Emergence of Nasal Carriage of ST80 and ST152 PVL+ Staphylococcus aureus Isolates from Livestock in Algeria
Abstract
:1. Introduction
2. Results
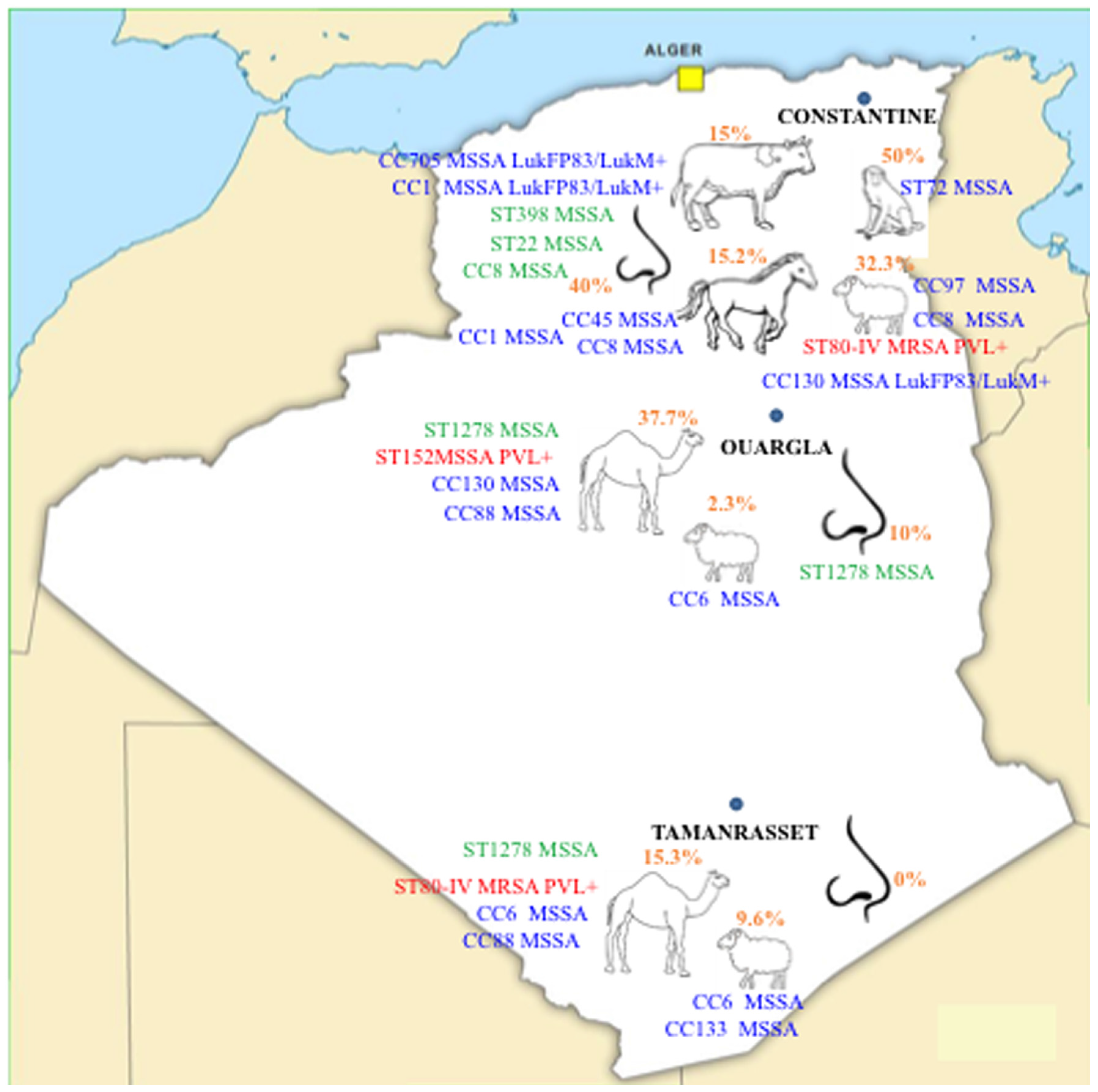
2.1. Prevalence of Nasal Colonization with S. aureus
2.2. Antimicrobial Resistance
2.3. CCs Distribution
2.4. Virulence Genes
2.5. Resistance Genes
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Study Population
5.2. Specimen Collection and Bacterial Isolation
5.3. Identification and Susceptibility Testing
5.4. Oligonucleotide DNA Arrays and Genotyping
5.5. Statistical Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- McCaig, L.F.; McDonald, L.C.; Mandal, S.; Jernigan, D.B. Staphylococcus aureus-associated skin and soft tissue infections in ambulatory care. Emerg. Infect. Dis. 2006, 12, 1715–1723. [Google Scholar] [CrossRef] [PubMed]
- Wertheim, H.F.; Melles, D.C.; Vos, M.C.; Van Leeuwen, W.; Van Belkum, A.; Verbrugh, H.A.; Nouwen, J.L. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect. Dis. 2005, 5, 751–762. [Google Scholar] [CrossRef]
- Iverson, S.A.; Brazil, A.M.; Ferguson, J.M.; Nelson, K.; Lautenbach, E.; Rankin, S.C.; Morris, D.O.; Davis, M.F. Anatomical patterns of colonization of pets with staphylococcal species in homes of people with methicillin-resistant Staphylococcus aureus (MRSA) skin or soft tissue infection (SSTI). Vet. Microbiol. 2015, 176, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Akoda, E.; Zhang, K. Methicillin-Resistant Staphylococcus aureus carriage among students at a historically black university: A Case Study. Int. J. Microbiol. 2013, 2013, 979734. [Google Scholar] [CrossRef] [PubMed]
- Woolhouse, M.; Ward, M.; Van Bunnik, B.; Farrar, J. Antimicrobial resistance in humans, livestock and the wider environment. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370, 20140083. [Google Scholar] [CrossRef] [PubMed]
- Graveland, H.; Wagenaar, J.A.; Bergs, K.; Heesterbeek, H.; Heederik, D. Persistence of livestock associated MRSA CC398 in humans is dependent on intensity of animal contact. PLoS ONE 2011, 6, e16830. [Google Scholar] [CrossRef] [PubMed]
- Abdulgader, S.M.; Shittu, A.O.; Nicol, M.P.; Kaba, M. Molecular epidemiology of methicillin-resistant Staphylococcus aureus in Africa: A systematic review. Front. Microbiol. 2015, 6, 348. [Google Scholar] [CrossRef] [PubMed]
- Gharsa, H.; Slama, K.B.; Lozano, C.; Gómez-Sanz, E.; Klibi, N.; Sallem, R.B.; Gómez, P.; Zarazaga, M.; Boudabous, A.; Torres, C. Prevalence, antibiotic resistance, virulence traits and genetic lineages of Staphylococcus aureus in healthy sheep in Tunisia. Vet. Microbiol. 2012, 156, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Raji, M.A.; Garaween, G.; Ehricht, R.; Monecke, S.; Shibl, A.M.; Senok, A. Genetic Characterization of Staphylococcus aureus isolated from retail meat in Riyadh, Saudi Arabia. Front. Microbiol. 2016, 7, 911. [Google Scholar] [CrossRef] [PubMed]
- Stegger, M.; Wirth, T.; Andersen, P.S.; Skov, R.L.; De Grassi, A.; Simões, P.M.; Tristan, A.; Petersen, A.; Aziz, M.; Kiil, K.; et al. Origin and evolution of European community-acquired methicillin-resistant Staphylococcus aureus. MBio 2014, 5, e01044-14. [Google Scholar] [CrossRef] [PubMed]
- Djahmi, N.; Messad, N.; Nedjai, S.; Moussaoui, A.; Mazouz, D.; Richard, J.L.; Sotto, A.; Lavigne, J.P. Molecular epidemiology of Staphylococcus aureus strains isolated from inpatients with infected diabetic foot ulcers in an Algerian University Hospital. Clin. Microbiol. Infect. 2013, 19, E398–E404. [Google Scholar] [CrossRef] [PubMed]
- Schaumburg, F.; Köck, R.; Friedrich, A.W.; Soulanoudjingar, S.; Ngoa, U.A.; von Eiff, C.; Issifou, S.; Kremsner, P.G.; Hermann, M.; Peters, G.; et al. Population structure of Staphylococcus aureus from remote African Babongo Pygmies. PLoS Negl. Trop. Dis. 2011, 5, e1150. [Google Scholar] [CrossRef] [PubMed]
- Lozano, C.; Gharsa, H.; Ben Slama, K.; Zarazaga, M.; Torres, C. Staphylococcus aureus in animals and food: Methicillin resistance, prevalence and population structure. A review in the African continent. Microorganisms 2016, 4, E12. [Google Scholar] [CrossRef] [PubMed]
- Al Laham, N.; Mediavilla, J.R.; Chen, L.; Abdelateef, N.; Elamreen, F.A.; Ginocchio, C.C.; Pierard, D.; Becker, K.; Kreiswirth, B.N. MRSA clonal complex 22 strains harboring toxic shock syndrome toxin (TSST-1) are endemic in the primary hospital in Gaza, Palestine. PLoS ONE 2015, 10, e0120008. [Google Scholar] [CrossRef] [PubMed]
- Biber, A.; Abuelaish, I.; Rahav, G.; Raz, M.; Cohen, L.; Valinsky, L.; Taran, D.; Goral, A.; Elhamdany, A.; Regev-Yochay, G.; et al. A typical hospital-acquired methicillin-resistant Staphylococcus aureus clone is widespread in the community in the Gaza strip. PLoS ONE 2012, 7, e42864. [Google Scholar] [CrossRef] [PubMed]
- Rasigade, J.P.; Laurent, F.; Lina, G.; Meugnier, H.; Bes, M.; Vandenesch, F.; Etienne, J.; Tristan, A. Global distribution and evolution of Panton-Valentine leukocidin-positive methicillin-susceptible Staphylococcus aureus, 1981–2007. J. Infect. Dis. 2010, 201, 1589–1597. [Google Scholar] [CrossRef] [PubMed]
- Schaumburg, F.; Pauly, M.; Anoh, E.; Mossoun, A.; Wiersma, L.; Schubert, G.; Flammen, A.; Alabi, A.S.; Muyembe-Tamfum, J.J.; Grobusch, M.P.; et al. Staphylococcus aureus complex from animals and humans in three remote African regions. Clin. Microbiol. Infect. 2015, 21, e1–e8. [Google Scholar] [CrossRef] [PubMed]
- Boss, R.; Cosandey, A.; Luini, M.; Artursson, K.; Bardiau, M.; Breitenwieser, F.; Hehenberger, E.; Lam, T.; Mansfeld, M.; Michel, A.; et al. Bovine Staphylococcus aureus: Subtyping, evolution, and zoonotic transfer. J. Dairy Sci. 2016, 99, 515–528. [Google Scholar] [CrossRef] [PubMed]
- Budd, K.E.; McCoy, F.; Monecke, S.; Cormican, P.; Mitchell, J.; Keane, O.M. Extensive genomic diversity among bovine-adapted Staphylococcus aureus: Evidence for a genomic rearrangement within CC97. PLoS ONE 2015, 10, e0134592. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burton, S.; Reid-Smith, R.; McClure, J.T.; Weese, J.S. Staphylococcus aureus colonization in healthy horses in Atlantic Canada. Can. Vet. J. 2008, 49, 797–799, PMCID:PMC2465786. [Google Scholar] [PubMed]
- Mai-siyama, I.B.; Okon, K.O.; Adamu, N.B.; Askira, U.M.; Isyaka, T.M.; Adamu, S.G. Methicillin-resistant Staphylococcus aureus (MRSA) colonization rate among ruminant animals slaughtered for human consumption and contact persons in Maiduguri, Nigeria. Afr. J. Microbiol. Res. 2014, 8, 2643–2649. [Google Scholar] [CrossRef]
- Schaumburg, F.; Mugisha, L.; Peck, B.; Becker, K.; Gillespie, T.R.; Peters, G.; Leendertz, F.H. Drug-resistant human Staphylococcus aureus in sanctuary apes pose a threat to endangered wild ape populations. Am. J. Primatol. 2012, 74, 1071–1075. [Google Scholar] [CrossRef] [PubMed]
- Jaradat, Z.; Al Aboudi, A.; Shatnawi, M.; Ababneh, Q. Staphylococcus aureus isolates from camels differ in coagulase production, genotype and methicillin resistance gene profiles. J. Microbiol. Biotechnol. Food Sci. 2013, 2, 2455–2461. [Google Scholar]
- Al-Thani, R.F.; Al-Ali, F. Incidences and antimicrobial susceptibility profile of Staphylococcus species isolated from animals in different Qatari farms. Afr. J. Microbiol. 2012, 6, 7454–7458. [Google Scholar] [CrossRef]
- Alzohairy, M.A. Colonization and antibiotic susceptibility pattern of methicillin resistance Staphylococcus aureus (MRSA) among farm animals in Saudi Arabia. Afr. J. Bacteriol. Res. 2011, 3, 63–68. [Google Scholar]
- Vautor, E.; Abadie, G.; Guibert, J.M.; Chevalier, N.; Pepin, M. Nasal carriage of Staphylococcus aureus in dairy sheep. Vet. Microbiol. 2005, 106, 235–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sung, J.M.; Lloyd, D.H.; Lindsay, J.A. Staphylococcus aureus host specificity: Comparative genomics of human versus animal isolates by multi-strain microarray. Microbiology 2008, 154, 1949–1959. [Google Scholar] [CrossRef] [PubMed]
- Herron-Olson, L.; Fitzgerald, J.R.; Musser, J.M.; Kapur, V. Molecular correlates of host specialization in Staphylococcus aureus. PLoS ONE 2007, 2, e1120. [Google Scholar] [CrossRef] [PubMed]
- Van Duijkeren, E.; Wolfhagen, M.J.; Heck, M.E.; Wannet, W.J. Transmission of a Panton-Valentine leukocidin-positive, methicillin-resistant Staphylococcus aureus strain between humans and a dog. J. Clin. Microbiol. 2005, 43, 6209–6211. [Google Scholar] [CrossRef] [PubMed]
- Fall, C.; Seck, A.; Richard, V.; Ndour, M.; Sembene, M.; Laurent, F.; Breurec, S. Epidemiology of Staphylococcus aureus in pigs and farmers in the largest farm in Dakar, Senegal. Foodborne Pathog. Dis. 2012, 9, 962–965. [Google Scholar] [CrossRef] [PubMed]
- Rainard, P.; Corrales, J.C.; Barrio, M.B.; Cochard, T.; Poutrel, B. Leucotoxic activities of Staphylococcus aureus strains isolated from cows, ewes, and goats with mastitis: Importance of LukM/LukF-PV leukotoxin. Clin. Diagn. Lab. Immunol. 2003, 10, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, M.; Ohta, T.; Uchiyama, I.; Baba, T.; Yuzawa, H.; Kobayashi, I.; Cui, L.; Oguchi, A.; Aoki, K.I.; Nagai, Y.; et al. Whole genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 2001, 357, 1225–1240. [Google Scholar] [CrossRef]
- Haveri, M.; Roslöf, A.; Rantala, L.; Pyörälä, S. Virulence genes of bovine Staphylococcus aureus from persistent and nonpersistent intramammary infections with different clinical characteristics. J. Appl. Microbiol. 2007, 103, 993–1000. [Google Scholar] [CrossRef] [PubMed]
- EUCAST-CASFM. Available online: http://www.sfm-microbiologie.org (accessed on 12 August 2017).
- Monecke, S.; Coombs, G.; Shore, A.C.; Coleman, D.C.; Akpaka, P.; Borg, M.; Chow, H.; Ip, M.; Jatzwauk, L.; Jonas, D.; et al. A field guide to pandemic, epidemic and sporadic clones of methicillin-resistant Staphylococcus aureus. PLoS ONE 2011, 6, e17936. [Google Scholar] [CrossRef] [PubMed]
| Antibiotics | Origin of Isolates | ||||||
|---|---|---|---|---|---|---|---|
| Humans (n = 27) n (%) | Horses (n = 11) n (%) | Camels (n = 41) n (%) | Cattle (n = 8) n (%) | Sheep (n = 30) n (%) | Monkeys (n = 1) n (%) | Total Animals (N = 91) n (%) | |
| Penicillin G | 24 (88.8) | 11 (100) | 14 (34.1) | 2 (25.0) | 16 (53.3) | 1 (100) | 44 (48.4) |
| Cefoxitin | 0 (0) | 0 (0) | 3 (7.3) | 0 (0) | 6 (20.0) | 0 (0) | 9 (9.9) |
| Erythromycin | 6 (22.2) | 0 (0) | 0 (0) | 0 (0) | 2 (6.7) | 0 (0) | 2 (2.2) |
| Ceftobiprole | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Clindamycin | 4 (14.8) | 0 (0) | 0 (0) | 0 (0) | 2 (6.7) | 0 (0) | 2 (2.2) |
| Quinupristin/Dalfopristin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Kanamycin | 3 (11.1) | 6 (54.5) | 3 (7.3) | 0 (0) | 3 (10.0) | 0 (0) | 12 (13.2) |
| Tobramycin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Gentamicin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Minocycline | 3 (11.1) | 6 (54.5) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 6 (6.6) |
| Ofloxacin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Fucidic acid | 0 (0) | 0 (0) | 2 (4.9) | 0 (0) | 0 (0) | 0 (0) | 2 (2.2) |
| Linezolid | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Fosfomycin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Rifampicin | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Cotrimoxazole | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| CC | Clone Assignment | Leukocidin Genes Content | Origin of Isolates | p | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Human (n = 12) n (%) | Horses (n = 5) n (%) | Camels (n = 23) n (%) | Cattle (n = 6) n (%) | Sheep (n = 19) n (%) | Monkeys (n = 1) n (%) | Total Animals (N = 54) n (%) | H vs A | |||
| CC130 | ST130 | - | 0 (0) | 0 (0) | 6 (26.1) | 0 (0) | 0 (0) | 0 (0) | 6 (11.1) | NS |
| ST130-MSSA | LukF-P83/LukM+ | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 3 (15.8) | 0 (0) | 3 (5.6) | NS | |
| CC1 | ST1-MSSA | - | 0 (0) | 2 (40.0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 2 (3.7) | NS |
| ST1-MSSA | LukF-P83/LukM+ | 0 (0) | 0 (0) | 0 (0) | 1 (16.7) | 0 (0) | 0 (0) | 1 (1.9) | NS | |
| ST1278-MSSA | - | 1 (8.3) | 0 (0) | 4 (17.4) | 0 (0) | 0 (0) | 0 (0) | 4 (7.4) | NS | |
| CC8 | ST8-MSSA | - | 4 (33.3) | 2 (40.0) | 0 (0) | 0 (0) | 4 (21.1) | 0 (0) | 6 (11.1) | NS |
| ST72-MSSA | - | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 1 | 1 (1.9) | NS | |
| CC6 | ST6-MSSA | - | 0 (0) | 0 (0) | 3 (13.0) | 0 (0) | 4 (21.1) | 0 (0) | 7 (13.0) | NS |
| CC80 | ST80-MRSA IV | LukS-PV/LukF-PV+ | 0 (0) | 0 (0) | 2 (8.7) | 0 (0) | 4 (21.1) | 0 (0) | 6 (11.1) | NS |
| CC88 | ST88-MSSA | - | 0 (0) | 0 (0) | 6 (26.1) | 0 (0) | 0 (0) | 0 (0) | 6 (11.1) | NS |
| CC97 | ST97-MSSA | - | 0 (0) | 0 (0) | 0 (0) | 1 (16.7) | 2 (10.5) | 0 (0) | 3 (5.6) | NS |
| CC15 | ST15-MSSA | - | 3 (25.0) | 0 (0) | 0 (0) | 2 (33.3) | 0 (0) | 0 (0) | 2 (3.7) | 0.038 |
| CC133 | ST133-MSSA | - | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 2 (10.5) | 0 (0) | 2 (3.7) | NS |
| CC705 | ST705 | LukF-P83/LukM+ | 0 (0) | 0 (0) | 0 (0) | 2 (33.3) | 0 (0) | 0 (0) | 2 (3.7) | NS |
| CC152 | ST152-MSSA | LukS-PV/LukF-PV+ | 0 (0) | 0 (0) | 1 (4.3) | 0 (0) | 0 (0) | 0 (0) | 1 (1.9) | NS |
| CC30 | ST30-MSSA | - | 1 (8.3) | 0 (0) | 1 (4.3) | 0 (0) | 0 (0) | 0 (0) | 1 (1.9) | NS |
| CC45 | ST45-MSSA | - | 0 (0) | 1 (20.0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 1 (1.9) | NS |
| CC398 | ST398-MSSA | - | 2 (16.7) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0.030 |
| CC22 | ST22-MSSA | - | 1 (8.3) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| Virulence Genes | Origin of Isolates | p | ||||||
|---|---|---|---|---|---|---|---|---|
| Human (n = 12) n (%) | Horses (n = 5) n (%) | Camels (n = 23) n (%) | Cattle (n = 6) n (%) | Sheep (n = 19) n (%) | Monkeys (n = 1) n (%) | Total Animals (N = 54) n (%) | H vs. A | |
| Virulence genes | ||||||||
| Enterotoxins | ||||||||
| sea | 0 (0) | 0 (0) | 0 (0) | 1 (16.7) | 0 (0) | 0 (0) | 1 (1.9) | NS |
| seb | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| egc cluster * | 2 (16.6) | 1 (20) | 1 (4.3) | 2 (33.3) | 0 (0) | 1 (100) | 5 (9.2) | NS |
| seg | 2 (16.6) | 1 (20) | 1 (4.3) | 2 (33.3) | 0 (0) | 1 (100) | 5 (9.2) | NS |
| seh | 1 (8.3) | 2 (40) | 4 (17.4) | 1 (16.7) | 0 (0) | 0 (0) | 7 (12.9) | NS |
| sek | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| seq | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| Other toxins | ||||||||
| tst | 2 (16.6) | 0 (0) | 3 (13) | 2 (33.3) | 7 (36.8) | 0 (0) | 12 (22.2) | NS |
| etA | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| etB | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| etD | 2 (16.6) | 0 (0) | 2 (8.7) | 0 (0) | 4 (21.0) | 0 (0) | 6 (11.1) | NS |
| edinB | 2 (16.6) | 0 (0) | 9 (39.1) | 0 (0) | 7 (36.8) | 0 (0) | 16 (29.6) | NS |
| lukS-PV/lukF-PV | 0 (0) | 0 (0) | 3 (13) | 0 (0) | 4 (21) | 0 (0) | 7 (13) | NS |
| lukDE | 12 (100) | 4 (80) | 10 (43.4) | 5 (83.3) | 10 (52.6) | 1 (100) | 30 (55.5) | 0.002 |
| Hemolysins | ||||||||
| hla | 11 (91.6) | 3 (60) | 21 (91.3) | 6 (100) | 19 (100) | 1 (100) | 50 (92.5) | NS |
| hld | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| hlgA | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| hlg | 5 (41.6) | 1 (20) | 7 (30.4) | 2 (33.3) | 4 (21) | 1 (100) | 15 (27.7) | NS |
| hlgv | 10 (83.3) | 4 (80) | 20 (86.9) | 6 (100) | 19 (100) | 1 (100) | 50 (92.5) | NS |
| MSCRAMMs | ||||||||
| bbp | 12 (100) | 5 (100) | 21 (91.3) | 4 (66.6) | 17 (89.4) | 1 (100) | 48 (88.8) | NS |
| cna | 3 (25) | 3 (60) | 8 (34.7) | 1 (16.7) | 4 (21) | 0 (0) | 16 (29.6) | NS |
| ebpS | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| clfA | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| clfB | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| fib | 7 (58.3) | 3 (60) | 21 (91.3) | 6 (100) | 19 (100) | 1 (100) | 50 (92.5) | 0.007 |
| fnbA | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| fnbB | 10 (83.3) | 4 (80) | 20 (86.9) | 4 (66.6) | 12 (63.1) | 1 (100) | 41 (75.9) | NS |
| Capsule components | ||||||||
| cap5 | 8 (66.6) | 2 (40) | 5 (21.7) | 1 (16.7) | 6 (31.5) | 1 (100) | 15 (27.7) | 0.017 |
| cap8 | 4 (33.3) | 3 (60) | 18 (78.2) | 5 (83.3) | 13 (68.4) | 0 (0) | 39 (72.2) | 0.017 |
| icaA | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| icaC | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| icaD | 12 (100) | 5 (100) | 23 (100) | 6 (100) | 19 (100) | 1 (100) | 54 (100) | NS |
| Other virulence factors | ||||||||
| chp | 7 (58.3) | 1 (20) | 0 (0) | 2 (33.3) | 0 (0) | 0 (0) | 3 (5.5) | 0.0001 |
| scn | 8 (66.6) | 1 (20) | 7 (30.4) | 2 (33.3) | 3 (15.8) | 0 (0) | 13 (24) | 0.012 |
| Accessory gene regulators | ||||||||
| agr1 | 7 (58.33) | 3 (60) | 4 (17.4) | 1 (16.7) | 12 (63.1) | 1 (100) | 21 (38.8) | NS |
| agr2 | 3 (25) | 0 (0) | 0 (0) | 4 (66.6) | 0 (0) | 0 (0) | 4 (7.4) | NS |
| agr3 | 2 (16.6) | 2 (40) | 19 (82.6) | 1 (16.7) | 7 (36.8) | 0 (0) | 29 (53.7) | 0.026 |
| agr4 | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| Resistance genes | ||||||||
| mecA | 0 (0) | 0 (0) | 2 (8.7) | 0 (0) | 4 (21.0) | 0 (0) | 6 (11.1) | NS |
| mecC | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| blaZ | 11 (91.6) | 5 (100) | 5 (21.7) | 2 (33.3) | 5 (26.3) | 1 (100) | 18 (33.3) | 0.001 |
| ermA | 1 (8.3) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| ermC | 1 (8.3) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| aacA-aphD | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | NS |
| tetM | 2 (16.6) | 2 (40) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 2 (3.7) | NS |
| tetK | 3 (25) | 4 (80) | 0 (0) | 2 (33.3) | 5 (26.3) | 0 (0) | 11 (20.3) | NS |
| fosB | 8 (66.6) | 2 (40) | 1 (4.3) | 2 (33.3) | 6 (31.5) | 1 (100) | 12 (22.2) | 0.004 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Agabou, A.; Ouchenane, Z.; Ngba Essebe, C.; Khemissi, S.; Chehboub, M.T.E.; Chehboub, I.B.; Sotto, A.; Dunyach-Remy, C.; Lavigne, J.-P. Emergence of Nasal Carriage of ST80 and ST152 PVL+ Staphylococcus aureus Isolates from Livestock in Algeria. Toxins 2017, 9, 303. https://doi.org/10.3390/toxins9100303
Agabou A, Ouchenane Z, Ngba Essebe C, Khemissi S, Chehboub MTE, Chehboub IB, Sotto A, Dunyach-Remy C, Lavigne J-P. Emergence of Nasal Carriage of ST80 and ST152 PVL+ Staphylococcus aureus Isolates from Livestock in Algeria. Toxins. 2017; 9(10):303. https://doi.org/10.3390/toxins9100303
Chicago/Turabian StyleAgabou, Amir, Zouleikha Ouchenane, Christelle Ngba Essebe, Salim Khemissi, Mohamed Tedj Eddine Chehboub, Ilyes Bey Chehboub, Albert Sotto, Catherine Dunyach-Remy, and Jean-Philippe Lavigne. 2017. "Emergence of Nasal Carriage of ST80 and ST152 PVL+ Staphylococcus aureus Isolates from Livestock in Algeria" Toxins 9, no. 10: 303. https://doi.org/10.3390/toxins9100303







