Genetic Divergence and Chemotype Diversity in the Fusarium Head Blight Pathogen Fusarium poae
Abstract
:1. Introduction
2. Results
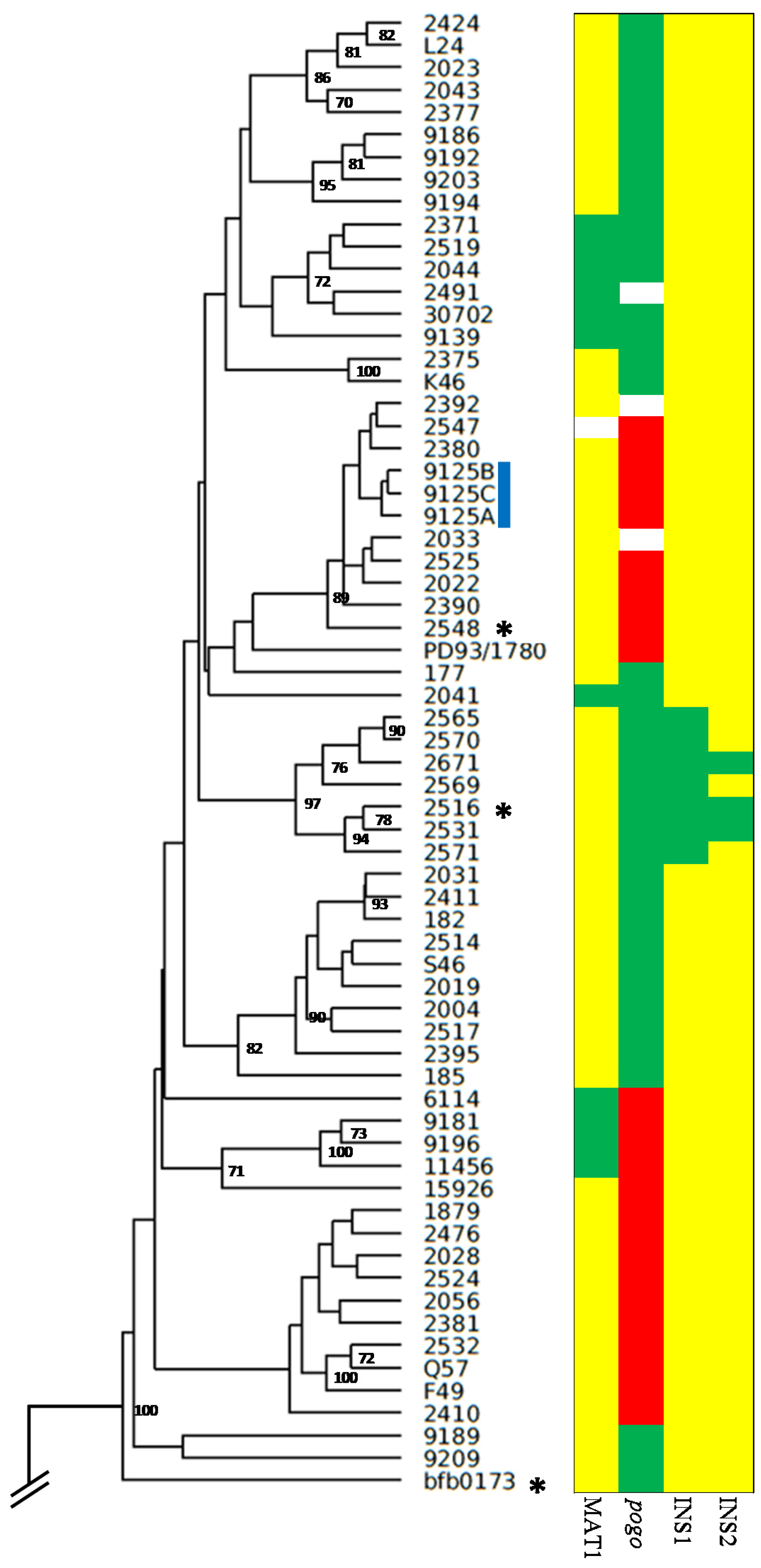
2.1. Genetic Diversity of Fusarium poae
2.2. Fusarium poae Likely Combines Sexual and Asexual Reproduction
2.3. Transposable Element Proliferation between Near-Clonal Isolates
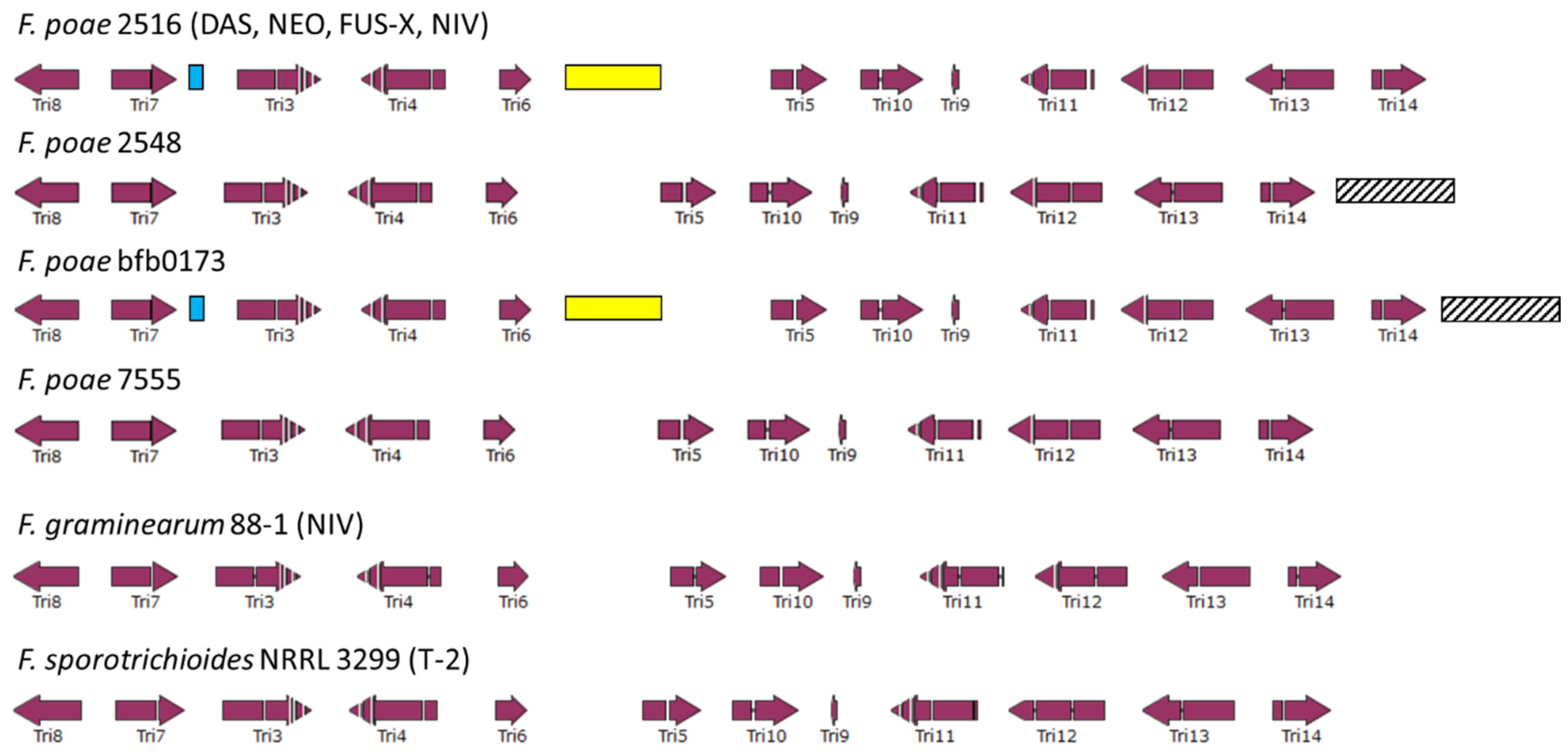
2.4. Assessing the Genetic Chemotype of F. poae Isolates
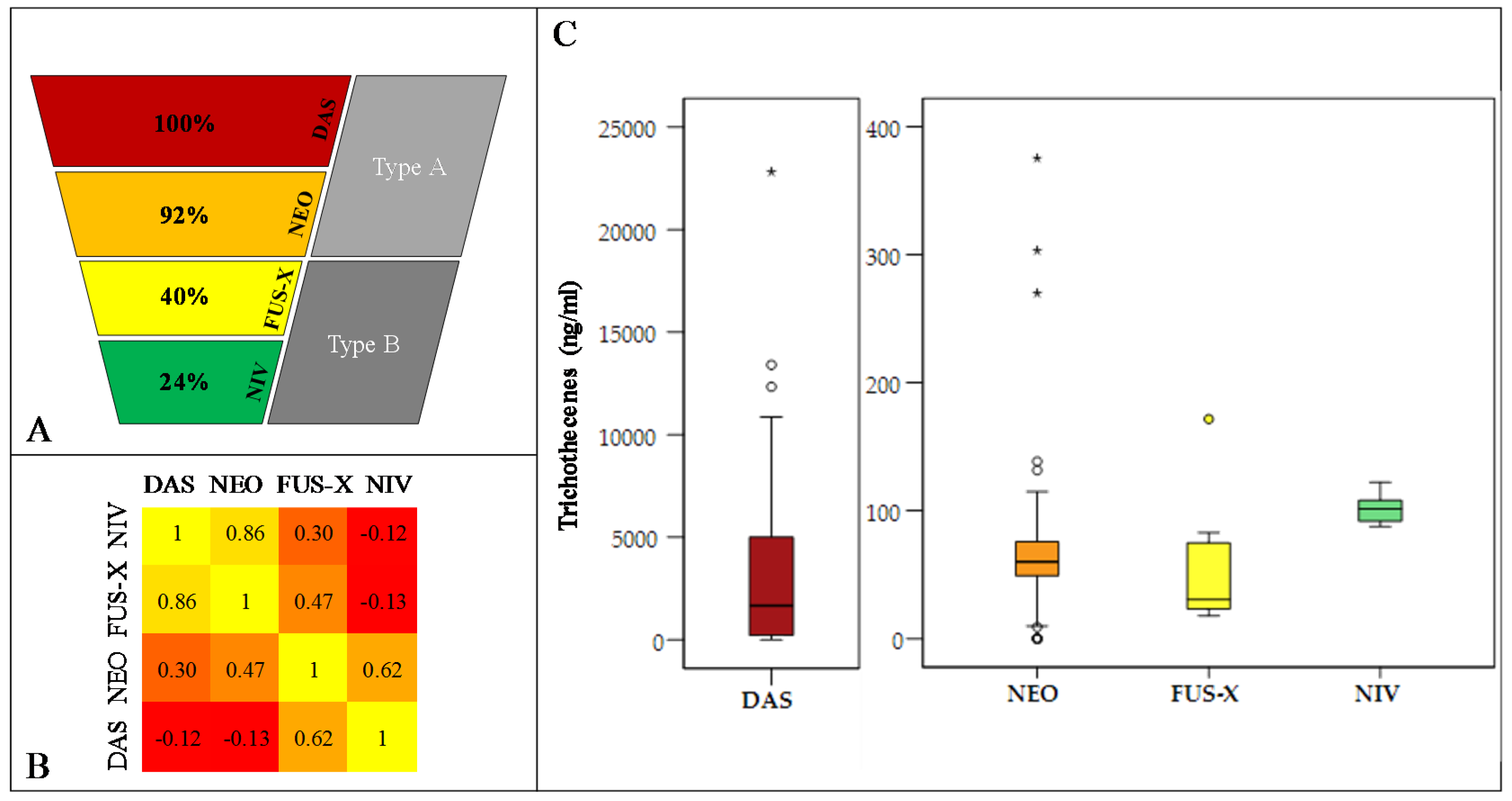
2.5. Assessing the Phenotypic Chemotype of F. poae Isolates
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Fusarium Collection
5.2. Phylogenetic Analyses
5.3. Fusarium poae Genome Data
5.4. Trichothecene Production Analyses
5.5. Diagnostic PCRs and Amplicon Sequencing
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Goswami, R.S.; Kistler, H.C. Heading for disaster: Fusarium graminearum on cereal crops. Mol. Plant Pathol. 2004, 5, 515–525. [Google Scholar] [CrossRef] [PubMed]
- Da Rocha, M.E.B.; Freire, F.D.O.; Maia, F.B.F.; Guedes, M.I.F.; Rondina, D. Mycotoxins and their effects on human and animal health. Food Control 2014, 36, 159–165. [Google Scholar] [CrossRef]
- Parry, D.W.; Jenkinson, P.; McLeod, L. Fusarium ear blight (scab) in small-grain cereals—A review. Plant Pathol. 1995, 44, 207–238. [Google Scholar] [CrossRef]
- Xu, X.M.; Parry, D.W.; Nicholson, P.; Thomsett, M.A.; Simpson, D.; Edwards, S.G.; Cooke, B.M.; Doohan, F.M.; Brennan, J.M.; Moretti, A.; et al. Predominance and association of pathogenic fungi causing Fusarium ear blight in wheat in four European countries. Eur. J. Plant Pathol. 2005, 112, 143–154. [Google Scholar] [CrossRef]
- Audenaert, K.; Van Broeck, R.; Bekaert, B.; De Witte, F.; Heremans, B.; Messens, K.; Hofte, M.; Haesaert, G. Fusarium head blight (fhb) in Flanders: Population diversity, inter-species associations and DON contamination in commercial winter wheat varieties. Eur. J. Plant Pathol. 2009, 125, 445–458. [Google Scholar] [CrossRef]
- Lindblad, M.; Gidlund, A.; Sulyok, M.; Borjesson, T.; Krska, R.; Olsen, M.; Fredlund, E. Deoxynivalenol and other selected Fusarium toxins in Swedish wheat—Occurrence and correlation to specific fusarium species. Int. J. Food Microbiol. 2013, 167, 284–291. [Google Scholar] [CrossRef] [PubMed]
- Covarelli, L.; Beccari, G.; Prodi, A.; Generotti, S.; Etruschi, F.; Juan, C.; Ferrer, E.; Manes, J. Fusarium species, chemotype characterisation and trichothecene contamination of durum and soft wheat in an area of central Italy. J. Sci. Food Agric. 2015, 95, 540–551. [Google Scholar] [CrossRef] [PubMed]
- Vanheule, A.; Audenaert, K.; De Boevre, M.; Landschoot, S.; Bekaert, B.; Munaut, F.; Eeckhout, M.; Hofte, M.; De Saeger, S.; Haesaert, G. The compositional mosaic of Fusarium species and their mycotoxins in unprocessed cereals, food and feed products in belgium. Int. J. Food Microbiol. 2014, 181, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Audenaert, K.; Monbaliu, S.; Deschuyffeleer, N.; Maene, P.; Vekeman, F.; Haesaert, G.; De Saeger, S.; Eeckhout, M. Neutralized electrolyzed water efficiently reduces fusarium spp. in vitro and on wheat kernels but can trigger deoxynivalenol (DON) biosynthesis. Food Control 2012, 23, 515–521. [Google Scholar] [CrossRef]
- McCormick, S.P.; Stanley, A.M.; Stover, N.A.; Alexander, N.J. Trichothecenes: From simple to complex mycotoxins. Toxins 2011, 3, 802–814. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Tokai, T.; Takahashi-Ando, N.; Ohsato, S.; Fujimura, M. Molecular and genetic studies of Fusarium trichothecene biosynthesis: Pathways, genes, and evolution. Biosci. Biotechnol. Biochem. 2007, 71, 2105–2123. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S.P.; Alexander, N.J.; Proctor, R.H. Heterologous expression of two trichothecene p450 genes in Fusarium verticillioides. Can. J. Microbiol. 2006, 52, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, Y.; Fukasaku, K.; Tanaka, T.; Matsui, Y.; Ueno, Y. Fusarium poae and Fusarium crookwellense, fungi responsible for the natural occurrence of nivalenol in hokkaido. Appl. Environ. Microbiol. 1993, 59, 3334–3338. [Google Scholar]
- Thrane, U.; Adler, A.; Clasen, P.E.; Galvano, F.; Langseth, W.; Logrieco, A.; Nielsen, K.F.; Ritieni, A. Diversity in metabolite production by Fusarium langsethiae, Fusarium poae, and Fusarium sporotrichioides. Int. J. Food Microbiol. 2004, 95, 257–266. [Google Scholar] [CrossRef] [PubMed]
- Vogelgsang, S.; Sulyok, M.; Baenziger, I.; Krska, R.; Schuhmacher, R.; Forrer, H.-R. Effect of fungal strain and cereal substrate on in vitro mycotoxin production by Fusarium poae and Fusarium avenaceum. Food Addit. Contam. 2008, 25, 745–757. [Google Scholar] [CrossRef] [PubMed]
- McDonald, B.A.; Linde, C. Pathogen population genetics, evolutionary potential, and durable resistance. Ann. Rev. Phytopath. 2002, 40, 349–379. [Google Scholar] [CrossRef] [PubMed]
- Kerenyi, Z.; Moretti, A.; Waalwijk, C.; Olah, B.; Hornok, L. Mating type sequences in asexually reproducing Fusarium species. Appl. Environ. Microbiol. 2004, 70, 4419–4423. [Google Scholar] [CrossRef] [PubMed]
- Tavanti, A.; Gow, N.A.R.; Maiden, M.C.J.; Odds, F.C.; Shaw, D.J. Genetic evidence for recombination in Candida albicans based on haplotype analysis. Fungal Genet. Biol. 2004, 41, 553–562. [Google Scholar] [CrossRef] [PubMed]
- Vanheule, A.; Audenaert, K.; Warris, S.; van de Geest, H.; Schijlen, E.; Hofte, M.; De Saeger, S.; Haesaert, G.; Waalwijk, C.; van der Lee, T. Living apart together: Crosstalk between the core and supernumerary genomes in a fungal plant pathogen. BMC Genom. 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Galagan, J.E.; Selker, E.U. RIP: The evolutionary cost of genome defense. Trends Genet. 2004, 20, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Dinolfo, M.I.; Castanares, E.; Stenglein, S.A. Characterization of a Fusarium poae world-wide collection by using molecular markers. Eur. J. Plant Pathol. 2014, 140, 119–132. [Google Scholar] [CrossRef]
- Kerenyi, Z.; Taborhegyi, E.; Pomazi, A.; Hornok, L. Variability amongst strains of Fusarium poae assessed by vegetative compatibility and RAPD polymorphism. Plant Pathol. 1997, 46, 882–889. [Google Scholar] [CrossRef]
- Somma, S.; Alvarez, C.; Ricci, V.; Ferracane, L.; Ritieni, A.; Logrieco, A.; Moretti, A. Trichothecene and beauvericin mycotoxin production and genetic variability in Fusarium poae isolated from wheat kernels from northern italy. Food Addit. Contam. 2010, 27, 729–737. [Google Scholar] [CrossRef] [PubMed]
- Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; Vandelee, T.; Hornes, M.; Frijters, A.; Pot, J.; Peleman, J.; Kuiper, M.; et al. AFLP—A new technique for DNA-fingerprinting. Nucl. Acids Res. 1995, 23, 4407–4414. [Google Scholar] [CrossRef] [PubMed]
- Dinolfo, M.I.; Stenglein, S.A.; Moreno, M.V.; Nicholson, P.; Jennings, P.; Salerno, G.L. ISSR markers detect high genetic variation among Fusarium poae isolates from Argentina and England. Eur. J. Plant Pathol. 2010, 127, 483–491. [Google Scholar] [CrossRef]
- Stenglein, S.A.; Rodriguero, M.S.; Chandler, E.; Jennings, P.; Salerno, G.L.; Nicholson, P. Phylogenetic relationships of Fusarium poae based on EF-1 alpha and mtssu sequences. Fungal Biol. 2010, 114, 96–106. [Google Scholar] [CrossRef] [PubMed]
- Kulik, T.; Pszczolkowska, A. Multilocus sequence analysis of Fusarium poae. J. Plant Pathol. 2011, 93, 119–126. [Google Scholar]
- Fekete, C.; Hornok, L. A repetitive DNA sequence associated with karyotype variability in Fusarium poae. Acta Phytopathol. Entomol. Hung. 1997, 32, 29–38. [Google Scholar]
- Li, Y.; van der Lee, T.A.J.; Evenhuis, A.; van den Bosch, G.B.M.; van Bekkum, P.J.; Forch, M.G.; van Gent-Pelzer, M.P.E.; van Raaij, H.M.G.; Jacobsen, E.; Huang, S.W.; et al. Population dynamics of Phytophthora infestans in the Netherlands reveals expansion and spread of dominant clonal lineages and virulence in sexual offspring. G3 Genes Genom. Genet. 2012, 2, 1529–1540. [Google Scholar] [CrossRef] [PubMed]
- Mueller, U.G.; Wolfenbarger, L.L. AFLP genotyping and fingerprinting. Trends Ecol. Evol. 1999, 14, 389–394. [Google Scholar] [CrossRef]
- Bonin, A.; Bellemain, E.; Bronken Eidesen, P.; Pompanon, F.; Brochmann, C.; Taberlet, P. How to track and assess genotyping errors in population genetics studies. Mol. Ecol. 2004, 13, 3261–3273. [Google Scholar] [CrossRef] [PubMed]
- Proctor, R.H.; McCormick, S.P.; Alexander, N.J.; Desjardins, A.E. Evidence that a secondary metabolic biosynthetic gene cluster has grown by gene relocation during evolution of the filamentous fungus Fusarium. Mol. Microbiol. 2009, 74, 1128–1142. [Google Scholar] [CrossRef] [PubMed]
- Dinolfo, M.I.; Barros, G.G.; Stenglein, S.A. Development of a PCR assay to detect the potential production of nivalenol in Fusarium poae. FEMS Microbiol. Lett. 2012, 332, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Varga, E.; Wiesenberger, G.; Hametner, C.; Ward, T.J.; Dong, Y.; Schöfbeck, D.; McCormick, S.; Broz, K.; Stückler, R.; Schuhmacher, R.; et al. New tricks of an old enemy: Isolates of Fusarium graminearum produce a type A trichothecene mycotoxin. Environ. Microbiol. 2015, 17, 2588–2600. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.; Niessen, L.; Vogel, R.F. AFLP analysis of Fusarium species in the section Sporotrichiella—Evidence for Fusarium langsethiae as a new species. Int. J. Food Microbiol. 2004, 95, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, R.; Torp, M.; Kosiak, B.; Holst-Jensen, A. Phylogeny and toxigenic potential is correlated in Fusarium species as revealed by partial translation elongation factor 1 alpha gene sequences. Mycol. Res. 2005, 109, 173–186. [Google Scholar] [CrossRef] [PubMed]
- Pasquali, M.; Migheli, Q. Genetic approaches to chemotype determination in type B-trichothecene producing fusaria. Int. J. Food Microbiol. 2014, 189, 164–182. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.W.; Dyer, R.B.; McCormick, S.P.; Kendra, D.F.; Plattner, R.D. Functional demarcation of the Fusarium core trichothecene gene cluster. Fungal Genet. Biol. 2004, 41, 454–462. [Google Scholar] [CrossRef] [PubMed]
- Cuomo, C.A.; Gueldener, U.; Xu, J.-R.; Trail, F.; Turgeon, B.G.; Di Pietro, A.; Walton, J.D.; Ma, L.-J.; Baker, S.E.; Rep, M.; et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 2007, 317, 1400–1402. [Google Scholar] [CrossRef] [PubMed]
- Ward, T.J.; Bielawski, J.P.; Kistler, H.C.; Sullivan, E.; O’Donnell, K. Ancestral polymorphism and adaptive evolution in the trichothecene mycotoxin gene cluster of phytopathogenic Fusarium. Proc. Natl. Acad. Sci. USA 2002, 99, 9278–9283. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, K.; Ward, T.J.; Aberra, D.; Kistler, H.C.; Aoki, T.; Orwig, N.; Kimura, M.; Bjornstad, A.; Klemsdal, S.S. Multilocus genotyping and molecular phylogenetics resolve a novel head blight pathogen within the Fusarium graminearum species complex from Ethiopia. Fungal Genet. Biol. 2008, 45, 1514–1522. [Google Scholar] [CrossRef] [PubMed]
- Feschotte, C. Opinion—Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 2008, 9, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Sinzelle, L.; Izsvak, Z.; Ivics, Z. Molecular domestication of transposable elements: From detrimental parasites to useful host genes. Cell. Mol. Life Sci. 2009, 66, 1073–1093. [Google Scholar] [CrossRef] [PubMed]
- Tag, A.G.; Garifullina, G.F.; Peplow, A.W.; Ake, C.; Phillips, T.D.; Hohn, T.M.; Beremand, M.N. A novel regulatory gene, tri10, controls trichothecene toxin production and gene expression. Appl. Environ. Microbiol. 2001, 67, 5294–5302. [Google Scholar] [CrossRef] [PubMed]
- Pasquali, M.; Cocco, E.; Guignard, C.; Hoffmann, L. The effect of agmatine on trichothecene type B and zearalenone production in Fusarium graminearum, F. culmorum and F. poae. PeerJ 2016, 4, e1672. [Google Scholar] [CrossRef] [PubMed]
- Vogelgsang, S.; Sulyok, M.; Hecker, A.; Jenny, E.; Krska, R.; Schuhmacher, R.; Forrer, H.-R. Toxigenicity and pathogenicity of Fusarium poae and Fusarium avenaceum on wheat. Eur. J. Plant Pathol. 2008, 122, 265–276. [Google Scholar] [CrossRef]
- Desjardins, A.E.; McCormick, S.P.; Appell, M. Structure-activity relationships of trichothecene toxins in an Arabidopsis thaliana leaf assay. J. Agric. Food Chem. 2007, 55, 6487–6492. [Google Scholar] [CrossRef] [PubMed]
- Placinta, C.; D’Mello, J.; Macdonald, A. A review of worldwide contamination of cereal grains and animal feed with Fusarium mycotoxins. Anim. Feed Sci. Technol. 1999, 78, 21–37. [Google Scholar] [CrossRef]
- Liu, W.Z.; Sundheim, L.; Langseth, W. Trichothecene production and the relationship to vegetative compatibility groups in Fusarium poae. Mycopathologia 1998, 140, 105–114. [Google Scholar] [CrossRef]
- Schollenberger, M.; Mueller, H.-M.; Liebscher, M.; Schlecker, C.; Berger, M.; Hermann, W. Accumulation kinetics of three scirpentriol-based toxins in oats inoculated in vitro with isolates of Fusarium sporotrichioides and Fusarium poae. Toxins 2011, 3, 442–452. [Google Scholar] [CrossRef] [PubMed]
- Parry, D.W.; Nicholson, P. Development of a pcr assay to detect Fusarium poae in wheat. Plant Pathol. 1996, 45, 383–391. [Google Scholar] [CrossRef]
- Scauflaire, J.; Gourgue, M.; Munaut, F. Fusarium temperatum sp. nov from maize, an emergent species closely related to Fusarium subglutinans. Mycologia 2011, 103, 586–597. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal w and clustal x version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Gardiner, D.M.; Kazan, K.; Manners, J.M. Nutrient profiling reveals potent inducers of trichothecene biosynthesis in Fusarium graminearum. Fungal Genet. Biol. 2009, 46, 604–613. [Google Scholar] [CrossRef] [PubMed]
- Spanjer, M.C.; Rensen, P.M.; Scholten, J.M. Lc-ms/ms multi-method for mycotoxins after single extraction, with validation data for peanut, pistachio, wheat, maize, cornflakes, raisins and figs. Food Addit. Contam. 2008, 25, 472–489. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D. PAUP*: Phylogenetic Analysis Using Parsimony, Version 4.0 b10; PAUP: Sunderland, MA, USA, 2003. [Google Scholar]
- Plotree, D.; Plotgram, D. Phylip-phylogeny inference package (version 3.2). Cladistics 1989, 5, 163–166. [Google Scholar]

| ID | Location | Host | Year | Reference | Mating Type | INS1 | INS2 | AFLP | DAS | NEO | FUS-X | NIV |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 175 | Aas, Norway | barley | 1996 | Sundheim L. unpubl. | MAT1-2 | - | - | - | 355 | 55.1 | ND | ND |
| 177 | Norway | wheat | 1996 | Sundheim L. unpubl. | MAT1-1 | - | - | + | NCA | |||
| 182 | Norway | barley | 1996 | Sundheim L. unpubl. | MAT1-1 | - | - | + | NCA | |||
| 185 | Norway | barley | 1996 | Sundheim L. unpubl. | MAT1-1 | - | - | + | ND | ND | ND | ND |
| 1879 | Bottelare, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 2160 | 66 | 25 | 88 |
| 2004 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 89 | ND | ND | ND |
| 2019 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 231 | 54 | ND | ND |
| 2022 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 1588 | 62 | 18 | ND |
| 2023 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 44 | 12 | ND | ND |
| 2028 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 3650 | 79 | 77 | 93 |
| 2031 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 109 | ND | ND | ND |
| 2033 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 1306 | 59 | ND | ND |
| 2041 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-2 | - | - | + | 60 | 12 | >LOD | 108 |
| 2043 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | 3971 | 72 | ND | ND |
| 2044 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-2 | - | - | + | 144 | 9 | ND | ND |
| 2056 | Zwevegem, Belgium | wheat | 2010 | this study | MAT1-1 | - | - | + | NCA | |||
| 2371 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-2 | - | - | + | 6923 | 101 | ND | ND |
| 2375 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 15 | 10 | ND | ND |
| 2377 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 31 | 11 | >LOD | 103 |
| 2380 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 1743 | 64 | ND | ND |
| 2381 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 222 | 54 | ND | ND |
| 2390 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 3027 | 67 | ND | ND |
| 2392 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 196 | 14 | ND | ND |
| 2395 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 193 | 55 | ND | ND |
| 2410 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 9562 | 84 | ND | ND |
| 2411 | Bottelare, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 238 | 54 | ND | ND |
| 2424 | Koksijde, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 2098 | 59 | ND | ND |
| 2476 | Poperinge, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 488 | 75 | 172 | 103 |
| 2491 | Poperinge, Belgium | wheat | 2011 | this study | MAT1-2 | - | - | + | 12,336 | 99 | ND | ND |
| 2514 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 2800 | 73 | 83 | 116 |
| 2516 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | + | + | + | 1126 | 57 | ND | ND |
| 2517 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 641 | 58 | ND | ND |
| 2519 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-2 | - | - | + | 9766 | 139 | ND | ND |
| 2521 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | + | - | - | ND | ND | ND | ND |
| 2524 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 380 | 17 | ND | ND |
| 2525 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 2195 | 61 | ND | ND |
| 2531 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | + | + | + | 803 | 59 | 22 | 92 |
| 2532 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 1940 | 66 | 25 | ND |
| 2547 | Zwevegem, Belgium | wheat | 2011 | this study | no amplicon | - | - | + | NCA | |||
| 2548 | Zwevegem, Belgium | wheat | 2011 | this study | MAT1-1 | - | - | + | 67 | ND | ND | ND |
| 2565 | Zuienkerke, Belgium | wheat | 2011 | this study | MAT1-1 | + | - | + | 69 | 18 | >LOD | 115 |
| 2569 | Zuienkerke, Belgium | wheat | 2011 | this study | MAT1-1 | + | - | + | 4910 | 44 | >LOD | 122 |
| 2570 | Zuienkerke, Belgium | wheat | 2011 | this study | MAT1-1 | + | - | + | 6630 | 66 | 25 | ND |
| 2571 | Zuienkerke, Belgium | wheat | 2011 | this study | MAT1-1 | + | - | + | NCA | |||
| 2671 | Linter, Belgium | wheat | 2011 | this study | MAT1-1 | + | + | + | 2457.8 | 75 | 37 | 100 |
| 6127 | Wageningen, The Netherlands | wheat | 1964 | MUCL | MAT1-2 | - | - | - | 10,864 | 303 | 75 | ND |
| 6114 | Denmark | barley | 1964 | MUCL, Hennebert G.L. unpubl. | MAT1-2 | - | - | + | 135 | 55 | ND | ND |
| 7555 | Heverlee, Belgium | wheat | 1965 | MUCL | MAT1-1 | - | - | - | 1067.8 | ND | ND | ND |
| 9125 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 5096 | 73 | ND | ND |
| 9139 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-2 | - | - | + | 8 | ND | ND | ND |
| 9181 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-2 | - | - | + | NCA | |||
| 9186 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 8024 | 77 | 24 | ND |
| 9189 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 6342 | 98 | ND | ND |
| 9192 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 10,415 | 132 | 46 | 91 |
| 9194 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 13,404 | 270 | 81 | ND |
| 9196 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-2 | - | - | + | 3610 | 115 | 34 | ND |
| 9203 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 115 | 55 | ND | ND |
| 9209 | Ferrara, Italy | wheat | 2005 | Somma et al. (2010) | MAT1-1 | - | - | + | 5502 | 115 | 28 | ND |
| 11456 | Heverlee, Belgium | barley | 1968 | MUCL, Meyer J.A. unpubl. | MAT1-2 | - | - | + | 264 | 20 | >LOD | 103 |
| 30702 | unknown | in vitro plant | 1990 | MUCL, Marchand D. unpubl. | MAT1-2 | - | - | + | 22,804 | 375 | 57 | ND |
| 15926 | Quebec, Canada | wheat | 1970 | MUCL, Hennebert G.L. unpubl. | MAT1-1 | - | - | + | NCA | |||
| 42824 | Belgium | wheat | 2000 | MUCL | MAT1-1 | - | - | - | 2120 | 73 | ND | ND |
| bfb0173 | China | barley | 2005 | Yang et al. (2008) | MAT1-1 | - | - | + | NCA | |||
| F49 | Ath, Belgium | maize | 2007 | MUCL (Scaufflaire J.) | MAT1-1 | - | - | + | 5720 | 76 | ND | ND |
| K46 | Ath, Belgium | maize | 2007 | MUCL (Scaufflaire J.) | MAT1-1 | - | - | + | 5339 | 100 | ND | ND |
| L24 | Buissenal, Belgium | maize | 2007 | MUCL (Scaufflaire J.) | MAT1-1 | - | - | + | 162 | 55 | ND | ND |
| Q57 | Buissenal, Belgium | maize | 2007 | MUCL (Scaufflaire J.) | MAT1-1 | - | - | + | 916 | 56 | ND | ND |
| S46 | Villeroux, Belgium | maize | 2007 | MUCL (Scaufflaire J.) | MAT1-1 | - | - | + | 1208 | 59 | ND | ND |
| PD93/1780 | The Netherlands | carnation | 2003 | WUR, Waalwijk et al. (2003) | MAT1-1 | - | - | + | 4008 | 66 | 23 | ND |
| n = 69 | 1 not determined 13 MAT1-2 55 MAT1-1 | 8 isolates containing INS1 | 3 isolates containing INS2 | AFLP for 64 isolates | chemotype for 61 isolates; no chemotype analysed for 8 isolates | |||||||
| # On the Core Genome of 2516 | # Present in 2516 but Not in 2531 (No Read Support) | |||
|---|---|---|---|---|
| Superfamily | Family | |||
| Retrotransposons | ||||
| RLG_Maggy | Gypsy/Ty3 like | 27 | 1 | |
| RLG_Skippy | Gypsy/Ty3 like | 5 | 2 | |
| DNA transposons | ||||
| DTF_Fot4 | Pogo | 1 | 0 | |
| DTF_Fot5-A | Pogo | 40 | 26 | |
| DTF_ESP4-B | Pogo | 12 | 0 | |
| DTF_Drogon | Pogo | 33 | 33 | |
| DTF_Viserion | Pogo | 8 | 3 | |
| DTM_Hop7 | Mutator | 8 | 8 | |
| DTM_Hop4 | Mutator | 1 | 0 | |
| Sum | 135 | 73 | ||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vanheule, A.; De Boevre, M.; Moretti, A.; Scauflaire, J.; Munaut, F.; De Saeger, S.; Bekaert, B.; Haesaert, G.; Waalwijk, C.; Van der Lee, T.; et al. Genetic Divergence and Chemotype Diversity in the Fusarium Head Blight Pathogen Fusarium poae. Toxins 2017, 9, 255. https://doi.org/10.3390/toxins9090255
Vanheule A, De Boevre M, Moretti A, Scauflaire J, Munaut F, De Saeger S, Bekaert B, Haesaert G, Waalwijk C, Van der Lee T, et al. Genetic Divergence and Chemotype Diversity in the Fusarium Head Blight Pathogen Fusarium poae. Toxins. 2017; 9(9):255. https://doi.org/10.3390/toxins9090255
Chicago/Turabian StyleVanheule, Adriaan, Marthe De Boevre, Antonio Moretti, Jonathan Scauflaire, Françoise Munaut, Sarah De Saeger, Boris Bekaert, Geert Haesaert, Cees Waalwijk, Theo Van der Lee, and et al. 2017. "Genetic Divergence and Chemotype Diversity in the Fusarium Head Blight Pathogen Fusarium poae" Toxins 9, no. 9: 255. https://doi.org/10.3390/toxins9090255









