Practical Dynamic Contrast Enhanced MRI in Small Animal Models of Cancer: Data Acquisition, Data Analysis, and Interpretation
Abstract
:1. Background and Motivation
2. Basic Theory of DCE-MRI
2.1. Calibrating the Concentration of Contrast Agent to Measured MRI Parameters
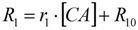
2.2. Classes of DCE-MRI Methods
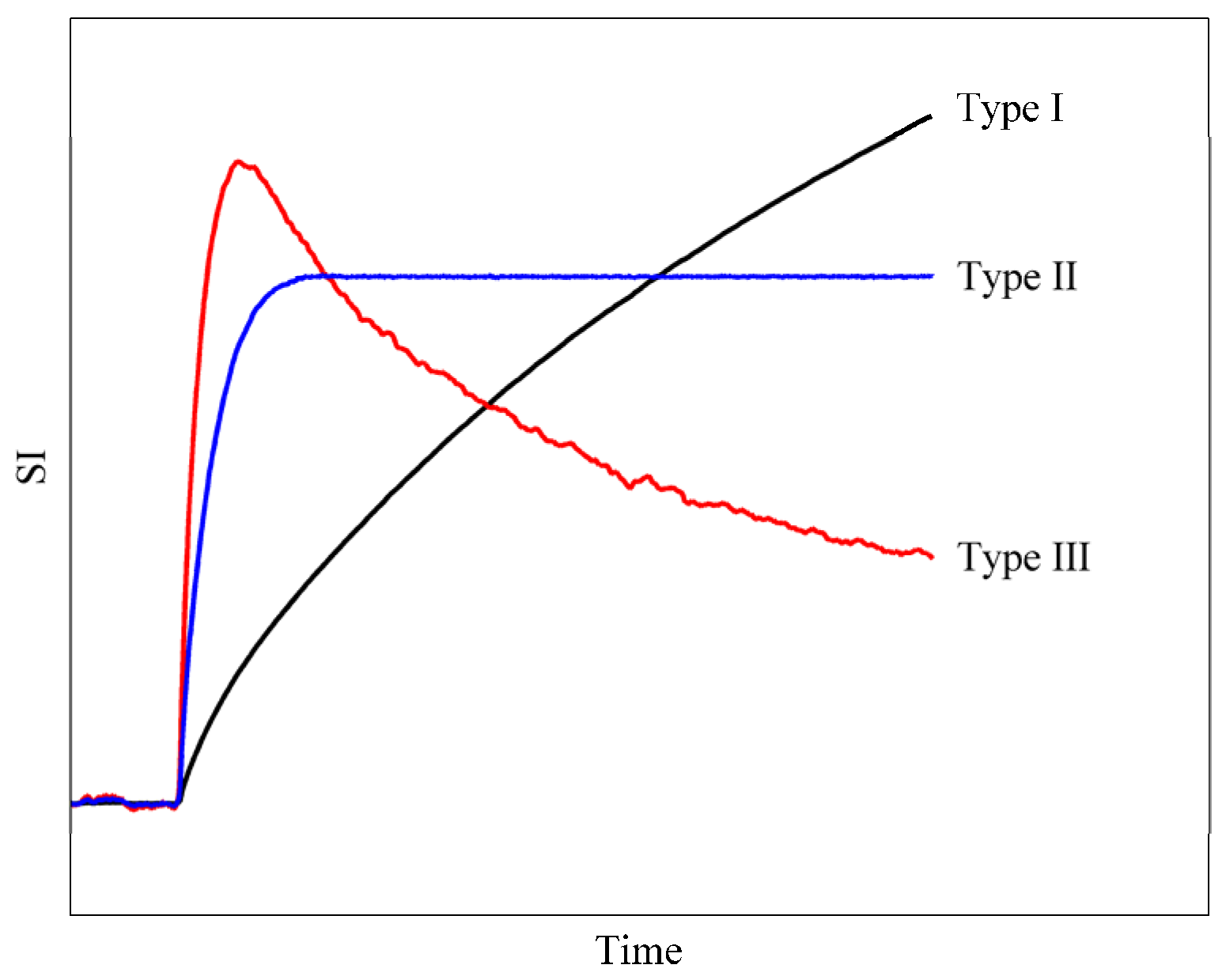
2.2.1. Qualitative Methods
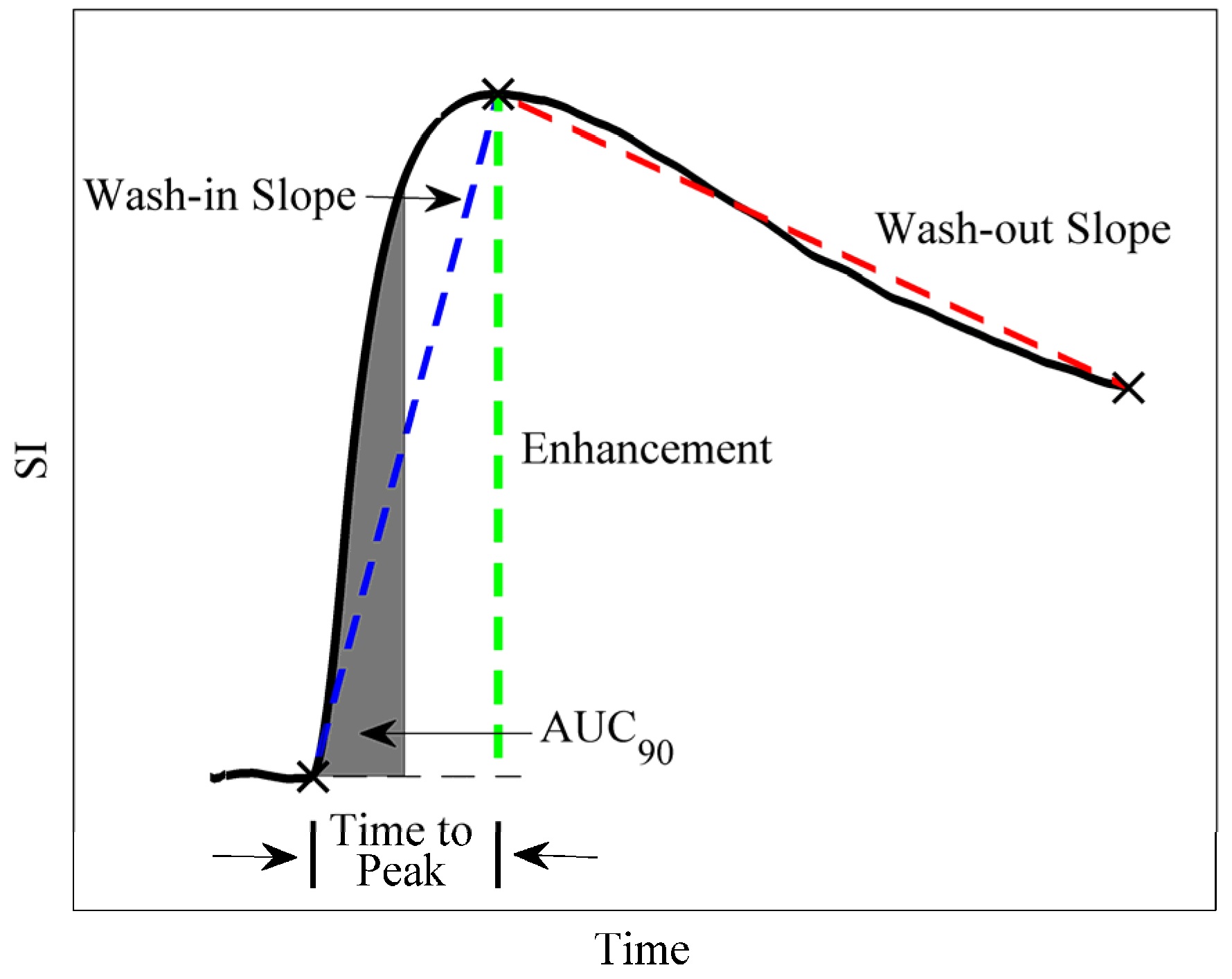
2.2.2. Semi-Quantitative Methods
2.2.3. Quantitative Methods
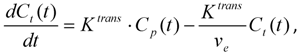
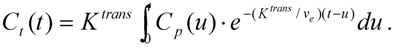
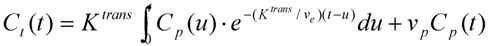
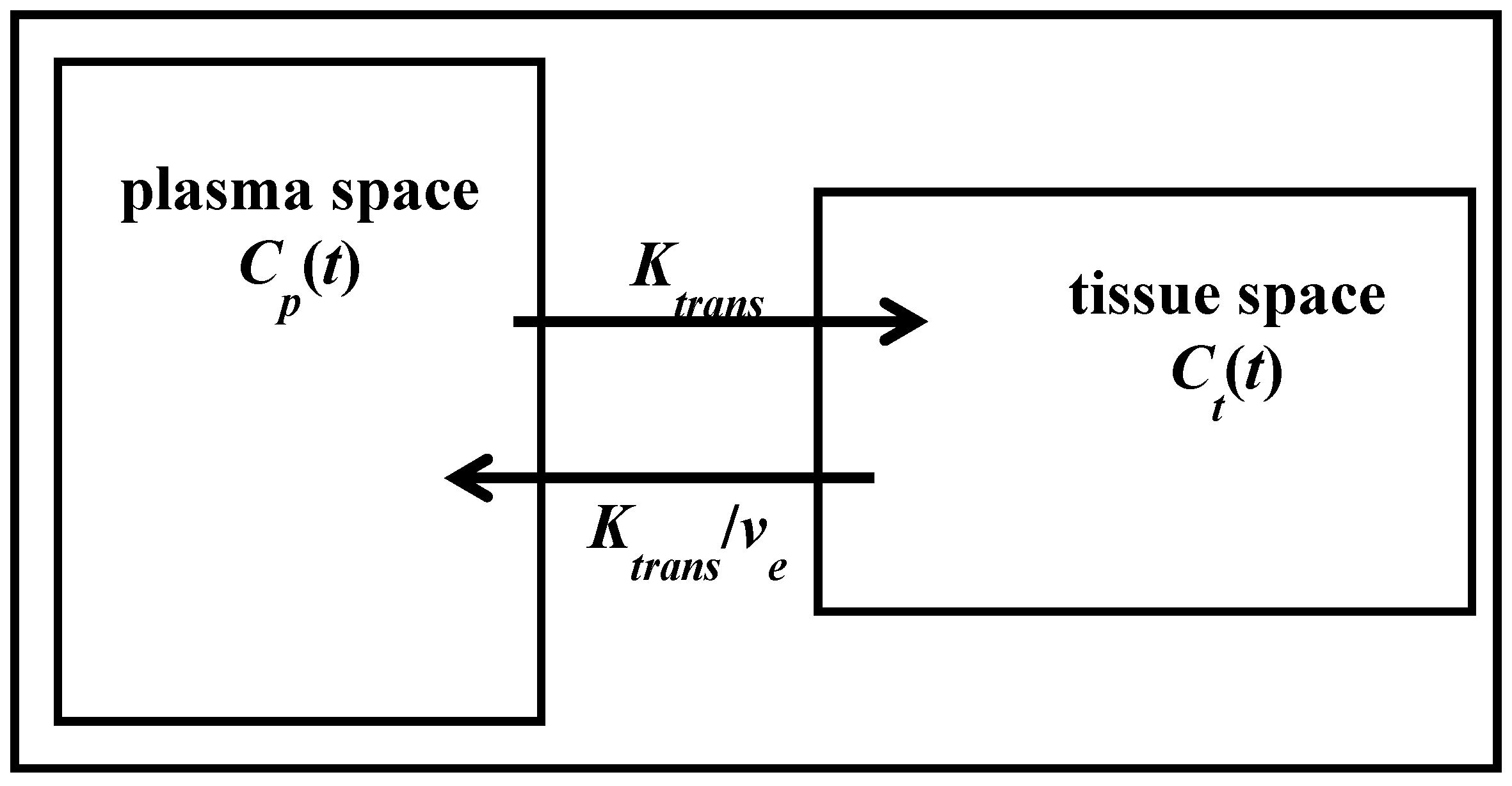
2.3. Pre-Contrast T1
2.4. Requirement of Fast T1 Imaging
| Author, year [reference] | Spatial resolution | Temporal resolution | Description |
|---|---|---|---|
| Loveless, 2012 [89] | in-plane = 0.27 mm2; matrix = 1282; slice thickness = 1 mm | temporal resolution = 25.6 s; TR/TE/α = 100 ms/2.82 ms/25° | Used a population average AIF since assessing heterogeneity from whole tumor volume was a priority. Study sacrificed temporal resolution for high spatial resolution. |
| Benjaminsen, 2004 [90] | in-plane = 0.5 mm × 0.2 mm; matrix = 256 × 128; slice thickness = 2 mm | temporal resolution = 27 s; TR/TE/α = 200 ms/3.6 ms/80° | Used blood sampling to determine AIF, and sacrificed temporal resolution for whole tumor volume coverage. Also used a population average AIF from the left ventricle of additional animals with different scan parameters to achieve a faster temporal resolution. |
| Kim, H 2011 [91] | in-plane = 0.23 mm2; matrix = 1282; slice thickness = 1 mm | temporal resolution = 58.8 s; TR/TE/α = 115 ms/3 ms/30° | Used a reference region analysis since spatial resolution and whole tumor volume coverage was a priority. Study sacrificed temporal resolution for high spatial resolution. |
| Li, 2010 [92] | in-plane = 0.35 mm2; matrix = 128 × 64; slice thickness = 1 mm | temporal resolution = 1.6 s; TR/TE/α = 25 ms/1.4 ms/20° | Used a fast gradient echo sequence to achieve high temporal resolution in order to collect individual AIFs from image data. Study sacrificed whole tumor volume coverage by only collecting three slices. |
| Skinner, 2012 [93] | in-plane = 0.25 mm2; matrix = 1282; slice thickness = 2 mm | temporal resolution = 1.9 s; TR/TE/α = 10 ms/2.1 ms/15° | Used individual AIFs for kinetic modeling. Study sacrificed whole tumor coverage (only collected central tumor slice) to achieve high temporal resolution. |
| Kim, J 2012 [94] | in-plane = 0.23 mm2; matrix = 1282; slice thickness = 2.5 mm | temporal resolution = 6.4 s; TR/TE/α = 67 ms/3 ms/70° | Used fast imaging sequence to achieve temporal resolution, however study sacrificed through-plane spatial resolution (2.5 mm) for whole tumor volume coverage. AIF was collected from image data for kinetic modeling, although 6.4 s might be too long to adequately sample the peak of the CA concentration curve. |
2.5. AIF

3. Specific Considerations for Small Animal Imaging
3.1. AIF Measurement
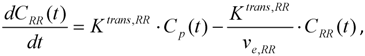
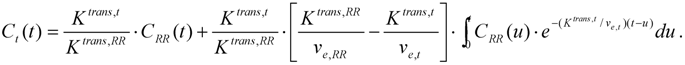
3.2. Reproducibility/Repeatability
| Author, year [reference] | Subject | Tissue | Parameter | 95% CI | Repeatability Index |
|---|---|---|---|---|---|
| Galbraith, 2002 [70] | Human | Tumor Muscle | Ktrans | (−16%)–(+19%) | 0.32 mL(blood)/mL(tissue)/min |
| kep | ±16% | 0.91 mL(blood)/mL(tissue)/min | |||
| ve | ±6% | 7.62 mL/mL | |||
| Ktrans | (−30%)–(+44%) | 0.61 mL(blood)/mL(tissue)/min | |||
| kep | ±61% | 1.28 mL(blood)/mL(tissue)/min | |||
| ve | ±13% | 5.71 mL/mL | |||
| Yankeelov, 2006 [109] | Mouse | Tumor Muscle | Ktrans | * | 0.222 mL(blood)/mL(tissue)/min |
| ve | 0.204 mL(blood)/mL(tissue)/min | ||||
| Ktrans | 0.197 mL/mL | ||||
| Barnes, 2012 [113] | Mouse | Tumor | Ktrans | ±14% | 0.073 mL(blood)/mL(tissue)/min |
| ve | ±8% | 0.113 mL/mL | |||
| Ktrans | ±21% | 0.075 mL(blood)/mL(tissue)/min | |||
| ve | ±5% | 0.069 mL/mL | |||
| vp | ±15% | 0.014 |
3.3. Animal Care and Monitoring
3.4. Planning a Small Animal DCE-MRI Study
4. Methods of Validating DCE-MRI Analyses
4.1. Histology
| Author, Year (reference) | Tissue of Interest | Histology Technique | MRI Parameter | MVD Correlation ( r2) | Statistical Significance( p value) |
|---|---|---|---|---|---|
| Cheng, 2007 [117] | Bladder tissue constructs | IHC-CD31 | AUC (60 s) | 0.784 | 0.003 |
| Ktrans | 0.4 | NS | |||
| vp | 0.696 | 0.012 | |||
| Ren, 2008 [118] | Prostate | IHC-CD34 | time to peak, Tm | −0.71 | <0.007 |
| Percent enhancement, % SI | 0.557 | <0.007 | |||
| Enhancement rate ( R = % SI/Tm) | 0.747 | <0.007 | |||
| Hulka, 1997 [119] | Breast cancer | IHC-factor VIII-related antigen | E∙F | 0.36 | <0.01 |
| Yao, 2008 [120] | Rectal cancer | IHC-CD34 | Ktrans | 0.495 | 0.026 |
| Haris, 2008 [121] | Brain tuberculomas | IHC-CD34 | Ktrans | 0.231 | 0.076 |
| Orth, 2007 [122] | Breast cancer xenografts | IHC-CD31 | Ktrans (Gadomer-17) | 0.13 | 0.659 |
| vb (Gadomer-17) | −0.081 | 0.782 | |||
| Ktrans (Magnevist) | 0.045 | 0.874 | |||
| vb (Magnevist) | −0.15 | 0.594 | |||
| Reitan, 2010 [123] | Osteosarcoma xenografts | Fluorescently-labeled Dextran | Ktrans | 0.93 | 0.04 |
4.2. Dynamic Contrast Enhanced Computed Tomography
4.3. Other Modalities
5. Relationships to Other Imaging Modalities
5.1. Possible Relationships between DCE-MRI and DW-MRI
5.2. Possible Relationships between DCE-MRI and Common PET Tracers
5.2.1. PET Imaging of Hypoxia
5.2.2. PET Imaging of Glycolysis
5.2.3. PET Imaging of Cell Proliferation
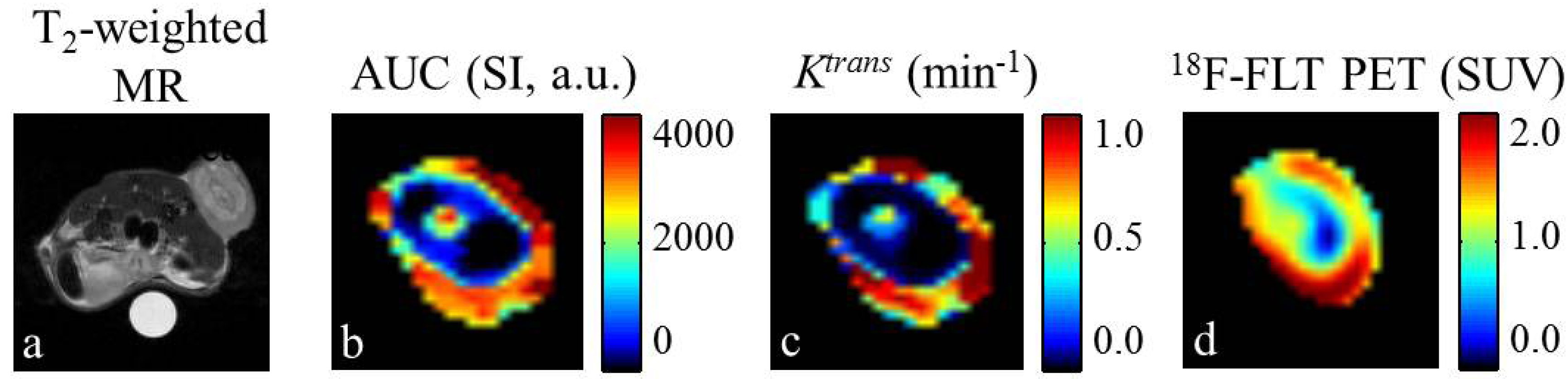
6. Limitations
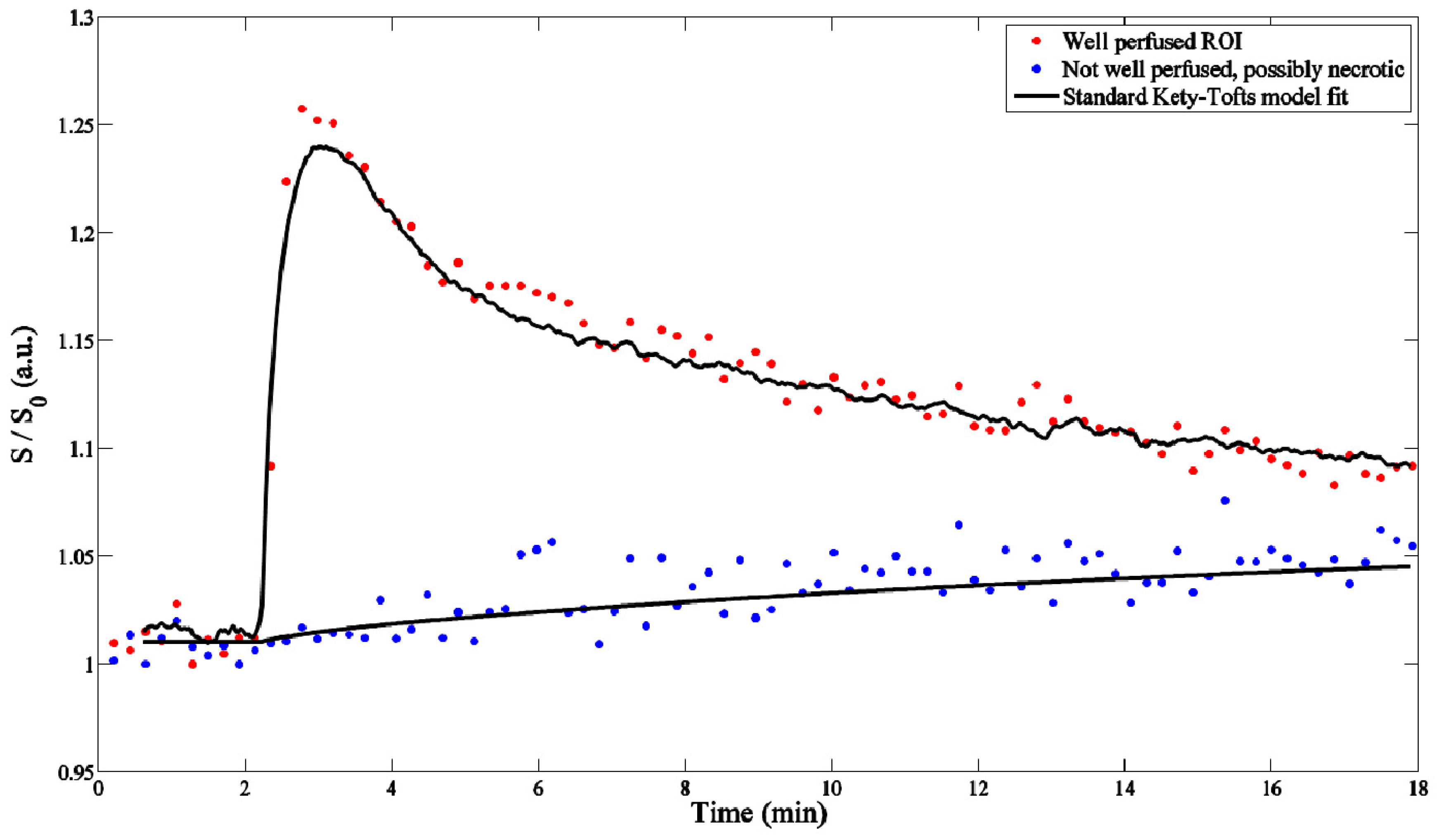
7. Summary
Acknowledgments
References
- Weis, S.M.; Cheresh, D.A. Tumor angiogenesis: Molecular pathways and therapeutic targets. Nat. Med. 2011, 17, 1359–1370. [Google Scholar]
- Ferrara, N.; Gerber, H.; LeCouter, J. The biology of vegf and its receptors. Nat. Med. 2003, 9, 669–676. [Google Scholar] [CrossRef]
- Young, R.; Reed, M. Anti-angiogenic therapy: Concept to clinic. Microcirculation 2012, 19, 115–125. [Google Scholar] [CrossRef]
- Vredenburgh, J.J.; Desjardins, A.; Herndon, J.E.; Dowell, J.M.; Reardon, D.A.; Quinn, J.A.; Rich, J.N.; Sathornsumetee, S.; Gururangan, S.; Wagner, M.; et al. Phase ii trial of bevacizumab and irinotecan in recurrent malignant glioma. Clin.Cancer Res. 2007, 13, 1253–1259. [Google Scholar] [CrossRef]
- Hurwitz, H.; Fehrenbacher, L.; Novotny, W.; Cartwright, T.; Hainsworth, J.; Heim, W.; Berlin, J.; Baron, A.; Griffing, S.; Holmgren, E.; et al. Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N. Engl. J. Med. 2004, 350, 2335–2342. [Google Scholar] [CrossRef]
- Miller, K.; Wang, M.; Gralow, J.; Dickler, M.; Cobleigh, M.; Perez, E.A.; Shenkier, T.; Cella, D.; Davidson, N.E. Paclitaxel plus bevacizumab versus paclitaxel alone for metastatic breast cancer. N. Engl. J. Med. 2007, 357, 2666–2676. [Google Scholar] [CrossRef]
- Saltz, L.B.; Clarke, S.; Diaz-Rubio, E.; Scheithauer, W.; Figer, A.; Wong, R.; Koski, S.; Lichinitser, M.; Yang, T.S.; Rivera, F.; et al. Bevacizumab in combination with oxaliplatin-based chemotherapy as first-line therapy in metastatic colorectal cancer: A randomized phase iii study. J. Clin. Oncol. 2008, 26, 2013–2019. [Google Scholar]
- Friedman, H.S.; Prados, M.D.; Wen, P.Y.; Mikkelsen, T.; Schiff, D.; Abrey, L.E.; Yung, W.K.; Paleologos, N.; Nicholas, M.K.; Jensen, R.; et al. Bevacizumab alone and in combination with irinotecan in recurrent glioblastoma. J. Clin. Oncol. 2009, 27, 4733–4740. [Google Scholar]
- Miles, D.W.; Chan, A.; Dirix, L.Y.; Cortes, J.; Pivot, X.; Tomczak, P.; Delozier, T.; Sohn, J.H.; Provencher, L.; Puglisi, F.; et al. Phase iii study of bevacizumab plus docetaxel compared with placebo plus docetaxel for the first-line treatment of human epidermal growth factor receptor 2-negative metastatic breast cancer. J. Clin. Oncol. 2010, 28, 3239–3247. [Google Scholar]
- Robert, N.J.; Dieras, V.; Glaspy, J.; Brufsky, A.M.; Bondarenko, I.; Lipatov, O.N.; Perez, E.A.; Yardley, D.A.; Chan, S.Y.; Zhou, X.; et al. Ribbon-1: Randomized, double-blind, placebo-controlled, phase iii trial of chemotherapy with or without bevacizumab for first-line treatment of human epidermal growth factor receptor 2-negative, locally recurrent or metastatic breast cancer. J. Clin. Oncol. 2011, 29, 1252–1260. [Google Scholar]
- Demetri, G.D.; van Oosterom, A.T.; Garrett, C.R.; Blackstein, M.E.; Shah, M.H.; Verweij, J.; McArthur, G.; Judson, I.R.; Heinrich, M.C.; Morgan, J.A.; et al. Efficacy and safety of sunitinib in patients with advanced gastrointestinal stromal tumour after failure of imatinib: A randomised controlled trial. Lancet 2006, 368, 1329–1338. [Google Scholar]
- Escudier, B.; Eisen, T.; Stadler, W.M.; Szczylik, C.; Oudard, S.; Siebels, M.; Negrier, S.; Chevreau, C.; Solska, E.; Desai, A.A.; et al. Sorafenib in advanced clear-cell renal-cell carcinoma. N. Engl. J. Med. 2007, 356, 125–134. [Google Scholar] [CrossRef]
- Llovet, J.M.; Ricci, S.; Mazzaferro, V.; Hilgard, P.; Gane, E.; Blanc, J.F.; de Oliveira, A.C.; Santoro, A.; Raoul, J.L.; Forner, A.; et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 2008, 359, 378–390. [Google Scholar]
- Motzer, R.J.; Hutson, T.E.; Tomczak, P.; Michaelson, M.D.; Bukowski, R.M.; Oudard, S.; Negrier, S.; Szczylik, C.; Pili, R.; Bjarnason, G.A.; et al. Overall survival and updated results for sunitinib compared with interferon alfa in patients with metastatic renal cell carcinoma. J. Clin. Oncol. 2009, 27, 3584–3590. [Google Scholar]
- Motzer, R.J.; Hutson, T.E.; Tomczak, P.; Michaelson, M.D.; Bukowski, R.M.; Rixe, O.; Oudard, S.; Negrier, S.; Szczylik, C.; Kim, S.T.; et al. Sunitinib versus interferon alfa in metastatic renal-cell carcinoma. N. Engl. J. Med. 2007, 356, 115–124. [Google Scholar]
- Tofts, P.S.; Brix, G.; Buckley, D.L.; Evelhoch, J.L.; Henderson, E.; Knopp, M.V.; Larsson, H.B.W.; Lee, T.Y.; Mayr, N.A.; Parker, G.J.M.; et al. Estimating kinetic parameters from dynamic contrast-enhanced t1-weighted mri of a diffusable tracer: Standardized quantities and symbols. J. Magn. Reson. Imag. 1999, 10, 223–232. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; Gore, J.C. Dynamic contrast enhanced magnetic resonance imaging in oncology: Theory, data, acquisition, analysis, and examples. Curr. Med. Imag. Rev. 2007, 3, 91–107. [Google Scholar] [CrossRef]
- Lee, L.; Sharma, S.; Morgan, B.; Allegrini, P.; Schnell, C.; Brueggen, J.; Cozens, R.; Horsfield, M.; Guenther, C.; Steward, W.; et al. Biomarkers for assessment of pharmacologic activity for a vascular endothelial growth factor (vegf) receptor inhibitor, ptk787/zk 222584 (ptk/zk): Translation of biological activity in a mouse melanoma metastasis model to phase i studies in patients with advanced colorectal cancer with liver metastases. Cancer Chemother. Pharmacol. 2006, 57, 761–771. [Google Scholar] [CrossRef]
- Beauregard, D.A.; Thelwall, P.E.; Chaplin, D.J.; Hill, S.A.; Adams, G.E.; Brindle, K.M. Magnetic resonance imaging and spectroscopy of combretastatin a4 prodrug-induced disruption of tumour perfusion and energetic status. Br. J. Cancer. 1998, 77, 1761–1767. [Google Scholar] [CrossRef]
- Maxwell, R.J.; Wilson, J.; Prise, V.E.; Vojnovic, B.; Rustin, G.J.; Lodge, M.A.; Tozer, G.M. Evaluation of the anti-vascular effects of combretastatin in rodent tumours by dynamic contrast enhanced mri. NMR Biomed. 2002, 15, 89–98. [Google Scholar] [CrossRef]
- Robinson, S.P.; McIntyre, D.J.O.; Checkley, D.; Tessier, J.J.; Howe, F.A.; Griffiths, J.R.; Ashton, S.E.; Ryan, A.J.; Blakey, D.C.; Waterton, J.C. Tumour dose response to the antivascular agent zd6126 assessed by magnetic resonance imaging. Br. J. Canc. 2003, 88, 1592–1597. [Google Scholar] [CrossRef]
- Chang, Y.C.; Yu, C.J.; Chen, C.M.; Hu, F.C.; Hsu, H.H.; Tseng, W.Y.I.; Ting-Fang Shih, T.; Yang, P.C.; Chih-Hsin Yang, J. Dynamic contrast-enhanced mri in advanced nonsmall-cell lung cancer patients treated with first-line bevacizumab, gemcitabine, and cisplatin. J. Magn. Reson. Imag. 2012, 36, 387–396. [Google Scholar] [CrossRef]
- Hoff, B.A.; Bhojani, M.S.; Rudge, J.; Chenevert, T.L.; Meyer, C.R.; Galbán, S.; Johnson, T.D.; Leopold, J.S.; Rehemtulla, A.; Ross, B.D.; et al. Dce and dw-mri monitoring of vascular disruption following vegf-trap treatment of a rat glioma model. NMR Biomed. 2012, 25, 935–942. [Google Scholar] [CrossRef]
- Hsu, C.Y.; Shen, Y.C.; Yu, C.W.; Hsu, C.; Hu, F.C.; Hsu, C.H.; Chen, B.B.; Wei, S.Y.; Cheng, A.L.; Shih, T.T.F. Dynamic contrast-enhanced magnetic resonance imaging biomarkers predict survival and response in hepatocellular carcinoma patients treated with sorafenib and metronomic tegafur/uracil. J. Hepatol. 2011, 55, 858–865. [Google Scholar] [CrossRef]
- Yopp, A.; Schwartz, L.; Kemeny, N.; Gultekin, D.; Gönen, M.; Bamboat, Z.; Shia, J.; Haviland, D.; D’Angelica, M.; Fong, Y.; et al. Antiangiogenic therapy for primary liver cancer: Correlation of changes in dynamic contrast-enhanced magnetic resonance imaging with tissue hypoxia markers and clinical response. Ann. Surg. Oncol. 2011, 18, 2192–2199. [Google Scholar] [CrossRef]
- Lemasson, B.; Christen, T.; Tizon, X.; Farion, R.; Fondraz, N.; Provent, P.; Segebarth, C.; Barbier, E.L.; Genne, P.; Duchamp, O.; et al. Assessment of multiparametric mri in a human glioma model to monitor cytotoxic and anti-angiogenic drug effects. NMR Biomed. 2011, 24, 473–482. [Google Scholar] [CrossRef]
- Heiss, W.D.; Raab, P.; Lanfermann, H. Multimodality assessment of brain tumors and tumor recurrence. J. Nucl. Med. 2011, 52, 1585–1600. [Google Scholar] [CrossRef]
- Jacobs, M.A.; Ouwerkerk, R.; Wolff, A.C.; Gabrielson, E.; Warzecha, H.; Jeter, S.; Bluemke, D.A.; Wahl, R.; Stearns, V. Monitoring of neoadjuvant chemotherapy using multiparametric, (2)(3)na sodium mr, and multimodality (pet/ct/mri) imaging in locally advanced breast cancer. Breast. Cancer. Res. Treat. 2011, 128, 119–126. [Google Scholar] [CrossRef]
- De Bruyne, S.; van Damme, N.; Smeets, P.; Ferdinande, L.; Ceelen, W.; Mertens, J.; van de Wiele, C.; Troisi, R.; Libbrecht, L.; Laurent, S.; et al. Value of dce-mri and fdg-pet/ct in the prediction of response to preoperative chemotherapy with bevacizumab for colorectal liver metastases. Br. J. Canc. 2012, 106, 1926–1933. [Google Scholar] [CrossRef]
- Viel, T.; Talasila, K.M.; Monfared, P.; Wang, J.; Jikeli, J.F.; Waerzeggers, Y.; Neumaier, B.; Backes, H.; Brekka, N.; Thorsen, F.; et al. Analysis of the growth dynamics of angiogenesis-dependent and angiogenesis-independent experimental glioblastomas by multimodal small-animal pet and mri. J. Nucl. Med. 2012, 53, 1135–1145. [Google Scholar] [CrossRef]
- Jansen, J.F.A.; Schöder, H.; Lee, N.Y.; Stambuk, H.E.; Wang, Y.; Fury, M.G.; Patel, S.G.; Pfister, D.G.; Shah, J.P.; Koutcher, J.A.; et al. Tumor metabolism and perfusion in head and neck squamous cell carcinoma: Pretreatment multimodality imaging with 1 h magnetic resonance spectroscopy, dynamic contrast-enhanced mri, and [18f] fdg-pet. Int.J. Radiat. Oncol. Biol. Phys. 2012, 82, 299–307. [Google Scholar] [CrossRef]
- Calamante, F. Perfusion mri using dynamic-susceptibility contrast mri: Quantification issues in patient studies. Top Magn. Reson. Imag. 2010, 21, 75–85. [Google Scholar] [CrossRef]
- Stanisz, G.J.; Henkelman, R.M. Gd-dtpa relaxivity depends on macromolecular content. Magn. Reson. Med. 2000, 44, 665–667. [Google Scholar] [CrossRef]
- Landis, C.S.; Li, X.; Telang, F.W.; Molina, P.E.; Palyka, I.; Vetek, G.; Springer, C.S. Equilibrium transcytolemmal water-exchange kinetics in skeletal muscle in vivo. Magn. Reson. Med. 1999, 42, 467–478. [Google Scholar] [CrossRef]
- Li, X.; Priest, R.A.; Woodward, W.J.; Tagge, I.J.; Siddiqui, F.; Huang, W.; Rooney, W.D.; Beer, T.M.; Garzotto, M.G.; Springer, C.S., Jr. Feasibility of shutter-speed dce-mri for improved prostate cancer detection. Magn. Reson. Med. 2012, 27, 24211. [Google Scholar]
- Li, X.; Priest, R.A.; Woodward, W.J.; Siddiqui, F.; Beer, T.M.; Garzotto, M.G.; Rooney, W.D.; Springer, C.S., Jr. Cell membrane water exchange effects in prostate dce-mri. J. Magn. Reson. 2012, 218, 77–85. [Google Scholar]
- Bains, L.J.; McGrath, D.M.; Naish, J.H.; Cheung, S.; Watson, Y.; Taylor, M.B.; Logue, J.P.; Parker, G.J.; Waterton, J.C.; Buckley, D.L. Tracer kinetic analysis of dynamic contrast-enhanced mri and ct bladder cancer data: A preliminary comparison to assess the magnitude of water exchange effects. Magn. Reson. Med. 2010, 64, 595–603. [Google Scholar]
- Buckley, D.L.; Kershaw, L.E.; Stanisz, G.J. Cellular-interstitial water exchange and its effect on the determination of contrast agent concentration in vivo: Dynamic contrast-enhanced mri of human internal obturator muscle. Magn. Reson. Med. 2008, 60, 1011–1019. [Google Scholar] [CrossRef]
- Caravan, P. Strategies for increasing the sensitivity of gadolinium based mri contrast agents. Chem. Soc. Rev. 2006, 35, 512–523. [Google Scholar] [CrossRef]
- Weinmann, H.J.; Brasch, R.C.; Press, W.R.; Wesbey, G.E. Characteristics of gadolinium-dtpa complex: A potential nmr contrast agent. AJR Am. J. Roentgenol. 1984, 142, 619–624. [Google Scholar]
- Kaewlai, R.; Abujudeh, H. Nephrogenic systemic fibrosis. Am. J. Roentgenol. 2012, 199, W17–W23. [Google Scholar] [CrossRef]
- Wermuth, P.J.; Jimenez, S.A. Gadolinium compounds signaling through tlr 4 and tlr 7 in normal human macrophages: Establishment of a proinflammatory phenotype and implications for the pathogenesis of nephrogenic systemic fibrosis. J. Immunol. 2012, 189, 318–327. [Google Scholar] [CrossRef]
- Wiesinger, B.; Kehlbach, R.; Hemsen, J.; Bebin, J.; Bantleon, R.; Schwenzer, N.; Spira, D.; Claussen, C.D.; Wiskirchen, J. Effects of magnetic resonance imaging contrast agents on human umbilical vein endothelial cells and evaluation of magnetic resonance imaging contrast media-triggered transforming growth factor-beta induction in dermal fibroblasts (hsf) as a model for nephrogenic systemic fibrosis. Invest. Radiol. 2011, 46, 71–76. [Google Scholar] [CrossRef]
- Del Galdo, F.; Wermuth, P.J.; Addya, S.; Fortina, P.; Jimenez, S.A. Nfkappab activation and stimulation of chemokine production in normal human macrophages by the gadolinium-based magnetic resonance contrast agent omniscan: Possible role in the pathogenesis of nephrogenic systemic fibrosis. Ann. Rheum. Dis. 2010, 69, 2024–2033. [Google Scholar] [CrossRef]
- Steger-Hartmann, T.; Raschke, M.; Riefke, B.; Pietsch, H.; Sieber, M.A.; Walter, J. The involvement of pro-inflammatory cytokines in nephrogenic systemic fibrosis-A mechanistic hypothesis based on preclinical results from a rat model treated with gadodiamide. Exp. Toxicol. Pathol. 2009, 61, 537–552. [Google Scholar] [CrossRef]
- Sieber, M.A.; Pietsch, H.; Walter, J.; Haider, W.; Frenzel, T.; Weinmann, H.J. A preclinical study to investigate the development of nephrogenic systemic fibrosis: A possible role for gadolinium-based contrast media. Invest. Radiol. 2008, 43, 65–75. [Google Scholar] [CrossRef]
- Rohrer, M.; Bauer, H.; Mintorovitch, J.; Requardt, M.; Weinmann, H.J. Comparison of magnetic properties of mri contrast media solutions at different magnetic field strengths. Invest. Radiol. 2005, 40, 715–724. [Google Scholar] [CrossRef]
- Yang, C.T.; Chuang, K.H. Gd(iii) chelates for mri contrast agents: From high relaxivity to "smart", from blood pool to blood-brain barrier permeable. Med.Chem.Comm. 2012, 3, 552–565. [Google Scholar] [CrossRef]
- Pathak, A.P.; Penet, M.F.; Bhujwalla, Z.M. MR molecular imaging of tumor vasculature and vascular targets. In Advances in genetics; Renata, P., Ed.; Academic Press, 2010; Volume 69, pp. 1–30. [Google Scholar]
- Barrett, T.; Kobayashi, H.; Brechbiel, M.; Choyke, P.L. Macromolecular mri contrast agents for imaging tumor angiogenesis. Eur. J. Radiol. 2006, 60, 353–366. [Google Scholar] [CrossRef]
- Kiessling, F.; Morgenstern, B.; Zhang, C. Contrast agents and applications to assess tumor angiogenesis in vivo by magnetic resonance imaging. Curr. Med. Chem. 2007, 14, 77–91. [Google Scholar] [CrossRef]
- Evelhoch, J.L. Key factors in the acquisition of contrast kinetic data for oncology. J. Magn. Reson. Imag. 1999, 10, 254–259. [Google Scholar] [CrossRef]
- McIntyre, D.J.; Robinson, S.P.; Howe, F.A.; Griffiths, J.R.; Ryan, A.J.; Blakey, D.C.; Peers, I.S.; Waterton, J.C. Single dose of the antivascular agent, zd6126 (n-acetylcolchinol-o-phosphate), reduces perfusion for at least 96 hours in the gh3 prolactinoma rat tumor model. Neoplasia 2004, 6, 150–157. [Google Scholar] [CrossRef]
- Galbraith, S.M.; Maxwell, R.J.; Lodge, M.A.; Tozer, G.M.; Wilson, J.; Taylor, N.J.; Stirling, J.J.; Sena, L.; Padhani, A.R.; Rustin, G.J. Combretastatin a4 phosphate has tumor antivascular activity in rat and man as demonstrated by dynamic magnetic resonance imaging. J. Clin. Oncol. 2003, 21, 2831–2842. [Google Scholar]
- Medved, M.; Karczmar, G.; Yang, C.; Dignam, J.; Gajewski, T.; Kindler, H.; Vokes, E.; Maceneany, P.; Mitchel, M.; Stadler, W. Semiquantitative analysis of dynamic contrast enhanced mri in cancer patients: Variability and changes in tumor tissue over time. J. Magn. Reson. Imag. 2004, 20, 122–128. [Google Scholar] [CrossRef]
- Mross, K.; Drevs, J.; Muller, M.; Medinger, M.; Marme, D.; Hennig, J.; Morgan, B.; Lebwohl, D.; Masson, E.; Ho, Y.Y.; et al. Phase i clinical and pharmacokinetic study of ptk/zk, a multiple vegf receptor inhibitor, in patients with liver metastases from solid tumours. Eur. J. Canc. 2005, 41, 1291–1299. [Google Scholar] [CrossRef]
- Hillman, G.G.; Singh-Gupta, V.; Zhang, H.; Al-Bashir, A.K.; Katkuri, Y.; Li, M.; Yunker, C.K.; Patel, A.D.; Abrams, J.; Haacke, E.M. Dynamic contrast-enhanced magnetic resonance imaging of vascular changes induced by sunitinib in papillary renal cell carcinoma xenograft tumors. Neoplasia 2009, 11, 910–920. [Google Scholar]
- Marzola, P.; Degrassi, A.; Calderan, L.; Farace, P.; Nicolato, E.; Crescimanno, C.; Sandri, M.; Giusti, A.; Pesenti, E.; Terron, A.; et al. Early antiangiogenic activity of su11248 evaluated in vivo by dynamic contrast-enhanced magnetic resonance imaging in an experimental model of colon carcinoma. Clin. Canc. Res. 2005, 11, 5827–5832. [Google Scholar] [CrossRef]
- Tang, J.S.; Choy, G.; Bernardo, M.; Thomasson, D.; Libutti, S.K.; Choyke, P.L. Dynamic contrast-enhanced magnetic resonance imaging in the assessment of early response to tumor necrosis factor alpha in a colon carcinoma model. Investig. Radiol. 2006, 41, 691–696. [Google Scholar] [CrossRef]
- Checkley, D.; Tessier, J.J.; Kendrew, J.; Waterton, J.C.; Wedge, S.R. Use of dynamic contrast-enhanced mri to evaluate acute treatment with zd6474, a vegf signalling inhibitor, in pc-3 prostate tumours. Br. J. Cancer 2003, 89, 1889–1895. [Google Scholar] [CrossRef]
- Kuhl, C.; Mielcareck, P.; Klaschik, S.; Leutner, C.; Wardelmann, E.; Gieseke, J.; Schild, H. Dynamic breast mr imaging: Are signal intensity time course data useful for differential diagnosis of enhancing lesions. Radiology 1999, 211, 101–110. [Google Scholar]
- Fischer, U.; Kopka, L.; Grabbe, E. Breast carcinoma: Effect of preoperative contrast-enhanced mr imaging on the therapuetic approach. Radiology 1999, 213, 881–888. [Google Scholar]
- Buadu, L.; Murakami, J.; Murayama, S.; Hashiguchi, N.; Sakai, S.; Masuda, K.; Toyoshima, S.; Kuroki, S.; Ohno, S. Breast lesions: Correlation of contrast medium enhancement patterns on mr images with histopathologic findings and tumor angiogenesis. Radiology 1996, 200, 639–649. [Google Scholar]
- Ei Khouli, R.H.; Jacobs, M.A.; Mezban, S.D.; Huang, P.; Kamel, I.R.; Macura, K.J.; Bluemke, D.A. Diffusion-weighted imaging improves the diagnostic accuracy of conventional 3.0-t breast mr imaging. Radiology 2010, 256, 64–73. [Google Scholar] [CrossRef]
- Yabuuchi, H.; Matsuo, Y.; Okafuji, T.; Kamitani, T.; Soeda, H.; Setoguchi, T.; Sakai, S.; Hatakenaka, M.; Kubo, M.; Sadanaga, N.; et al. Enhanced mass on contrast-enhanced breast mr imaging: Lesion characterization using combination of dynamic contrast-enhanced and diffusion-weighted mr images. J. Magn. Reson.Imag. 2008, 28, 1157–1165. [Google Scholar] [CrossRef]
- Qayyum, A.; Birdwell, R.L.; Daniel, B.L.; Nowels, K.W.; Jeffrey, S.S.; Agoston, T.A.; Herfkens, R.J. Mr imaging features of infiltrating lobular carcinoma of the breast: Histopathologic correlation. Am. J. Roentgenol. 2002, 178, 1227–1232. [Google Scholar]
- Weinstein, S.P.; Orel, S.G.; Heller, R.; Reynolds, C.; Czerniecki, B.; Solin, L.J.; Schnall, M. Mr imaging of the breast in patients with invasive lobular carcinoma. Am. J. Roentgenol. 2001, 176, 399–406. [Google Scholar]
- Orel, S.G.; Schnall, M.D.; LiVolsi, V.A.; Troupin, R.H. Suspicious breast lesions: Mr imaging with radiologic-pathologic correlation. Radiology 1994, 190, 485–493. [Google Scholar]
- Buadu, L.D.; Murakami, J.; Murayama, S.; Hashiguchi, N.; Sakai, S.; Toyoshima, S.; Masuda, K.; Kuroki, S.; Ohno, S. Patterns of peripheral enhancement in breast masses: Correlation of findings on contrast medium enhanced mri with histologic features and tumor angiogenesis. J. Comput. Assist. Tomo. 1997, 21, 421–430. [Google Scholar] [CrossRef]
- Galbraith, S.M.; Lodge, M.A.; Taylor, N.J.; Rustin, G.J.S.; Bentzen, S.; Stirling, J.J.; Padhani, A.R. Reproducibility of dynamic contrast-enhanced mri in human muscle and tumours: Comparison of quantitative and semi-quantitative analysis. NMR Biomed. 2002, 15, 132–142. [Google Scholar] [CrossRef]
- Kety, S.S. The theory and applications of the exchange of inert gas at the lungs and tissues. Pharmacol. Rev. 1951, 3, 1–41. [Google Scholar]
- Shames, D.M.; Kuwatsuru, R.; Vexler, V.; Mühler, A.; Brasch, R.C. Measurement of capillary permeability to macromolecules by dynamic magnetic resonance imaging: A quantitative noninvasive technique. Magn. Reson. Med. 1993, 29, 616–622. [Google Scholar] [CrossRef]
- Faranesh, A.Z.; Kraitchman, D.L.; McVeigh, E.R. Measurement of kinetic parameters in skeletal muscle by magnetic resonance imaging with an intravascular agent. Magn. Reson. Med. 2006, 55, 1114–1123. [Google Scholar] [CrossRef]
- Daldrup, H.; Shames, D.; Wendland, M.; Okuhata, Y.; Link, T.; Rosenau, W.; Lu, Y.; Brasch, R. Correlation of dynamic contrast-enhanced mr imaging with histologic tumor grade: Comparison of macromolecular and small-molecular contrast media. Am. J. Roentgenol. 1998, 171, 941–949. [Google Scholar]
- Bradley, D.P.; Tessier, J.L.; Checkley, D.; Kuribayashi, H.; Waterton, J.C.; Kendrew, J.; Wedge, S.R. Effects of azd2171 and vandetanib (zd6474, zactima) on haemodynamic variables in an sw620 human colon tumour model: An investigation using dynamic contrast-enhanced mri and the rapid clearance blood pool contrast agent, p792 (gadomelitol). NMR Biomed. 2008, 21, 42–52. [Google Scholar] [CrossRef]
- Wedge, S.R.; Kendrew, J.; Hennequin, L.F.; Valentine, P.J.; Barry, S.T.; Brave, S.R.; Smith, N.R.; James, N.H.; Dukes, M.; Curwen, J.O.; et al. Azd2171: A highly potent, orally bioavailable, vascular endothelial growth factor receptor-2 tyrosine kinase inhibitor for the treatment of cancer. Canc. Res. 2005, 65, 4389–4400. [Google Scholar]
- Luo, Y.; Jiang, F.; Cole, T.; Hradil, V.; Reuter, D.; Chakravartty, A.; Albert, D.; Davidsen, S.; Cox, B.; McKeegan, E.; et al. A novel multi-targeted tyrosine kinase inhibitor, linifanib (abt-869), produces functional and structural changes in tumor vasculature in an orthotopic rat glioma model. Cancer Chemotherapy and Pharmacology 2012, 69, 911–921. [Google Scholar] [CrossRef]
- Nielsen, T.; Murata, R.; Maxwell, R.J.; Stødkilde-Jørgensen, H.; Østergaard, L.; Horsman, M.R. Preclinical studies to predict efficacy of vascular changes induced by combretastatin a-4 disodium phosphate in patients. International Journal of Radiation Oncology*Biology*Physics 2008, 70, 859–866. [Google Scholar] [CrossRef]
- Kingsley, P.B. Methods of measuring spin-lattice (t1) relaxation times: An annotated bibliography. Concepts Magn. Reson. 1999, 11, 243–276. [Google Scholar] [CrossRef]
- Guilfoyle, D.N.; Dyakin, V.V.; O’Shea, J.; Pell, G.S.; Helpern, J.A. Quantitative measurements of proton spin-lattice (t1) and spin-spin (t2) relaxation times in the mouse brain at 7.0 t. Magn. Reson. Med. 2003, 49, 576–580. [Google Scholar] [CrossRef]
- Loveless, M.E.; Halliday, J.; Liess, C.; Xu, L.; Dortch, R.D.; Whisenant, J.; Waterton, J.C.; Gore, J.C.; Yankeelov, T.E. A quantitative comparison of the influence of individual versus population-derived vascular input functions on dynamic contrast enhanced-mri in small animals. Magn. Reson. Med. 2012, 67, 226–236. [Google Scholar] [CrossRef]
- Haase, A.; Frahm, J.; Matthaei, D.; Hanicke, W.; Merboldt, K.D. Flash imaging. Rapid nmr imaging using low flip-angle pulses. J. Magn. Reson. 1986, 67, 258–266. [Google Scholar]
- Haacke, M.; Brown, R.; Thompson, M.; Venkatesan, R. Magnetic Resonance Imaging: Physical Principles and Sequence Design; Wiley-Liss: Hoboken, NJ, USA, 1999. [Google Scholar]
- Loveless, M.E.; Whisenant, J.G.; Wilson, K.; Lyshchik, A.; Sinha, T.K.; Gore, J.C.; Yankeelov, T.E. Coregistration of ultrasonography and magnetic resonance imaging with a preliminary investigation of the spatial colocalization of vascular endothelial growth factor receptor 2 expression and tumor perfusion in a murine tumor model. Mol. Imag. 2009, 8, 187–198. [Google Scholar]
- Yankeelov, T.E.; Niermann, K.J.; Huamani, J.; Kim, D.W.; Quarles, C.C.; Fleischer, A.C.; Hallahan, D.E.; Price, R.R.; Gore, J.C. Correlation between estimates of tumor perfusion from microbubble contrast-enhanced sonography and dynamic contrast-enhanced magnetic resonance imaging. J. Ultrasound. Med. 2006, 25, 487–497. [Google Scholar]
- Yankeelov, T.E.; Gore, J.C. Dynamic contrast enhanced magnetic resonance imaging in oncology: Theory, data acquisition, analysis, and examples. Curr. Med. Imag. Rev. 2009, 3, 91–107. [Google Scholar] [CrossRef]
- Hornak, J.P.; Szumowski, J.; Bryant, R.G. Magnetic field mapping. Magn. Reson. Med. 1988, 6, 158–163. [Google Scholar] [CrossRef]
- Yarnykh, V.L. Actual flip-angle imaging in the pulsed steady state: A method for rapid three-dimensional mapping of the transmitted radiofrequency field. Magn. Reson. Med. 2007, 57, 192–200. [Google Scholar] [CrossRef]
- Loveless, M.E.; Lawson, D.; Collins, M.; Nadella, M.V.; Reimer, C.; Huszar, D.; Halliday, J.; Waterton, J.C.; Gore, J.C.; Yankeelov, T.E. Comparisons of the efficacy of a jak 1/2 inhibitor (azd 1480) with a vegf signaling inhibitor (cediranib) and sham treatments in mouse tumors using dce-mri, dw-mri, and histology. Neoplasia 2012, 14, 54–64. [Google Scholar]
- Benjaminsen, I.C.; Graff, B.A.; Brurberg, K.G.; Rofstad, E.K. Assessment of tumor blood perfusion by high-resolution dynamic contrast-enhanced mri: A preclinical study of human melanoma xenografts. Magn. Reson. Med. 2004, 52, 269–276. [Google Scholar] [CrossRef]
- Kim, H.; Folks, K.; Guo, L.; Stockard, C.; Fineberg, N.; Grizzle, W.; George, J.; Buchsbaum, D.; Morgan, D.; Zinn, K. Dce-mri detects early vascular response in breast tumor xenografts following anti-dr5 therapy. Mol. Imag. Biol. 2011, 13, 94–103. [Google Scholar] [CrossRef]
- Li, X.; Rooney, W.D.; Várallyay, C.G.; Gahramanov, S.; Muldoon, L.L.; Goodman, J.A.; Tagge, I.J.; Selzer, A.H.; Pike, M.M.; Neuwelt, E.A.; et al. Dynamic-contrast-enhanced-mri with extravasating contrast reagent: Rat cerebral glioma blood volume determination. J. Magn. Reson. 2010, 206, 190–199. [Google Scholar]
- Skinner, J.; Yankeelov, T.E.; Peterson, T.; Does, M. Comparison of dynamic contrast enhanced mri and quantitative spect in a rat glioma model. Contrast Media Mol. Imag. 2012, in press. [Google Scholar]
- Kim, J.H.; Im, G.H.; Yang, J.; Choi, D.; Lee, W.J.; Lee, J.H. Quantitative dynamic contrast-enhanced mri for mouse models using automatic detection of the arterial input function. NMR Biomed. 2012, 25, 674–684. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; Luci, J.J.; Lepage, M.; Li, R.; Debusk, L.; Lin, P.C.; Price, R.R.; Gore, J.C. Quantitative pharmacokinetic analysis of dce-mri data without an arterial input function: A reference region model. Magn. Reson. Imag. 2005, 23, 519–529. [Google Scholar] [CrossRef]
- Fritz-Hansen, T.; Rostrup, E.; Larsson, H.B.; Sondergaard, L.; Ring, P.; Henriksen, O. Measurement of the arterial concentration of gd-dtpa using mri: A step toward quantitative perfusion imaging. Magn. Reson. Med. 1996, 36, 225–231. [Google Scholar] [CrossRef]
- Van Osch, M.J.P.; Vonken, E.-j.P.A.; Viergever, M.A.; van der Grond, J.; Bakker, C.J.G. Measuring the arterial input function with gradient echo sequences. Magn. Reson. Med. 2003, 49, 1067–1076. [Google Scholar] [CrossRef]
- Port, R.E.; Knopp, M.V.; Hoffmann, U.; Milker-Zabel, S.; Brix, G. Multicompartment analysis of gadolinium chelate kinetics: Blood-tissue exchange in mammary tumors as monitored by dynamic mr imaging. J. Magn. Reson. Med. 1999, 10, 233–241. [Google Scholar]
- Port, R.E.; Knopp, M.V.; Brix, G. Dynamic contrast-enhanced mri using gd-dtpa: Interindividual variability of the arterial input function and consequences for the assessment of kinetics in tumors. Magn. Reson. Med. 2001, 45, 1030–1038. [Google Scholar] [CrossRef]
- Parker, G.J.; Barker, G.J.; Tofts, P.S. Accurate multislice gradient echo t (1) measurement in the presence of non-ideal rf pulse shape and rf field nonuniformity. Magn. Reson. Med. 2001, 45, 838–845. [Google Scholar] [CrossRef]
- Parker, G.J.M.; Roberts, C.; Macdonald, A.; Buonaccorsi, G.A.; Cheung, S.; Buckley, D.L.; Jackson, A.; Watson, Y.; Davies, K.; Jayson, G.C. Experimentally-derived functional form for a population-averaged high-temporal-resolution arterial input function for dynamic contrast-enhanced mri. Magn. Reson. Med. 2006, 56, 993–1000. [Google Scholar] [CrossRef]
- McGrath, D.M.; Bradley, D.P.; Tessier, J.L.; Lacey, T.; Taylor, C.J.; Parker, G.J.M. Comparison of model-based arterial input functions for dynamic contrast-enhanced mri in tumor bearing rats. Magn. Reson. Med. 2009, 61, 1173–1184. [Google Scholar] [CrossRef]
- Li, X.; Welch, E.B.; Arlinghaus, L.R.; Chakravarthy, A.B.; Xu, L.; Farley, J.; Loveless, M.E.; Mayer, I.A.; Kelley, M.C.; Meszoely, I.M.; et al. A novel aif tracking method and comparison of dce-mri parameters using individual and population-based aifs in human breast cancer. Phys. Med. Biol. 2011, 56, 5753–5769. [Google Scholar] [CrossRef]
- Pickup, S.; Zhou, R.; Glickson, J. Mri estimation of the arterial input function in mice. Acad. Radiol. 2003, 10, 963–968. [Google Scholar] [CrossRef]
- Kovar, D.A.; Lewis, M.; Karczmar, G.S. A new method for imaging perfusion and contrast extraction fraction: Input functions derived from reference tissues. J. Magn. Reson. Imag. 1998, 8, 1126–1134. [Google Scholar] [CrossRef]
- Yang, C.; Karczmar, G.S.; Medved, M.; Stadler, W.M. Estimating the arterial input function using two reference tissues in dynamic contrast-enhanced mri studies: Fundamental concepts and simulations. Magn. Reson. Med. 2004, 52, 1110–1117. [Google Scholar] [CrossRef]
- Heisen, M.; Fan, X.; Buurman, J.; van Riel, V.; Karczmar, G.; ter Haar Romeny, B. The use of a reference tissue arterial input function with low-temporal-resolution dce-mri data. Phys. Med. Biol. 2010, 55, 4871–4883. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; Cron, G.O.; Addison, C.L.; Wallace, J.C.; Wilkins, R.C.; Pappas, B.A.; Santyr, G.E.; Gore, J.C. Comparison of a reference region model with direct measurement of an aif in the analysis of dce-mri data. Magn. Reson. Med. 2007, 57, 353–361. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; DeBusk, L.M.; Billheimer, D.D.; Luci, J.J.; Lin, P.C.; Price, R.R.; Gore, J.C. Repeatability of a reference region model for analysis of murine dce-mri data at 7 t. J. Magn. Reson. Imag. 2006, 24, 1140–1147. [Google Scholar] [CrossRef]
- Walker-Samuel, S.; Parker, C.C.; Leach, M.O.; Collins, D.J. Reproducibility of reference tissue quantification of dynamic contrast-enhanced data: Comparison with a fixed vascular input function. Phys. Med. Biol. 2007, 52, 75–89. [Google Scholar] [CrossRef]
- Padhani, A.R.; Hayes, C.; Landau, S.; Leach, M.O. Reproducibility of quantitative dynamic mri or normal human tissues. NMR Biomed. 2001, 15, 143–153. [Google Scholar]
- Morgan, B.; Utting, J.F.; Higginson, A.; Thomas, A.L.; Steward, W.P.; Horsfield, M.A. A simple, reproducible method for monitoring the treatment of tumours using dynamic contrast-enhanced mr imaging. Br. J. Cancer 2006, 94, 1420–1427. [Google Scholar] [CrossRef]
- Barnes, S.; Whisenant, J.; Loveless, M.; Ayers, G.; Yankeelov, T. Assessing the reproducibility of dynamic contrast enhanced magnetic resonance imaging in a murine model of breast cancer. Magn. Reson. Med. 2012. [Google Scholar] [CrossRef]
- Fueger, B.J.; Czernin, J.; Hildebrandt, I.; Tran, C.; Halpern, B.S.; Stout, D.; Phelps, M.E.; Weber, W.A. Impact of animal handling on the results of 18f-fdg pet studies in mice. J. Nucl. Med. 2006, 47, 999–1006. [Google Scholar]
- Norris, D.G. High field human imaging. J. Magn. Reson. Imag. 2003, 18, 519–529. [Google Scholar] [CrossRef]
- Padhani, A.R.; Liu, G.; Koh, D.M.; Chenevert, T.L.; Thoeny, H.C.; Takahara, T.; Dzik-Jurasz, A.; Ross, B.D.; Van Cauteren, M.; Collins, D.; et al. Diffusion-weighted magnetic resonance imaging as a cancer biomarker: Consensus and recommendations. Neoplasia 2009, 11, 102–125. [Google Scholar]
- Cheng, H.-L.M.; Wallis, C.; Shou, Z.; Farhat, W.A. Quantifying angiogenesis in vegf-enhanced tissue-engineered bladder constructs by dynamic contrast-enhanced mri using contrast agents of different molecular weights. J. Magn. Reson. Imag. 2007, 25, 137–145. [Google Scholar] [CrossRef]
- Ren, J.; Huan, Y.; Wang, H.; Chang, Y.J.; Zhao, H.T.; Ge, Y.L.; Liu, Y.; Yang, Y. Dynamic contrast-enhanced mri of benign prostatic hyperplasia and prostatic carcinoma: Correlation with angiogenesis. Clin. Radiol. 2008, 63, 153–159. [Google Scholar] [CrossRef]
- Hulka, C.A.; Edmister, W.B.; Smith, B.L.; Tan, L.; Sgroi, D.C.; Campbell, T.; Kopans, D.B.; Weisskoff, R.M. Dynamic echo-planar imaging of the breast: Experience in diagnosing breast carcinoma and correlation with tumor angiogenesis. Radiology 1997, 205, 837–842. [Google Scholar]
- Yao, W.W.; Zhang, H.; Ding, B.; Fu, T.; Jia, H.; Pang, L.; Song, L.; Xu, W.; Song, Q.; Chen, K.; et al. Rectal cancer: 3 d dynamic contrast-enhanced mri; correlation with microvascular density and clinicopathological features. Radiol. Med. 2011, 116, 366–374. [Google Scholar] [CrossRef]
- Haris, M.; Husain, N.; Singh, A.; Awasthi, R.; Singh Rathore, R.K.; Husain, M.; Gupta, R.K. Dynamic contrast-enhanced (dce) derived transfer coefficient (ktrans) is a surrogate marker of matrix metalloproteinase 9 (mmp-9) expression in brain tuberculomas. J. Magn. Reson. Imag. 2008, 28, 588–597. [Google Scholar] [CrossRef]
- Orth, R.C.; Bankson, J.; Price, R.; Jackson, E.F. Comparison of single-tracer and dual-tracer pharmacokinetic modeling of dynamic contrast-enhanced mri data using low, medium, and high molecular weight contrast agents. Magn. Reson. Med. 2007, 58, 705–716. [Google Scholar] [CrossRef]
- Reitan, N.K.; Thuen, M.; Goa, P.E.; Davies, C.D.L. Characterization of tumor microvascular structure and permeability: Comparison between magnetic resonance imaging and intravital confocal imaging. J. Biomed. Optics 2010, 15, 036004. [Google Scholar] [CrossRef]
- Mayr, N.A.; Hawighorst, H.; Yuh, W.T.C.; Essig, M.; Magnotta, V.A.; Knopp, M.V. Mr microcirculation assessment in cervical cancer: Correlations with histomorphological tumor markers and clinical outcome. J. Magn. Reson. Imag. 1999, 10, 267–276. [Google Scholar] [CrossRef]
- Egeland, T.A.M.; Simonsen, T.G.; Gaustad, J.V.; Gulliksrud, K.; Ellingsen, C.; Rofstad, E.K. Dynamic contrast-enhanced magnetic resonance imaging of tumors: Preclinical validation of parametric images. Radiat. Res. 2009, 172, 339–347. [Google Scholar] [CrossRef]
- Aref, M.; Chaudhari, A.R.; Bailey, K.L.; Aref, S.; Wiener, E.C. Comparison of tumor histology to dynamic contrast enhanced magnetic resonance imaging-based physiological estimates. Magn. Reson. Imag. 2008, 26, 1279–1293. [Google Scholar] [CrossRef]
- Sinha, T.K.; Khatib-Shahidi, S.; Yankeelov, T.E.; Mapara, K.; Ehtesham, M.; Cornett, D.S.; Dawant, B.M.; Caprioli, R.M.; Gore, J.C. Integrating spatially resolved three-dimensional maldi ims with in vivo magnetic resonance imaging. Nat. Meth. 2008, 5, 57–59. [Google Scholar] [CrossRef]
- Meyer, C.R.; Moffat, B.A.; Kuszpit, K.K.; Bland, P.L.; McKeever, P.E.; Johnson, T.D.; Chenevert, T.L.; Rehemtulla, A.; Ross, B.D. A methodology for registration of a histological slide and in vivo mri volume based on optimizing mutual information. Mol. Imag. 2006, 5, 16–23. [Google Scholar]
- Zanzonico, P.B. Broad-spectrum multi-modality image registration: From pet, ct, and mri to autoradiography, microscopy, and beyond. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2006, 1, 1584–1588. [Google Scholar]
- Naish, J.H.; McGrath, D.M.; Bains, L.J.; Passera, K.; Roberts, C.; Watson, Y.; Cheung, S.; Taylor, M.B.; Logue, J.P.; Buckley, D.L.; et al. Comparison of dynamic contrast-enhanced mri and dynamic contrast-enhanced ct biomarkers in bladder cancer. Magn. Reson. Med. 2011, 66, 219–226. [Google Scholar] [CrossRef]
- Korporaal, J.G.; van den Berg, C.A.; van Osch, M.J.; Groenendaal, G.; van Vulpen, M.; van der Heide, U.A. Phase-based arterial input function measurements in the femoral arteries for quantification of dynamic contrast-enhanced (dce) mri and comparison with dce-ct. Magn. Reson. Med. 2011, 66, 1267–1274. [Google Scholar] [CrossRef]
- Kierkels, R.G.J.; Backes, W.H.; Janssen, M.H.M.; Buijsen, J.; Beets-Tan, R.G.H.; Lambin, P.; Lammering, G.; Oellers, M.C.; Aerts, H.J.W.L. Comparison between perfusion computed tomography and dynamic contrast-enhanced magnetic resonance imaging in rectal cancer. Int. J. Radiat. Oncol. Biol. Phys. 2010, 77, 400–408. [Google Scholar] [CrossRef]
- Ng, C.S.; Waterton, J.C.; Kundra, V.; Brammer, D.; Ravoori, M.; Han, L.; Wei, W.; Klumpp, S.; Johnson, V.E.; Jackson, E.F. Reproducibility and comparison of dce-mri and dce-ct perfusion parameters in a rat tumor model. Tech. Canc. Res. Treat. 2012, 11, 279–288. [Google Scholar]
- Donahue, K.M.; Weisskoff, R.M.; Parmelee, D.J.; Callahan, R.J.; Wilkinson, R.A.; Mandeville, J.B.; Rosen, B.R. Dynamic gd-dtpa enhanced mri measurement of tissue cell volume fraction. Magn. Reson. Med. 1995, 34, 423–432. [Google Scholar] [CrossRef]
- Pellerin, M.; Yankeelov, T.E.; Lepage, M. Incorporating contrast agent diffusion into the analysis of dce-mri data. Magn. Reson. Med. 2007, 58, 1124–1134. [Google Scholar] [CrossRef]
- Einstein, A. Über die von der molekularkinetischen theorie der wärme geforderte bewegung von in ruhenden flüssigkeiten suspendierten teilchen. Ann. Phys. 1905, 322, 549–560. [Google Scholar] [CrossRef]
- Anderson, A.W.; Xie, J.; Pizzonia, J.; Bronen, R.A.; Spencer, D.D.; Gore, J.C. Effects of cell volume fraction changes on apparent diffusion in human cells. Magn. Reson. Imag. 2000, 18, 689–695. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; Lepage, M.; Chakravarthy, A.; Broome, E.E.; Niermann, K.J.; Kelley, M.C.; Meszoely, I.; Mayer, I.A.; Herman, C.R.; McManus, K.; et al. Integration of quantitative dce-mri and adc mapping to monitor treatment response in human breast cancer: Initial results. Magn. Reson. Imag. 2007, 25, 1–13. [Google Scholar]
- Mills, S.J.; Soh, C.; Rose, C.J.; Cheung, S.; Zhao, S.; Parker, G.J.M.; Jackson, A. Candidate biomarkers of extravascular extracellular space: A direct comparison of apparent diffusion coefficient and dynamic contrast-enhanced mr imaging-Derived measurement of the volume of the extravascular extracellular space in glioblastoma multiforme. Am. J. Neuroradiol. 2010, 31, 549–553. [Google Scholar] [CrossRef]
- Arlinghaus, L.R.; Li, X.; Rahman, A.R.; Welch, E.B.; Xu, L.; Gore, J.C.; Yankeelov, T.E. On the relationship between the apparent diffusion coefficient and extravascular extracellular volume fraction in human breast cancer. Magn. Reson. Imag. 2011, 29, 630–638. [Google Scholar] [CrossRef]
- Dunphy, M.P.S.; Lewis, J.S. Radiopharmaceuticals in preclinical and clinical development for monitoring of therapy with pet. J. Nucl. Med. 2009, 50, 106–121. [Google Scholar] [CrossRef]
- Dearling, J.; Lewis, J.; Mullen, G.; Welch, M.; Blower, P. Copper bis (thiosemicarbazone) complexes as hypoxia imaging agents: Structure-activity relationships. J. Biol. Inorg. Chem. 2002, 7, 249–259. [Google Scholar] [CrossRef]
- Lewis, J.S.; Welch, M.J. Pet imaging of hypoxia. Q. J. Nucl. Med. Mol. Imag. 2001, 45, 183–188. [Google Scholar]
- Cho, H.; Ackerstaff, E.; Carlin, S.; Lupu, M.E.; Wang, Y.; Rizwan, A.; O’Donoghue, J.; Ling, C.C.; Humm, J.L.; Zanzonico, P.B.; et al. Noninvasive multimodality imaging of the tumor microenvironment: Registered dynamic magnetic resonance imaging and positron emission tomography studies of a preclinical tumor model of tumor hypoxia. Neoplasia 2009, 11, 247–259. [Google Scholar]
- Coleman, C.N. Hypoxia in tumors: A paradigm for the approach to biochemical and physiologic heterogeneity. J. Natl. Canc. Inst. 1988, 80, 310–317. [Google Scholar] [CrossRef]
- Jansen, J.F.; Schoder, H.; Lee, N.Y.; Wang, Y.; Pfister, D.G.; Fury, M.G.; Stambuk, H.E.; Humm, J.L.; Koutcher, J.A.; Shukla-Dave, A. Noninvasive assessment of tumor microenvironment using dynamic contrast-enhanced magnetic resonance imaging and 18 f-fluoromisonidazole positron emission tomography imaging in neck nodal metastases. Int. J. Radiat. Oncol. Biol. Phys. 2010, 77, 1403–1410. [Google Scholar] [CrossRef]
- Shaw, R.J. Glucose metabolism and cancer. Curr. Opin. Cell. Biol. 2006, 18, 598–608. [Google Scholar] [CrossRef]
- Metz, S.; Ganter, C.; Lorenzen, S.; van Marwick, S.; Herrmann, K.; Lordick, F.; Nekolla, S.G.; Rummeny, E.J.; Wester, H.J.; Brix, G.; et al. Phenotyping of tumor biology in patients by multimodality multiparametric imaging: Relationship of microcirculation, αvβ3 expression, and glucose metabolism. J. Nucl. Med. 2010, 51, 1691–1698. [Google Scholar] [CrossRef]
- Partridge, S.C.; Vanantwerp, R.K.; Doot, R.K.; Chai, X.; Kurland, B.F.; Eby, P.R.; Specht, J.M.; Dunnwald, L.K.; Schubert, E.K.; Lehman, C.D.; et al. Association between serial dynamic contrast-enhanced mri and dynamic 18 f-fdg pet measures in patients undergoing neoadjuvant chemotherapy for locally advanced breast cancer. J. Mag. Reson. Imag. 2010, 32, 1124–1131. [Google Scholar] [CrossRef]
- Huang, J.; Chunta, J.L.; Amin, M.; Lee, D.Y.; Grills, I.S.; Wong, C.Y.; Yan, D.; Marples, B.; Martinez, A.A.; Wilson, G.D. Detailed characterization of the early response of head-neck cancer xenografts to irradiation using (18) f-fdg-pet imaging. Int. J. Radiat. Oncol. Biol. Phys. 2012, 11, 11. [Google Scholar]
- Mani, S.; Chen, Y.; Arlinghaus, L.; Li, X.; Chakravarthy, A.; Bhave, S.; Welch, E.B.; Levy, M.; Yankeelov, T.E. Early prediction of the response of breast tumors to neoadjuvant chemotherapy using quantitative mri and machine learning. AMIA Annu. Symp. Proc. 2011, 868–877. [Google Scholar]
- Cheebsumon, P.; Velasquez, L.; Hoekstra, C.; Hayes, W.; Kloet, R.; Hoetjes, N.; Smit, E.; Hoekstra, O.; Lammertsma, A.; Boellaard, R. Measuring response to therapy using fdg pet: Semi-quantitative and full kinetic analysis. Eur. J. Nucl. Med. Mol. Imag. 2011, 38, 832–842. [Google Scholar] [CrossRef]
- Braren, R.; Altomonte, J.; Settles, M.; Neff, F.; Esposito, I.; Ebert, O.; Schwaiger, M.; Rummeny, E.; Steingoetter, A. Validation of preclinical multiparametric imaging for prediction of necrosis in hepatocellular carcinoma after embolization. J. Hepatol. 2011, 55, 1034–1040. [Google Scholar] [CrossRef]
- Sala, E.; Kataoka, M.Y.; Priest, A.N.; Gill, A.B.; McLean, M.A.; Joubert, I.; Graves, M.J.; Crawford, R.A.F.; Jimenez-Linan, M.; Earl, H.M.; et al. Advanced ovarian cancer: Multiparametric mr imaging demonstrates response- and metastasis-specific effects. Radiology 2012, 263, 149–159. [Google Scholar] [CrossRef]
- Brix, G.; Semmler, W.; Port, R.E.; Schad, L.R.; Layer, G.; Lorenz, W.J. Pharmacokinetic parameters in cns gd-dtpa enhanced mr imaging. J. Comput. Assist. Tomo. 1991, 15, 621–628. [Google Scholar] [CrossRef]
- Larsson, H.B.W.; Stubgard, M.; Frederiksen, J.L.; Jensen, M.; Henriksen, O.; Paulson, O.B. Quantitation of blood-brain barrier defect by magnetic resonance imaging and gadolinium-dtpa in patients with multiple sclerosis and brain tumors. Magn. Reson. Med. 1990, 16, 117–131. [Google Scholar] [CrossRef]
- Tofts, P.S.; Kermode, A.G. Measurement of the blood-brain barrier permeability and leakage space using dynamic mr imaging. 1. Fundamental concepts. Magn. Reson. Med. 1991, 17, 357–367. [Google Scholar] [CrossRef]
- Donahue, M.; Blakeley, J.; Zhou, J.; Pomper, M.; Laterra, J.; van Zijl, P. Evaluation of human brain tumor heterogeneity using mutliple t 1-based mri signal weighting approaches. Magn. Reson. Med. 2008, 59, 336–344. [Google Scholar] [CrossRef]
- Jia, G.; O’Dell, C.; Heverhagen, J.; Xang, X.; Liang, J.; Jacko, R.; Sammet, S.; Pellas, T.; Cole, P.; Knopp, M.V. Colorectal liver metastases: Contrast agent diffusion coefficient for quantification of contrast enhancement heterogeneity at mr imaging. Radiology 2008, 248, 901–909. [Google Scholar] [CrossRef]
- Landis, C.; Li, X.; Telang, F.; Coderre, J.; Micca, P.; Rooney, W.; Latour, L.; Vetek, G.; Palyka, I.; Springer, C.S., Jr. Determination of the mri contrast agent concentration time course in vivo following bolus injection: Effect of equilibrium transcytolemmal water exchange. Magn. Reson. Med. 2000, 44, 563–574. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; Rooney, W.D.; Huang, W.; Dyke, J.P.; Li, X.; Tudorica, A.; Lee, J.H.; Koutcher, J.A.; Springer, C.S. Evidence for shutter-speed variation in cr bolus-tracking studies of human pathology. NMR Biomed. 2005, 18, 173–185. [Google Scholar] [CrossRef]
- Yankeelov, T.E.; Rooney, W.D.; Li, X.; Springer, C.S., Jr. Variation of the relaxographic “shutter-speed” for transcytolemmal water exchange affects the cr bolus-tracking curve shape. Magn. Reson. Med. 2003, 50, 1151–1169. [Google Scholar] [CrossRef]
- Li, X.; Huang, W.; Yankeelov, T.E.; Tudorica, A.; Rooney, W.D.; Springer, C.S. Shutter-speed analysis of contrast reagent bolus-tracking data: Preliminary observations in benign and malignant breast disease. Magn. Reson. Med. 2005, 53, 724–729. [Google Scholar] [CrossRef]
- Zhou, R.; Pickup, S.; Yankeelov, T.E.; Springer, C.S., Jr.; Glickson, J.D. Simultaneous measurement of arterial input function and tumor pharmacokinetics in mice by dynamic contrast enhanced imaging: Effects of transcytolemmal water exchange. Magn. Reson. Med. 2004, 52, 248–257. [Google Scholar] [CrossRef]
- Donahue, K.M.; Weisskoff, R.M.; Burstein, D. Water diffusion and exchange as they influence contrast enhancement. J. Magn. Reson. Imag. 1997, 7, 102–110. [Google Scholar] [CrossRef]
- McConnell, H.M. Reaction rates by nuclear magnetic resonance. J. Chem. Phys. 1958, 28, 430–431. [Google Scholar] [CrossRef]
- Woessner, D.E. Nuclear transfer effects in nuclear magnetic resonance pulse experiments. J. Chem. Phys. 1961, 35, 41–48. [Google Scholar] [CrossRef]
- Huang, W.; Tudorica, L.A.; Li, X.; Thakur, S.B.; Chen, Y.; Morris, E.A.; Tagge, I.J.; Korenblit, M.E.; Rooney, W.D.; Koutcher, J.A.; et al. Discrimination of benign and malignant breast lesions by using shutter-speed dynamic contrast-enhanced mr imaging. Radiology 2011, 261, 394–403. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Barnes, S.L.; Whisenant, J.G.; Loveless, M.E.; Yankeelov, T.E. Practical Dynamic Contrast Enhanced MRI in Small Animal Models of Cancer: Data Acquisition, Data Analysis, and Interpretation. Pharmaceutics 2012, 4, 442-478. https://doi.org/10.3390/pharmaceutics4030442
Barnes SL, Whisenant JG, Loveless ME, Yankeelov TE. Practical Dynamic Contrast Enhanced MRI in Small Animal Models of Cancer: Data Acquisition, Data Analysis, and Interpretation. Pharmaceutics. 2012; 4(3):442-478. https://doi.org/10.3390/pharmaceutics4030442
Chicago/Turabian StyleBarnes, Stephanie L., Jennifer G. Whisenant, Mary E. Loveless, and Thomas E. Yankeelov. 2012. "Practical Dynamic Contrast Enhanced MRI in Small Animal Models of Cancer: Data Acquisition, Data Analysis, and Interpretation" Pharmaceutics 4, no. 3: 442-478. https://doi.org/10.3390/pharmaceutics4030442
APA StyleBarnes, S. L., Whisenant, J. G., Loveless, M. E., & Yankeelov, T. E. (2012). Practical Dynamic Contrast Enhanced MRI in Small Animal Models of Cancer: Data Acquisition, Data Analysis, and Interpretation. Pharmaceutics, 4(3), 442-478. https://doi.org/10.3390/pharmaceutics4030442




