Genetic Relationships, Carbendazim Sensitivity and Mycotoxin Production of the Fusarium Graminearum Populations from Maize, Wheat and Rice in Eastern China
Abstract
:1. Introduction
2. Results and Discussion
2.1. Species Composition
| Host | County | Numbers | F. asiaticum | F. graminearum | ||
|---|---|---|---|---|---|---|
| 3ADON | 15ADON | NIV | 15ADON | |||
| Rice | Rudong | 6 | 4 | 2 | ||
| Jiangdu | 5 | 3 | 1 | 1 | ||
| Dongtai | 5 | 2 | 2 | 1 | ||
| Xinghua | 5 | 4 | 1 | |||
| Baoying | 4 | 2 | 2 | |||
| Sheyang | 5 | 3 | 2 | |||
| Subtotal | 30 | 18 | 5 | 7 | ||
| Maize | Qidong | 4 | 1 | 1 | 2 | |
| Huoqiu | 3 | 1 | 1 | 1 | ||
| Dafeng | 4 | 1 | 2 | 1 | ||
| Suixi | 3 | 1 | 2 | |||
| Lingbi | 4 | 2 | 2 | |||
| Siyang | 5 | 5 | ||||
| Sheyang | 3 | 3 | ||||
| Xinyi | 3 | 3 | ||||
| Peixian | 4 | 4 | ||||
| Subtotal | 33 | 4 | 4 | 3 | 23 | |
| Wheat | Huoqiu | 4 | 2 | 1 | 1 | |
| Rudong | 5 | 3 | 2 | |||
| Lixin | 4 | 2 | 2 | |||
| Huaiyuan | 4 | 3 | 1 | |||
| Xinghua | 4 | 4 | ||||
| Sixian | 5 | 4 | 1 | |||
| Shuyang | 4 | 3 | 1 | |||
| Tongshan | 4 | 4 | ||||
| Subtotal | 34 | 25 | 8 | 1 | ||
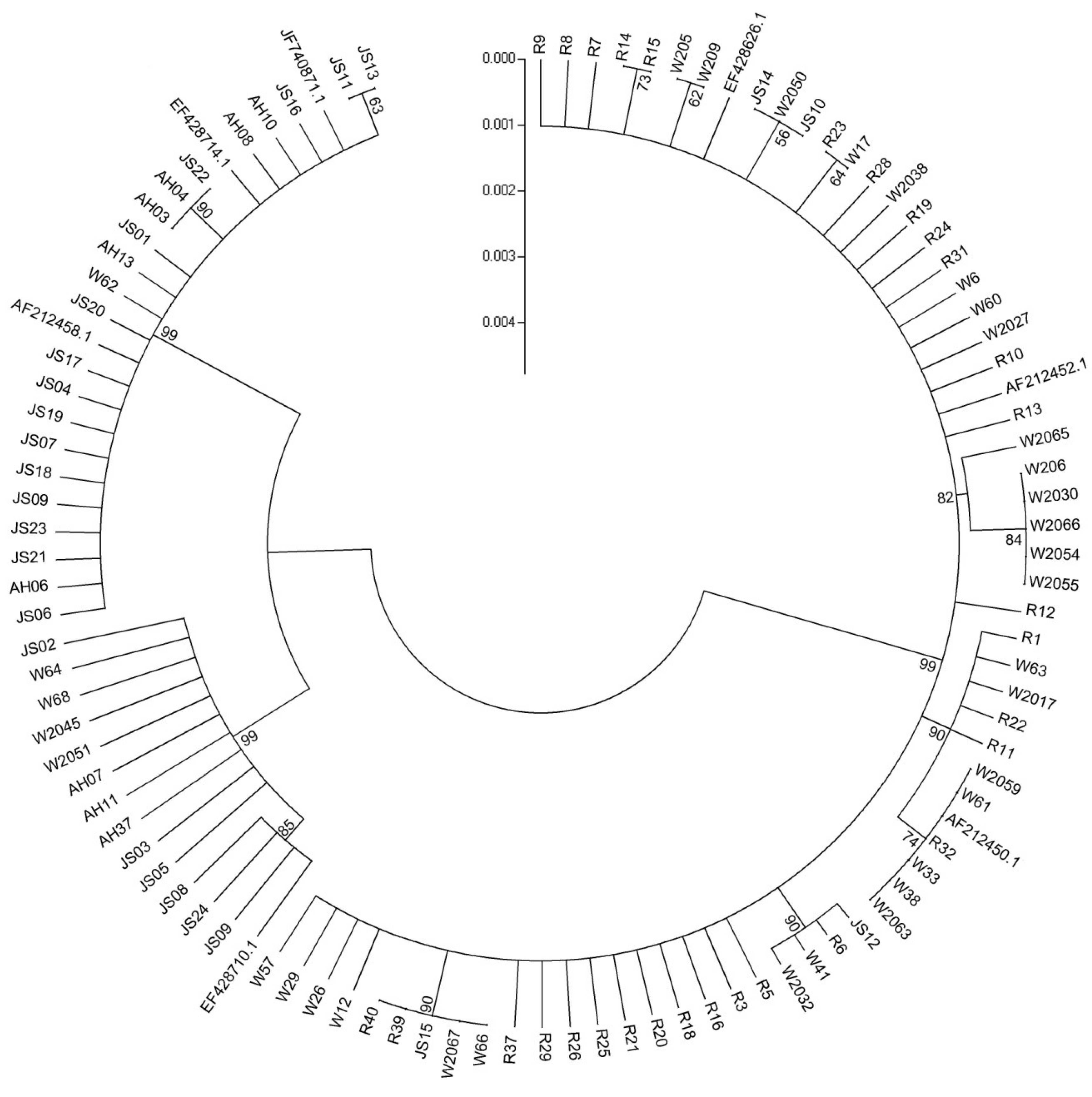
2.2. Trichothecene Chemotypes Determination
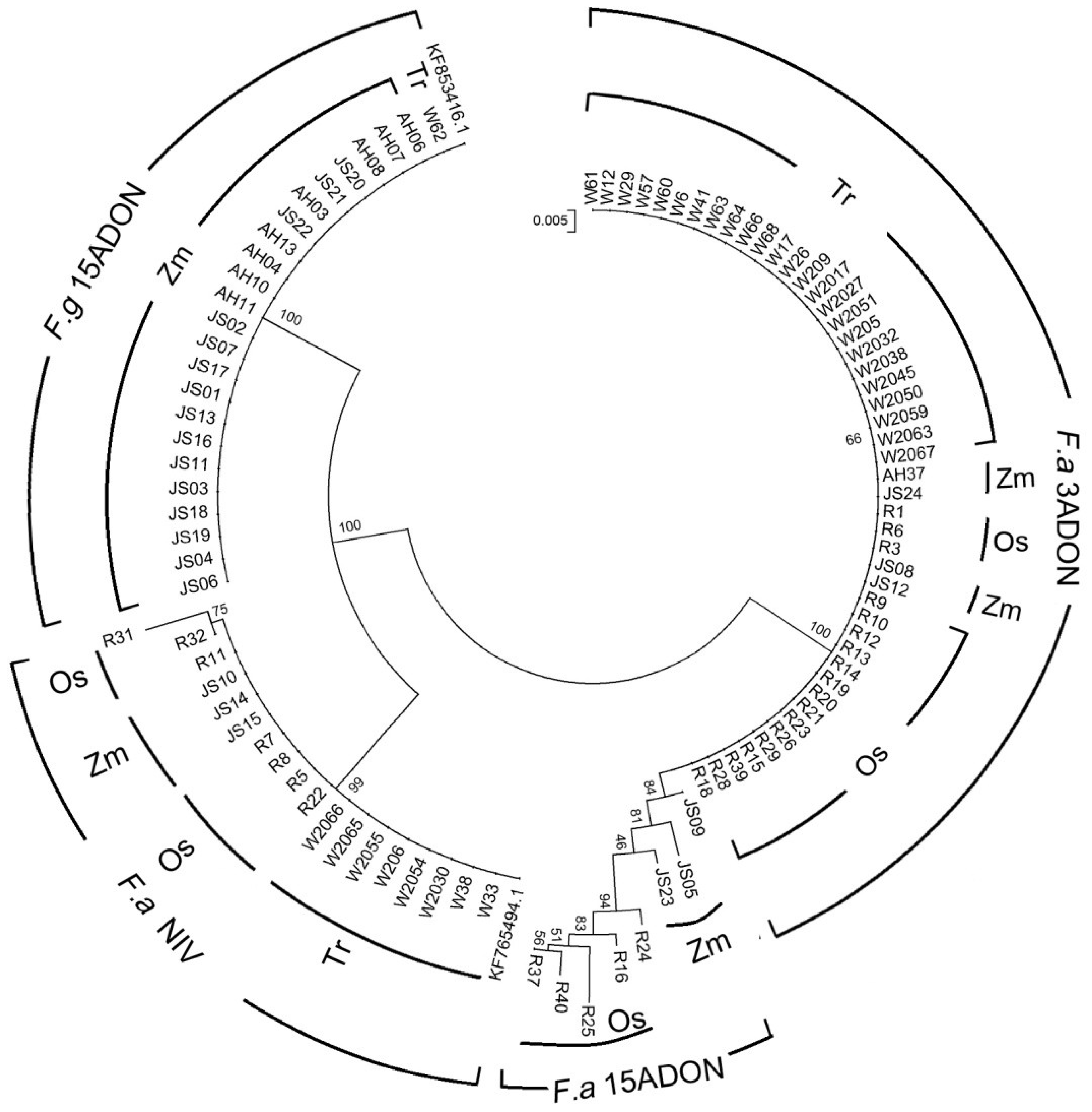
2.3. Phylogenetic Analysis
2.4. Sensitivity of the Fusarium Isolates to Carbendazim

2.5. Chemical Analysis of DON and ZEN
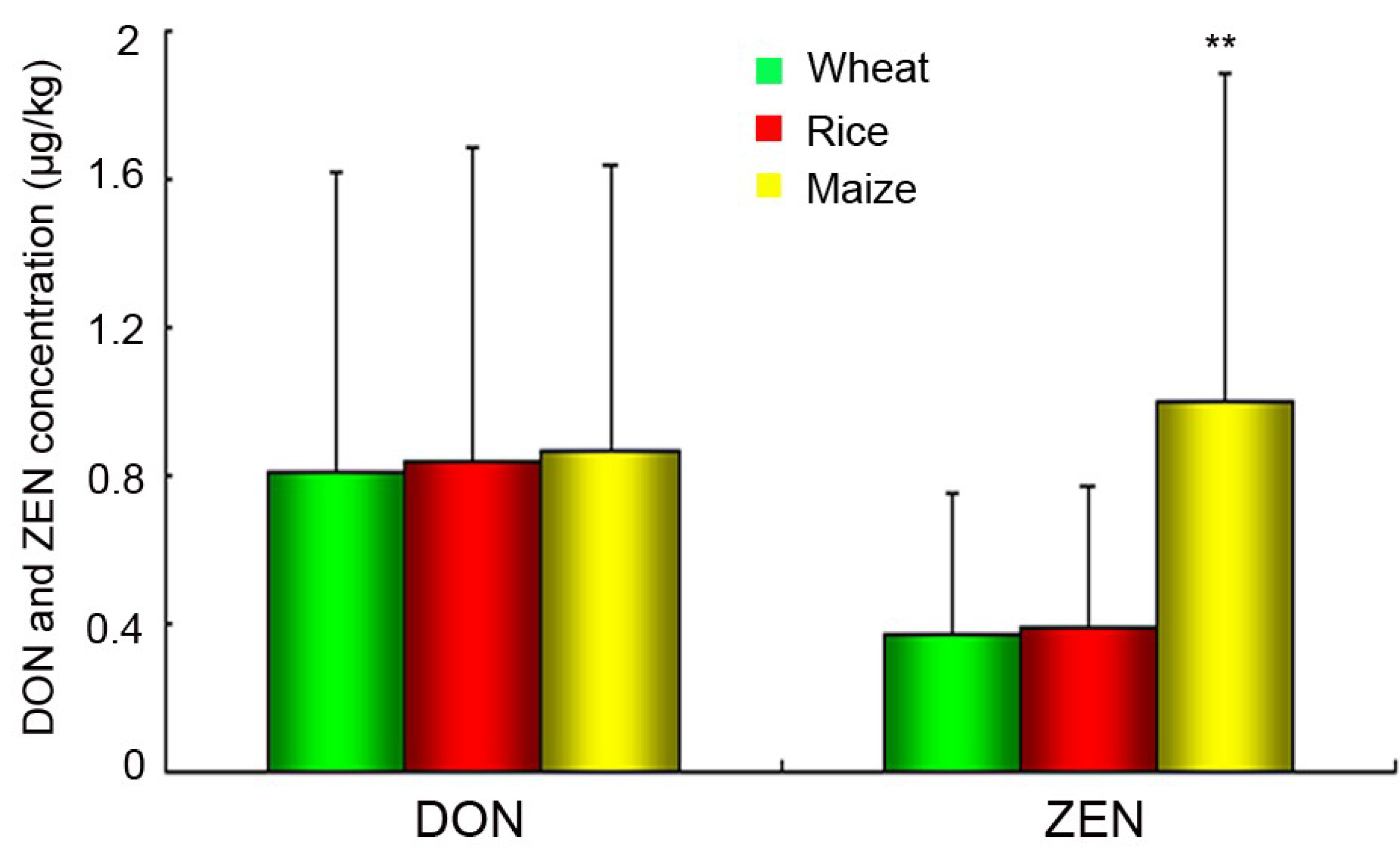
2.6. Discussion
3. Experimental Section
3.1. Fungal Isolates
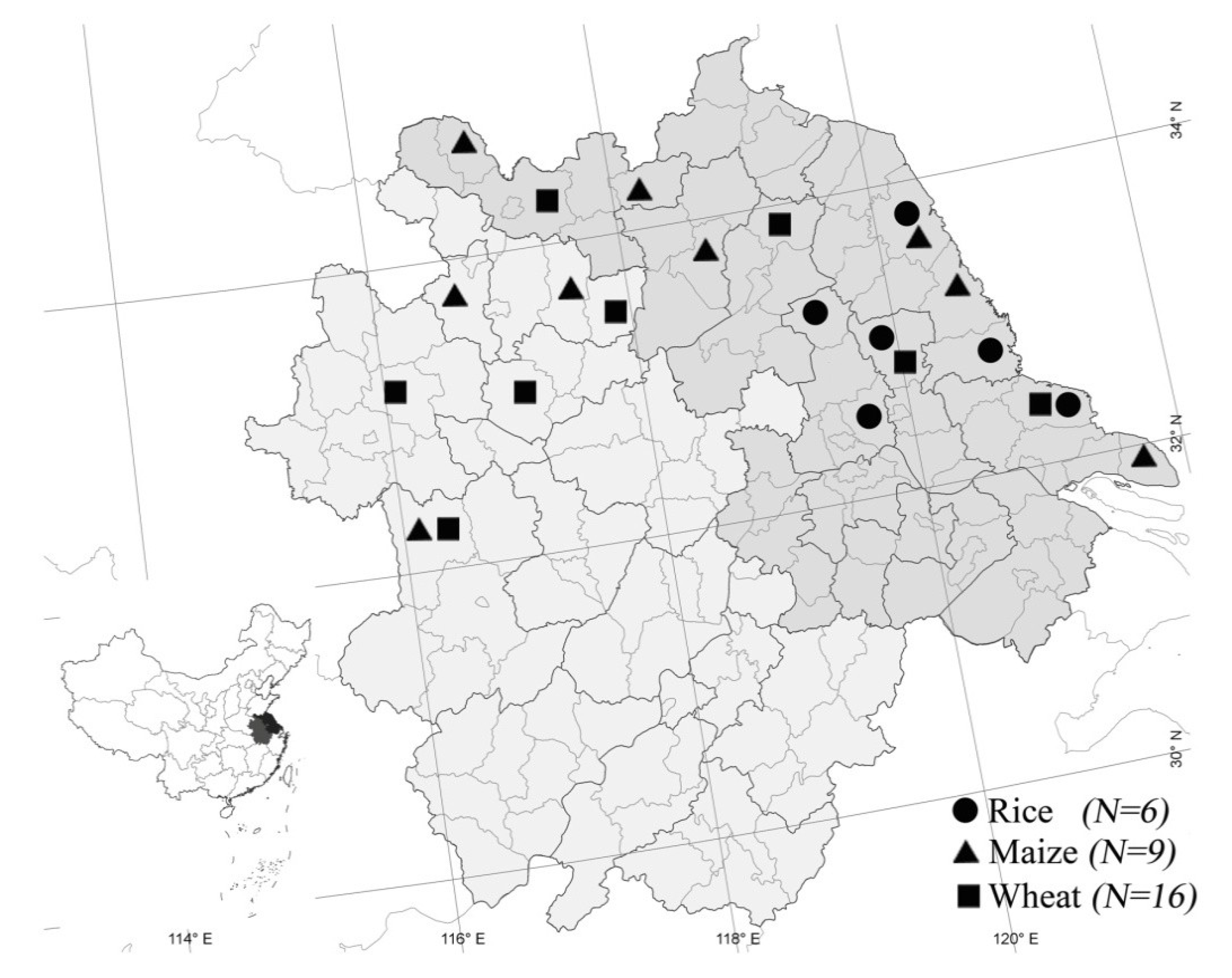
3.2. Identification of F. graminearum and Trichothecene Genotype
3.3. DNA Sequencing
3.4. Sequence Analysis and Phylogeny Reconstruction
3.5. In Vitro Carbendazim Sensitivity
3.6. Production of ZEN and DON
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Parry, D.W.; Jenkinson, P.; McLeod, L. Fusarium ear blight (scab) in small grain cereals—A review. Plant pathol. 1995, 44, 207–238. [Google Scholar] [CrossRef]
- Desjardins, A.E.; Proctor, R.H. Genetic diversity and trichothecene chemotypes of the Fusarium graminearum clade isolated from maize in Nepal and identification of a putative new lineage. Fungal Biol. 2011, 115, 38–48. [Google Scholar] [CrossRef]
- McMullen, M.; Jones, R.; Gallenberg, D. Scab of wheat and barley: A re-emerging disease of devastating impact. Plant. Dis. 1997, 81, 1340–1348. [Google Scholar] [CrossRef]
- Miller, J.D.; Greenhalgh, R.; Wang, Y.; Lu, M. Trichothecene chemotypes of three Fusarium species. Mycologia 1991, 83, 121–130. [Google Scholar] [CrossRef]
- Peraica, M.; Radic, B.; Lucic, A.; Pavlovic, M. Toxic effects of mycotoxins in humans. Bull. World Health Organ. 1999, 77, 754–766. [Google Scholar]
- Bennett, J.W.; Klich, M. Mycotoxins. Clin. Microbiol. Rev. 2003, 16, 497–516. [Google Scholar]
- Pestka, J.J. Deoxynivalenol: Mechanisms of action, human exposure, and toxicological relevance. Arch. Toxicol. 2010, 84, 663–679. [Google Scholar] [CrossRef]
- Jansen, C.; von Wettstein, D.; Schafer, W.; Kogel, K.H.; Felk, A.; Maier, F.J. Infection patterns in barley and wheat spikes inoculated with wild-type and trichodiene synthase gene disrupted Fusarium graminearum. Proc. Natl. Acad. Sci. USA 2005, 102, 16892–16897. [Google Scholar] [CrossRef]
- Proctor, R.H.; Hohn, T.M.; McCormick, S.P. Reduced virulence of Gibberella. zeae caused by disruption of a trichothecene toxin biosynthetic gene. Mol. Plant Microbe Inter. 1995, 8, 593–601. [Google Scholar] [CrossRef]
- Kuiper-Goodman, T.; Scott, P.M.; Watanabe, H. Risk assessment of the mycotoxin zearalenone. Regul. Toxicol. Pharm. 1987, 7, 253–306. [Google Scholar] [CrossRef]
- Desjardins, A.E.; Proctor, R.H. Biochemistry and Genetics of Fusarium Toxins. In Fusarium Paul E. Nelson Memorial Symposium; American Phytopathological Society Press: St. Paul, MN, USA, 2001. [Google Scholar]
- O’Donnell, K.; Kistler, H.C.; Tacke, B.K.; Casper, H.H. Gene genealogies reveal global phylogeographic structure and reproductive isolation among lineages of Fusarium graminearum, the fungus causing wheat scab. Proc. Natl. Acad. Sci. USA 2000, 97, 7905–7910. [Google Scholar]
- O’Donnell, K.; Ward, T.J.; Geiser, D.M.; Kistler, H.C.; Aoki, T. Genealogical concordance between the mating type locus and seven other nuclear genes supports formal recognition of nine phylogenetically distinct species within the Fusarium graminearum clade. Fungal Genet. Biol. 2004, 41, 600–623. [Google Scholar] [CrossRef]
- O’Donnell, K.; Ward, T.J.; Aberra, D.; Kistler, H.C.; Aoki, T.; Orwig, N.; Kimura, M.; Bjørnstad, Å.; Klemsdal, S.S. Multilocus genotyping and molecular phylogenetics resolve a novel head blight pathogen within the Fusarium graminearum species complex from Ethiopia. Fungal Genet. Biol. 2008, 45, 1514–1522. [Google Scholar] [CrossRef]
- Sarver, B.A.J.; Ward, T.J.; Gale, L.R.; Karen, B.; Kistler, H.C.; Takayuki, A.; Nicholson, J.; Carter, J.; O’Donnell, K. Novel Fusarium head blight pathogens from Nepal and Louisiana revealed by multilocus genealogical concordance. Fungal Genet. Biol. 2011, 48, 1096–1107. [Google Scholar] [CrossRef]
- Starkey, D.E.; Ward, T.J.; Aoki, T.; Gale, L.H.; Kistler, H.C.; Geiser, D.M.; Suga, H.; Varga, J.; Tóth, B.; O’Donnell, K. Global molecular surveillance reveals novel Fusarium head blight species and trichothecene toxin diversity. Fungal Genet. Biol. 2007, 44, 1191–1204. [Google Scholar] [CrossRef]
- Xu, X.M.; Nicholson, P. Community ecology of fungal pathogens causing wheat head blight. Annu. Rev. Phytopathol. 2009, 47, 83–103. [Google Scholar] [CrossRef]
- Gale, L.R.; Chen, L.F.; Hernick, C.A.; Takamura, K.; Kistler, H.C. Population analysis of Fusarium graminearum from wheat fields in eastern China. Phytopathology 2002, 92, 1315–1322. [Google Scholar] [CrossRef]
- Suga, H.; Karugia, G.W.; Ward, T.; Gale, L.R.; Tomimura, K.; Nakajima, T.; Miyasaka, A.; Koizumi, S.; Kageyama, K.; Hyakumachi, M. Molecular characterization of the Fusarium graminearum species complex in Japan. Phytopathology 2008, 98, 159–166. [Google Scholar] [CrossRef]
- Lee, J.; Chang, I.Y.; Kim, H.; Yun, S.H.; Leslie, J.F.; Lee, Y.W. Genetic diversity and fitness of Fusarium graminearum populations from rice in Korea. Appl. Environ. Microb. 2009, 75, 3289–3295. [Google Scholar] [CrossRef]
- Lee, J.; Kim, H.; Jeon, J.J.; Kim, H.S.; Zeller, K.A.; Carter, L.L.; Leslie, J.F.; Lee, Y.W. Population structure of and mycotoxin production by Fusarium graminearum from maize in South Korea. Appl. Environ. Microb. 2012, 78, 2161–2167. [Google Scholar] [CrossRef]
- Qu, B.; Li, H.P.; Zhang, J.B.; Xu, Y.B.; Huang, T.; Wu, A.B.; Zhao, C.S.; Carter, J.; Nicholson, P.; Liao, Y.C. Geographic distribution and genetic diversity of Fusarium graminearum and F. asiaticum on wheat spikes throughout China. Plant Pathol. 2008, 57, 15–24. [Google Scholar]
- Desjardins, A.E. Fusarium. Mycotoxins: Chemistry, Genetics, and Biology; APS Press: St. Paul, MN, USA, 2006. [Google Scholar]
- Ryu, J.C.; Ohtsubo, K.; Izumiyama, N.; Nakamura, K.; Tanaka, T.; Yamamura, H.; Ueno, Y. The acute and chronic toxicities of nivalenol in mice. Fund. Appl. Toxicol. 1988, 11, 38–47. [Google Scholar] [CrossRef]
- Ward, T.J.; Bielawski, J.P.; Kistler, H.C.; Sullivan, E.; O’Donnell, K. Ancestral polymorphism and adaptive evolution in the trichothecene mycotoxin gene cluster of phytopathogenic Fusarium. Proc. Natl. Acad. Sci. USA 2002, 99, 9278–9283. [Google Scholar] [CrossRef]
- Desjardins, A.E.; Manandhar, H.K.; Plattner, R.D.; Manandhar, G.G.; Poling, S.M.; Maragos, C.M. Fusarium species from Nepalese rice and production of mycotoxins and gibberellic acid by selected species. Appl. Environ. Microb. 2000, 66, 1020–1025. [Google Scholar] [CrossRef]
- Lee, T.; Oh, D.W.; Kim, H.S.; Lee, J.; Kim, Y.H.; Yun, S.H.; Lee, Y.W. Identification of deoxynivalenol and nivalenol producing chemotypes of Gibberella. zeae by using PCR. Appl. Environ. Microb. 2001, 67, 2966–2972. [Google Scholar] [CrossRef]
- Jennings, P.; Coates, M.E.; Turner, J.A.; Chandler, E.A.; Nicholson, P. Determination of deoxynivalenol and nivalenol chemotypes of Fusarium culmorum isolates from England and Wales by PCR assay. Plant Pathol. 2004, 53, 182–190. [Google Scholar] [CrossRef]
- Gale, L.R.; Harrison, S.A.; Ward, T.J.; O’Donnell, K.; Milus, E.A.; Gale, S.W.; Kistler, H.C. Nivalenol-Type populations of Fusarium graminearum and F. asiaticum are prevalent on wheat in Southern Louisiana. Phytopathology 2011, 101, 124–134. [Google Scholar]
- Kristensen, R.; Torp, M.; Kosiak, B.; Holst-Jensen, A. Phylogeny and toxigenic potential is correlated in Fusarium species as revealed by partial translation elongation factor 1 α gene sequences. Mycol. Res. 2005, 109, 173–186. [Google Scholar] [CrossRef]
- O’Donnell, K.; Kistler, H.C.; Cigelnik, E.; Ploetz, R.C. Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proc. Natl. Acad. Sci. USA 1998, 95, 2044–2049. [Google Scholar] [CrossRef]
- Stępień, Ł. The use of Fusarium secondary metabolite biosynthetic genes in chemotypic and phylogenetic studies. Crit. Rev. Microbiol. 2013, 402, 176–185. [Google Scholar]
- Stępień, Ł.; Gromadzka, K.; Chełkowski, J. Polymorphism of mycotoxin biosynthetic genes among Fusarium equiseti isolates from Italy and Poland. J. Appl. Genet. 2012, 53, 227–236. [Google Scholar] [CrossRef]
- Yang, L.; van der Lee, T.; Yang, X.; Yu, D.; Waalwijk, C. Fusarium populations on Chinese barley show a dramatic gradient in mycotoxin profiles. Phytopathology 2008, 98, 719–727. [Google Scholar] [CrossRef]
- Zhang, J.B.; Li, H.P.; Dang, F.J.; Qu, B.; Xu, Y.B.; Zhao, C.S.; Liao, Y.C. Determination of the trichothecene mycotoxin chemotypes and associated geographical distribution and phylogenetic species of the Fusarium graminearum clade from China. Mycol. Res. 2007, 111, 967–975. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, Z.; van der Lee, T.; Chen, W.Q.; Xu, J.S.; Yang, L.; Yu, D.; Waalwijk, C.; Feng, J. Population genetic analyses of Fusarium asiaticum populations from barley suggest a recent shift favoring 3ADON producers in southern China. Phytopathology 2010, 100, 328–336. [Google Scholar] [CrossRef]
- Zhang, H.; van der Lee, T.; Waalwijk, C.; Chen, W.Q.; Xu, J.; Xu, J.S.; Zhang, Y.; Feng, J. Population Analysis of the Fusarium graminearum Species Complex from Wheat in China Show a Shift to More Aggressive Isolates. PLoS One 2012, 7, e31722. [Google Scholar]
- Kushiro, M.; Saitoh, H.; Sugiura, Y.; Aoki, T.; Kawamoto, S.; Sato, T. Experimental infection of Fusarium proliferatum in Oryza sativa plants; fumonisin B1 production and survival rate in grains. Int. J. Food Microbiol. 2012, 156, 204–208. [Google Scholar] [CrossRef]
- Carter, J.P.; Rezanoor, H.N.; Desjardins, A.E.; Nicholson, P. Variation in Fusarium graminearum isolates from Nepal associated with their host of origin. Plant Pathol. 2000, 49, 452–460. [Google Scholar] [CrossRef]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual; Blackwell Professional: Ames, IA, USA, 2006. [Google Scholar]
- Müllenborn, C.; Steiner, U.; Ludwig, M.; Oerke, E.C. Effect of fungicides on the complex of Fusarium species and saprophytic fungi colonizing wheat kernels. Eur. J. Plant Pathol. 2008, 120, 157–166. [Google Scholar] [CrossRef]
- Yuan, S.; Zhou, M. A major gene for resistance to carbendazim, in field strains of Gibberella zeae. Can. J. Plant Pathol. 2005, 27, 58–63. [Google Scholar] [CrossRef]
- Qiu, J.B.; Xu, J.H.; Shi, J.R. Molecular characterization of the Fusarium graminearum species complex in Eastern China. Eur. J. Plant Pathol. 2014, 139, 811–823. [Google Scholar] [CrossRef]
- Karugia, G.W.; Suga, H.; Gale, L.R.; Nakajima, T.; Ueda, A.; Hyakumachi, M. Population structure of Fusarium asiaticum from two Japanese regions and eastern China. J. Gen. Plant Pathol. 2009, 75, 110–118. [Google Scholar] [CrossRef]
- Zeller, K.A.; Bowden, R.L.; Leslie, J.F. Diversity of epidemic populations of Gibberella zeae from small quadrats in Kansas and North Dakota. Phytopathology 2003, 93, 874–880. [Google Scholar] [CrossRef]
- Burlakoti, R.R.; Neate, S.M.; Adhikari, T.B.; Gyawali, S.; Salas, B.; Steffenson, B.J.; Schwarz, P.B. Trichothecene profiling and population genetic analysis of Gibberella zeae from barley in North Dakota and Minnesota. Phytopathology 2011, 101, 687–695. [Google Scholar] [CrossRef]
- Talas, F.; Parzies, H.K.; Miedaner, T. Diversity in genetic structure and chemotype composition of Fusarium graminearum sensu stricto populations causing wheat head blight in individual fields in Germany. Eur. J. Plant Pathol. 2011, 131, 39–48. [Google Scholar] [CrossRef]
- Ramirez, M.L.; Reynoso, M.M.; Farnochi, M.C.; Torres, A.M.; Leslie, J.F.; Chulze, S.N. Population genetic structure of Gibberella zeae isolated from wheat in Argentina. Food Addit. Contam. 2007, 24, 1115–1120. [Google Scholar] [CrossRef]
- Shen, C.M.; Hu, Y.C.; Sun, H.Y.; Li, W.; Guo, J.H.; Chen, H.G. Geographic distribution of trichothecene chemotypes of the Fusarium graminearum species complex in major winter wheat production areas of China. Plant Dis. 2012, 96, 1172–1178. [Google Scholar] [CrossRef]
- Ndoye, M.; Zhang, J.B.; Wang, J.H.; Gong, A.D.; Li, H.P.; Qu, B.; Li, S.J.; Liao, Y.C. Nivalenol and 15-acetyldeoxynivalenol Chemotypes of Fusarium graminearum Clade Species are Prevalent on Maize throughout China. J. Phytopathol. 2012, 160, 519–524. [Google Scholar] [CrossRef]
- Boutigny, A.L.; Ward, T.J.; Ballois, N.; Iancu, G.; Ioos, R. Diversity of the Fusarium graminearum species complex on French cereals. Eur. J. Plant Pathol. 2014, 138, 133–148. [Google Scholar] [CrossRef]
- Boutigny, A.L.; Ward, T.J.; van Coller, G.J.; Flett, B.; Lamprecht, S.C.; O’Donnell, K.; Viljoen, A. Analysis of the Fusarium graminearum species complex from wheat, barley and maize in South Africa provides evidence of species-specific differences in host preference. Fungal Genet. Biol. 2011, 48, 914–920. [Google Scholar] [CrossRef]
- Desjardins, A.E.; Busman, M.; Manandhar, G.; Jarosz, A.M.; Manandhar, H.K.; Proctor, R.H. Gibberella ear rot of maize (Zea. mays) in Nepal: Distribution of the mycotoxins nivalenol and deoxynivalenol in naturally and experimentally infected maize. J. Agric. Food Chem. 2008, 56, 5428–5436. [Google Scholar] [CrossRef]
- Burlakoti, R.R.; Ali, S.; Secor, G.A.; Neate, S.M.; McMullen, M.P.; Adhikari, T.B. Genetic relationships among populations of Gibberella zeae from barley, wheat, potato, and sugar beet in the Upper Midwest of the United States. Phytopathology 2008, 98, 969–976. [Google Scholar] [CrossRef]
- Wang, X.; Cui, Y.; Fan, F.; Song, Y.; Ren, J.; Meng, Q.; Xu, W.; Jiang, L. Phylogenetic, carbendazim sensitivity and mycotoxin genotype analyses of Fusarium graminearum complex species isolated from wheat Fusarium Head Blight in China. J. Phytopathol. 2010, 158, 576–578. [Google Scholar] [CrossRef]
- Hardy, A.; Silva-Fernandes, A.; Speijers, G.; Hans, R.; Delcour, M.P.; Kuiper, H.; Führ, F.; Carere, A.; Richard-Molard, D.; Thomas, M. Opinion on the Relationship between the Use of Plant Protection Products on Food Plants and the Occurrence of Mycotoxins in Foods; European Commission, Health and Consumer Protection Directorate-General: Brussels, Belgium, 1999; Scientific Committee on Plants; SCP/RESI/063-Final. [Google Scholar]
- D’Mello, J.P.F.; Macdonald, A.M.C.; Briere, L. Mycotoxin production in a carbendazim-resistant strain of Fusarium sporotrichioides. Mycotoxin. Res. 2000, 16, 101–111. [Google Scholar] [CrossRef]
- D’Mello, J.P.F.; MacDonald, A.M.C.; Postel, D.; Dijksma, W.T.P.; Dujardin, A.; Plactina, C.M. Pesticide use and mycotoxin production in Fusarium and Aspergillus. phytopathogens. Eur. J. Plant Pathol. 1998, 104, 741–751. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Yu, J.J.; Zhang, Y.N.; Zhang, X.; Cheng, C.J.; Wang, J.X.; Hollomon, D.W.; Fan, P.S.; Zhou, M.G. Effect of carbendazim resistance on trichothecene production and aggressiveness of Fusarium graminearum. Mol. Plant Microbe Inter. 2009, 22, 1143–1150. [Google Scholar] [CrossRef]
- Kalampokis, I.; Doukas, E.G.; Markoglou, A.N. Effect of DMI-resistance on mycotoxin production and fitness parameters of Fusarium graminearum. In Proceedings of International Congress of Postharvest Pathology, Leida, Spain, 11–14 April 2011.
- Markoglou, A.N.; Vitoratos, A.G.; Doukas, E.G.; Ziogas, B.N. Phytopathogenic and mycotoxigenic characterization of laboratory mutant strains of Fusarium verticillioides resistant to demethylation inhibiting fungicides. In Proceedings of the DPG-BCPC 3rd International Symposium-Plant Protection and Plant Health in Europe, Berlin, Germany, 14–16 May 2009; Feldmann, F., Alford, D.V., Eds.; p. 195.
- Becher, R.; Hettwer, U.; Karlovsky, P.; Deising, H.B.; Wirsel, S.G.R. Adaptation of Fusarium graminearum to tebuconazole yielded descendants diverging for levels of fitness, fungicide resistance, virulence, and mycotoxin production. Phytopathology 2010, 100, 444–453. [Google Scholar] [CrossRef]
- Nicholson, P.; Simpson, D.R.; Weston, G.; Rezanoor, H.N.; Lees, A.K.; Parry, D.W.; Joyce, D. Detection and quantification of Fusarium culmorum and Fusarium graminearum in cereals using PCR assays. Physiol. Mol. Plant Pathol. 1998, 53, 17–37. [Google Scholar] [CrossRef]
- Li, H.P.; Wu, A.B.; Zhao, C.S.; Scholten, O.; Löffler, H.; Liao, Y.C. Development of a generic PCR detection of deoxynivalenol and nivalenol chemotypes of Fusarium graminearum. FEMS Microbiol. Lett. 2005, 243, 505–511. [Google Scholar] [CrossRef]
- Wang, J.H.; Zhang, J.B.; Chen, F.F.; Li, H.P.; Ndoye, M.; Liao, Y.C. A multiplex PCR assay for genetic chemotyping of toxigenic Fusarium graminearum and wheat grains for 3-acetyldeoxynivalenol, 15-acetyldeoxynivalenol and nivalenol mycotoxins. J. Food Agric. Environ. 2012, 10, 505–511. [Google Scholar]
- Alexander, N.J.; Hohn, T.M.; McCormick, S.P. The Tri11 gene of Fusarium sporotrichioides encodes a cytochrome P-450 monooxygenase required for C-15 hydroxylation in trichothecene biosynthesis. Appl. Environ. Microb. 1998, 64, 221–225. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acid Res. 1994, 2222, 4673–4680. [Google Scholar] [CrossRef]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Qiu, J.B.; Xu, J.Q.; Yu, J.J.; Bi, C.W.; Chen, C.J.; Zhou, M.G. Localisation of the benzimidazole fungicide binding site of Gibberella zeae β2-tubulin studied by site-directed mutagenesis. Pest. Manag. Sci. 2011, 67, 191–198. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Qiu, J.; Shi, J. Genetic Relationships, Carbendazim Sensitivity and Mycotoxin Production of the Fusarium Graminearum Populations from Maize, Wheat and Rice in Eastern China. Toxins 2014, 6, 2291-2309. https://doi.org/10.3390/toxins6082291
Qiu J, Shi J. Genetic Relationships, Carbendazim Sensitivity and Mycotoxin Production of the Fusarium Graminearum Populations from Maize, Wheat and Rice in Eastern China. Toxins. 2014; 6(8):2291-2309. https://doi.org/10.3390/toxins6082291
Chicago/Turabian StyleQiu, Jianbo, and Jianrong Shi. 2014. "Genetic Relationships, Carbendazim Sensitivity and Mycotoxin Production of the Fusarium Graminearum Populations from Maize, Wheat and Rice in Eastern China" Toxins 6, no. 8: 2291-2309. https://doi.org/10.3390/toxins6082291




