A Deeper Examination of Thorellius atrox Scorpion Venom Components with Omic Technologies
Abstract
:1. Introduction
2. Results and Discussion
2.1. RNA Extraction, RNA-Seq and Transcriptome Assembly
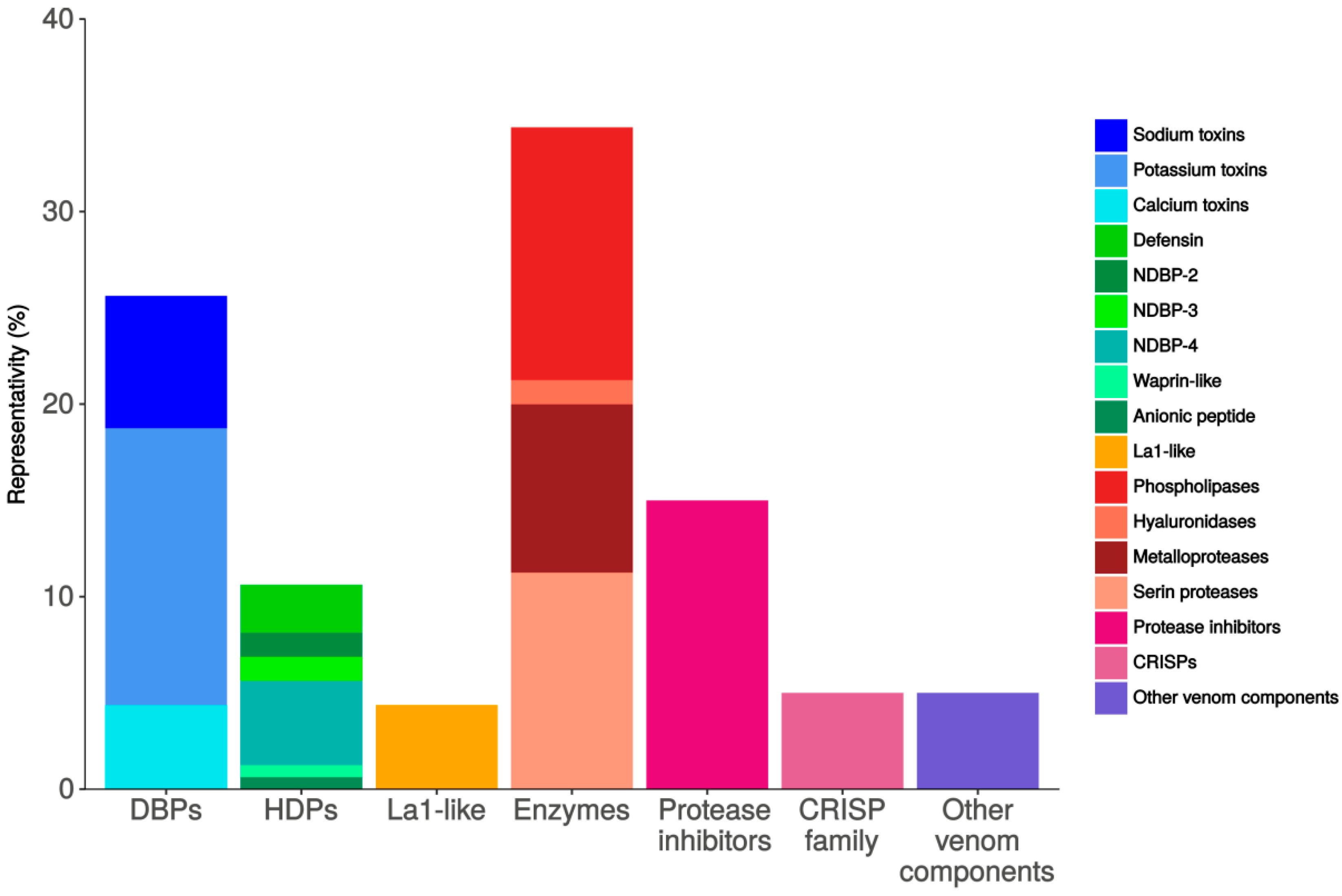
2.2. Transcriptome Analysis
2.3. Transcript Nomenclature
2.4. DBPs
2.4.1. Toxins Acting on Voltage-Gated Sodium Channels
2.4.2. Toxins Acting on Potassium Channels
2.4.3. Toxins Acting on Calcium Channels
2.5. Host Defense Peptides (HDPs)
2.6. Enzymes
2.7. Protease Inhibitors
2.8. Other Venom Components
2.8.1. La1-Like Peptides
2.8.2. CRISP Family
2.8.3. Other Undefined Venom Components
3. Mass Spectrometry Analysis
4. Conclusions
5. Material and Methods
5.1. Biological Material
5.2. Extraction of Total RNA from Venom Glands, RNA-Seq and Transcriptome Assembly
5.3. Mass Spectrometry Analysis
5.3.1. Nanoscale Liquid Chromatography-Mass Spectrometry
5.3.2. Molecular Mass Fingerprinting
5.3.3. Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS)
5.3.4. MS Data Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lourenço, W.R. The scorpion families and their geographical distribution. J. Venom. Anim. Toxins 2001, 7, 3–23. [Google Scholar] [CrossRef]
- Santibáñez-López, C.E.; Francke, O.F.; Ureta, C.; Possani, L.D. Scorpions from Mexico: From species diversity to venom complexity. Toxins 2016, 8, 2. [Google Scholar] [CrossRef] [PubMed]
- González-Santillán, E.; Prendini, L. Phylogeny of the North American Vaejovid scorpion subfamily Syntropinae Kraepelin, 1905, based on morphology, mitochondrial and nuclear DNA. Cladistics 2015, 31, 341–405. [Google Scholar] [CrossRef]
- Santibáñez-López, C.E.; Possani, L.D. Overview of the Knottin scorpion toxin-like peptides in scorpion venoms: Insights on their classification and evolution. Toxicon 2015, 107, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Ortiz, E.; Gurrola, G.B.; Schwartz, E.F.; Possani, L.D. Scorpion venom components as potential candidates for drug development. Toxicon 2015, 93, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Shi, W.; Zeng, X.C.; Ge, F.; Yang, M.; Nie, Y.; Bao, A.; Wu, S.; Guoji, E. Unique diversity of the venom peptides from the scorpion Androctonus bicolor revealed by transcriptomic and proteomic analysis. J. Proteom. 2015, 128, 231–250. [Google Scholar] [CrossRef] [PubMed]
- Santibáñez-López, C.E.; Cid-Uribe, J.I.; Batista, C.V.F.; Ortiz, E.; Possani, L.D. Venom gland transcriptomic and proteomic analyses of the enigmatic scorpion Superstitionia donensis (Scorpiones: Superstitioniidae), with insights on the evolution of its venom components. Toxins 2016, 8, 367. [Google Scholar] [CrossRef] [PubMed]
- Santibáñez-López, C.E.; Cid-Uribe, J.I.; Zamudio, F.Z.; Batista, C.V.F.; Ortiz, E.; Possani, L.D. Venom gland transcriptomic and venom proteomic analyses of the scorpion Megacormus gertschi Díaz-Najera, 1966 (Scorpiones: Euscorpiidae: Megacorminae). Toxicon 2017, 133, 95–109. [Google Scholar] [CrossRef] [PubMed]
- Jimenez-Vargas, J.M.; Quintero-Hernández, V.; Gonzalez-Morales, L.; Ortiz, E.; Possani, L.D. Design and expression of recombinant toxins from Mexican scorpions of the genus Centruroides for production of antivenoms. Toxicon 2017, 128, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Jimenez-Vargas, J.M.; Restano-Cassulini, R.; Quintero-Hernandez, V.; Gurrola, G.B.; Possani, L.D. Recombinant expression of the toxic peptide ErgTx1 and role of Met35 on its stability and function. Peptides 2011, 32, 560–567. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Carreto, S.; Quintero-Hernández, V.; Jiménez-Vargas, J.M.; Corzo, G.; Possani, L.D.; Becerril, B.; Ortiz, E. Gene cloning and functional characterization of four novel antimicrobial-like peptides from scorpions of the family Vaejovidae. Peptides 2012, 34, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Quintero-Hernández, V.; Ramírez-Carreto, S.; Romero-Gutiérrez, M.T.; Valdez-Velázquez, L.L.; Becerril, B.; Possani, L.D.; Ortiz, E. Transcriptome analysis of scorpion species belonging to the Vaejovis genus. PLoS ONE 2015, 10, e0117188. [Google Scholar] [CrossRef] [PubMed]
- Soleglad, M.E.; Fet, V. Contributions to scorpion systematics. III. Subfamilies Smeringurinae and Syntropinae (Scorpiones: Vaejovidae). Euscorpius 2008, 71, 1–115. [Google Scholar]
- Quintero-Hernández, V.; Jiménez-Vargas, J.M.; Gurrola, G.B.; Valdivia, H.H.; Possani, L.D. Scorpion venom components that affect ion-channels function. Toxicon 2013, 76, 328–342. [Google Scholar] [CrossRef] [PubMed]
- Cao, Z.; Di, Z.; Wu, Y.; Li, W. Overview of scorpion species from China and their toxins. Toxins 2014, 6, 796–815. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.; Zeng, X.C.; Zeng, X.; Nie, Y.; Zhang, L.; Wu, S.; Bao, A. Transcriptomic analysis of the venom glands from the scorpion Hadogenes troglodytes revealed unique and extremely high diversity of the venom peptides. J. Proteom. 2017, 150, 40–62. [Google Scholar] [CrossRef] [PubMed]
- Possani, L.D.; Becerril, B.; Delepierre, M.; Tytgat, J. Scorpion toxins specific for Na+-channels. Eur. J. Biochem. 1999, 264, 287–300. [Google Scholar] [CrossRef] [PubMed]
- Couraud, F.; Jover, E.; Dubois, J.M.; Rochat, H. Two types of scorpion toxin receptor sites, one related to the activation, the other to the inactivation of the action potential sodium channel. Toxicon 1982, 20, 9–16. [Google Scholar] [CrossRef]
- De la Vega, R.C.R.; Possani, L.D. Novel paradigms on scorpion toxins that affects the activating mechanism of sodium channels. Toxicon 2007, 49, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Mille, B.G.; Peigneur, S.; Predel, R.; Tytgat, J. Trancriptomic approach reveals the molecular diversity of Hottentotta conspersus (Buthidae) venom. Toxicon 2015, 99, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Soleglad, M.E.; Fet, V. The Systematics of the Scorpion Subfamily Uroctoninae (Scorpiones: Chactidae). Rev. Ibérica Aracnol. 2004, 10, 81–128. [Google Scholar]
- Valdez-Cruz, N.A.; Batista, C.V.F.; Zamudio, F.Z.; Bosmans, F.; Tytgat, J.; Possani, L.D. Phaiodotoxin, a novel structural class of insect-toxin isolated from the venom of the Mexican scorpion Anuroctonus phaiodactylus. Eur. J. Biochem. 2004, 271, 4753–4761. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Ma, Y.; He, Y.; Di, Z.; Wu, Y.; Cao, Z.; Li, W. Comparative venom gland transcriptome analysis of the scorpion Lychas mucronatus reveals intraspecific toxic gene diversity and new venomous components. BMC Genom. 2010, 11, 452. [Google Scholar] [CrossRef]
- Srairi-Abid, N.; Guijarro, J.I.; Benkhalifa, R.; Mantegazza, M.; Cheikh, A.; Ben Aissa, M.; Haumont, P.-Y.; Delepierre, M.; El Ayeb, M. A new type of scorpion Na+-channel-toxin-like polypeptide active on K+ channels. Biochem. J. 2005, 388, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Luna-Ramírez, K.; Quintero-Hernández, V.; Juárez-González, V.R.; Possani, L.D. Whole transcriptome of the venom gland from Urodacus yaschenkoi scorpion. PLoS ONE 2015, 10, e0127883. [Google Scholar] [CrossRef] [PubMed]
- Tytgat, J.; Chandy, K.G.; Garcia, M.L.; Gutman, G.A.; Martin-Eauclaire, M.F.; van der Walt, J.J.; Possani, L.D. A unified nomenclature for short-chain peptides isolated from scorpion venoms: Alpha-KTx molecular subfamilies. Trends Pharmacol. Sci. 1999, 20, 444–447. [Google Scholar] [CrossRef]
- Ramirez, K.L.; Jimenez Vargas, J.M. Scorpine-like peptides. Single Cell Biol. 2016, 5, 2–4. [Google Scholar] [CrossRef]
- Possani Postay, L.D.; Gurrola-Briones, G.; Salas-Castillo, S.P.; Ferreira Batista, C.V.; Varga, Z.S.; Panyi, G.; Gáspár, R. VM23 and VM24, Two Scorpion Peptides that Block Human T-Lymphocyte Potassium Channels (Sub-Type Kv 1.3) with High Selectivity and Decrease the In Vivo DTH-Responsesin Rats. Patent WO2008139243A1, 20 November 2008. [Google Scholar]
- Conde, R.; Zamudio, F.Z.; Rodr, M.H.; Possani, L.D. Scorpine, an anti-malaria and anti-bacterial agent purifed from scorpion venom. FEBS Lett. 2000, 471, 165–168. [Google Scholar] [CrossRef]
- Zhu, S.; Tytgat, J. The scorpine family of defensins: gene structure, alternative polyadenylation and fold recognition. Cell. Mol. Life Sci. 2004, 61, 1751–1763. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, E.; Diego-garcia, E.; Rodríguez de la Vega, R.; Possani, L.D. Transcriptome analysis of the venom gland of the Mexican scorpion Hadrurus gertschi (Arachnida: Scorpiones). BMC Genom. 2007, 8, 119. [Google Scholar] [CrossRef] [PubMed]
- Chagot, B.; Pimentel, C.; Dai, L.; Pil, J.; Tytgat, J.; Nakajima, T.; Corzo, G.; Darbon, H.; Ferrat, G. An unusual fold for potassium channel blockers: NMR structure of three toxins from the scorpion Opisthacanthus madagascariensis. Biochem. J. 2005, 388, 263–271. [Google Scholar] [CrossRef] [PubMed]
- Vandendriessche, T.; Kopljar, I.; Jenkins, D.P.; Diego-Garcia, E.; Abdel-Mottaleb, Y.; Vermassen, E.; Clynen, E.; Schoofs, L.; Wulff, H.; Snyders, D.; et al. Purification, molecular cloning and functional characterization of HelaTx1 (Heterometrus laoticus): The first member of a new k-KTX subfamily. Biochem. Pharmacol. 2012, 83, 1307–1317. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Luo, F.; Feng, J.; Yang, W.; Zeng, D.; Zhao, R.; Cao, Z.; Liu, M.; Li, W.; Jiang, L.; et al. Genomic and structural characterization of Kunitz-Type peptide LmKTT-1a highlights diversity and evolution of scorpion potassium channel toxins. PLoS ONE 2013, 8, e60201. [Google Scholar] [CrossRef] [PubMed]
- Chuang, R.S.; Jaffe, H.; Cribbs, L.; Perez-Reyes, E.; Swartz, K.J. Inhibition of T-type voltage-gated calcium channels by a new scorpion toxin. Nat. Neurosci. 1998, 1, 668–674. [Google Scholar] [CrossRef] [PubMed]
- Sidach, S.S.; Mintz, I.M. Kurtoxin, a gating modifier of neuronal high- and low-threshold ca channels. J. Neurosci. 2002, 22, 2023–2034. [Google Scholar]
- Olamendi-Portugal, T.; Garcia, B.I.; Lopez-Gonzalez, I.; Van Der Walt, J.; Dyason, K.; Ulens, C.; Tytgat, J.; Felix, R.; Darszon, A.; Possani, L.D. Two new scorpion toxins that target voltage-gated Ca2+ and Na+ channels. Biochem. Biophys. Res. Commun. 2002, 299, 562–568. [Google Scholar] [CrossRef]
- López-González, I.; Olamendi-Portugal, T.; De La Vega-Beltrán, J.L.; Van Der Walt, J.; Dyason, K.; Possani, L.D.; Felix, R.; Darszon, A. Scorpion toxins that block T-type Ca2+ channels in spermatogenic cells inhibit the sperm acrosome reaction. Biochem. Biophys. Res. Commun. 2003, 300, 408–414. [Google Scholar] [CrossRef]
- Schwartz, E.F.; Michelle Capes, E.; Diego-García, E.; Zamudio, F.Z.; Fuentes, O.; Possani, L.D.; Valdivia, H.H. Characterization of hadrucalcin, a peptide from Hadrurus gertschi scorpion venom with pharmacological activity on ryanodine receptors. Br. J. Pharmacol. 2009, 157, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Xiao, L.; Gurrola, G.B.; Zhang, J.; Valdivia, C.R.; SanMartin, M.; Zamudio, F.Z.; Zhang, L.; Possani, L.D.; Valdivia, H.H. Structure-function relationships of peptides forming the calcin family of ryanodine receptor ligands. J. Gen. Physiol. 2016, 147, 375–394. [Google Scholar] [CrossRef] [PubMed]
- Vargas-Jaimes, L.; Xiao, L.; Zhang, J.; Possani, L.D.; Valdivia, H.H.; Quintero-Hernandez, V. Recombinant expression of Intrepicalcin from the scorpion Vaejovis intrepidus and its effect on skeletal ryanodine receptors. Biochim. Biophys. Acta 2017, 1861, 936–946. [Google Scholar] [CrossRef] [PubMed]
- Matsushita, N.; Miyashita, M.; Sakai, A.; Nakagawa, Y.; Miyagawa, H. Purification and characterization of a novel short-chain insecticidal toxin with two disulfide bridges from the venom of the scorpion Liocheles australasiae. Toxicon 2007, 50, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.J.; Vetter, I.; Lewis, R.J.; Peigneur, S.; Tytgat, J.; Lam, A.; Gallant, E.M.; Beard, N.A.; Alewood, P.F.; Dulhunty, A.F. Multiple actions of phi-LITX-Lw1a on ryanodine receptors reveal a functional link between scorpion DDH and ICK toxins. Proc. Natl. Acad. Sci. USA 2013, 110, 8906–8911. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.E. Agatoxins: Ion channel specific toxins from the american funnel web spider, Agelenopsis aperta. Toxicon 2004, 43, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Hancock, R.E.W.; Haney, E.F.; Gill, E.E. The immunology of host defence peptides: Beyond antimicrobial activity. Nat. Rev. Immunol. 2016, 16, 321–334. [Google Scholar] [CrossRef] [PubMed]
- Steinstraesser, L.; Kraneburg, U.; Jacobsen, F.; Al-Benna, S. Host defense peptides and their antimicrobial-immunomodulatory duality. Immunobiology 2011, 216, 322–333. [Google Scholar] [CrossRef] [PubMed]
- Cuperus, T.; Coorens, M.; van Dijk, A.; Haagsman, H.P. Avian host defense peptides. Dev. Comp. Immunol. 2013, 41, 352–369. [Google Scholar] [CrossRef] [PubMed]
- Oppenheim, J.J.; Biragyn, A.; Kwak, L.W.; Yang, D. Roles of antimicrobial peptides such as defensins in innate and adaptive immunity. Ann. Rheum. Dis 2003, 62, ii17–ii21. [Google Scholar] [CrossRef] [PubMed]
- Wilmes, M.; Sahl, H.G. Defensin-based anti-infective strategies. Int. J. Med. Microbiol. 2014, 304, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Xie, Z.; Zhang, Q.; Li, Y.; Yang, F.; Chen, Z.; Li, W.; Cao, Z.; Wu, Y. Scorpion potassium channel-blocking defensin highlights a functional link with neurotoxin. J. Biol. Chem. 2016, 291, 7097–7106. [Google Scholar] [CrossRef] [PubMed]
- Brogden, K.A. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nat. Rev. Microbiol. 2005, 3, 238–250. [Google Scholar] [CrossRef] [PubMed]
- Almaaytah, A.; Albalas, Q. Scorpion venom peptides with no disulfide bridges: A review. Peptides 2014, 51, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Aponte, C.A.; Silva-Sanchez, J.; Quintero-Hernández, V.; Rodríguez-Romero, A.; Balderas, C.; Possani, L.D.; Gurrola, G.B. Vejovine, a new antibiotic from the scorpion venom of Vaejovis mexicanus. Toxicon 2011, 57, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Nie, Y.; Zeng, X.C.; Cao, H.; Zhang, L.; Zhou, L.; Yang, Y.; Luo, X.; Liu, Y. Genomic and functional characterization of three new venom peptides from the scorpion Heterometrus spinifer. Peptides 2014, 53, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Melo, M.M.A.; Daniele-Silva, A.; Teixeira, D.G.; Estrela, A.B.; Melo, K.R.T.; Oliveira, V.S.; Rocha, H.A.O.; Ferreira, L.D.S.; Pontes, D.L.; Lima, J.P.M.S.; et al. Structure and in vitro activities of a Copper II-chelating anionic peptide from the venom of the scorpion Tityus stigmurus. Peptides 2017, 94, 91–98. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, U.C.; Candido, D.M.; Coronado Dorce, V.A.; Junqueira-De-Azevedo, I.D.L.M. The transcriptome recipe for the venom cocktail of Tityus bahiensis scorpion. Toxicon 2015, 95, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Harris, F.; Dennison, S.; Phoenix, D. Anionic antimicrobial peptides from eukaryotic organisms. Curr. Protein Pept. Sci. 2009, 10, 585–606. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Wang, Y.; Wei, L.; Ye, H.; Liu, H.; Wang, L.; Liu, R.; Li, D.; Lai, R. Snake venom-like waprin from the frog of Ceratophrys calcarata contains antimicrobial function. Gene 2013, 514, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, K.; Kikuchi, T.; Endo, Y.; Usui, K.; Takahashi, M.; Shibata, N.; Kusakabe, T.; Xin, H.; Hoshi, S.; Miki, M.; et al. Mouse SWAM1 and SWAM2 are antibacterial proteins composed of a single whey acidic protein motif. J. Immunol. 2003, 170, 1973–1979. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, P.L.; Fletcher, M.D.; Weninger, K.; Anderson, T.E.; Martin, B.M. Vesicle-associated membrane protein (VAMP) cleavage by a new metalloprotease from the Brazilian scorpion Tityus serrulatus. J. Biol. Chem. 2010, 285, 7405–7416. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, P.B. The contribution of proteinase inhibitors to immune defense. Trends Immunol. 2001, 22, 47–52. [Google Scholar] [CrossRef]
- Miyashita, M.; Otsuki, J.; Hanai, Y.; Nakagawa, Y.; Miyagawa, H. Characterization of peptide components in the venom of the scorpion Liocheles australasiae (Hemiscorpiidae). Toxicon 2007, 50, 428–437. [Google Scholar] [CrossRef] [PubMed]
- Martinez, G.; Hograindleur, J.P.; Voisin, S.; Nahed, R.A.; Aziz, T.M.A.E.; Escoffier, J.; Bessonnat, J.; Fovet, C.M.; Waard, M.D.; Hennebicq, S.; et al. Spermaurin, an La1-like peptide from the venom of the scorpion Scorpio maurus palmatus, improves sperm motility and fertilization in different mammalian species. Mol. Hum. Reprod. 2017, 23, 116–131. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, G.M.; Roelants, K.; O’Bryan, M.K. The CAP superfamily: Cysteine-rich secretory proteins, antigen 5, and pathogenesis-related 1 proteins—Roles in reproduction, cancer, and immune defense. Endocr. Rev. 2008, 29, 865–897. [Google Scholar] [CrossRef] [PubMed]
- Adade, C.M.; Carvalho, A.L.O.; Tomaz, M.A.; Costa, T.F.R.; Godinho, J.L.; Melo, P.A.; Lima, A.P.C.A.; Rodrigues, J.C.F.; Zingali, R.B.; Souto-Padrón, T. Crovirin, a snake venom Cysteine-Rich Secretory Protein (CRISP) with promising activity against Trypanosomes and Leishmania. PLoS Negl. Trop. Dis. 2014, 8, e3252. [Google Scholar] [CrossRef] [PubMed]
- Valdez-Velázquez, L.L.; Quintero-Hernández, V.; Romero-Gutiérrez, M.T.; Coronas, F.I.V.; Possani, L.D. Mass fingerprinting of the venom and transcriptome of venom gland of scorpion Centruroides tecomanus. PLoS ONE 2013, 8, e66486. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]






| Species Code | Meaning | Family Code | Meaning | Subtype Code | Meaning | Example |
|---|---|---|---|---|---|---|
| Tat | T. atrox | NaT | Na-channel | Alp | Alpha-Na Toxin | TatNaTAlp01 |
| Toxins | Bet | Beta-Na Toxin | TatNaTBet01 | |||
| KTx | K-channel Toxins | Alp | Alpha-K Toxin | TatKTxAlp01 | ||
| Bet | Beta-K Toxin | TatKTxBet01 | ||||
| Kap | Kappa-K Toxin | TatKTxKap01 | ||||
| Del | Delta-K Toxin | TatKTxDel01 | ||||
| Scr | Scorpin-like | TatKTxScr01 | ||||
| CaT | Ca-channel Toxins | Clc | Calcin | TatCaTClc01 | ||
| Lio | Liotoxin-like | TatCaTLio01 | ||||
| Ome | Omegascorpin | TatCaTOme01 | ||||
| HDP | Host Defense Peptides | Def | Defensin | TatHDPDef01 | ||
| ND1–5 | NDBPs families 1–5 | TatHDPND201 | ||||
| Ani | Anionic peptide | TatHDPAni01 | ||||
| Wap | Waprin-like | TatHDPWap01 | ||||
| Enz | Enzymes | PA2 | Phospholipase A2 | TatEnzPA201 | ||
| PLB | Phospholipase B | TatEnzPLB01 | ||||
| PLD | Phospholipase D | TatEnzPLD01 | ||||
| SeP | Serine protease | TatEnzSeP01 | ||||
| MtP | Metalloprotease | TatEnzMtP01 | ||||
| Hya | Hyaluronidase | TatEnzHya01 | ||||
| Pin | Protease | Srp | Serpin-like | TatPInSrp01 | ||
| Inhibitors | Kun | Kunitz-type | TatPInKun01 | |||
| Oth | Other | La1 | La1-like | TatOthLa101 | ||
| venom | CRI | CRISP | TatOthCRI01 | |||
| components | Und | Undefined | TatOthUnd01 |
| ID | NDBp Family | Length of the Mature Peptide | Hydrophobicity | Hydrophobic Moment | Charge |
|---|---|---|---|---|---|
| TatHDPND202 | NDBP-2 | 47 | 0.263 | 0.065 | +6 |
| TatHDPND201 | NDBP-2 | 43 | 0.212 | 0.089 | +6 |
| TatHDPND301 | NDBP-3 | 19 | 0.903 | 0.377 | +2 |
| TatHDPND302 | NDBP-3 | 25 | 0.495 | 0.327 | +4 |
| TatHDPND405 | NDBP-4 | 13 | 0.819 | 0.606 | +1 |
| TatHDPND406 | NDBP-4 | 13 | 0.742 | 0.458 | +1 |
| TatHDPND407 | NDBP-4 | 13 | 0.746 | 0.456 | +1 |
| TatHDPND402 | NDBP-4 | 13 | 0.778 | 0.779 | 0 |
| TatHDPND403 | NDBP-4 | 13 | 0.752 | 0.792 | +1 |
| TatHDPND401 | NDBP-4 | 13 | 0.793 | 0.595 | +1 |
| RT 1 (min) | MW 2 (Da) | RT (min) | MW (Da) |
|---|---|---|---|
| 1–20 | 1462.7, 2057.24, 2117.68, 2265.06, 2796.27, 3111.96, 9115.86, 10663.93, 11,123.28, | 140–160 | 1944.15, 2645.50, 2815.60, 6330.03, 6473.90, 6714.40, 7438.62, 7639.27, 7843.00, 8049.12, 8213.16, 8829.81, 8950.11,9535.2 |
| 20–40 | 1076.62, 1205.68, 1212.80, 1673.85, 1817.88, 3427.38, 3499.92, 3586.92, 3878.10, 4197.53, 12,306.36 | 160–180 | 1337.72, 1497.81, 2193.06, 2248.28, 2347.32, 3338.30, 7040.46, 7956.10, 8201.97, 8727.14 |
| 40–60 | 1331.64, 1799.04, 1886.82, 2333.32, 2411.36, 2447.40, 2592.26, 3777.63, 3945.62, 5813.52 | 180–200 | 1296.10, 2151.20, 4171.38, 4302.42, 4389.42, 4697.56, 4762.08, 6195.66 |
| 60–80 | 2377.16, 2850.1, 2944.70, 3606.60, 4485.10, 4595.04, 5279.52, 5654.40 | 200–220 | 10,039.5, 13,729.41, 14,079.03 |
| 80–100 | 3332.90, 3535.47, 3718.60, 3787.85, 4113.96, 4125.80, 4204.00, 4279.05, 4290.36, 5196.42, 5756.56, 7011.33, 7123.96, 7236.99, 8126.40, 8328.51 | 220–240 | 1828.00, 6554.31, 6750.45, 6946.57, 7269.84, 8272.50, 10,545.20, 12,430.9, 13,591.92, 13,815.51, 14,614.72 |
| 100–120 | 3223.80, 3243.80, 3569.92, 3767.15, 4250.67, 8468.54, 8581.60, 8716.70, 9056.88, 9490.25 | 240–260 | 3821.44, 5409.48, 10,882.9, 16,915.41 |
| 120–140 | 1198.64, 1648.86, 3267.39, 4036.16, 4348.40, 4561.84, 4815.2 | 260–290 | 2038.11, 3347.5, 4505.55, 4791.65, 4949.7, 8355.48, 11,174.46, 11,833.92, 11,847.44, 11,899.27, 13,891.59, 14,257.71, 14,705.56, 14,741.70 |
| Sodium toxins | |||
| Transcriptome ID | Theoretical Mass | Experimental Mass | RT Range |
| TatNaTBet03 | 5196.79 | 5196.42 | 80–100 |
| TatNaTBet08 | 6195.85 | 6195.66 | 180–200 |
| Potassium toxins | |||
| TatKTxAlp10 | 3607.43 | 3606.60 | 60–80 |
| TatKTxAlp12 | 4114.86 | 4113.96 | 100–120 |
| Calcium toxins | |||
| TatCaTClc01 | 3788.48 | 3787.85 | 80–100 |
| Transcriptome ID | Score | Coverage | Protein Type | Accession Number of the Reference Protein |
|---|---|---|---|---|
| comp8310_c0_seq1 | 46.06 | 19.1% | Allatostatins-like | XP_013775495 |
| comp32030_c1_seq1 | 28.07 | 34.8% | Angiotensin-converting enzyme | XP_013773749 |
| comp32030_c2_seq1 | 32.73 | 7.7% | Angiotensin-converting enzyme | XP_013773749 |
| comp33161_c0_seq1 | 535.88 | 24.5% | Angiotensin-converting enzyme | XP_013773749 |
| comp33725_c0_seq1 | 65.33 | 16.8% | Angiotensin-converting enzyme | XP_013773749 |
| comp33936_c0_seq1 | 64.74 | 13.1% | Angiotensin-converting enzyme | XP_013773749 |
| TatCaTClc01 | 88.83 | 24.2% | Calcium toxin. Calcin | A0A1L4BJ42 |
| comp32319_c0_seq1 | 18.83 | 7.56% | Ectonucleoside triphosphate diphosphohydrolase 2-like | XP_013778001 |
| comp881_c0_seq1 | 452.55 | 18.7% | Elastase-like protein | CAX51421 |
| TatHDPND201 | 513.12 | 46.7% | HDP. NDBP-2 family | F1AWB0 |
| TatHDPND301 | 22.46 | 94.7% | HDP. NDBP-3 family | ALG64974 |
| ViVlp1 | 762.84 | 28.6% | HDP. NDBP-2 family | AGK88593 |
| ViAMP1 | 188.70 | 70.8% | HDP. NDBP-3 family | ALG64975 |
| TaHDPND401 | 254.77 | 61.5% | HDP. NDBP-4 family | I0DEB5 |
| TatHDPND403 | 37.65 | 100% | HDP. NDBP-4 family | I0DEB5 |
| ViCT2 | 882.53 | 76.9% | HDP. NDBP-4 family | I0DEB3 |
| TatEnzHya01 | 161.40 | 39.1% | Hyaluronidase | API81375 |
| comp15335_c0_seq1 | 92.74 | 10.3% | Hypothetical protein | CAX51393 |
| comp30560_c0_seq1 | 67.45 | 9.1% | Hypothetical protein | AEX09195 |
| comp31101_c0_seq1 | 103.49 | 29.1% | Hypothetical protein (allergen type) | CAX51409 |
| comp30730_c0_seq1 | 16.64 | 6.5% | Hypothetical protein RvY_03950 | GAU91754 |
| ViLa1lp1 | 47.91 | 63.3% | La1-like | AOF40216 |
| TatOthLa101 | 469.44 | 45.5% | La1-like | AOF40202 |
| comp34524_c0_seq1 | 74.23 | 11.5% | Metalloproteinase | XP_009865190 |
| TatEnzMtp04 | 31.25 | 18.8% | Metalloproteinase | AMO02513 |
| comp32637_c0_seq1 | 828.12 | 42% | Nucleotidase | XP_013774694 |
| comp26928_c1_seq1 | 214.51 | 23.2% | Other venom components | N/A |
| comp27809_c1_seq1 | 1255.98 | 24.4% | Other venom components | N/A |
| comp30392_c0_seq1 | 34.49 | 32.9% | Other venom components | CAX51433 |
| comp32982_c0_seq3 | 20.83 | 13.7% | Other venom components | N/A |
| comp43100_c0_seq1 | 70.29 | 15.3% | Other venom components | N/A |
| comp31198_c0_seq1 | 20.19 | 3.13% | Other venom components | N/A |
| TatEnzPA201 | 1616.89 | 45.5% | Phospholipase A2 | API81339 |
| TatEnzPA213 | 253.38 | 27% | Phospholipase A2 | API81335 |
| TatEnzPA215 | 877.33 | 50.2% | Phospholipase A2 | API81335 |
| TatEnzPA202 | 94.96 | 31.3% | Phospholipase A2 | API81335 |
| comp20627_c0_seq1 | 14.64 | 0.9% | Protein kinase C-binding protein NELL2-like | XP_022243213 |
| TatOthCRI06 | 24.43 | 15.9% | Putative cysteine-rich protein | JAT91149 |
| TatOthCRI07 | 10.69 | 23.1% | Putative cysteine-rich protein | API81352 |
| comp30427_c0_seq1 | 16.87 | 2.27% | Steryl-sulfatase-like isoform | XP_0193859 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Romero-Gutierrez, T.; Peguero-Sanchez, E.; Cevallos, M.A.; Batista, C.V.F.; Ortiz, E.; Possani, L.D. A Deeper Examination of Thorellius atrox Scorpion Venom Components with Omic Technologies. Toxins 2017, 9, 399. https://doi.org/10.3390/toxins9120399
Romero-Gutierrez T, Peguero-Sanchez E, Cevallos MA, Batista CVF, Ortiz E, Possani LD. A Deeper Examination of Thorellius atrox Scorpion Venom Components with Omic Technologies. Toxins. 2017; 9(12):399. https://doi.org/10.3390/toxins9120399
Chicago/Turabian StyleRomero-Gutierrez, Teresa, Esteban Peguero-Sanchez, Miguel A. Cevallos, Cesar V. F. Batista, Ernesto Ortiz, and Lourival D. Possani. 2017. "A Deeper Examination of Thorellius atrox Scorpion Venom Components with Omic Technologies" Toxins 9, no. 12: 399. https://doi.org/10.3390/toxins9120399







