio, V.; Dini, L. Timing the multiple cell death pathways initiated by Rose Bengal acetate photodynamic therapy. Cell Death Dis. 2011, 2, e169. [Google Scholar] [CrossRef]
- Panzarini, E.; Tenuzzo, B.; Palazzo, F.; Chionna, A.; Dini, L. Apoptosis induction and mitochondria alteration in human HeLa tumor cells by photoproducts of Rose Bengal acetate. J. Photochem. Photobiol. B 2006, 83, 39–47. [Google Scholar] [CrossRef]
- Panzarini, E.; Tenuzzo, B.; Dini, L. Photodynamic therapy-induced apoptosis of HeLa cells. Ann. NY Acad. Sci. 2009, 1171, 617–626. [Google Scholar] [CrossRef]
- Kessel, D.; Castelli, M. Evidence that bcl-2 is the target of three photosensitizers that induce a rapid apoptotic response. Photochem. Photobiol. 2001, 74, 318–322. [Google Scholar] [CrossRef]
- Weyergang, A.; Berg, K.; Kaalhus, O.; Peng, Q.; Selbo, P.K. Photodynamic therapy targets the mTOR signaling network in vitro and in vivo. Mol. Pharm. 2009, 6, 255–264. [Google Scholar]
- Herd, H.L.; Malugin, A.; Ghandehari, H. Silica nanoconstruct cellular toleration threshold in vitro. J. Control. Release 2011, 153, 40–48. [Google Scholar] [CrossRef]
- Li, H.; Li, Y.; Jiao, J.; Hu, H.M. Alpha-alumina nanoparticles induce efficient autophagy-dependent cross-presentation and potent antitumour response. Nat. Nanotechnol. 2011, 6, 645–650. [Google Scholar]
- Monick, M.M.; Powers, L.S.; Walters, K.; Lovan, N.; Zhang, M.; Gerke, A.; Hansdottir, S.; Hunninghake, G.W. Identification of an autophagy defect in smokers’ alveolar macrophages. J. Immunol. 2010, 185, 5425–5435. [Google Scholar] [CrossRef]
- Seleverstov, O.; Zabirnyk, O.; Zscharnack, M.; Bulavina, L.; Nowicki, M.; Heinrich, J.M.; Yezhelyev, M.; Emmrich, F.; O’Regan, R.; Bader, A. Quantum dots for human mesenchymal stem cells labeling. A size-dependent autophagy activation. Nano Lett. 2006, 6, 2826–2832. [Google Scholar] [CrossRef]
- Yokoyama, T.; Tam, J.; Kuroda, S.; Scott, A.W.; Aaron, J.; Larson, T.; Shanker, M.; Correa, A.M.; Kondo, S.; Roth, J.A.; et al. EGFR-targeted hybrid plasmonic magnetic nanoparticles synergistically induce autophagy and apoptosis in non-small cell lung cancer cells. PLoS One 2011, 6, e25507. [Google Scholar]
- Harhaji, L.; Isakovic, A.; Raicevic, N.; Markovic, Z.; Todorovic-Markovic, B.; Nikolic, N.; Vranjes-Djuric, S.; Markovic, I.; Trajkovic, V. Multiple mechanisms underlying the anticancer action of nanocrystalline fullerene. Eur. J. Pharmacol. 2007, 568, 89–98. [Google Scholar] [CrossRef]
- Wu, Y.N.; Yang, L.X.; Shi, X.Y.; Li, I.C.; Biazik, J.M.; Ratinac, K.R.; Chen, D.H.; Thordarson, P.; Shieh, D.B.; Braet, F. The selective growth inhibition of oral cancer by iron core-gold shell nanoparticles through mitochondria-mediated autophagy. Biomaterials 2011, 32, 4565–4573. [Google Scholar] [CrossRef]
- Khan, M.I.; Mohammad, A.; Patil, G.; Naqvi, S.A.; Chauhan, L.K.; Ahmad, I. Induction of ROS, mitochondrial damage and autophagy in lung epithelial cancer cells by iron oxide nanoparticles. Biomaterials 2012, 33, 1477–1488. [Google Scholar] [CrossRef]
- Li, C.; Liu, H.; Sun, Y.; Wang, H.; Guo, F.; Rao, S.; Deng, J.; Zhang, Y.; Miao, Y.; Guo, C.; et al. PAMAM nanoparticles promote acute lung injury by inducing autophagic cell death through the Akt-TSC2-mTOR signaling pathway. J. Mol. Cell Biol. 2009, 1, 37–45. [Google Scholar] [CrossRef]
- Liu, H.L.; Zhang, Y.L.; Yang, N.; Zhang, Y.X.; Liu, X.Q.; Li, C.G.; Zhao, Y.; Wang, Y.G.; Zhang, G.G.; Yang, P.; et al. A functionalized single-walled carbon nanotube-induced autophagic cell death in human lung cells through Akt-TSC2-mTOR signaling. Cell Death Dis. 2011, 2, e159. [Google Scholar] [CrossRef]
- Cirstea, D.; Hideshima, T.; Rodig, S.; Santo, L.; Pozzi, S.; Vallet, S.; Ikeda, H.; Perrone, G.; Gorgun, G.; Patel, K.; et al. Dual inhibition of akt/mammalian target of rapamycin pathway by nanoparticle albumin-bound-rapamycin and perifosine induces antitumor activity in multiple myeloma. Mol. Cancer Ther. 2010, 9, 963–975. [Google Scholar] [CrossRef]
- Zhang, Q.; Yang, W.; Man, N.; Zheng, F.; Shen, Y.; Sun, K.; Li, Y.; Wen, L.P. Autophagy-mediated chemosensitization in cancer cells by fullerene C60 nanocrystal. Autophagy 2009, 5, 1107–1117. [Google Scholar] [CrossRef]
- Calzolai, L.; Franchini, F.; Gilliland, D.; Rossi, F. Protein-nanoparticle interaction: Identification of the ubiquitin—Gold nanoparticle interaction site. Nano Lett. 2010, 10, 3101–3105. [Google Scholar] [CrossRef]
- Ryter, S.W.; Nakahira, K.; Haspel, J.A.; Choi, A.M. Autophagy in pulmonary diseases. Annu. Rev. Physiol. 2012, 74, 377–401. [Google Scholar] [CrossRef]
- Seleverstov, O.; Phang, J.M.; Zabirnyk, O. Semiconductor nanocrystals in autophagy research: Methodology improvement at nanosized scale. Methods Enzymol. 2009, 452, 277–296. [Google Scholar]
- Joshi, P.; Chakraborti, S.; Ramirez-Vick, J.E.; Ansari, Z.A.; Shanker, V.; Chakrabarti, P.; Singh, S.P. The anticancer activity of chloroquine-gold nanoparticles against MCF-7 breast cancer cells. Colloids Surf. B Biointerfaces 2012, 95, 195–200. [Google Scholar] [CrossRef]
- Wei, P.; Zhang, L.; Lu, Y.; Man, N.; Wen, L. C60(Nd) nanoparticles enhance chemotherapeutic susceptibility of cancer cells by modulation of autophagy. Nanotechnology 2010, 21, 495101. [Google Scholar] [CrossRef]
- Salvador-Morales, C.; Gao, W.; Ghatalia, P.; Murshed, F.; Aizu, W.; Langer, R.; Farokhzad, O.C. Multifunctional nanoparticles for prostate cancer therapy. Exp. Rev. Anticancer Ther. 2009, 9, 211–221. [Google Scholar] [CrossRef]
- Veale, D.; Kerr, N.; Gibson, G.J.; Harris, A.L. Characterization of epidermal growth factor receptor in primary human non-small cell lung cancer. Cancer Res. 1989, 49, 1313–1317. [Google Scholar]
- Pao, W.; Miller, V.A. Epidermal growth factor receptor mutations, small-molecule kinase inhibitors, and non-small-cell lung cancer: Current knowledge and future directions. J. Clin. Oncol. 2005, 23, 2556–2568. [Google Scholar] [CrossRef]
- Cappuzzo, F.; Hirsch, F.R.; Rossi, E.; Bartolini, S.; Ceresoli, G.L.; Bemis, L.; Haney, J.; Witta, S.; Danenberg, K.; Domenichini, I.; et al. Epidermal growth factor receptor gene and protein and gefitinib sensitivity in non-small-cell lung cancer. J. Natl. Cancer Inst. 2005, 97, 643–655. [Google Scholar] [CrossRef]
- Paez, J.G.; Jänne, P.A.; Lee, J.C.; Tracy, S.; Greulich, H.; Gabriel, S.; Herman, P.; Kaye, F.J.; Lindeman, N.; Boggon, T.J.; et al. EGFR mutations in lung cancer: Correlation with clinical response to gefitinib therapy. Science 2004, 304, 1497–1500. [Google Scholar] [CrossRef]
- Sayes, C.; Fortner, J.; Lyon, D.; Boyd, A.; Ausman, K.; Tao, Y.; Sitharaman, B.; Wilson, L.; West, J.; Colvin, V.L. The differential cytotoxicity of water soluble fullerenes. Nano Lett. 2004, 4, 1881–1887. [Google Scholar] [CrossRef]
- Isakovic, A.; Markovic, Z.; Todorovic-Markovic, B.; Nikolic, N.; Vranjes-Djuric, S.; Mirkovic, M.; Dramicanin, M.; Harhaji, L.; Raicevic, N.; Nikolic, Z.; et al. Distinct cytotoxic mechanisms of pristine versus hydroxylated fullerene. Toxicol. Sci. 2006, 91, 173–183. [Google Scholar] [CrossRef]
- Giese, A.; Bjerkvig, R.; Berens, M.E.; Westphal, M. Cost of migration: Invasion of malignant gliomas and implications for treatment. J. Clin. Oncol. 2003, 21, 1624–1636. [Google Scholar] [CrossRef]
- Finn, O.J. Cancer vaccines: Between the idea and the reality. Nat. Rev. Immunol. 2003, 3, 630–641. [Google Scholar] [CrossRef]
- Amaravadi, R.K.; Thompson, C.B. The roles of therapy-induced autophagy and necrosis in cancer treatment. Clin. Cancer Res. 2007, 13, 7271–7279. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Panzarini, E.; Inguscio, V.; Tenuzzo, B.A.; Carata, E.; Dini, L. Nanomaterials and Autophagy: New Insights in Cancer Treatment. Cancers 2013, 5, 296-319. https://doi.org/10.3390/cancers5010296
Panzarini E, Inguscio V, Tenuzzo BA, Carata E, Dini L. Nanomaterials and Autophagy: New Insights in Cancer Treatment. Cancers. 2013; 5(1):296-319. https://doi.org/10.3390/cancers5010296
Chicago/Turabian StylePanzarini, Elisa, Valentina Inguscio, Bernardetta Anna Tenuzzo, Elisabetta Carata, and Luciana Dini. 2013. "Nanomaterials and Autophagy: New Insights in Cancer Treatment" Cancers 5, no. 1: 296-319. https://doi.org/10.3390/cancers5010296





