UV-Sensitivity of Shiga Toxin-Converting Bacteriophage Virions Φ24B, 933W, P22, P27 and P32
Abstract
:1. Introduction
2. Results and Discussion
| Bacteriophage | Survival of Cells in Infected Culture (% of Survivors) | |
|---|---|---|
| No UV | 50 J/m2 UV | |
| λ papa | 31 ± 6 | 30 ± 6 |
| Φ24B | 36 ± 8 | 88 ± 7 * |
| 933W | 34 ± 1 | 39 ± 2 |
| P22 | 44 ± 7 | 68 ± 6 * |
| P27 | 43 ± 2 | 93 ± 1 * |
| P32 | 37 ± 7 | 63 ± 9 * |

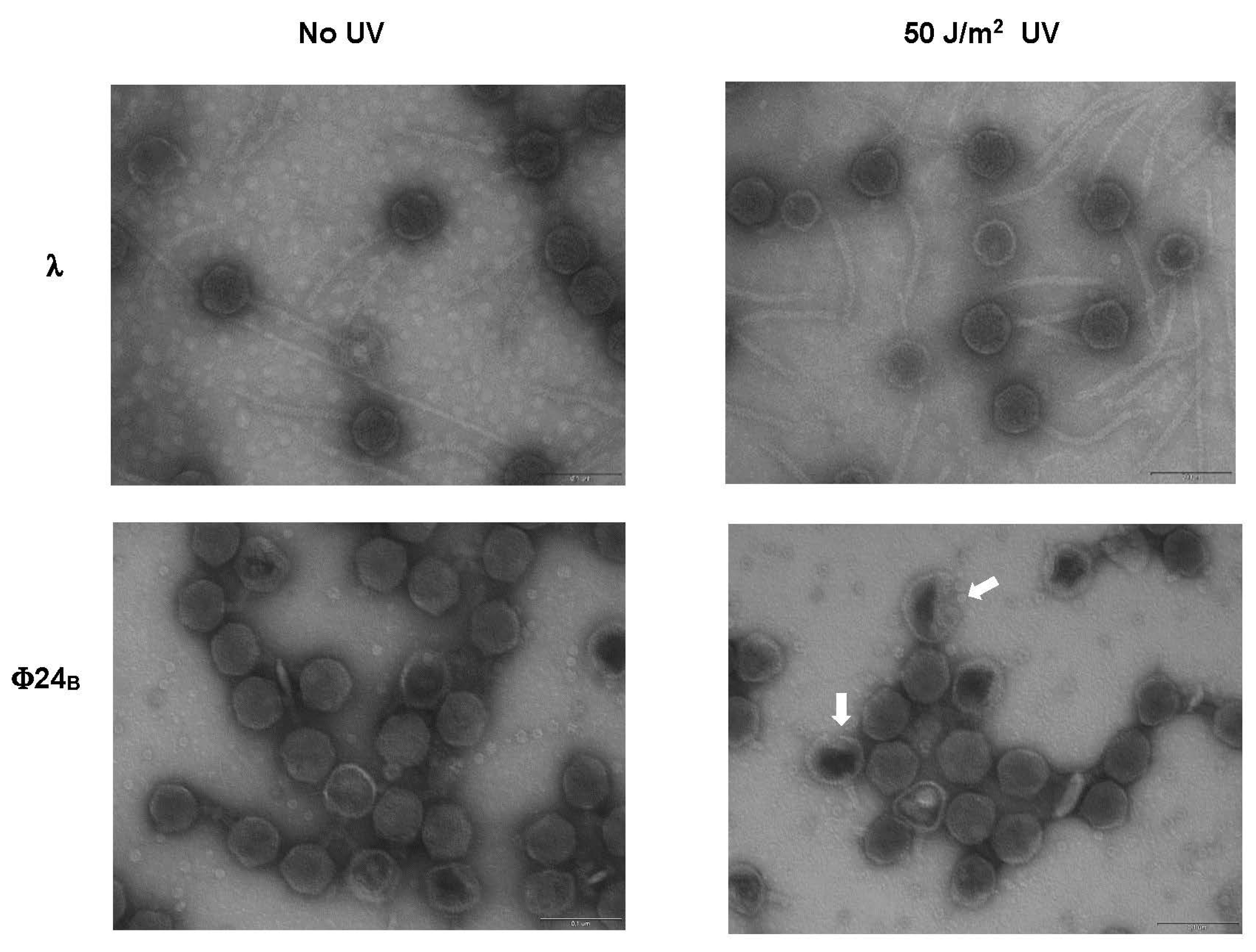
| Bacteriophage and Conditions | Fractions of Different Kinds of Virions (%) | ||
|---|---|---|---|
| Normal | Phage Ghosts | Untypical, with Partially Damaged Heads | |
| λ papa (non-irradiated) | 93 | 7 | 0 |
| λ papa (UV-irradiated) | 89 | 11 | 0 |
| Φ24B (non-irradiated) | 91 | 9 | 0 |
| Φ24B (UV-irradiated) | 71 | 23 | 6 |
3. Experimental Section
3.1. Bacteria and Bacteriophages
3.2. Evaluation of Bacterial Survival after Bacteriophage Infection
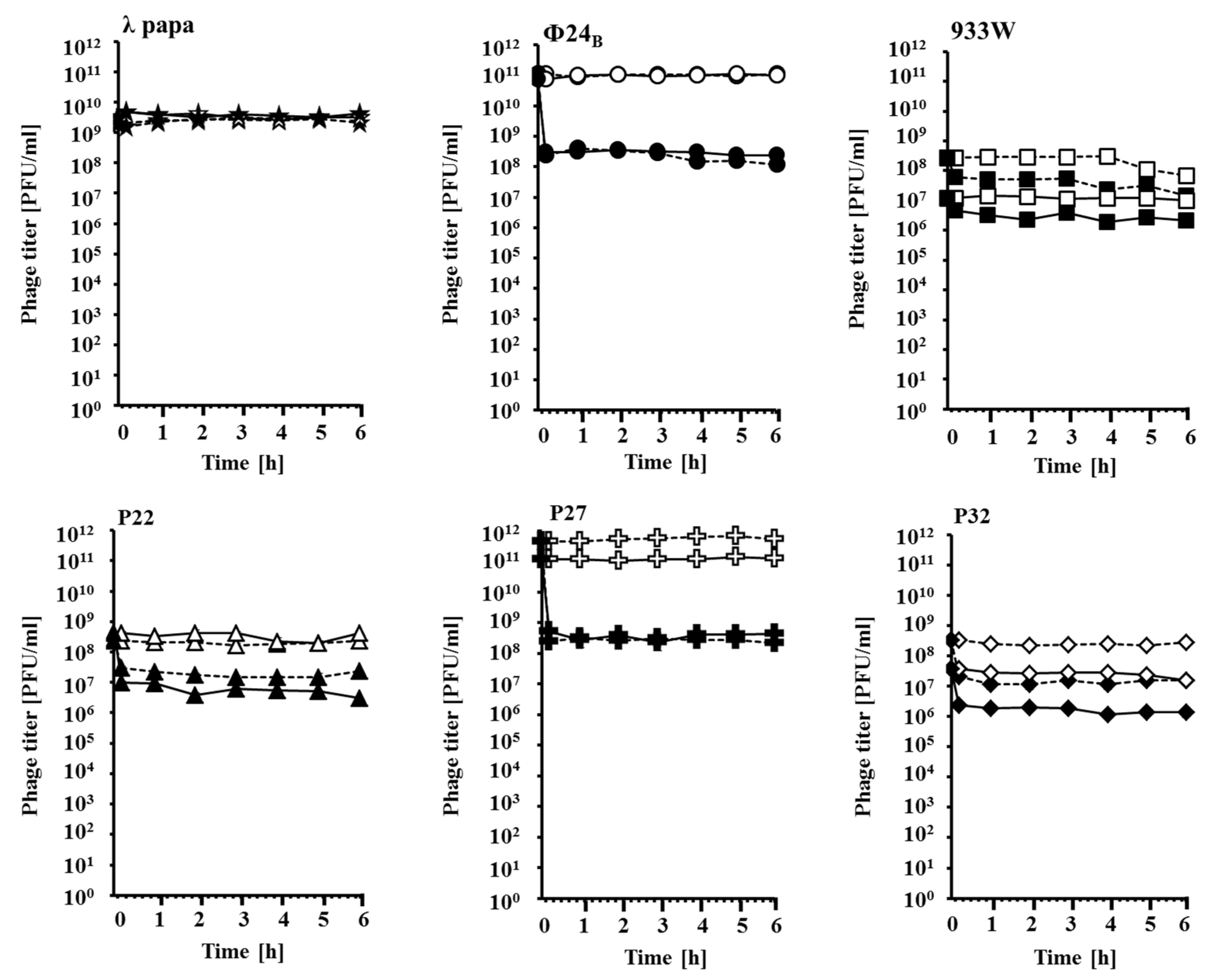
3.3. Estimation of Phage Lysate Stability
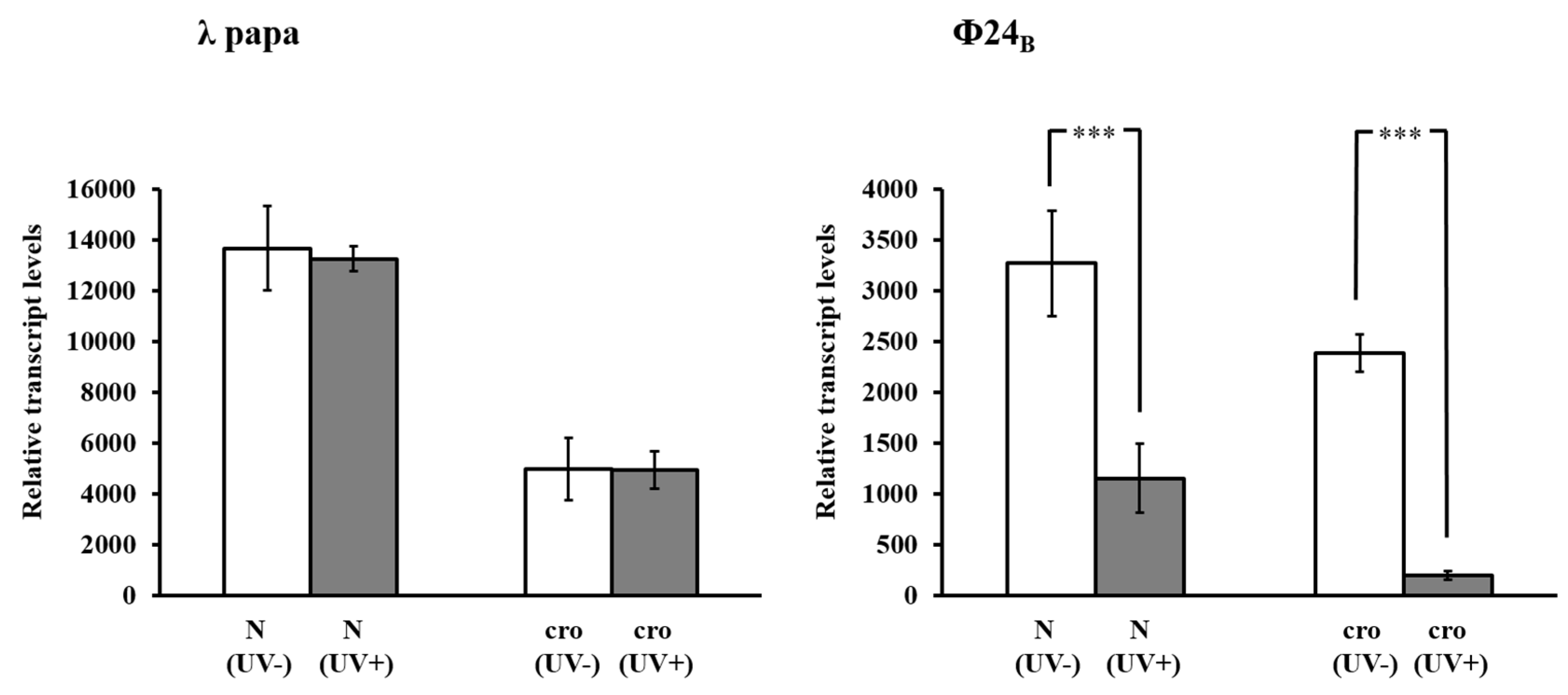
3.4. Measurement of Relative DNA Contents
3.5. Electron Microscopy
3.6. Bacteriophage Infection and cDNA Sample Preparation
3.7. Real-Time PCR Assay and Data Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gyles, C.L. Shiga toxin-producing Escherichia coli: An overview. J. Animal Sci. 2007, 6 (Suppl. 13), 45–62. [Google Scholar] [CrossRef] [PubMed]
- Hunt, J.M. Shiga toxin-producing Escherichia coli (STEC). Clin. Lab. Med. 2010, 30, 21–45. [Google Scholar] [CrossRef] [PubMed]
- Obrig, T.G.; Moran, T.P.; Brown, J.E. The mode of action of Shiga toxin on peptide elongation of eukaryotic protein synthesis. Biochem. J. 1987, 244, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Endo, Y.; Tsurugi, K.; Yutsudo, T.; Takeda, Y.; Ogasawara, T.; Igarashi, K. Site of action of a Vero toxin (VT2) from Escherichia coli O157:H7 and of Shiga toxin on eukaryotic ribosomes. RNA N-glycosidase activity of the toxins. Eur. J. Biochem. 1988, 171, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Razzaq, S. Hemolytic uremic syndrome: An emerging health risk. Am. Fam. Physician 2006, 74, 991–996. [Google Scholar] [PubMed]
- Bloch, S.K.; Felczykowska, A.; Nejman-Faleńczyk, B. Escherichia coli O104:H4 outbreak-have we learnt a lesson from it? Acta Biochim. Pol. 2012, 59, 483–488. [Google Scholar] [PubMed]
- Karch, H.; Denamur, E.; Dobrindt, U.; Finlay, B.B.; Hengge, R.; Johannes, L.; Ron, E.Z.; Tønjum, T.; Sansonetti, P.J.; Vicente, M. The enemy within us: Lessons from the 2011 European Escherichia coli O104:H4 outbreak. EMBO Mol. Med. 2012, 4, 841–848. [Google Scholar] [CrossRef] [PubMed]
- Werber, D.; Krause, G.; Frank, C.; Fruth, A.; Flieger, A.; Mielke, M.; Schaade, L.; Stark, K. Outbreaks of virulent diarrheagenic Escherichia coli—Are we in control? BMC Med. 2012, 10. [Google Scholar] [CrossRef] [PubMed]
- Allison, H.E. Stx-phages: Drivers and mediators of the evolution of STEC and STEC-like pathogens. Future Microbiol. 2007, 2, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Łoś, J.M.; Łoś, M.; Węgrzyn, G. Bacteriophages carrying Shiga toxin genes: Genomic variations, detection and potential treatment of pathogenic bacteria. Future Microbiol. 2011, 6, 909–924. [Google Scholar] [CrossRef] [PubMed]
- Ptashne, M. A Genetic Switch: Phage Lambda Revisited, 3rd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2004; pp. 1–154. [Google Scholar]
- Węgrzyn, G.; Węgrzyn, A. Genetic switches during bacteriophage lambda development. Prog. Nucleic Acid Res. Mol. Biol. 2005, 79, 1–48. [Google Scholar] [PubMed]
- Węgrzyn, G.; Licznerska, K.; Węgrzyn, A. Phage λ-new insights into regulatory circuits. Adv. Virus Res. 2012, 82, 155–178. [Google Scholar] [PubMed]
- Łoś, J.M.; Łoś, M.; Węgrzyn, A.; Węgrzyn, G. Altruism of Shiga toxin-producing Escherichia coli: Recent hypothesis versus experimental results. Front. Cell. Infect. Microbiol. 2013, 2. [Google Scholar] [CrossRef] [PubMed]
- Riley, L.M.; Veses-Garcia, M.; Hillman, J.D.; Handfield, M.; McCarthy, A.J.; Allison, H.E. Identification of genes expressed in cultures of E. coli lysogens carrying the Shiga toxin-encoding prophage Φ24B. BMC Microbiol. 2012, 12. [Google Scholar] [CrossRef] [PubMed]
- Imamovic, L.; Muniesa, M. Characterizing RecA-independent induction of Shiga toxin2-encoding phages by EDTA treatment. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.L.; James, C.E.; Sergeant, M.J.; Yaxian, Y.; Saunders, J.R.; McCarthy, A.J.; Allison, H.E. Short-tailed Stx phages exploit the conserved YaeT protein to disseminate Shiga toxin genes among enterobacteria. J. Bacteriol. 2007, 189, 7223–7233. [Google Scholar] [CrossRef] [PubMed]
- Yue, W.F.; Du, M.; Zhu, M.J. High temperature in combination with UV irradiation enhances horizontal transfer of stx2 gene from E. coli O157:H7 to non-pathogenic E. coli. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Muniesa, M.; Jofre, J. Abundance in sewage of bacteriophages that infect Escherichia coli O157:H7 and that carry the Shiga toxin 2 gene. Appl. Environ. Microbiol. 1998, 64, 2443–2448. [Google Scholar] [PubMed]
- Muniesa, M.; Lucena, F.; Jofre, J. Comparative survival of free Shiga toxin 2-encoding phages and Escherichia coli strains outside the gut. Appl. Environ. Microbiol. 1999, 65, 5615–5618. [Google Scholar] [PubMed]
- Johannessen, G.S.; James, C.E.; Allison, H.E.; Smith, D.L.; Saunders, J.R.; McCarthy, A.J. Survival of a Shiga toxin-encoding bacteriophage in a compost model. FEMS Microbiol. Lett. 2005, 245, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Dumke, R.; Schröter-Bobsin, U.; Jacobs, E.; Röske, I. Detection of phages carrying the Shiga toxin 1 and 2 genes in waste water and river water samples. Lett. Appl. Microbiol. 2006, 42, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Imamovic, L.; Ballesté, E.; Jofre, J.; Muniesa, M. Quantification of Shiga toxin-converting bacteriophages in wastewater and in fecal samples by real-time quantitative PCR. Appl. Environ. Microbiol. 2010, 76, 5693–5701. [Google Scholar] [CrossRef] [PubMed]
- Rooks, D.J.; Yan, Y.; McDonald, J.E.; Woodward, M.J.; McCarthy, A.J.; Allison, H.E. Development and validation of a qPCR-based method for quantifying Shiga toxin-encoding and other lambdoid bacteriophages. Environ. Microbiol. 2010, 12, 1194–1204. [Google Scholar] [CrossRef] [PubMed]
- Imamovic, L.; Muniesa, M. Quantification and evaluation of infectivity of Shiga toxin-encoding bacteriophages in beef and salad. Appl. Environ. Microbiol. 2011, 77, 3536–3540. [Google Scholar] [CrossRef] [PubMed]
- Imamovic, L.; Serra-Moreno, R.; Jofre, J.; Muniesa, M. Quantification of Shiga toxin 2-encoding bacteriophages, by real-time PCR and correlation with phage infectivity. J. Appl. Microbiol. 2010, 108, 1105–1114. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, H.W. Bacteriophage electron microscopy. Adv. Virus Res. 2012, 82, 1–32. [Google Scholar] [PubMed]
- Ackermann, H.W.; Prangishvili, D. Prokaryote viruses studied by electron microscopy. Arch. Virol. 2012, 157, 1843–1849. [Google Scholar] [CrossRef] [PubMed]
- Duckworth, D.H. Biological activity of bacteriophage ghosts and “take-over” of host functions by bacteriophage. Bacteriol. Rev. 1970, 34, 344–363. [Google Scholar] [PubMed]
- Allué-Guardia, A.; Jofre, J.; Muniesa, M. Stability and infectivity of cytolethal distending toxin type V gene-carrying bacteriophages in a water mesocosm and under different inactivation conditions. Appl. Environ. Microbiol. 2012, 78, 5818–5823. [Google Scholar] [CrossRef] [PubMed]
- Clark, E.M.; Wright, H.; Lennon, K.A.; Craik, V.A.; Clark, J.R.; March, J.B. Inactivation of recombinant bacteriophages lambda by use of chemical agents and UV radiation. Appl. Environ. Microbiol. 2012, 78, 3033–3036. [Google Scholar] [CrossRef] [PubMed]
- Theitler, D.J.; Nasser, A.; Gerchman, Y.; Kribus, A.; Mamane, H. Synergistic effect of heat and solar UV on DNA damage and water disinfection of E. coli and bacteriophages MS2. J. Water Health 2012, 10, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Guglielmotti, D.M.; Mercanti, D.J.; Reinheimer, J.A.; Quiberoni Adel, L. Review: Efficiency of physical and chemical treatments on the inactivation of dairy bacteriophages. Front. Microbiol. 2012, 2. [Google Scholar] [CrossRef] [PubMed]
- Allué-Guardia, A.; Martínez-Castillo, A.; Muniesa, M. Persistence of infectious Shiga toxin-encoding bacteriophages after disinfection treatments. Appl. Environ. Microbiol. 2014, 80, 2142–2149. [Google Scholar] [CrossRef] [PubMed]
- Diston, D.; Ebdon, J.E.; Taylor, H.D. Inactivation of bacteriophages infecting Bacteroides strain GB124 using UV-B radiation. Photochem. Photobiol. 2014, 90, 622–627. [Google Scholar] [CrossRef] [PubMed]
- Wigginton, K.R. Virus inactivation mechanisms: Impact of disinfectants on virus function and structural integrity. Environ. Sci. Technol. 2012, 46, 12069–12078. [Google Scholar] [CrossRef] [PubMed]
- Jensen, K.F. The Escherichia coli K-12 “wild types” W3110 and MG1655 have an rph frameshift mutation that leads to pyrimidine starvation due to low pyrE expression levels. J. Bacteriol. 1993, 175, 3401–3407. [Google Scholar] [PubMed]
- Allison, H.E.; Sergeant, M.J.; James, C.E.; Saunders, J.R.; Smith, D.L.; Sharp, R.J.; Marks, T.S.; McCarthy, A.J. Immunity profiles of wild-type and recombinant Shiga-like toxin-encoding bacteriophages and characterization of novel double lysogens. Infect. Immun. 2003, 71, 3409–3418. [Google Scholar] [CrossRef] [PubMed]
- Gamage, S.D.; Patton, A.K.; Hanson, J.F.; Weiss, A.A. Diversity and host range of Shiga toxin-encoding phage. Infect. Immun. 2004, 72, 7131–7139. [Google Scholar] [CrossRef] [PubMed]
- Nejman-Faleńczyk, B.; Bloch, S.; Licznerska, K.; Dydecka, A.; Felczykowska, A.; Topka, G.; Węgrzyn, A.; Węgrzyn, G. A small, microRNA-size, ribonucleic acid regulating gene expression and development of Shiga toxin-converting bacteriophage Φ24Β. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Nowicki, D.; Kobiela, W.; Węgrzyn, A.; Wegrzyn, G.; Szalewska-Pałasz, A. ppGpp-dependent negative control of DNA replication of Shiga toxin-converting bacteriophages in Escherichia coli. J. Bacteriol. 2013, 195, 5007–5015. [Google Scholar] [CrossRef] [PubMed]
- Bloch, S.; Nejman-Faleńczyk, B.; Łoś, J.M.; Barańska, S.; Łepek, K.; Felczykowska, A.; Łoś, M.; Węgrzyn, G.; Węgrzyn, A. Genes from the exo-xis region of λ and Shiga toxin-converting bacteriophages influence lysogenization and prophage induction. Arch. Microbiol. 2013, 195, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Czajkowski, R.; Ozymko, Z.; de Jager, V.; Siwinska, J.; Smolarska, A.; Ossowicki, A.; Narajczyk, M.; Łojkowska, E. Genomic, proteomic and morphological characterization of two novel broad host lytic bacteriophages ΦPD10.3 and ΦPD23.1 infecting pectinolytic Pectobacterium. spp. and Dickeya. spp. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Bloch, S.; Nejman-Faleńczyk, B.; Dydecka, A.; Łoś, J.M.; Felczykowska, A.; Węgrzyn, A.; Węgrzyn, G. Different expression patterns of genes from the exo-xis region of bacteriophage λ and Shiga toxin-converting bacteriophage Ф24B following infection or prophage induction in Escherichia coli. PLoS ONE 2014, 13. [Google Scholar] [CrossRef] [PubMed]
- Nowicki, D.; Bloch, S.; Nejman-Faleńczyk, B.; Szalewska-Palasz, A.; Węgrzyn, A.; Węgrzyn, G. Defects in RNA polyadenylation impair both lysogenization by and lytic development of Shiga toxin-converting bacteriophages. J. Gen. Virol. 2015, 96, 1957–1968. [Google Scholar] [CrossRef] [PubMed]
- Strauch, E.; Hammerl, J.A.; Konietzny, A.; Schneiker-Bekel, S.; Arnold, W.; Goesmann, A.; Pühler, A.; Beutin, L. Bacteriophage 2851 is a prototype phage for dissemination of the Shiga toxin variant gene 2c in Escherichia coli O157:H7. Infect. Immun. 2008, 76, 5466–5477. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bloch, S.; Nejman-Faleńczyk, B.; Topka, G.; Dydecka, A.; Licznerska, K.; Narajczyk, M.; Necel, A.; Węgrzyn, A.; Węgrzyn, G. UV-Sensitivity of Shiga Toxin-Converting Bacteriophage Virions Φ24B, 933W, P22, P27 and P32. Toxins 2015, 7, 3727-3739. https://doi.org/10.3390/toxins7093727
Bloch S, Nejman-Faleńczyk B, Topka G, Dydecka A, Licznerska K, Narajczyk M, Necel A, Węgrzyn A, Węgrzyn G. UV-Sensitivity of Shiga Toxin-Converting Bacteriophage Virions Φ24B, 933W, P22, P27 and P32. Toxins. 2015; 7(9):3727-3739. https://doi.org/10.3390/toxins7093727
Chicago/Turabian StyleBloch, Sylwia, Bożena Nejman-Faleńczyk, Gracja Topka, Aleksandra Dydecka, Katarzyna Licznerska, Magdalena Narajczyk, Agnieszka Necel, Alicja Węgrzyn, and Grzegorz Węgrzyn. 2015. "UV-Sensitivity of Shiga Toxin-Converting Bacteriophage Virions Φ24B, 933W, P22, P27 and P32" Toxins 7, no. 9: 3727-3739. https://doi.org/10.3390/toxins7093727







