1. Introduction
Throughout history, some of the natural products most commonly used and abused by humankind have been the fungus-derived ergot alkaloids (reviewed in [
1]). These alkaloids are named after the resting structures (ergots, also called sclerotia), produced by
Claviceps species on ears of grasses. The ergots are common contaminants of cereals that must be diligently removed to prevent poisoning of food and feed grain supplies. The ergot alkaloids are a diverse group of compounds, and depending on the prominent forms that may occur in contaminated grain, they can cause outbreaks of either convulsive or gangrenous ergotism with devastating results. Ergot alkaloids have also been used for millennia to aid childbirth, and for birth control, treatment of migraines and, recently, treatment of parkinsonism and other CNS disorders. They also are source material for the semisynthetic drug, LSD, by far the most hallucinogenic substance known. In addition to ergot alkaloids,
Claviceps species can also produce a second class of neurotropic alkaloids, the indole-diterpenes (“tremorgens”), known to cause staggers in livestock [
2]. Given the importance and notoriety of
Claviceps species, it is fitting that they lend their name to the Clavicipitaceae, a large and interesting family of fungi that is surprisingly diverse in host interactions and natural products.
The Clavicipitaceae (Hypocreales, Pezizomycotina, Ascomycota) include pathogens, parasites and symbionts of plants, invertebrate animals, or other fungi, and can produce four different classes of neurotropic alkaloids. Alkaloid structures, and pathways currently deemed likely for three of these classes are shown in
Figure 1. The aforementioned ergot alkaloids and indole-diterpenes are produced by many Clavicipitaceae, but are also known from fungi in several other orders of the Pezizomycotina. In contrast, two other alkaloid classes, the lolines and peramine, have a much more restricted distribution and more specific activity against invertebrates. Lolines are potent insecticidal alkaloids [
3] that have been found in numerous cool-season grasses (Poaceae, subfamily Poöideae), as well as a spattering of dicotyledonous plants [
4]. The reason for their occurrence in the Poöideae has been determined: They are synthesized by symbiotic Clavicipitaceae in the monophyletic “epichloae” (which includes
Epichloë and
Neotyphodium species). Similarly, the insect feeding deterrent, peramine, is synthesized by many epichloae, and is known only from cool-season grasses possessing these symbionts. There is ample evidence that many epichloae behave as mutualistic symbionts, and that their capability to make alkaloids that combat herbivores amounts to a major host benefit in mutualism [
5].
Figure 1.
Structures and pathways for alkaloids produced by epichloae (adapted from [
6]). Summaries of pathways are shown for indole-diterpenes (
A), ergot alkaloids (
B) and lolines (
C) with structures of major forms of the alkaloids found in grasses symbiotic with epichloae. Panel (
D) shows the structure of a fourth protective alkaloid, peramine, also produced by many epichloae.
Figure 1.
Structures and pathways for alkaloids produced by epichloae (adapted from [
6]). Summaries of pathways are shown for indole-diterpenes (
A), ergot alkaloids (
B) and lolines (
C) with structures of major forms of the alkaloids found in grasses symbiotic with epichloae. Panel (
D) shows the structure of a fourth protective alkaloid, peramine, also produced by many epichloae.
The epichloae grow systemically in plant aerial tissues by an endobiotic growth habit whereby fungal hyphae invade intercellular spaces and grow as adjacent host cells elongate [
7]. Generally there is no obvious external growth from vegetative host tissues, but when host plants produce flowering tillers there is a developmental “decision” as to how the fungus responds. In some cases, every flowering tiller exhibits the epichloid fruiting structure (stroma), which surrounds the immature inflorescence and halts its maturation (choke disease). Ascospores (meiotic spores), and conceivably conidia (mitotic spores) produced on the stromata, can mediate infection of new plants (horizontal transmission) [
8,
9,
10]. In other cases no fruiting structure appears, inflorescences develop normally, and the fungus grows benignly in developing embryos, thereby ensuring transmission from a mother plant to its progeny (vertical transmission). Many epichloae use both transmission strategies, choking some tillers while allowing others to develop normally. However, those epichloae that fail to produce stromata are strictly asexual and—as far as is known—only transmit vertically. The majority of such apparently asexual species described to date have been characterized genetically as interspecific hybrids [
11,
12].
Phylogenetic analysis of housekeeping genes [
11,
12], along with genome size estimates [
13], suggest that such hybrid species have retained most or all of the genome content contributed by two or, sometimes, three ancestral species. Thus, hybridization resulted in diploid or triploid epichloae, whereas the sexual species are haploid. Since there is no evidence that such polyploid hybrids can form easily [
14,
15], it seems likely that hybridization often leads to increased benefits to the fungus and the host grass, for which there is recent empirical evidence [
16,
17]. Selosse and Schardl [
18] suggest several possible benefits, and one that is relevant to this paper is pyramiding of alkaloid genes for enhanced host protection.
In a survey of epichloae for
in symbio production of lolines, ergot alkaloids and peramine, most sexual species were found to lack detectable levels, whereas most asexual species produced one or more of these alkaloids [
19]. Since that survey, techniques have advanced to detect a broader range of ergot alkaloids, indole-diterpenes and lolines, including intermediates, spur products and end products [
4,
20,
21]. Furthermore, most or all of the genes have been identified for biosynthesis of each of the four alkaloid classes [
6]. These advances make it much easier to predict alkaloid profiles on a genetic basis [
22,
23], and to follow up these predictions with comprehensive chemotyping.
Surveys of epichloae for the gene clusters encoding biosynthetic enzymes for lolines, (
LOL), ergot alkaloids (
EAS), and indole-diterpenes (
IDT) have led to interesting findings. Most epichloae, both sexual and asexual, have genes for one or more of these classes of protective alkaloids [
24]. Furthermore, there is considerable chemotypic variation among isolates of several sexual epichloae, though many isolates lack key genes required to make any of these three kinds of alkaloid. In contrast, most asexual epichloae surveyed to date have genes for one or more of these alkaloids, although there is also chemotypic diversity within some hybrid species. There have been fewer surveys for
perA, which is required for peramine production, but the limited results so far suggest that peramine is very common, though not universal, among isolates of both sexual and asexual epichloae.
In this study we survey and sequence selected LOL, EAS and IDT genes from representatives of most known nonhybrid species of epichloae, both sexual and asexual, and from several asexual hybrids, in order to identify the sources of these genes in the hybrids. We find evidence that strict vertical transmission may often serve as a selective filter for strains that produce protective alkaloids.
Table 1.
Species of Epichloë and Neotyphodium in this study, and their most abundant alkaloids.
Table 1.
Species of Epichloë and Neotyphodium in this study, and their most abundant alkaloids.
| | | | | EA a | | IDT a | | LOL a | |
|---|
| Species and variety or genotype | Isolate b | Reference or PI | Host c | Pre-dicted | Observed | Pre-dicted | Ob-served | Pre-dicted | Observed |
|---|
| Epichloë amarillans | E57 = ATCC 200744 | [6] | Agrostis hyemalis | – | – | – | nt | NANL | NANL |
| E. amarillans | E4668 | This study | Agrostis hyemalis | ERV | – | – | nt | – | nt |
| E. baconii | E1031 = ATCC 200745 | [25] | Calamagrostis villosa | – | nt | – | nt | – | nt |
| E. brachyelytri | E4804 | [6] | Brachyelytrum erectum | CC | nt | – | nt | AcAP | AcAP |
| E. bromicola | E502 = ATCC 200750 | [6] | Bromus erectus | – | nt | – | nt | – | – |
| E. canadensis | e4815 | | Elymus canadensis | ERV | ERV CC | – | nt | NANL | NANL |
| E. canadensis | CWR5 | [26] | Elymus canadensis | ERV | ERV CC | – | nt | NANL | NANL |
| E. canadensis | CWR34 | [26] | Elymus canadensis | CC | CC | – | nt | AcAP | AcAP |
| E. elymi | E56 = ATCC 201551 | [6] | Elymus virginicus | CC | CC | – | nt | – | – |
| E. festucae | E2368 | [6] | Festuca rubra and Lolium giganteum (ascospore isolate) | ERV | – | – | nt | NFL | NFL NML NAL NANL |
| E. festucae | Fl1 | [6] | Festuca trachyphylla | ERV | ERV CC | LTB | LTB | – | – |
| E. glyceriae | E277 | [6] | Glyceria striata | ERV | – | – | nt | AcAP | AcAP |
| E. “mollis” | E3601 = AL9924 | This study | Holcus mollis | ERV | nt | – | nt | – | nt |
| E. poae | E4646 | This study | Poa nemoralis | – | nt | – | nt | – | nt |
| E. poae | E5819 | [6] | Poa nemoralis | ERV | nt | – | nt | – | nt |
| E. typhina | E8 = ATCC 200736 | [6] | Lolium perenne | – | nt | – | nt | – | – |
| FaTG-2 G2 | NFe45079 | [23] | Lolium sp. (6x) | ERV | ERV CC | LTB | LTB TD | – | nt |
| FaTG-2 G3 | NFe45115 | [23] | Lolium sp. (6x) | ERV | ERV CC | TD | TD | – | nt |
| FaTG-3 | NFe1100 | This study | Lolium sp. (6x) | – | – | TD | nt | NFL | NFL |
| FaTG-3 | e4074 | [11] | Lolium sp. (6x) | – | – | – | nt | NFL | NFL NML NAL |
| FaTG-4 | e4305 | PI 598863 | Lolium sp. (10x) | ERV | nt | TD | nt | – | – |
| Neotyphodium aotearoae | e899 = MYA-1229 | [27] | Echinopogon ovatus | – | nt | TD | nt | NFL | NFL |
| N. chisosum | e3609 = ATCC 64037 | [28] | Achnatherum eminens | – | nt | – | nt | NFL | nt |
| N. coenophialum | e19 = ATCC 90664 | [11] | Lolium arundinaceum | ERV | ERV CC | – | nt | NFL | NFL NML NAL NANL |
| N. coenophialum | e4163 | PI 422777 | Lolium sp. (4x) | ERV | ERV | TD | nt | NFL | NFL NML NAL NANL |
| N. coenophialum | e4309 | PI 598903 | Lolium arundinaceum | – | – | TD | nt | NFL | NANL |
| N. funkii | e4096 | [28] | Achnatherum robustum | CC | CC | TD | nt | – | – |
| N. gansuense var. inebrians | e818 = MYA-1228 | [6,28] | Achnatherum inebrians | EN LAH | EN LAA LAH | – | nt | – | – |
| N. gansuense | e7080 | [6] | Achnatherum inebrians | – | nt | PAX | PAX | – | nt |
| N. occultans | non-culturable | [29] | Lolium spp. (2x) | – | nt | PAX | nt | NFL | NFL NML NAL NANL |
| N. siegelii | e915 = ATCC 74483 | [11,30] | Lolium pratense | – | – | PAX | nt | NFL | NFL NML NAL NANL |
| N. uncinatum | e167 = CBS 102646 | [6] | Lolium pratense | – | – | – | nt | NFL | NFL NML NAL NANL |
| N. lolii × E. typhina | Lp1 = e144 | [11] | Lolium perenne | ERV | ERV CC | TD | TD | – | – |
| PauTG-2 | e55 | [11] | Poa autumnalis | nt | nt | nt | nt | NFL | NFL NML NANL |
2. Results
2.1. Genome Assemblies and Identification of Alkaloid Genes
Isolates of
Epichloë and
Neotyphodium species are listed in
Table 1, and statistics for genome assemblies are given in
Table 2. For genomes with comparable fold-coverage and comparable average read length, the quality of the assemblies depended mainly on the genome size and ploidy. Considering its polyploid nature and large genome size, an exceptionally good assembly was obtained for the
N. coenophialum e4163 genome, in which three of the four alkaloid gene clusters were contained within individual contigs of greater than 150 kb (
Figure 2). One of these contigs, which contained the
IDT cluster, had a telomeric repeat beginning 565 bp downstream of the
idtK stop codon. Both the
EAS1 and
EAS2 clusters were assembled on separate contigs, and the very close relationship of all
EAS2 genes with the
E. festucae EAS genes indicated that the clusters remained intact according to their ancestral relationships. Although the
N. coenophialum e4163 assembly showed no indication of telomeres linked to the
EAS clusters, in the assembly of
N. coenophialum strain e19, both
lpsB1 and
lpsB2 were on contigs with telomere repeats at the ends. In both cases, the telomeres were downstream of the
lpsB gene. The
N. coenophialumLOL clusters were not completely assembled, but were arranged in three contigs of the e4163 assembly. One contig contained
lolF and flanking (downstream) housekeeping genes, another contig contained eight genes,
lolC-lolE, arranged similarly to those in
N. uncinatum [
31], and the third contig contained
lolN and
lolM. The arrangement of the
LOL genes and housekeeping genes flanking
lolF was consistent with their arrangement in the genome of
E. festucae E2368 [
6].
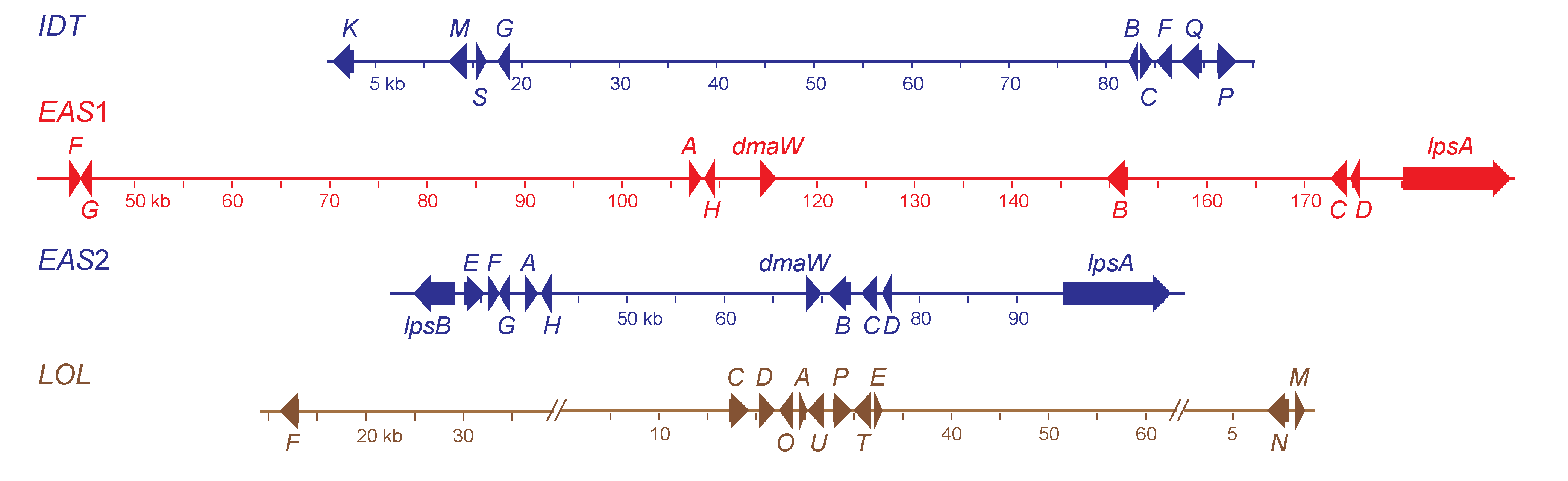
Figure 2.
Structures of the alkaloid biosynthesis gene clusters in
Neotyphodium coenophialum e4163. Maps are color coded according to ancestral clade of origin (see
Figure 3). The indole-diterpene (
IDT) gene cluster was from clade II (
E. festucae). Each gene in the
IDT cluster (accession number KC970578) is designated by its final letter, where
K =
idtK,
etc. Two ergot-alkaloid gene clusters were identified, one from clade V (
EAS1, accession number KC989569), and the other from clade II (
EAS2, accession number KC989570). Genes in the
EAS clusters are abbreviated as follows:
A = easA,
B = cloA,
C = easC,
D = easD,
E = easE,
F = easF,
G = easG,
H = easH. Other
EAS cluster gene names are given in full. The loline alkaloid (
LOL, accession numbers KC990458, KC990457 and KC990459) gene cluster was from clade Ib (
E. poae). Each gene in the
LOL cluster is designated by its final letter, where
F =
lolF,
etc.
Figure 2.
Structures of the alkaloid biosynthesis gene clusters in
Neotyphodium coenophialum e4163. Maps are color coded according to ancestral clade of origin (see
Figure 3). The indole-diterpene (
IDT) gene cluster was from clade II (
E. festucae). Each gene in the
IDT cluster (accession number KC970578) is designated by its final letter, where
K =
idtK,
etc. Two ergot-alkaloid gene clusters were identified, one from clade V (
EAS1, accession number KC989569), and the other from clade II (
EAS2, accession number KC989570). Genes in the
EAS clusters are abbreviated as follows:
A = easA,
B = cloA,
C = easC,
D = easD,
E = easE,
F = easF,
G = easG,
H = easH. Other
EAS cluster gene names are given in full. The loline alkaloid (
LOL, accession numbers KC990458, KC990457 and KC990459) gene cluster was from clade Ib (
E. poae). Each gene in the
LOL cluster is designated by its final letter, where
F =
lolF,
etc.
Table 2.
Assembly statistics for genomes of Epichloë and Neotyphodium species.
Table 2.
Assembly statistics for genomes of Epichloë and Neotyphodium species.
| Strain | Genome assembly size (Mb) | Endophyte ploidy | Fold-coverage | Number of contigs | Contig N50 (bp) a |
|---|
| E8 | 41.3 | 1 | 21.2 | 2253 | 42,567 |
| e19 | 95.4 | 3 | 34.4 | 8808 | 25,663 |
| E56 | 31.8 | 1 | 25.6 | 5206 | 32,484 |
| E57 | 38.0 | 1 | 27.7 | 1871 | 49,298 |
| Lp1 | 51.1 | 2 | 7.6 | 27,822 | 2228 |
| E277 | 49.3 | 1 | 27.7 | 2658 | 42,649 |
| e818 | 29.8 | 1 | 87.3 | 2519 | 39,993 |
| Fl1 | 34.9 | 1 | 52.3 | 1277 | 84,986 |
| e899 | 34.3 | 1 | 26.3 | 1848 | 56,460 |
| E1031 | 38.0 | 1 | 18.5 | 3358 | 34,415 |
| E2368 | 34.7 | 1 | 27.8 | 1316 | 87,544 |
| E3601 | 36.1 | 1 | 28.0 | 890 | 97,803 |
| e3609 | 108.7 | 3 | 10.2 | 32,104 | 4981 |
| e4096 | 61.2 | 2 | 29.5 | 6100 | 20,015 |
| e4163 | 97.7 | 3 | 32.5 | 4349 | 46,910 |
| e4305 | 76.6 | 2 | 21.9 | 17,406 | 7250 |
| e4309 | 81.8 | 3 | 14.6 | 34,254 | 3494 |
| E4668 | 40.7 | 1 | 25.0 | 1851 | 45,395 |
| E4804 | 44.1 | 1 | 24.6 | 4542 | 20,998 |
| e4815 | 63.3 | 2 | 13.7 | 21,592 | 4186 |
| E5819 | 34.0 | 1 | 28.6 | 2072 | 36,475 |
| E7080 | 39.5 | 1 | 41.5 | 1307 | 56,237 |
For other genomes, most alkaloid genes assembled well, although whole clusters were not generally assembled on single contigs. In cases where two copies of an alkaloid gene were present, the copies often failed to assemble individually over their whole length. Instead, regions with very high similarity tended to collapse into single short contigs. Nevertheless, inspection of the reads revealed polymorphisms indicative of the two gene copies. These situations arose in polyploid (hybrid) genomes with relatively low fold-coverage, affecting the assemblies of EAS genes in E. canadensis e4815 and LOL genes in N. chisosum e3609. To deconvolute the multiple gene copies we inspected the aligned reads in the .ace files generated by the Newbler assembler. For those strains lacking comprehensive genome sequence data, fragments of tubB and alkaloid genes were PCR-amplified and sequenced from template DNA derived from cultures or symbiotic plant tissue.
2.2. Endophyte Species Relationships and Hybrid Origins of Asexual Epichloae
Figure 3 presents a phylogeny based on the first three introns of the beta-tubulin gene,
tubB, relating representatives of sexual
Epichloë and other nonhybrids to the hybrid species in this study. Clades that include contributors to hybrid genomes are numbered I through VI, in accordance with the mating population designations used prior to formal naming of the sexual species [
32]. In some cases these clades correspond to individual sexual species, such as
E. festucae (clade II),
E. elymi (clade III) and
E. amarillans (clade IV). In contrast, clade I has an especially wide diversity, includes isolates from a broad range of grasses within the Poöideae, and encompasses four named
Epichloë species. Members of three of the clade I species,
E. typhina,
E. clarkii and
E. poae, are interfertile, whereas the fourth,
E. sylvatica, apparently is not interfertile with
E. typhina [
25]. Because of their importance in the origins of hybrid species, a major clade of
E. typhina has been designated as clade Ia, and those in the newly describe
E. poae are in clade Ib [
10]. The
E. bromicola clade (clade VI) includes
E. yangzii [
33], which may be considered a synonym of
E. bromicola [
34,
35].
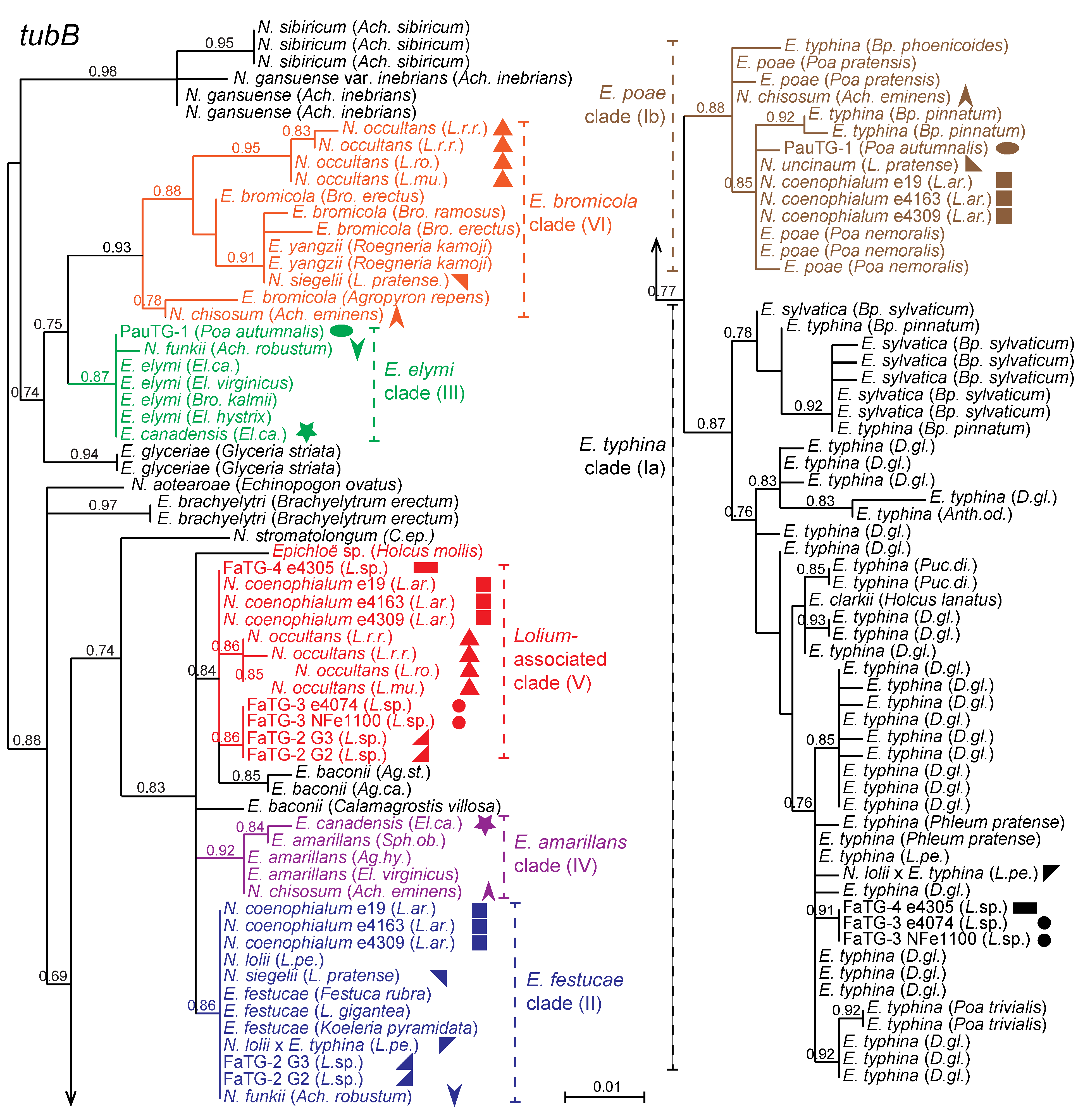
Figure 3.
Phylogeny of
tubB genes from all known haploid (non-hybrid) and ten hybrid epichloae. Where hybrids have multiple gene copies from different ancestral species, the different copies are indicated by symbols of the same shape with different colors. Phylogenetic clades contributing to hybrids are color coded and numbered Ia, Ib, II, III, IV, V, and VI.
Supplementary Table S1 contains the
tubB accession numbers.
Figure 3.
Phylogeny of
tubB genes from all known haploid (non-hybrid) and ten hybrid epichloae. Where hybrids have multiple gene copies from different ancestral species, the different copies are indicated by symbols of the same shape with different colors. Phylogenetic clades contributing to hybrids are color coded and numbered Ia, Ib, II, III, IV, V, and VI.
Supplementary Table S1 contains the
tubB accession numbers.
Hybrid endophytes tend to have multiple copies of housekeeping genes [
11]. With the exception of
N. uncinatum, all of the hybrid endophytes in this study had two or three
tubB genes, sequences of which were in distinct clades (
Figure 3). In past studies, the phylogenies of
tubB and genes for translation elongation factor 1-α (
tefA) and γ-actin (
actG) supported the same hybrid origins for each of these species [
11,
28]. In general, the hybrids are asexual, although some asexual epichloae appear to be haploid and not hybrid in origin.
Most of the broad-leaf members of
Festuca/Lolium/Schedonorus grass complex, such as tall fescue, meadow fescue and perennial ryegrass, have asexual endophytes, which in turn are restricted to individual grass species (see
Table 1). One species,
Neotyphodium lolii, is an asexual derivative of
E. festucae (clade II) that is by far the most common symbiont of
Lolium perenne (perennial ryegrass). This grass also can host
Epichloë typhina, as well as an asexual
N. lolii (clade II) ×
E. typhina (clade Ia) hybrid (exemplified by isolate Lp1). Perennial ryegrass is closely related to other diploid species including
Lolium pratense (=
Schedonorus pratensis; meadow fescue) and the various species of annual ryegrass. The most common endophyte of meadow fescue is
N. uncinatum, a hybrid of
E. bromicola (clade VI) and
E. poae [
10]. A second endophyte from meadow fescue is
N. siegelii, a hybrid of
E. bromicola (clade VI) and
E. festucae (clade II).
Neotyphodium occultans is the most common endophyte in the annual ryegrass species,
Lolium multiflorum,
L. temulentum,
L. canariense, and
L. rigidum. This endophyte has an
E. bromicola (clade VI) genome along with a second genome from the “
Lolium-associated endophyte” clade that is most closely related to
E. baconii isolates from
Agrostis species (clade V). Genomes from clade V appear in four other hybrid epichloae, all endophytes of grasses in the
Lolium arundinaceum (=
Schedonorus arundinaceus; tall fescue) species complex. The most complex hybrid,
N. coenophialum, has genomes from clade V,
E. festucae (clade II) and
E. poae (clade Ib), and has been found in hexaploid (6x) tall fescue from northern Europe and from the Atlas Mountains of North Africa [
36]. Two undescribed hybrid species, designated FaTG-2 and FaTG-3, have been isolated from Mediterranean 6x tall fescue, and two chemotype variants, G2 and G3, have been described previously [
23]. Another hybrid species that we designate FaTG-4 (referred to as FaTG-3-like in Ekanayake
et al. [
37]) was isolated from a decaploid (10x) tall fescue from Morocco. FaTG-2 is a clade V × clade II hybrid. FaTG-3 and FaTG-4 are clade V × clade Ia hybrids, distinguishable by variations in their clade V gene sequences and alkaloid profiles.
Genome sequences were also obtained from two endophytes of
Achnatherum species (tribe Stipeae). The
tubB and other phylogenies (
Figure 3,
Table 3) indicated that
N. chisosum from
Ach. eminens and
N. funkii from
Ach. robustum had different origins [
28]. Interestingly, each of these endophyte species was a hybrid with ancestors related both to North American species (
E. elymi clade III,
E. amarillans clade IV) and Eurasian species (
E. bromicola clade VI,
E. poae clade Ib, and
E. festucae II). These results suggest that the ancestral species previously had overlapping geographical ranges.
Table 3.
Origins of housekeeping and alkaloid gene loci in hybrids a.
Table 3.
Origins of housekeeping and alkaloid gene loci in hybrids a.
| Species | Isolate | tefA | tubB | EAS | IDT | LOL |
|---|
| E. canadensis | e4815 | III,IV | III,IV | III,IV | – | IV |
| E. canadensis | CWR34 | III,IV | III,IV | III | – | IV |
| N. lolii × E. typhina | Lp1 | Ia,II | Ia,II | II | II | – |
| FaTG-2 G2 | NFe45079 | II,V | II,V | II,V | II,V | – |
| FaTG-2 G3 | NFe45115 | II,V | II,V | V | V | – |
| FaTG-3 | NFe1100 | Ia,V | Ia,V | – | V | Ia |
| FaTG-4 | e4305 | Ia,V | Ia,V | V | V | – |
| N. chisosum | e3609 | Ib,IV,VI | Ib,IV,VI | – | – | IV,VI |
| N. coenophialum | e19 | Ib,II,V | Ib,II,V | II,V | (II) | Ib |
| N. coenophialum | e4163 | Ib,II,V | Ib,II,V | II,V | II | Ib |
| N. coenophialum | e4309 | Ib,II,V | Ib,II,V | – | II | Ib |
| N. funkii | e4096 | II,III | II,III | III | II | – |
| N. occultans | various | V,VI | V,VI | – | V,(VI) | VI |
| N. siegelii | e915 | II,VI | II,VI | – | II,(VI) | VI |
| N. uncinatum | e167 | Ib | VI | – | – | Ib,VI |
| PauTG-1 | e55 | Ib,III | Ib,III | – | – | Ib |
Phylogenetic analysis of genes from
E. canadensis, an endophyte of the host grass
Elymus canadensis (tribe Triticeae), supported hybrid origin from two North American ancestors,
E. elymi (clade III) and
E. amarillans (clade IV) (
Figure 3) [
26]. Both of these ancestral species have been isolated from the related host,
Elymus virginicus [
11], although only
E. elymi has so far been observed to fruit on
Elymus species.
2.3. IDT Gene Profiles and Origins in Hybrid Epichloae
Origins of the
IDT genes in the hybrid species were evident from phylogenetic analysis. The
idtM and
idtP phylogenies are provided in
Figure 4, as examples. The
E. festucae ancestors of
N. coenophialum strains e4309 and e4163,
N. funkii e4096, and
N. lolii ×
E. typhina Lp1, provided each of these hybrids with a complete complement of
IDT genes except for the lolitrem B-specifying genes,
ltmE and
ltmJ. Therefore, these hybrids may produce terpendoles or other indole-diterpenes, and apart from Lp1 [
22] their chemotype profiles have yet to be determined. A third
N. coenophialum strain, e19, had only
idtP, which also was apparently derived from the
E. festucae ancestor. None of these three hybrid species had
IDT genes from ancestors other than
E. festucae.
In contrast,
N. siegelii appeared to have acquired most
IDT genes (at least
idtG,
idtC,
idtB,
idtS,
idtM,
idtF, and
idtK) from its
E. festucae ancestor, and
idtP from another source, possibly but not definitively identified as itsclade VI ancestor (
Figure 4). Also in the
N. siegelii idtF gene a single base deletion was identified that should cause a frame shift and early termination, rendering the encoded protein nonfunctional. An identical mutation has been identified in
E. festucae isolates Fg1 (host
Festuca glauca) and Frc7 (host
Festuca rubra subsp.
commutata), which produce mainly lolitrem K and the pathway intermediate, paxilline, respectively [
22].
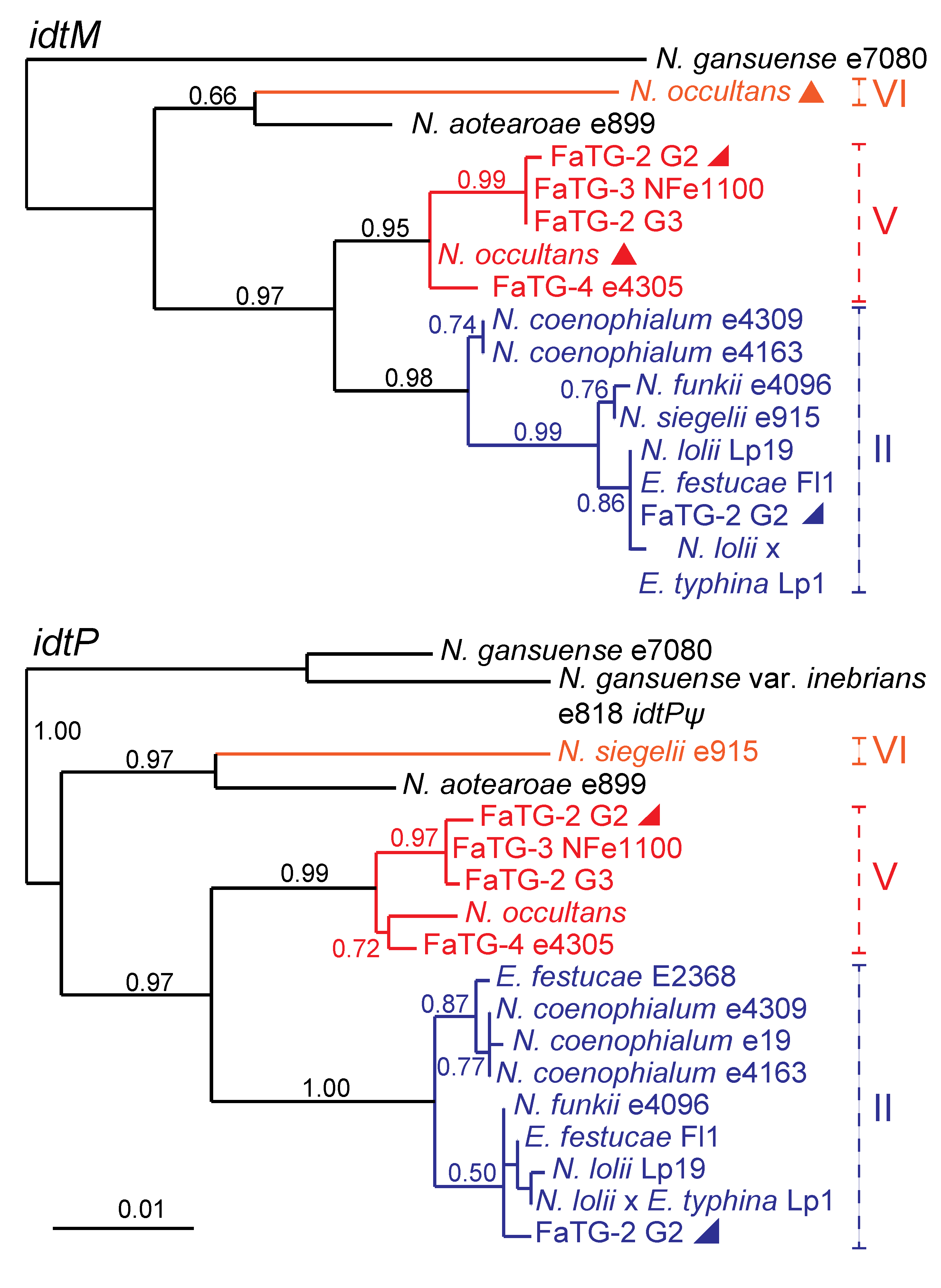
Figure 4.
Phylogeny of two
IDT genes,
idtM and
idtP in epichloae. Inactive genes (pseudogenes) are labeled ψ. Clades and multiple gene copies are labeled as in
Figure 3.
Supplementary Table S2 contains the
IDT accession numbers.
Figure 4.
Phylogeny of two
IDT genes,
idtM and
idtP in epichloae. Inactive genes (pseudogenes) are labeled ψ. Clades and multiple gene copies are labeled as in
Figure 3.
Supplementary Table S2 contains the
IDT accession numbers.
Neotyphodium occultans and three other distinct hybrid species, tentatively designated FaTG-2 (variants G2 and G3), FaTG-3, and FaTG-4, seemed likely to derive their
IDT genes from the clade Vancestor. In addition,
N. occultans had two copies of
idtC and
idtM, where the second copy seemed likely to be derived from
E. bromicola (clade VI). Also, no
idtK gene was identified in
N. occultans. However, comprehensive analysis of alkaloid clusters from
N. occultans was difficult because this species is not culturable, and we could not rule out the possibility that
idtK and clade VI copies of other
IDT genes failed to yield PCR products due to differential primer specificity. FaTG-2 genotype G2 isolates, which produce lolitrems [
23], also had
IDT gene copies derived from clades II and V, and their
ltmE and
ltmJ genes were from clade II (
E. festucae).
Results of phylogenetic analysis of all
IDT genes for all isolates with sequenced genomes (listed in
Table 2) were consistent with their hybrid ancestors, with the sole exception of
N. aotearoae e899, an asexual but nonhybrid epichloid endophyte. Phylogenetic relationships of five
IDT orthologs,
idtG,
idtC,
idtM,
idtS, and
idtP, and two pseudogenes,
idtB and
idtF, were consistent with the relationships of
N. aotearoae housekeeping genes such as
tubB (see, for example,
Figure 4). Four other genes, which we tentatively designated
idtB-like,
idtF-like,
idtK-like and
idtQ-like, were embedded in AT-rich repetitive sequences, and none assembled on the same contigs as the five
IDT gene orthologs. These four
idt-likegenes in
N. aotearoae were highly divergent from their closest homologs in other epichloae, suggesting that they may be xenologs (homologs resulting from horizontal gene transfer) or ancient paralogs.
2.4. EAS Gene Profiles and Origins in Hybrid Epichloae
The
EAS genes of hybrid epichloae, like the
IDT and
LOL genes, were derived from ancestors identifiable by comparing the
EAS and housekeeping gene phylogenies. Comprehensive phylogenetic analysis of all
EAS genes for all isolates with sequenced genomes (see
Table 2) indicated relationships consistent with their hybrid origins. As an example,
dmaW and
cloA phylogenies are given in
Figure 5. (For this analysis, the genes from
N. gansuense var.
inebrians E818 were omitted because its
EAS cluster appears from phylogenetic and structural analysis to be much more similar to those of
Periglandula and
Balansia species than to those so far identified in all other epichloae.) Interestingly, paralogous
dmaW genes were identified in two sexual strains,
Epichloë sp. E3601 (from the host grass
Holcus mollis) and
E. baconii E1031 (from
Calamagrostis villosa). These paralogs were not located in
EAS clusters, and though closely related to each other, they were very divergent from
EAS-cluster
dmaW genes. Nevertheless, they grouped within the clade of
dmaW genes from epichloae (data not shown), and for that reason appeared to have arisen from the genes encoding authentic dimethylallyltryptophan synthase. Another surprising result was the close relationship of
E. poaeEAS genes to those of
E. festucae, in stark contrast to the housekeeping gene relationships.
Clades II, III, IV and V all contributed
EAS clusters to hybrids (
Figure 5). Half of the 12 hybrid species investigated had
EAS clusters, and each of three hybrid species,
E. canadensis,
N. coenophialum and FaTG-2 (G2), had
EAS clusters from two different ancestors. Interestingly,
E. canadensis e4815 contained a complete
EAS cluster from the
E. amarillans ancestor but only four
EAS genes (
dmaW,
easE,
easF and
easC) from the
E. elymi ancestor, while the other
E. canadensis isolate CWR34 only contained the same four
E. elymi EAS genes (
Table 3). This was in keeping with the composition of
EAS clusters identified in isolates of
E. elymi (E56) and
E. amarillans (E4668).
Figure 5.
Phylogeny of two
EAS genes,
dmaW and
cloA. Clades and multiple gene copies are labeled as in
Figure 3.
Supplementary Table S2 contains the
EAS accession numbers.
Figure 5.
Phylogeny of two
EAS genes,
dmaW and
cloA. Clades and multiple gene copies are labeled as in
Figure 3.
Supplementary Table S2 contains the
EAS accession numbers.
2.5. LOL Gene Profiles and Origins in Hybrid Epichloae
The relationships of loline biosynthesis genes in hybrids were entirely consistent with the ancestral genome contributions as discerned from the
tubB phylogeny. This was true for
lolC,
lolN, and
lolP (
Figure 6) for all loline producers listed in
Table 3, and for all
LOL genes for all isolates with sequenced genomes (see
Table 2).
Figure 6.
Phylogeny of three
LOL genes,
lolC,
lolN, and
lolP. Pseudogenes of
lolC and
lolP are labeled
lolCψ and
lolPψ, respectively. Clades and multiple gene copies are labeled as in
Figure 3. The
lolN gene of FaTG-3 was not sequenced.
Supplementary Table S2 contains the
LOL accession numbers.
Figure 6.
Phylogeny of three
LOL genes,
lolC,
lolN, and
lolP. Pseudogenes of
lolC and
lolP are labeled
lolCψ and
lolPψ, respectively. Clades and multiple gene copies are labeled as in
Figure 3. The
lolN gene of FaTG-3 was not sequenced.
Supplementary Table S2 contains the
LOL accession numbers.
Even though no
LOL genes were identified in two ancestral species,
E. bromicola and
E. poae, common relationships of
LOL genes in several hybrids indicated their origin from these species or close relatives (
Figure 6). The number of hybrid species with
LOL clusters from
E. bromicola (four) or
E. poae (three) was surprising given the apparent rarity of
LOL in the corresponding sexual species. In a survey of 10 plants of three species with
E. bromicola endophytes, no loline alkaloid producer was identified [
19], and in a survey of 50
Poa nemoralis plants with
E. poae, we found no
LOL genes. Nevertheless, contributions of
E. bromicola and
E. poae ancestors could be inferred from the
LOL gene phylogenies (
Figure 6).
LOL clusters in four hybrids,
N. chisosum (
LOL1),
N. occultans,
N. siegelii and
N. uncinatum (
LOL1), all had closely related
LOL clusters, which were inferred, therefore, to be derived from
E. bromicola ancestors. Similarly, closely related
LOL clusters in
N. coenophialum,
N. uncinatum (
LOL2) and the
E. poae ×
E. elymi hybrid from
Poa autumnalis were inferred to be from
E. poae. The
lolC gene in
Neotyphodium sp. FaTG-3 was related to the inferred
E. poaelolC genes and to
lolC from
N. aotearoae, consistent with the
E. typhina ancestor of FaTG-3. However, as was the case for
E. bromicola and
E. poae, no loline alkaloid producers [
19] and no
LOL genes were found in surveyed populations of
E. typhina. The fourth ancestral contributor of a
LOL cluster was
E. amarillans (or a close relative), with which the
LOL cluster in
E. canadensis and the
LOL2 cluster in
N. chisosum grouped in clade IV.
Diversity of loline alkaloid profiles related to the presence or absence of certain late-pathway genes. Those that had the full suite of 11 LOL genes generally produced NFL and most or all of the other lolines including NANL, NML, loline, and AcAP. (The one exception is e4309, discussed later.) Furthermore, strains that lacked or had inactivating mutations in one late pathway LOL gene also had no functional copy of the downstream LOL genes. Hence, NANL producers E. amarillans E57, E. canadensis e4815, and E. glyceriae E277 lacked functional copies of lolN, lolM and lolP. Furthermore, E. brachyelytri E4804 and E. canadensis CWR34, which accumulated AcAP, lacked or had inactivating mutations in lolO as well as lolN, lolM and lolP. The only exception we observed to this pattern was N. coenophialum e4309, which had no obvious defect in any of the LOL genes and produced NANL, but no NFL. The basis for this chemotypic difference between e4309 and the NFL-producing N. coenophialum strains, such as e19 and e4163, is under investigation.
In every hybrid strain with two LOL clusters, one of the clusters was incomplete. In N. uncinatum, the LOL1 cluster from clade VI (E. bromicola) had all genes required for production of NFL, whereas the cluster from clade Ib lacked functional forms of three genes (lolN, lolM, and lolP) that were apparently required to take NANL to NFL. The clade Ib cluster had a remnant lolP, indicating that the capability to produce NFL may have been lost in the E. poae ancestor, rather than acquired in other clades. In support of this hypothesis, N. coenophialum e19 and e4163 both had LOL clusters from clade Ib that were complete, and both strains made NFL. The sequences of LOL2 genes in N. uncinatum were identical, or nearly so, to those of N. coenophialum.
Similarly, N. chisosum had two LOL clusters. The cluster from clade VI was also complete in this strain, but the cluster from clade IV had an inactivating deletion in lolP, and lacked lolN and lolM. The deletion in lolP was identical to that in E. amarillans E57, consistent with the LOL1 copy originating from the clade IV ancestor.
2.6. Alkaloid Cluster Compositions and Late-Pathway Gene Losses
Table 3 shows the clades of origin of the
EAS,
IDT, and
LOL clusters in species and strains of hybrid epichloae.
Table 4 lists the origins of individual genes in hybrids that have
EAS,
IDT, or
LOL genes from more than one ancestor. In all of these cases, different ancestors contributed the genes for different alkaloids. Interestingly, genome sequencing consistently revealed that when two different ancestors contributed genes for the same class of alkaloids, those clusters always differed from each other in the presence or absence late-pathway genes. For example, FaTG-2 G2 strain NFe45079 had
IDT genes from clades II and V. Although the IDT cluster originating from the
E. festucae (clade II) ancestor had the two additional
IDT genes for the final steps in lolitrem B biosynthesis (
ltmE and
ltmJ), there was no indication of such genes in clade V. The G3 genotype of FaTG-2 (strain NFe45115) lacked the clade II
IDT genes, in keeping with the observation that it produces terpendoles but not lolitrem B [
23]. This example illustrates that variation in homologous loci can lead to diversification of alkaloid profiles when clusters are subsequently lost.
Table 4.
Alkaloid biosynthesis gene origins in selected hybrid endophyte strains a.
Table 4.
Alkaloid biosynthesis gene origins in selected hybrid endophyte strains a.
| | E. can. e4815 | N. chi. e3609 | FaTG-2 G2 | N. coe. e19 | N. coe. e4163 | N. sie. e915 | N. unc. e167 |
|---|
| Ergot alkaloids | ERV CC | nt | ERV | ERV CC | ERV | – | – |
| dmaW | III, IV | – | II,V | II,V | II,V | – | – |
| easF | III, IV | – | II,V | II,V | II,V | – | – |
| easC | III, IV | – | II,V | II,V | II,V | – | – |
| easE | III, IV | – | II,V | II,V | II | – | – |
| easD | IV | – | II,V | II,V | II,V | – | – |
| easA | IV | – | II,V | II,V | II,V | – | – |
| easG | IV | – | II,V | II,V | II,V | – | – |
| cloA | IV | – | II,V | II,V | II,V | – | – |
| lpsB | IV | – | + | IIΨ,V | II | – | – |
| lpsA | IV | – | + | II,V | II,V | – | – |
| easH | IV | – | II,V | II,V | II,V | – | – |
| Indole-diterpenes | – | – | TD, LTB | – | nt | nt | – |
| idtG | – | – | + | – | II | II | – |
| idtC | – | – | II,V | – | II | II | – |
| idtM | – | – | II,V | – | II | II | – |
| idtB | – | – | II,V | – | II | II | – |
| idtS | – | – | II,V | – | II | II | – |
| idtP | – | – | II,V | II | II | VI | – |
| idtQ | – | – | II,V | – | II | + | – |
| idtF | – | – | II,V | – | II | IIΨ | – |
| idtK | – | – | II,V | – | II | II | – |
| idtE | – | – | II | – | – | – | – |
| idtJ | – | – | II | – | – | – | – |
| Lolines | NANL | nt | – | NFL NAL NANL NML | NFL NAL NANL NML | NFL NAL NANL NML | NFL NAL NANL NML |
| lolC | IV | IV,VI | – | Ib | Ib | VI | Ib,VI |
| lolF | IV | IV,VI | – | Ib | Ib | VI | Ib,VI |
| lolD | IV | IV,VI | – | Ib | Ib | VI | Ib,VI |
| lolT | IV | IV,VI | – | Ib | Ib | + | Ib,VI |
| lolU | IV | IV,VI | – | Ib | Ib | + | Ib,VI |
| lolO | IV | IV,VI | – | Ib | Ib | + | Ib,VI |
| lolE | IV | IV,VI | – | Ib | Ib | VI | Ib,VI |
| lolN | – | VI | – | Ib | Ib | VI | VI |
| lolM | – | VI | – | Ib | Ib | + | VI |
| lolP | IV∆ | IV∆,VI | – | Ib | Ib | VI | Ib∆,VI |
| lolA | IV | IV,VI | – | Ib | Ib | + | Ib,VI |
The EAS and LOL clusters in hybrids showed a similar pattern. Both N. chisosum and N. uncinatum had one complete LOL gene cluster and a second cluster that lacked the three genes for conversion of NANL to NFL. Likewise, there were three examples of strains with two EAS clusters where only one had all genes known to be required for ergovaline production. In E. canadensis e4815 the clade IV EAS cluster was complete, but the clade III EAS cluster had only the four functional genes for biosynthesis of chanoclavine. Since the clade IV EAS cluster is absent in strain CWR34, plants with this strain accumulate chanoclavine but not ergovaline. Similarly, in N. coenophialum e4163, the EAS1 cluster lacked easA and lpsB, whereas in another N. coenophialum strain, e19, EAS1 was complete, and the EAS2 cluster had a defective lpsB gene.
Based on the cluster maps in E. festucae and N. coenophialum e4163 (where the cluster sequences were best assembled), many of the genes for late pathway steps were located at the peripheries of the respective alkaloid gene clusters, and genes at such locations were often the ones that were absent from redundant clusters in hybrids. Specifically, lpsB and easE were at one end of the EAS cluster, lolN and lolM were at one end of the LOL cluster, ltmE and ltmJ were at one end of IDT, and idtK was at the other end of IDT. The exception was lolP, which was in the LOL core near early pathway genes. The observation that in all five examples of redundant alkaloid gene clusters (one for IDT,one for LOL and three for EAS) redundant genes for late pathway steps were absent raises the possibility that this pattern was determined by natural selection.
3. Discussion
The occurrence of alkaloid biosynthesis loci in the hybrid epichloae, as well as the relatively high levels of alkaloids that they produce in symbio [
38,
39], fits with theoretical predictions of the evolution of mutualism in heritable symbioses. The great majority of
Epichloë and
Neotyphodium species (epichloae) are capable of vertical, maternal-line transmission. Horizontal transmission in this group of fungi has so far been documented only for those that produce external stromata, manifesting abundant mitotic spores (conidia) or meiotic spores (ascospores) that can mediate infection of new plants or seeds [
8,
9,
10]. Only one hybrid species,
Epichloë liyangensis, has ever been observed to produce stromata [
40], so it appears that the hybrids are generally dependent on vertical transmission. Such a situation is expected to select for mutualism because the fitness of a vertically transmitted symbiont links positively to host fitness [
41,
42]. A common benefit conferred by epichloae to their hosts is the production of protective alkaloids that can deter, impair or even kill invertebrate and vertebrate herbivores [
1,
3,
4,
22,
43,
44,
45,
46,
47,
48,
49,
50,
51,
52,
53]. Whenever a hybrid forms, its fitness may depend on the alkaloid biosynthesis capabilities that it has obtained from its ancestors. Such selection on hybrids may explain a curious finding of this study; that is, surveys of
E. bromicola,
E. poae, and
E. typhina have failed to turn up isolates with
LOL genes, and in sequenced genomes of those species there was no indication of remnant
LOL genes of any sort. Yet, these three sexual species or close relatives were the apparent donors of
LOL clusters to five of the six loline-producing hybrid species in this study. Thus, it appears that selection has served as a highly effective filter for hybrid epichloae with alkaloid production capabilities.
The contrast of apparently low frequencies of
LOL genes in
E. bromicola,
E. poae, and
E. typhina, versus the existence of several hybrids that possess
LOL genes from one or more of these species, begs the question whether hybrid, asexual epichloae have more tendency to produce protective alkaloids than do sexual non-hybrids. The vast majority of hybrid epichloae surveyed to date have alkaloid biosynthesis genes. For instance, of 32 isolates from 24 hybrid species surveyed earlier, 26 isolates (81%) from 20 species have
EAS,
IDT or
LOL genes [
24]. In contrast, 53 isolates from ten sexual species have been surveyed, of which 23 isolates (43%) from eight species test positive for the determinant gene in each cluster. This is a slight downward revision from the number of positives reported previously in sexual isolates [
24], but follow-up investigations indicated that at least four cases of positive tests were due to paralogs that are probably not involved in those alkaloid pathways (for example, the
dmaW paralog in
E. baconii E1031). Although these surveys cannot be considered unbiased or random, they are suggestive that selection for alkaloid production might be stronger on epichloae that are strictly vertically transmitted, and such a hypothesis is worth further investigation.
It is intriguing that whenever hybrids had two homologous gene clusters for an alkaloid, one of the clusters lacked some late pathway genes. This was true of all such hybrids for which, by genome sequencing, we could conduct a comprehensive analysis of all of the alkaloid gene clusters. We can speculate on why this pattern is so common. One possibility is that it arose simply by chance and might simply reflect the frequent occurrence of various forms of those clusters in the ancestors of the hybrids. In fact, the alkaloid gene clusters of epichloae seem especially suited to gene losses because of their extensive tracts of interspersed repeats [
6]. In the case of
N. coenophialum, it appears that two ancestors contributed complete
EAS clusters, and as lineages then diverged, some lost certain genes, others lost different genes, and in some strains (e.g., e4309) the
EAS genes were altogether lost. These gene losses may just reflect an absence of selection to maintain them, and in the cases of strains e19 and e4163, the remaining complete cluster complements the defects. But, if these situations are simply random, why was there a tendency for late-pathway genes to be lost?
An intriguing alternative reason behind the selective loss of redundant late-pathway genes could be that the resulting alkaloid profiles provide added benefit to the host grass and, by extension, to the symbiont (again, because of reliance on vertical transmission). Panaccione [
54] hypothesized that alkaloid pathways may be less than maximally efficient to allow for accumulation of a variety of pathway intermediates. These intermediates may differ in biological activities, so that combinations of bioactive compounds may confer additive or synergistic activities against herbivores and parasites. Reduplication of an alkaloid gene cluster by hybridization may lead to increased efficiency, thereby reducing levels of intermediates and spur products relative to the pathway end product. In contrast, eliminating late pathway genes from just one of two homologous clusters in a genome might cause a reduced efficiency of the late pathway steps relative to early pathway steps, resulting in a more complex mixture of alkaloids. In support of this hypothesis, most plants with endophytes predicted to produce ergovaline have intermediates as well, particularly chanoclavine [
54]. Likewise, NFL producers have several loline alkaloid intermediates, often with substantial levels of NANL [
55,
56,
57]. And, the indole-diterpene pathway is no exception, with numerous intermediates and spur products evident in plants with endophytes that make lolitrem B [
22]. Therefore, future surveys should also be designed to address the question whether certain alkaloid profiles, including abundant expression of intermediates and spur products, might be selectively favored over simpler profiles dominated by pathway end products.