Phylogenetic Study of Polyketide Synthases and Nonribosomal Peptide Synthetases Involved in the Biosynthesis of Mycotoxins
Abstract
:1. Introduction
2. Results and Discussion
| Mycotoxin | Code | Fungal species | PKS | NRPS | Hybrid PKS-NRPS |
|---|---|---|---|---|---|
| AFLATOXIN | PksA | Aspergillus nomius | Q5VD79 | ||
| AFLATOXIN | PksA | Aspergillus flavus | AAS89999 | ||
| AFLATOXIN | PksL1 | Aspergillus oryzae | BAC45240 | ||
| AFLATOXIN | PksL1 | Aspergillus sojae | AAU08792 | ||
| AFLATOXIN | PksL1 | Aspergillus parasiticus | AAC41675 | ||
| AF-TOXIN | AFT9-1 | Alternaria alternata | BAD97694 | ||
| ALTERNARIOL | PksJ | Alternaria alternata | AFN68301 | ||
| ALTERNARIOL | PksH | Alternaria alternata | AFN68299 | ||
| BEAUVERICIN | BEAS | Beauveria bassiana | ACI30655 | ||
| BEAUVERICIN | putative | Fusarium oxysporum | EGU75688 | ||
| BIKAVERIN | Pks4 | Gibberella fujikuroi | CAB92399 | ||
| CERCOSPORIN | Cercospora nicotianae | AAT69682 | |||
| CERCOSPORIN | CTB1 | Cercospora coffeicola | ADO14690 | ||
| CITRININ | PksCT | Monascus purpureus | AAY33862 | ||
| CITRININ | Trichophyton tonsurans | EGD97507 | |||
| CITRININ | Coccidioides immitis | EJB11047 | |||
| CITRININ | putative | Aspergillus fumigatus | XP_746971 | ||
| CITRININ | putative | Neosartorya fischeri | EAW17018 | ||
| CYCLOPIAZONIC ACID | CpaA | Aspergillus flavus | BAI43678 | ||
| CYCLOPIAZONIC ACID | CpaA | Aspergillus oryzae | BAG82673 | ||
| CHETOGLOBOSIN A | CheA | Penicillium expansum | CAO91861 | ||
| CYTOCHALASIN E | CcsA | Aspergillus clavatus | XM_001270542 | ||
| DOTHISTROMIN | PksA | Mycosphaerella pini | AAZ95017 | ||
| ENNIATIN | Esyn1 | Fusarium equiseti | CAA79245 | ||
| ENNIATIN | Fusarium verticillioides | FVEG_09993 | |||
| EQUISETIN | EqiS | Fusarium heterosporum | Q5SBL2 | ||
| ERGOT ALKALOIDS | CPPS1 | Claviceps purpurea | CAB39315 | ||
| ERGOT ALKALOIDS | CPPS2 | Claviceps purpurea | AJ439610 | ||
| ERGOT ALKALOIDS | CPPS3 | Claviceps purpurea | AJ884677 | ||
| ERGOT ALKALOIDS | CPPS4 | Claviceps purpurea | AJ884678 | ||
| FUMONISIN | FUM1 | Fusarium oxysporum | ACB12550 | ||
| FUMONISIN | FUM1 | Gibberella moniliformis | AF155773 | ||
| FUMONISIN | FUM1-like | Aspergillus niger | CAK43811 | ||
| FUMONISIN | putative | Cochliobolus heterostrophus | AAR90266 | ||
| FUMONISIN | FUM10 | Fusarium oxysporum | ACB12556 | ||
| FUMONISIN | FUM10- like | Aspergillus niger | XP001389107 | ||
| FUMONISIN | FUM14 | Fusarium oxysporum | ACB12559 | ||
| FUMONISIN | FUM14 | Fusarium verticillioides | AAN74817 | ||
| FUMONISIN | FUM14-like | Aspergillus niger | XP_001389112 | ||
| FUMONISIN | putative | Trichoderma atroviride | EHK44571 | ||
| FUSARIN C | FusA | Fusarium fujikuroi | AFP73394 | ||
| FUSARIN C | FusS | Gibberella moniliformis | AAT28740 | ||
| FUSARIN C | PKS10 | Fusarium pseudograminearum | EKJ71911 | ||
| GLIOTOXIN | GliP | Aspergillus fumigatus | AAW03307 | ||
| GLIOTOXIN | GliP | Neosartorya fischeri | XP_001258083 | ||
| GLIOTOXIN | GliP-like | Aspergillus flavus | XP_002380016 | ||
| GLIOTOXIN | GliP-like | Trichoderma virens | EHK22005 | ||
| GLIOTOXIN | putative | Zymoseptoria tritici | XP_003854720 | ||
| HC TOXIN | HTS1 | Cochliobolus carbonum | AAA33023 | ||
| HC TOXIN | Penicillium digitatum | EKV17246 | |||
| LOVASTATIN | LovF | Aspergillus terreus | AAD34559 | ||
| LOVASTATIN | LovB | Aspergillus terreus | Q9Y8A5 | ||
| LOVASTATIN | Magnaporthe oryzae | EHA58179 | |||
| LOVASTATIN | LovB-like | Aspergillus fumigatus | XP_751268 | ||
| LOVASTATIN | putative | Neosartorya fischeri | XP_001262312 | ||
| LOVASTATIN | putative | Fusarium fujikuroi | CAC44633 | ||
| LOVASTATIN | putative | Aspergillus flavus | XP_002385159 | ||
| LOVASTATIN | putative | Aspergillus clavatus | XP_001270376 | ||
| 6-METHYL SALICILIC AC | AtX | Aspergillus terreus | BAA20102.2 | ||
| 6-METHYL SALICILIC AC | MsaS | Aspergillus clavatus | XP_001273093 | ||
| 6-METHYL SALICILIC AC | 6-MSAS | Penicillium griseofulvum | P22367 | ||
| NAPHTHOPYRONE | At4 | Aspergillus terreus | BAB88689 | ||
| NAPHTHOPYRONE | YWA1 | Arthroderma otae | XP_002842704 | ||
| NAPHTHOPYRONE | PksP/Alb1 | Aspergillus fumigatus | AAC39471 | ||
| NAPHTHOPYRONE | WA | Aspergillus nidulans | Q03149 | ||
| OCHRATOXIN A | otapksPN/otanpsPN | Penicillium nordicum | AAP33839 | AAS98174 | |
| OCHRATOXIN A | Aspergillus ochraceus | AAP32477/AAT92023 | |||
| OCHRATOXIN A | Aspergillus niger | XP_001397313 | CAK42678 | ||
| OCHRATOXIN A | Aspergillus carbonarius | PI73482 * | PI 132610* | ||
| OCHRATOXIN A | Aspergillus carbonarius | CAQ16344 | |||
| OCHRATOXIN A | Aoks1 | Aspergillus westerdijkiae | AY583209 | ||
| OCHRATOXIN A | putative | Penicillium verrucosum | AEN14493 | ||
| OCHRATOXIN A | putative | Aspergillus fumigatus | XP748589 | ||
| OCHRATOXIN A | putative | Aspergillus flavus | XP_002375804 | ||
| OCHRATOXIN A | putative | Neurospora crassa | XP955820 | ||
| OCHRATOXIN A | putative | Gibberella zeae | XP390793 | ||
| PATULIN | Penicillium expansum | DQ084387 | |||
| PENICILLIN | Penicillium chrysogenum | P26046 | |||
| PENICILLIN | AcvA | Aspergillus nidulans | CBF84349 | ||
| PM-TOXIN | PKS1 | Mycosphaerella zeae-maydis | AY495642 | ||
| PSEUROTIN A | PsoA | Aspergillus fumigatus | ABS87601 | ||
| SIDEROPHORE | SidC | Aspergillus fumigatus | EAL91050 | ||
| SIDEROPHORE | SidC | Aspergillus nidulans | CBF89140 | ||
| SIRODESMIN | SirP | Leptospheria maculans | AAS92545 | ||
| STERIGMATOCYSTIN | AflC | Aspergillus ochraceoroseus | ACH72912 | ||
| STERIGMATOCYSTIN | PksST | Aspergillus nidulans | AAA81586 | ||
| T-TOXIN | PKS1 | Cochliobolus heterostrophus | AAB08104 | ||
| T-TOXIN | PKS2 | Cochliobolus heterostrophus | AY495643 | ||
| ZEARALENON | PKS13 | Gibberella zeae | ABB90282 | ||
| ZEARALENON | PKS4 | Gibberella zeae | ABB90283 |
2.1. Analyses of KS and AT Domains of PKSs


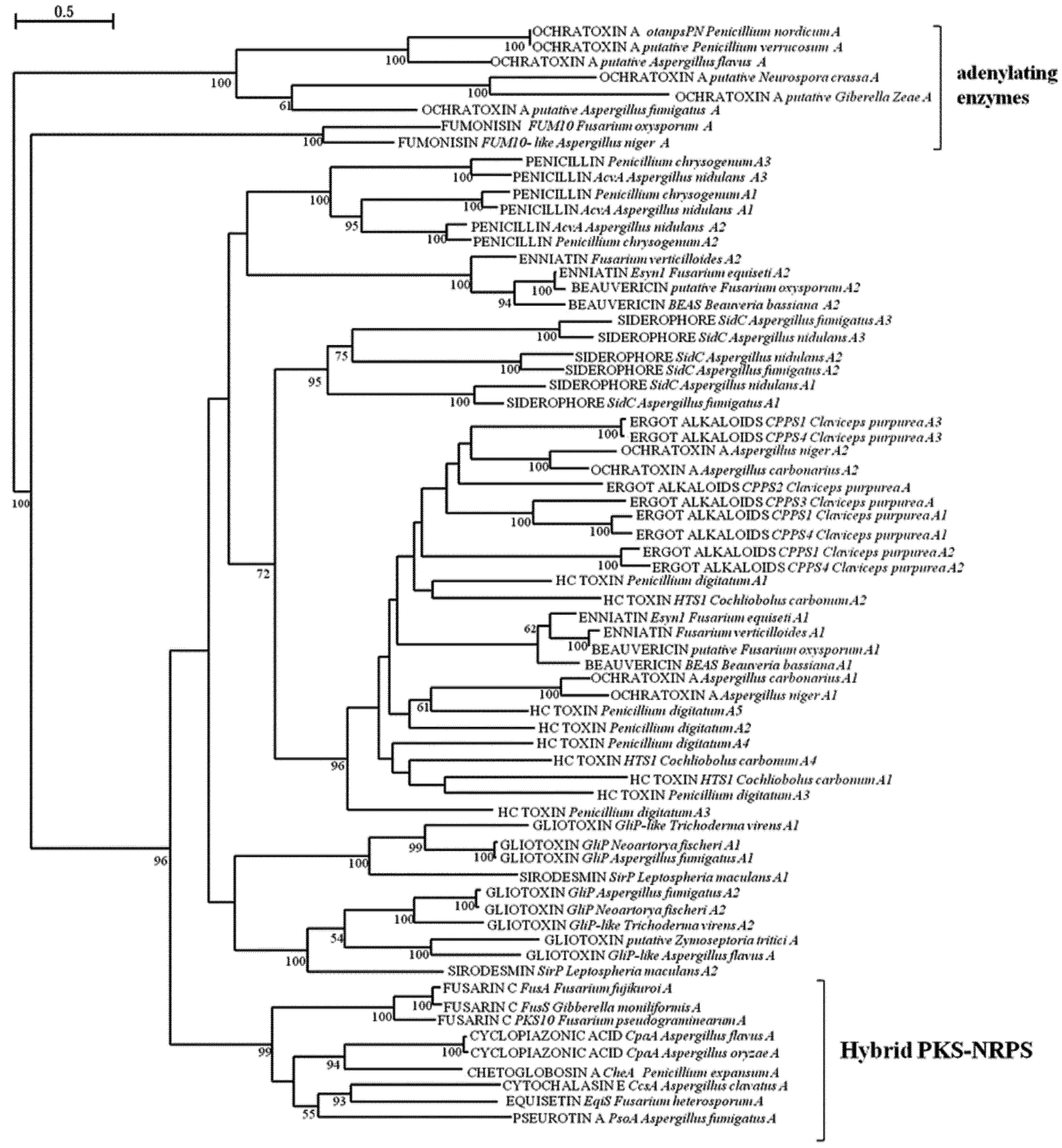
2.2. Analysis of A and C Domains of NRPSs

3. Experimental Section
3.1. Identification of PKS and NRPS Involved in Mycotoxin Biosynthesis
3.2. Identification of Enzymatic Domains
3.3. Multiple Sequence Alignment and Phylogenetic Analysis
4. Conclusion
References
- Finking, R.; Marahiel, M.A. Biosynthesis of nonribosomal peptides. Annu. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [CrossRef]
- Hertweck, C. Hidden biosynthetic treasures brought to light. Nat. Chem. Biol. 2009, 5, 450–452. [Google Scholar] [CrossRef]
- Fischbach, M.A.; Walsh, C.T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: Logic, machinery, and mechanism. Chem. Rev. 2006, 106, 3468–3496. [Google Scholar] [CrossRef]
- Schwarzer, D.; Marahiel, M.A. Multimodular biocatalysts for natural product assembly. Naturwissenschaften 2001, 88, 93–101. [Google Scholar] [CrossRef]
- Staunton, J.; Weissman, K.J. Polyketide biosynthesis: A millennium review. Nat. Prod. Rep. 2001, 18, 380–416. [Google Scholar] [CrossRef]
- Cox, R.J.; Simpson, T.J. Complex enzymes in microbial natural product biosynthesis, part B: Polyketides, aminocoumarins and carbohydrates. Chapter3: Fungal type I polyketide synthases. Methods Enzymol. 2009, 459, 49–98. [Google Scholar]
- Rix, U.; Fischer, C.; Remsing, L.L.; Rohr, J. Modification of post-PKS tailoring steps through combinatorial biosynthesis. Nat. Prod. Rep. 2002, 19, 542–580. [Google Scholar] [CrossRef]
- Meier, J.L.; Burkart, M.D. Complex Enzymes in Microbial Natural Product Biosynthesis, Part A: Overview Articles and Peptides. Chapter 9. Synthetic probes for polyketide and nonribosomal peptide biosynthetic enzymes. Methods Enzymol. 2009, 458, 219–254. [Google Scholar]
- Amoutzias, G.D.; Van de Peer, Y.; Mossialos, D. Evolution and taxonomic distribution of nonribosomal peptide and polyketide synthases. Future Microbiol. 2008, 3, 361–370. [Google Scholar] [CrossRef]
- Haynes, S.W.; Challis, G.L. Non-linear enzymatic logic in natural product modular mega-synthases and -synthetases. Curr. Opin. Drug. Discov. Devel. 2007, 10, 203–218. [Google Scholar]
- Hoffmeister, D.; Keller, N.P. Natural products of filamentous fungi: Enzymes, genes, and their regulation. Nat. Prod. Rep. 2007, 24, 393–416. [Google Scholar] [CrossRef]
- Kopp, F.; Marahiel, M.A. Macrocyclization strategies in polyketide and nonribosomal peptide biosynthesis. Nat. Prod. Rep. 2007, 24, 735–749. [Google Scholar] [CrossRef]
- Tanovic, A.; Samel, S.A.; Essen, L.O.; Marahiel, M.A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 2008, 321, 659–663. [Google Scholar] [CrossRef]
- Von Döhren, H. A survey of nonribosomal peptide synthetase (NRPS) genes in Aspergillus nidulans. Fungal Genet. Biol. 2009, 46, S45–S52. [Google Scholar] [CrossRef]
- Strieker, M.; Tanović, A.; Marahiel, M.A. Nonribosomal peptide synthetases: Structures and dynamics. Curr. Opin. Struct. Biol. 2010, 20, 234–240. [Google Scholar] [CrossRef]
- Evans, B.S.; Robinson, S.J.; Kelleher, N.L. Surveys of non-ribosomal peptide and polyketide assembly lines in fungi and prospects for their analysis in vitro and in vivo. Fungal Genet. Biol. 2011, 48, 49–61. [Google Scholar] [CrossRef]
- Eisfeld, K. Non-ribosomal peptide synthetases of fungi. In Physiology and Genetics: Selected Basic and Applied Aspects, 1st; Anke, T., Weber, D., Eds.; Springer: Heidelberg, Germany, 2009; Volume 15, pp. 305–330. [Google Scholar]
- Du, L.; Sánchez, C.; Shen, B. Hybrid peptide-polyketide natural products: Biosynthesis and prospects toward engineering novel molecules. Metab. Eng. 2001, 3, 78–95. [Google Scholar] [CrossRef]
- Jestoi, M. Emerging fusarium-mycotoxins fusaproliferin, beauvericin, enniatins, and moniliformin: A review. Crit. Rev. Food Sci. Nutr. 2008, 48, 21–49. [Google Scholar]
- Keller, N.P.; Hohn, T.M. Metabolic pathway gene clusters in filamentous fungi. Fungal Genet. Biol. 1997, 21, 17–29. [Google Scholar] [CrossRef]
- Bok, J.W.; Hoffmeister, D.; Maggio-Hall, L.A.; Murillo, R.; Glasner, J.D.; Keller, N.P. Genomic mining for Aspergillus natural products. Chem. Biol. 2006, 13, 31–37. [Google Scholar] [CrossRef]
- Winter, J.M.; Behnken, S.; Hertweck, C. Genomics-inspired discovery of natural products. Curr. Opin. Chem. Biol. 2011, 15, 22–31. [Google Scholar] [CrossRef]
- Wong, S.; Wolfe, K.H. Birth of a metabolic gene cluster in yeast by adaptive gene relocation. Nat. Genet. 2005, 37, 777–782. [Google Scholar] [CrossRef]
- Walton, J.D. Horizontal gene transfer and the evolution of secondary metabolite gene clusters in fungi: An hypothesis. Fungal Genet. Biol. 2000, 30, 167–171. [Google Scholar]
- Khaldi, N.; Collemare, J.; Lebrun, M.H.; Wolfe, K.H. Evidence for horizontal transfer of a secondary metabolite gene cluster between fungi. Genome Biol. 2008, 9, R18. [Google Scholar] [CrossRef]
- Keller, N.P.; Turner, G.; Bennett, J.W. Fungal secondary metabolism - from biochemistry to genomics. Nat. Rev. Microbiol. 2005, 3, 937–947. [Google Scholar] [CrossRef]
- Piel, J.; Höfer, I.; Hui, D. Evidence for a symbiosis island involved in horizontal acquisition of pederin biosynthetic capabilities by the bacterial symbiont of Paederus fuscipes beetles. J. Bacteriol. 2004, 186, 1280–1286. [Google Scholar] [CrossRef]
- Eisen, J.A.; Fraser, C.M. Phylogenomics: Intersection of evolution and genomics. Science 2003, 300, 1706–1707. [Google Scholar] [CrossRef]
- Kroken, S.; Glass, N.L.; Taylor, J.W.; Yoder, O.C.; Turgeon, B.G. Phylogenomic analysis of type I polyketide synthase genes in pathogenic and saprobic ascomycetes. Proc. Natl. Acad. Sci. USA 2003, 100, 15670–15675. [Google Scholar] [CrossRef]
- Schmitt, I.; Lumbsch, H.T. Ancient horizontal gene transfer from bacteria enhances biosynthetic capabilities of fungi. PLoS One 2009, 4, e4437. [Google Scholar] [CrossRef]
- Ziemert, N.; Podell, S.; Penn, K.; Badger, J.H.; Allen, E.; Jensen, P.R. The natural product domain seeker NaPDoS: A phylogeny based bioinformatic tool to classify secondary metabolite gene diversity. PLoS One 2012, 7, e34064. [Google Scholar]
- Bushley, K.E.; Turgeon, B.G. Phylogenomics reveals subfamilies of fungal nonribosomal peptide synthetases and their evolutionary relationships. BMC Evol. Biol. 2010, 10, 26. [Google Scholar]
- Nguyen, T.; Ishida, K.; Jenke-Kodama, H.; Dittmann, E.; Gurgui, C.; Hochmuth, T.; Taudien, S.; Platzer, M.; Hertweck, C.; Piel, J. Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat. Biotechnol. 2008, 26, 225–233. [Google Scholar]
- Cramer, R.A., Jr.; Stajich, J.E.; Yamanaka, Y.; Dietrich, F.S.; Steinbach, W.J.; Perfect, J.R. Phylogenomic analysis of non-ribosomal peptide synthetases in the genus Aspergillus. Gene 2006, 383, 24–32. [Google Scholar] [CrossRef]
- Tobiasen, C.; Aahman, J.; Ravnholt, K.S.; Bjerrum, M.J.; Grell, M.N.; Giese, H. Nonribosomal peptide synthetase (NPS) genes in Fusarium graminearum, F. culmorum and F. pseudograminearium and identification of NPS2 as the producer of ferricrocin. Curr. Genet. 2007, 51, 43–58. [Google Scholar]
- Varga, J.; Rigó, K.; Kocsubé, S.; Farkas, B.; Pál, K. Diversity of polyketide synthase gene sequences in Aspergillus species. Res. Microbiol. 2003, 154, 593–600. [Google Scholar] [CrossRef]
- Shimizu, T.; Kinoshita, H.; Ishihara, S.; Sakai, K.; Nagai, S.; Nihira, T. Polyketide synthase gene responsible for citrinin biosynthesis in Monascus purpureus. Appl. Environ. Microbiol. 2005, 71, 3453–3457. [Google Scholar] [CrossRef]
- Yu, J.; Chang, P.K.; Ehrlich, K.C.; Cary, J.W.; Bhatnagar, D.; Cleveland, T.E.; Payne, G.A.; Linz, J.E.; Woloshuk, C.P.; Bennett, J.W. Clustered pathway genes in aflatoxin biosynthesis. Appl. Environ. Microbiol. 2004, 70, 1253–1262. [Google Scholar] [CrossRef]
- Crawford, J.M.; Vagstad, A.L.; Whitworth, K.P.; Townsend, C.A. Synthetic strategy of nonreducing iterative polyketide synthases and the origin of the classical “Starter Unit Effect”. ChemBioChem 2008, 9, 1019–1023. [Google Scholar]
- Cary, J.W.; Ehrlich, K.C.; Beltz, S.B.; Harris-Coward, P.; Klich, M.A. Characterization of the Aspergillus ochraceoroseus aflatoxin/sterigmatocystin biosynthetic gene cluster. Mycologia 2009, 101, 352–362. [Google Scholar] [CrossRef]
- Bradshaw, R.E.; Jin, H.; Morgan, B.S.; Schwelm, A.; Teddy, O.R.; Young, C.A.; Zhang, S. A polyketide synthase gene required for biosynthesis of the aflatoxin-like toxin, dothistromin. Mycopathologia 2006, 161, 283–294. [Google Scholar] [CrossRef]
- Chang, P.K.; Matsushima, K.; Takahashi, T.; Yu, J.; Abe, K.; Bhatnagar, D.; Yuan, G.F.; Koyama, Y.; Cleveland, T.E. Understanding nonaflatoxigenicity of Aspergillus sojae: A windfall of aflatoxin biosynthesis research. Appl. Microbiol. Biotechnol. 2007, 76, 977–984. [Google Scholar] [CrossRef]
- Tominaga, M.; Lee, Y.H.; Hayashi, R.; Suzuki, Y.; Yamada, O.; Sakamoto, K.; Gotoh, K.; Akita, O. Molecular analysis of an inactive aflatoxin biosynthesis gene cluster in Aspergillus oryzae RIB strains. Appl. Environ. Microbiol. 2006, 72, 484–490. [Google Scholar] [CrossRef]
- Choquer, M.; Dekkers, K.L.; Chen, H.Q.; Cao, L.; Ueng, P.P.; Daub, M.E.; Chung, K.R. The CTB1 gene encoding a fungal polyketide synthase is required for cercosporin biosynthesis and fungal virulence of Cercospora nicotianae. Mol. Plant Microbe Interact. 2005, 18, 468–476. [Google Scholar] [CrossRef]
- Baker, S. Aspergillus Genomics and DHN-melanin Conidial Pigmentation. In Aspergillus in the Genomic Era; Varga, J., Samson, R.A., Eds.; Wageningen Academic Publishers: Wageningen, The Netherlands, 2008; pp. 73–82. [Google Scholar]
- Kim, Y.T.; Lee, Y.R.; Jin, J.; Han, K.H.; Kim, H.; Kim, J.C.; Lee, T.; Yun, S.H.; Lee, Y.W. Two different polyketide synthase genes are required for synthesis of zearalenone in Gibberella zeae. Mol. Microbiol. 2005, 58, 1102–1113. [Google Scholar] [CrossRef]
- Gaffoor, I.; Trail, F. Characterization of two polyketide synthase genes involved in zearalenone biosynthesis in Gibberella zeae. Appl. Environ. Microbiol. 2006, 72, 1793–1799. [Google Scholar] [CrossRef]
- Baker, S.E.; Kroken, S.; Inderbitzin, P.; Asvarak, T.; Li, B.Y.; Shi, L.; Yoder, O.C.; Turgeon, B.G. Two polyketide synthase-encoding genes are required for biosynthesis of the polyketide virulence factor, T-toxin, by Cochliobolus heterostrophus. Mol. Plant Microbe Interact. 2006, 19, 139–149. [Google Scholar] [CrossRef]
- Yun, S.H.; Turgeon, B.G.; Yoder, O.C. REMI-induced mutants of Mycosphaerella zeae-maydis lacking the polyketide PM-toxin are deficient in pathogenesis to corn. Physiol. Mol. Plant P. 1998, 52, 53–66. [Google Scholar] [CrossRef]
- Saha, D.; Fetzner, R.; Burkhardt, B.; Podlech, J.; Metzler, M.; Dang, H.; Lawrence, C.; Fischer, R. Identification of a polyketide synthase required for alternariol (aoh) and alternariol-9-methyl ether (ame) formation in Alternaria alternata. PLoS One 2012, 7, e40564. [Google Scholar]
- Kennedy, J.; Auclair, K.; Kendrew, S.G.; Park, C.; Vederas, J.C.; Hutchinson, C.R. Modulation of polyketide synthase activity by accessory proteins during lovastatin biosynthesis. Science 1999, 284, 1368–1372. [Google Scholar] [CrossRef]
- Xie, X.; Watanabe, K.; Wojcicki, W.A.; Wang, C.C.; Tang, Y. Biosynthesis of lovastatin analogs with a broadly specific acyltransferase. Chem. Biol. 2006, 13, 1161–1169. [Google Scholar] [CrossRef]
- Bartók, T.; Szécsi, A.; Szekeres, A.; Mesterházy, A.; Bartók, M. Detection of new fumonisin mycotoxins and fumonisin-like compounds by reversed-phase high-performance liquid chromatography/electrospray ionization ion trap mass spectrometry. Rapid Commun. Mass Spectrom. 2006, 20, 2447–2462. [Google Scholar] [CrossRef]
- Proctor, R.H.; Desjardins, A.E.; Plattner, R.D.; Hohn, T.M. A polyketide synthase gene required for biosynthesis of fumonisin mycotoxins in Gibberella fujikuroi mating population A. Fungal Genet. Biol. 1999, 27, 100–112. [Google Scholar] [CrossRef]
- Baker, S.E. Aspergillus niger genomics: Past, present and into the future. Med. Mycol. 2006, 44, S17–S21. [Google Scholar] [CrossRef]
- Pel, H.J.; de Winde, J.H.; Archer, D.B.; Dyer, P.S.; Hofmann, G.; Schaap, P.J.; Turner, G.; de Vries, R.P.; Albang, R.; Albermann, K.; et al. Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat. Biotechnol. 2007, 25, 221–231. [Google Scholar]
- Frisvad, J.C.; Smedsgaard, J.; Samson, R.A.; Larsen, T.O.; Thrane, U. Fumonisin B2 production by Aspergillus niger. J. Agric. Food Chem. 2007, 55, 9727–9732. [Google Scholar] [CrossRef]
- Månsson, M.; Klejnstrup, M.L.; Phipps, R.K.; Nielsen, K.F.; Frisvad, J.C.; Gotfredsen, C.H.; Larsen, T.O. Isolation and NMR characterization of fumonisin B2 and a new fumonisin B6 from Aspergillus niger. J. Agric. Food Chem. 2010, 58, 949–953. [Google Scholar] [CrossRef]
- Samson, R.A.; Hoekstra, E.S.; Frisvad, J.C. Introduction to Food- and Airborne Fungi, 7th ed; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2004. [Google Scholar]
- Noonim, P.; Mahakarnchanakul, W.; Nielsen, K.F.; Frisvad, J.C.; Samson, R.A. Fumonisin B2 production by Aspergillus niger in Thai coffee beans. Food Addit. Contam. 2009, 26, 94–100. [Google Scholar] [CrossRef]
- Chang, P.-K.; Ehrlich, K.C.; Fujii, I. Cyclopiazonic acid biosynthesis of Aspergillus flavus and Aspergillus oryzae. Toxins 2009, 1, 74–99. [Google Scholar] [CrossRef]
- Song, Z.; Cox, R.J.; Lazarus, C.M.; Simpson, T.J. Fusarin C biosynthesis in Fusarium moniliforme and Fusarium venenatum. ChemBioChem 2004, 5, 1196–1203. [Google Scholar] [CrossRef]
- Maiya, S.; Grundmann, A.; Li, X.; Li, S.M.; Turner, G. Identification of a hybrid PKS/NRPS required for pseurotin A biosynthesis in the human pathogen Aspergillus fumigatus. ChemBioChem 2007, 8, 1736–1743. [Google Scholar] [CrossRef]
- Schümann, J.; Hertweck, C. Molecular basis of cytochalasan biosynthesis in fungi: Gene cluster analysis and evidence for the involvement of a PKS-NRPS hybrid synthase by RNA silencing. J. Am. Chem. Soc. 2007, 129, 9564–9565. [Google Scholar] [CrossRef]
- Qiao, K.; Chooi, Y.H.; Tang, Y. Identification and engineering of the cytochalasin gene cluster from Aspergillus clavatus NRRL 1. Metab Eng. 2011, 13, 723–732. [Google Scholar] [CrossRef]
- Sims, J.W.; Fillmore, J.P.; Warner, D.D.; Schmidt, E.W. Equisetin biosynthesis in Fusarium heterosporum. Chem. Commun. 2005, 2, 186–188. [Google Scholar]
- White, S.; O'Callaghan, J.; Dobson, A.D. Cloning and molecular characterization of Penicillium expansum genes upregulated under conditions permissive for patulin biosynthesis. FEMS Microbiol. Lett. 2006, 255, 17–26. [Google Scholar] [CrossRef]
- O'Callaghan, J.; Caddick, M.X.; Dobson, A.D. A polyketide synthase gene required for ochratoxin A biosynthesis in Aspergillus ochraceus. Microbiology 2003, 149, 3485–3491. [Google Scholar] [CrossRef]
- Bacha, N.; Atoui, A.; Mathieu, F.; Liboz, T.; Lebrihi, A. Aspergillus westerdijkiae polyketide synthase gene "aoks1" is involved in the biosynthesis of ochratoxin A. Fungal Genet. Biol. 2009, 46, 77–84. [Google Scholar] [CrossRef] [Green Version]
- Karolewiez, A.; Geisen, R. Cloning a part of the ochratoxin A biosynthetic gene cluster of Penicillium nordicum and characterization of the ochratoxin polyketide synthase gene. Syst. Appl. Microbiol. 2005, 28, 588–595. [Google Scholar] [CrossRef]
- Gallo, A.; Perrone, G.; Solfrizzo, M.; Epifani, F.; Abbas, A.; Dobson, A.D.; Mulè, G. Characterisation of a pks gene which is expressed during ochratoxin A production by Aspergillus carbonarius. Int. J. Food Microbiol. 2009, 129, 8–15. [Google Scholar] [CrossRef]
- Proctor, R.H.; Brown, D.W.; Plattner, R.D.; Desjardins, A.E. Co-expression of 15 contiguous genes delineates a fumonisin biosynthetic gene cluster in Gibberella moniliformis. Fungal Genet. Biol. 2003, 38, 237–249. [Google Scholar] [CrossRef]
- Butchko, R.A.; Plattner, R.D.; Proctor, R.H. Deletion analysis of FUM genes involved in tricarballylic ester formation during fumonisin biosynthesis. J. Agric. Food Chem. 2006, 54, 9398–9404. [Google Scholar]
- Zaleta-Rivera, K.; Xu, C.; Yu, F.; Butchko, R.A.; Proctor, R.H.; Hidalgo-Lara, M.E.; Raza, A.; Dussault, P.H.; Du, L. A bidomain nonribosomal peptide synthetase encoded by FUM14 catalyzes the formation of tricarballylic esters in the biosynthesis of fumonisins. Biochemistry 2006, 45, 2561–2569. [Google Scholar] [CrossRef]
- Cramer, R.A., Jr.; Gamcsik, M.P.; Brooking, R.M.; Najvar, L.K.; Kirkpatrick, W.R.; Patterson, T.F.; Balibar, C.J.; Graybill, J.R.; Perfect, J.R.; Abraham, S.N.; et al. Disruption of a nonribosomal peptide synthetase in Aspergillus fumigatus eliminates gliotoxin production. Eukaryot. Cell 2006, 5, 972–980. [Google Scholar] [CrossRef]
- Gardiner, D.M.; Cozijnsen, A.J.; Wilson, L.M.; Pedras, M.S.; Howlett, B.J. The sirodesmin biosynthetic gene cluster of the plant pathogenic fungus Leptosphaeria maculans. Mol. Microbiol. 2004, 53, 1307–1318. [Google Scholar] [CrossRef]
- Wenzel, S.C.; Meiser, P.; Binz, T.M.; Mahmud, T.; Müller, R. Nonribosomal peptide biosynthesis: Point mutations and module skipping lead to chemical diversity. Angew. Chem. Int. Ed. Engl. 2006, 45, 2296–2301. [Google Scholar] [CrossRef]
- Glinski, M.; Urbanke, C.; Hornbogen, T.; Zocher, R. Enniatin synthetase is a monomer with extended structure: Evidence for an intramolecular reaction mechanism. Arch. Microbiol. 2002, 178, 267–273. [Google Scholar] [CrossRef]
- Scott-Craig, J.S.; Panaccione, D.G.; Pocard, J.A.; Walton, J.D. The cyclic peptide synthetase catalyzing HC-toxin production in the filamentous fungus Cochliobolus carbonum is encoded by a 15.7-kilobase open reading frame. J. Biol. Chem. 1992, 267, 26044–26049. [Google Scholar]
- Gallo, A.; Bruno, K.S.; Solfrizzo, M.; Perrone, G.; Mulè, G.; Visconti, A.; Baker, S.E. New insight into the ochratoxin A biosynthetic pathway through deletion of a nonribosomal peptide synthetase gene in Aspergillus carbonarius. Appl. Environ. Microbiol. 2012, 78, 8208–8218. [Google Scholar]
- Correia, T.; Grammel, N.; Ortel, I.; Keller, U.; Tudzynski, P. Molecular cloning and analysis of the ergopeptine assembly system in the ergot fungus Claviceps purpurea. Chem. Biol. 2003, 10, 1281–1292. [Google Scholar] [CrossRef]
- Haarmann, T.; Machado, C.; Lübbe, Y.; Correia, T.; Schardl, C.L.; Panaccione, D.G.; Tudzynski, P. The ergot alkaloid gene cluster in Claviceps purpurea: Extension of the cluster sequence and intra species evolution. Phytochemistry 2005, 66, 1312–1320. [Google Scholar] [CrossRef]
- Aharonowitz, Y.; Cohen, G.; Martin, J.F. Penicillin and cephalosporin biosynthetic genes: Structure, organization, regulation, and evolution. Annu. Rev. Microbiol. 1992, 46, 461–495. [Google Scholar] [CrossRef]
- Brakhage, A.A.; Al-Abdallah, Q.; Tüncher, A.; Spröte, P. Evolution of beta-lactam biosynthesis genes and recruitment of trans-acting factors. Phytochemistry 2005, 66, 1200–1210. [Google Scholar] [CrossRef]
- Liras, P.; Martín, J.F. Gene clusters for beta-lactam antibiotics and control of their expression: Why have clusters evolved, and from where did they originate? Int. Microbiol. 2006, 9, 9–19. [Google Scholar]
- Haas, H.; Eisendle, M.; Turgeon, B.G. Siderophores in fungal physiology and virulence. Annu. Rev. Phytopathol. 2008, 46, 149–187. [Google Scholar] [CrossRef]
- Bateman, A.; Coin, L.; Durbin, R.; Finn, R.D.; Hollich, V.; Griffiths-Jones, S.; Khanna, A.; Marshall, M.; Moxon, S.; Sonnhammer, E.L.; Studholme, D.J.; Yeats, C.; Eddy, S.R. The Pfam protein families database. Nucleic Acids Res. 2004, 32, D138–D141. [Google Scholar] [CrossRef]
- Haft, D.H.; Selengut, J.D.; White, O. The TIGRFAMs database of protein families. Nucleic Acids Res. 2003, 31, 371–373. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Stamatakis, A.; Hoover, P.; Rougemont, J. A rapid bootstrap algorithm for the RAxML Web servers. Syst. Biol. 2008, 57, 758–771. [Google Scholar] [CrossRef]
- Dereeper, A.; Guignon, V.; Blanc, G.; Audic, S.; Buffet, S.; Chevenet, F.; Dufayard, J.F.; Guindon, S.; Lefort, V.; Lescot, M.; et al. Phylogeny.ft: Robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008, 36, W465–W469. [Google Scholar] [CrossRef] [Green Version]
- Perrière, G.; Gouy, M. WWW-Query: An on-line retrieval system for biological sequence banks. Biochemie 1996, 78, 364–369. [Google Scholar] [CrossRef]
- Marcet-Houben, M.; Gabaldón, T. Acquisition of prokaryotic genes by fungal genomes. Trends Genet. 2010, 26, 5–8. [Google Scholar] [CrossRef]
- Rolland, T.; Neuvéglise, C.; Sacerdot, C.; Dujon, B. Insertion of horizontally transferred genes within conserved syntenic regions of yeast genomes. PLoS One 2009, 4, e6515. [Google Scholar]
- Slot, J.C.; Rokas, A. Horizontal transfer of a large and highly toxic secondary metabolic gene cluster between fungi. Curr. Biol. 2011, 21, 134–139. [Google Scholar] [CrossRef]
- Geisen, R.; Schmidt-Heydt, M. Physiological and Molecular Aspects of Ochratoxin A Biosynthesis. In Physiology and Genetics. The Mycota XV; Anke, T., Weber, D., Eds.; Springer-Verlag: Heidelberg, Germany, 2009; pp. 353–376. [Google Scholar]
- Rausch, C.; Hoof, I.; Weber, T.; Wohlleben, W.; Huson, D.H. Phylogenetic analysis of condensation domains in NRPS sheds light on their functional evolution. BMC Evol. Biol. 2007, 7, 78. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gallo, A.; Ferrara, M.; Perrone, G. Phylogenetic Study of Polyketide Synthases and Nonribosomal Peptide Synthetases Involved in the Biosynthesis of Mycotoxins. Toxins 2013, 5, 717-742. https://doi.org/10.3390/toxins5040717
Gallo A, Ferrara M, Perrone G. Phylogenetic Study of Polyketide Synthases and Nonribosomal Peptide Synthetases Involved in the Biosynthesis of Mycotoxins. Toxins. 2013; 5(4):717-742. https://doi.org/10.3390/toxins5040717
Chicago/Turabian StyleGallo, Antonia, Massimo Ferrara, and Giancarlo Perrone. 2013. "Phylogenetic Study of Polyketide Synthases and Nonribosomal Peptide Synthetases Involved in the Biosynthesis of Mycotoxins" Toxins 5, no. 4: 717-742. https://doi.org/10.3390/toxins5040717
APA StyleGallo, A., Ferrara, M., & Perrone, G. (2013). Phylogenetic Study of Polyketide Synthases and Nonribosomal Peptide Synthetases Involved in the Biosynthesis of Mycotoxins. Toxins, 5(4), 717-742. https://doi.org/10.3390/toxins5040717







