Phage Biodiversity in Artisanal Cheese Wheys Reflects the Complexity of the Fermentation Process
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacteria and Phages
2.2. Phage Screening
2.3. Selection of Phage Isolates for Genome Sequencing
2.4. DNA Preparation and Genome Sequencing
2.5. Genbank Accession Numbers
2.6. Thermal Inactivation Assays
3. Results
3.1. Phage Survey
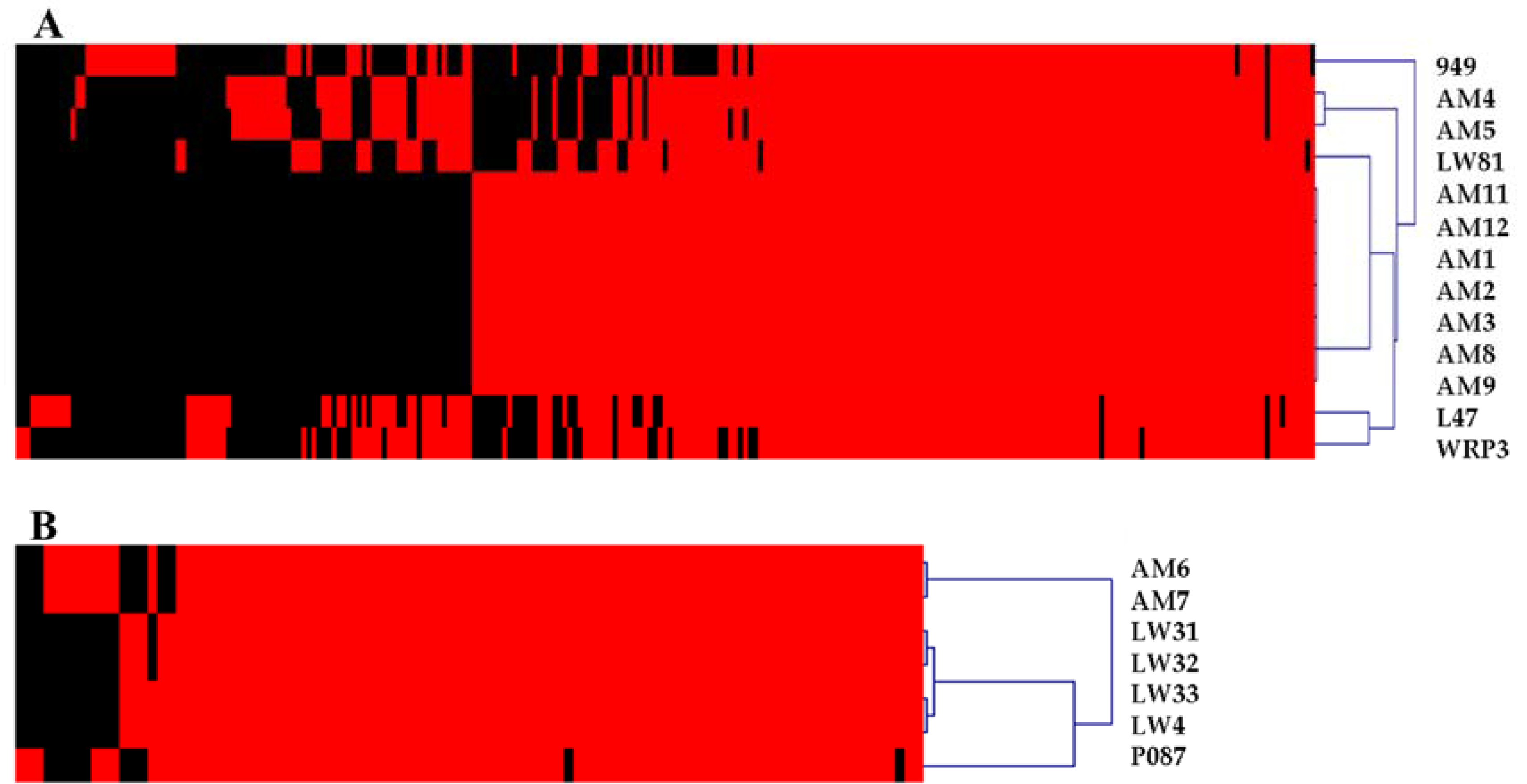
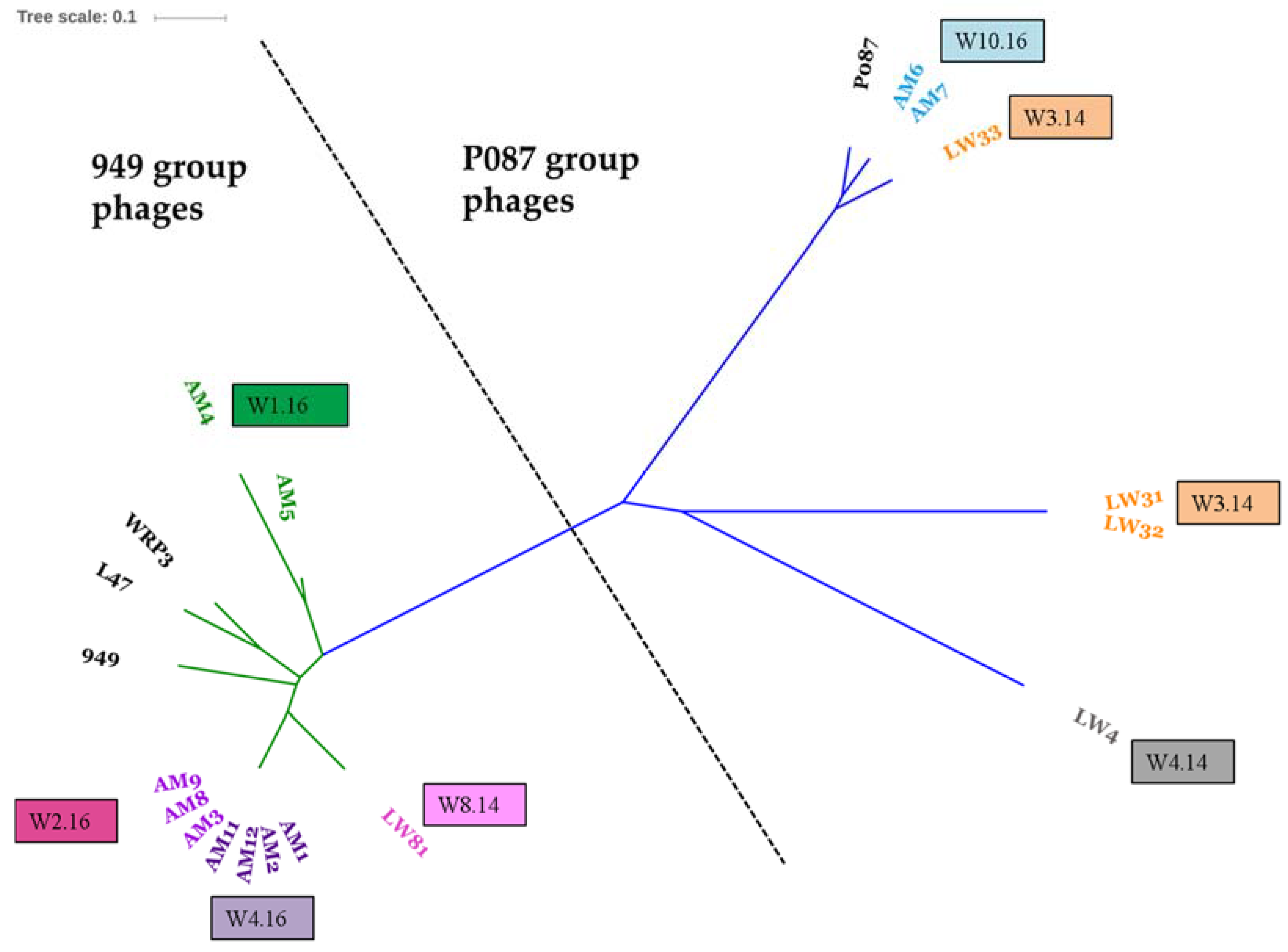
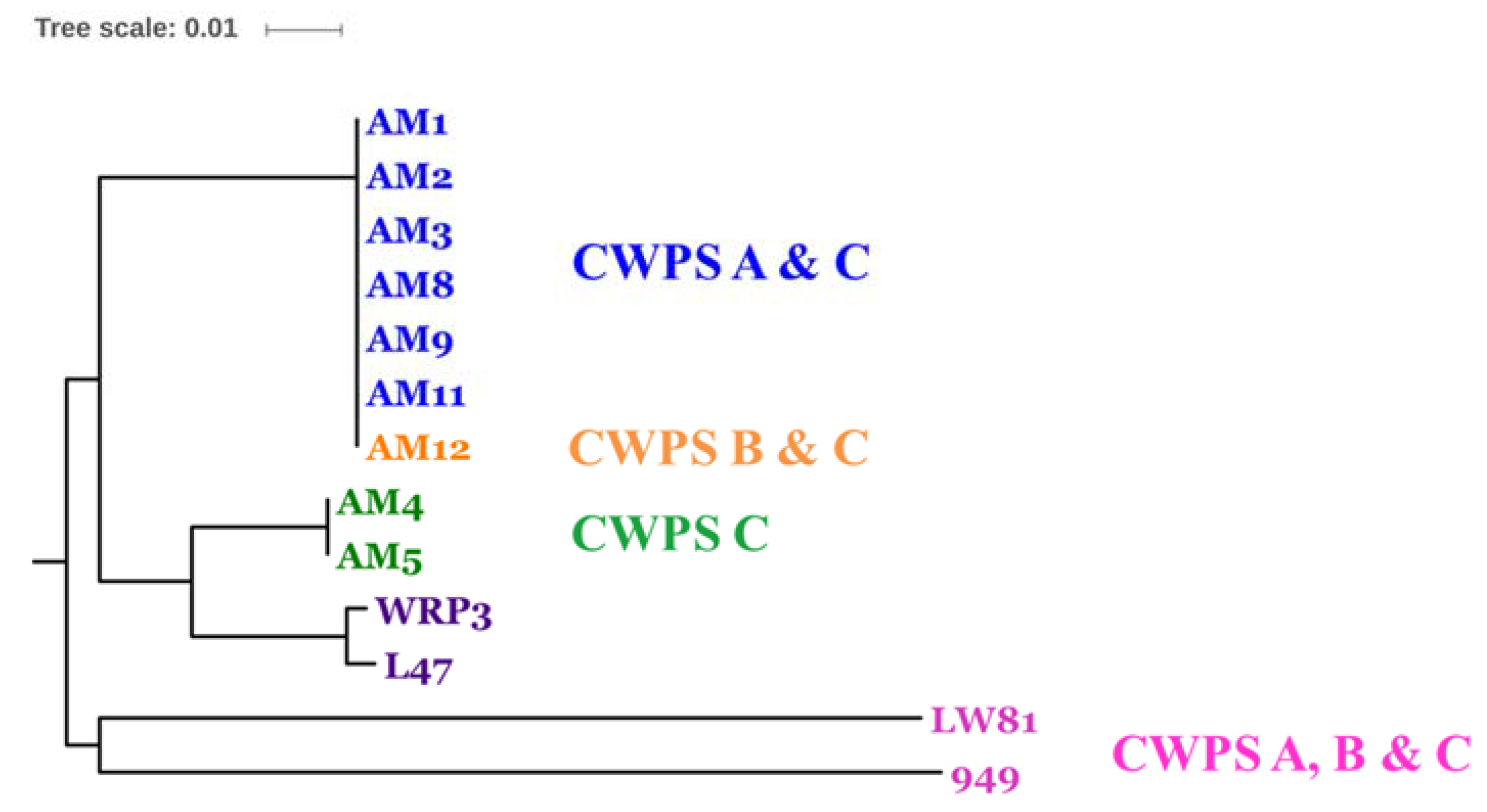
3.2. Diversity Assessment of the Phage Collection
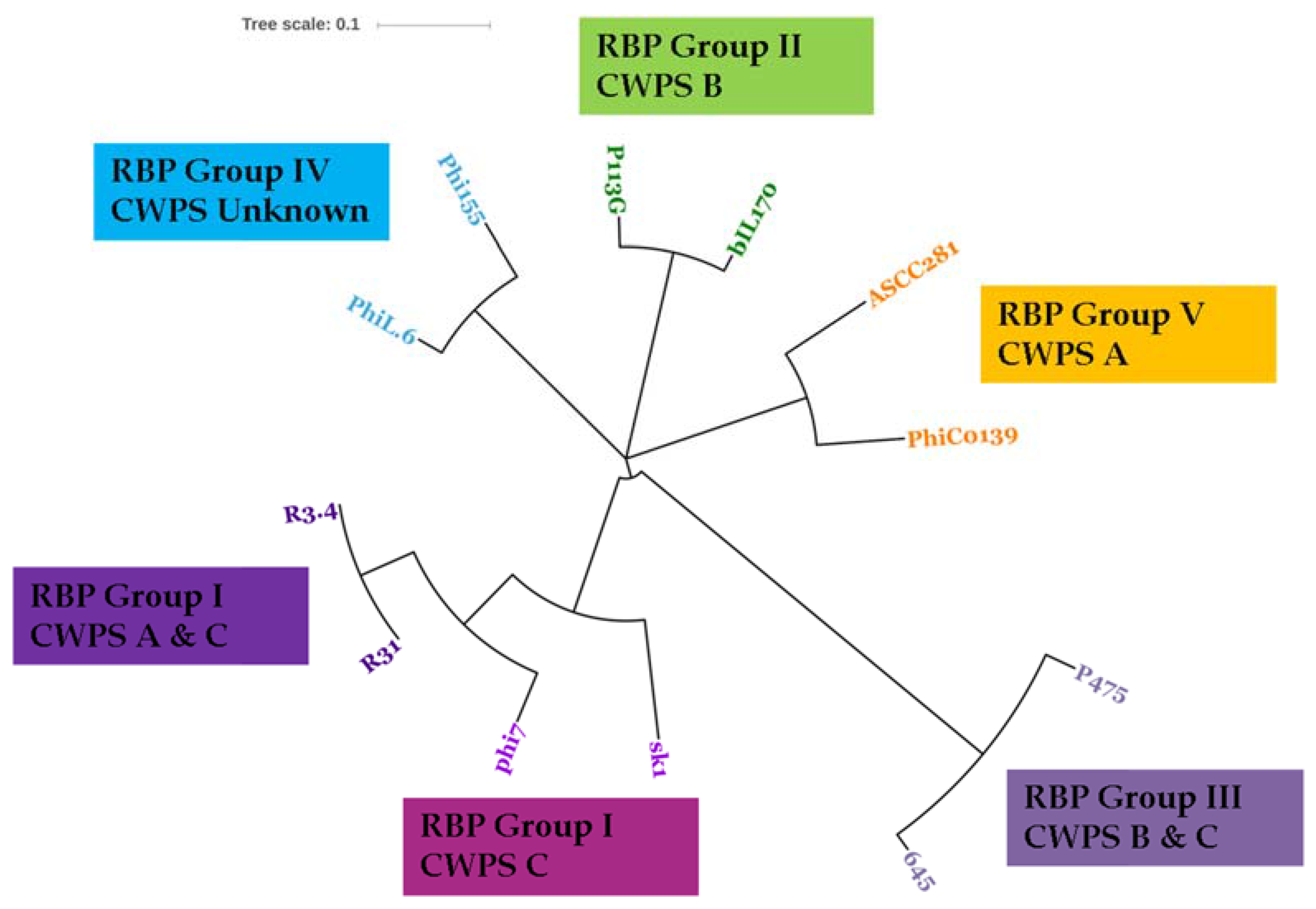
3.3. Phage-Host Interactions of the Isolated Phages
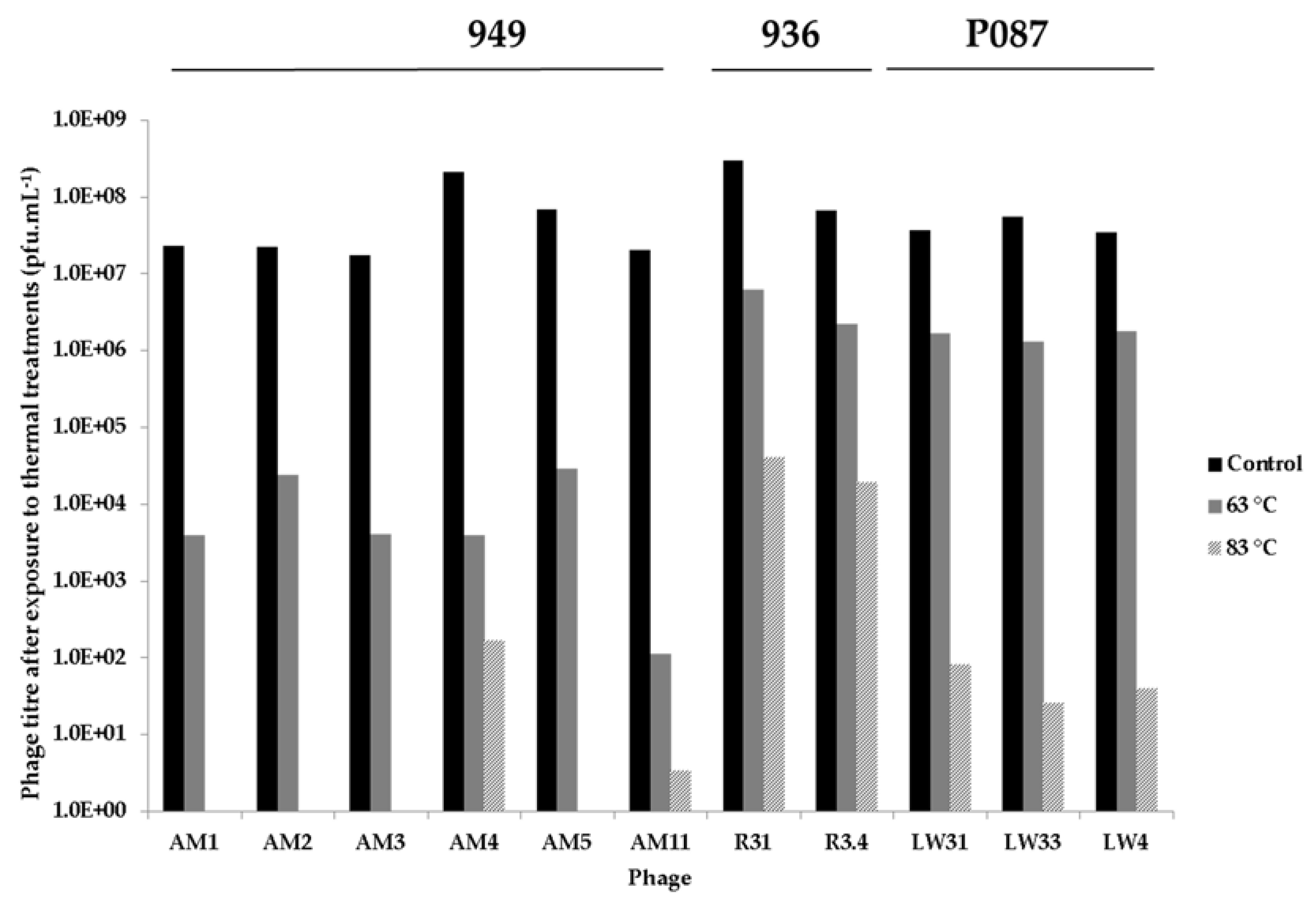
3.4. Thermal Inactivation of Phages
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bokulich, N.A.; Mills, D.A. Facility-specific “house” microbiome drives microbial landscapes of artisan cheese-making plants. Appl. Environ. Microbiol. 2013, 79, 5214–5223. [Google Scholar] [CrossRef] [PubMed]
- Scatassa, M.L.; Cardamone, C.; Miraglia, V.; Lazzara, F.; Fiorenza, G.; Macaluso, G.; Arcuri, L.; Settanni, L.; Mancuso, I. Characterisation of the microflora contaminating the wooden vats used for traditional sicilian cheese production. Ital. J. Food. Saf. 2015, 4, 4509. [Google Scholar] [CrossRef] [PubMed]
- Scatassa, M.L.; Gaglio, R.; Macaluso, G.; Francesca, N.; Randazzo, W.; Cardamone, C.; Di Grigoli, A.; Moschetti, G.; Settanni, L. Transfer, composition and technological characterization of the lactic acid bacterial populations of the wooden vats used to produce traditional stretched cheeses. Food Microbiol. 2015, 52, 31–41. [Google Scholar] [CrossRef] [PubMed]
- Deveau, H.; Labrie, S.J.; Chopin, M.C.; Moineau, S. Biodiversity and classification of lactococcal phages. Appl. Environ. Microbiol. 2006, 72, 4338–4346. [Google Scholar] [CrossRef] [PubMed]
- Szczepanska, A.K.; Hejnowicz, M.S.; Kolakowski, P.; Bardowski, J. Biodiversity of Lactococcus lactis bacteriophages in Polish dairy environment. Acta Biochim. Pol. 2007, 54, 151–158. [Google Scholar] [PubMed]
- Raiski, A.; Belyasova, N. Biodiversity of Lactococcus lactis bacteriophages in the Republic of Belarus. Int. J. Food Microbiol. 2009, 130, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.; Royer, B.; Mahony, J.; Hoyles, L.; Heller, K.; Neve, H.; Bonestroo, M.; Nauta, A.; van Sinderen, D. Biodiversity of lactococcal bacteriophages isolated from 3 Gouda-type cheese-producing plants. J. Dairy Sci. 2013, 96, 4945–4957. [Google Scholar] [CrossRef] [PubMed]
- Moineau, S.; Pandian, S.; Klaenhammer, T.R. Restriction/modification systems and restriction endonucleases are more effective on lactococcal bacteriophages that have emerged recently in the dairy industry. Appl. Environ. Microbiol. 1993, 59, 197–202. [Google Scholar] [PubMed]
- Jarvis, A.W. Differentiation of lactic streptococcal phages into phage species by DNA-DNA homology. Appl. Environ. Microbiol. 1984, 47, 343–349. [Google Scholar] [PubMed]
- Kotsonis, S.E.; Powell, I.B.; Pillidge, C.J.; Limsowtin, G.K.; Hillier, A.J.; Davidson, B.E. Characterization and genomic analysis of phage asccphi28, a phage of the family Podoviridae infecting Lactococcus lactis. Appl. Environ. Microbiol. 2008, 74, 3453–3460. [Google Scholar] [CrossRef] [PubMed]
- Mahony, J.; Randazzo, W.; Neve, H.; Settanni, L.; van Sinderen, D. Lactococcal 949 group phages recognize a carbohydrate receptor on the host cell surface. Appl. Environ. Microbiol. 2015, 81, 3299–3305. [Google Scholar] [CrossRef] [PubMed]
- Williamson, K.I.; Bertaud, W.S. A new bacteriophage active against a lactic Streptococcus. J. Bacteriol. 1951, 61, 643–645. [Google Scholar] [PubMed]
- Samson, J.E.; Moineau, S. Characterization of Lactococcus lactis phage 949 and comparison with other lactococcal phages. Appl. Environ. Microbiol. 2010, 76, 6843–6852. [Google Scholar] [CrossRef] [PubMed]
- Cavanagh, D.; Guinane, C.M.; Neve, H.; Coffey, A.; Ross, R.P.; Fitzgerald, G.F.; McAuliffe, O. Phages of non-dairy lactococci: Isolation and characterization of phiL47, a phage infecting the grass isolate Lactococcus lactis ssp. cremoris DPC6860. Front. Microbiol. 2014, 4, 417. [Google Scholar] [CrossRef] [PubMed]
- Braun, V.; Hertwig, S.; Neve, H.; Geis, A.; Teuber, M. Taxonomic differentiation of bacteriophages of Lactococcus lactis by electron microscopy, DNA-DNA hybridization, and protein profiles. J. Gen. Microbiol. 1989, 135, 2551–2560. [Google Scholar] [CrossRef]
- Villion, M.; Chopin, M.C.; Deveau, H.; Ehrlich, S.D.; Moineau, S.; Chopin, A. P087, a lactococcal phage with a morphogenesis module similar to an Enterococcus faecalis prophage. Virology 2009, 388, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Sciara, G.; Blangy, S.; Siponen, M.; Mc Grath, S.; van Sinderen, D.; Tegoni, M.; Cambillau, C.; Campanacci, V. A topological model of the baseplate of lactococcal phage Tuc2009. J. Biol. Chem. 2008, 283, 2716–2723. [Google Scholar] [CrossRef] [PubMed]
- Bebeacua, C.; Lai, L.; Vegge, C.S.; Brondsted, L.; van Heel, M.; Veesler, D.; Cambillau, C. Visualizing a complete Siphoviridae member by single-particle electron microscopy: The structure of lactococcal phage TP901-1. J. Virol. 2013, 87, 1061–1068. [Google Scholar] [CrossRef] [PubMed]
- Veesler, D.; Spinelli, S.; Mahony, J.; Lichiere, J.; Blangy, S.; Bricogne, G.; Legrand, P.; Ortiz-Lombardia, M.; Campanacci, V.; van Sinderen, D.; et al. Structure of the phage TP901-1 1.8 MDa baseplate suggests an alternative host adhesion mechanism. Proc. Natl. Acad. Sci. USA 2012, 109, 8954–8958. [Google Scholar] [CrossRef] [PubMed]
- Bebeacua, C.; Bron, P.; Lai, L.; Vegge, C.S.; Brondsted, L.; Spinelli, S.; Campanacci, V.; Veesler, D.; van Heel, M.; Cambillau, C. Structure and molecular assignment of lactococcal phage TP901-1 baseplate. J. Biol. Chem. 2010, 285, 39079–39086. [Google Scholar] [CrossRef] [PubMed]
- Mahony, J.; Martel, B.; Tremblay, D.M.; Neve, H.; Heller, K.J.; Moineau, S.; van Sinderen, D. Identification of a new P335 subgroup through molecular analysis of lactococcal phages Q33 and BM13. Appl. Environ. Microbiol. 2013, 79, 4401–4409. [Google Scholar] [CrossRef] [PubMed]
- Kot, W.; Neve, H.; Vogensen, F.K.; Heller, K.J.; Sorensen, S.J.; Hansen, L.H. Complete genome sequences of four novel Lactococcus lactis phages distantly related to the rare 1706 phage species. Genome Announc. 2014, 2, e00265-14. [Google Scholar] [CrossRef] [PubMed]
- Garneau, J.E.; Tremblay, D.M.; Moineau, S. Characterization of 1706, a virulent phage from Lactococcus lactis with similarities to prophages from other Firmicutes. Virology 2008, 373, 298–309. [Google Scholar] [CrossRef] [PubMed]
- Fortier, L.C.; Bransi, A.; Moineau, S. Genome sequence and global gene expression of q54, a new phage species linking the 936 and c2 phage species of Lactococcus lactis. J. Bacteriol. 2006, 188, 6101–6114. [Google Scholar] [CrossRef] [PubMed]
- Chopin, A.; Deveau, H.; Ehrlich, S.D.; Moineau, S.; Chopin, M.C. KSY1, a lactococcal phage with a T7-like transcription. Virology 2007, 365, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dupuis, M.E.; Moineau, S. Genome organization and characterization of the virulent lactococcal phage 1358 and its similarities to Listeria phages. Appl. Environ. Microbiol. 2010, 76, 1623–1632. [Google Scholar] [CrossRef] [PubMed]
- Lillehaug, D. An improved plaque assay for poor plaque-producing temperate lactococcal bacteriophages. J. Appl. Microbiol. 1997, 83, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Bolotin, A.; Wincker, P.; Mauger, S.; Jaillon, O.; Malarme, K.; Weissenbach, J.; Ehrlich, S.D.; Sorokin, A. The complete genome sequence of the lactic acid bacterium Lactococcus lactis ssp. lactis IL1403. Genome Res. 2001, 11, 731–753. [Google Scholar] [CrossRef] [PubMed]
- Mahony, J.; Kot, W.; Murphy, J.; Ainsworth, S.; Neve, H.; Hansen, L.H.; Heller, K.J.; Sorensen, S.J.; Hammer, K.; Cambillau, C.; et al. Investigation of the relationship between lactococcal host cell wall polysaccharide genotype and 936 phage receptor binding protein phylogeny. Appl. Environ. Microbiol. 2013, 79, 4385–4392. [Google Scholar] [CrossRef] [PubMed]
- Dupont, K.; Vogensen, F.K.; Josephsen, J. Detection of lactococcal 936-species bacteriophages in whey by magnetic capture hybridization PCR targeting a variable region of receptor-binding protein genes. J. Appl. Microbiol. 2005, 98, 1001–1009. [Google Scholar] [CrossRef] [PubMed]
- Labrie, S.; Moineau, S. Multiplex PCR for detection and identification of lactococcal bacteriophages. Appl. Environ. Microbiol. 2000, 66, 987–994. [Google Scholar] [CrossRef] [PubMed]
- Chevreux, B.; Pfisterer, T.; Drescher, B.; Driesel, A.J.; Muller, W.E.; Wetter, T.; Suhai, S. Using the Miraest assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res. 2004, 14, 1147–1159. [Google Scholar] [CrossRef] [PubMed]
- Hyatt, D.; Chen, G.L.; Locascio, P.F.; Land, M.L.; Larimer, F.W.; Hauser, L.J. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010, 11, 119. [Google Scholar] [CrossRef] [PubMed]
- Gish, W.; States, D.J. Identification of protein coding regions by database similarity search. Nat. Genet. 1993, 3, 266–272. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and psi-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Lugli, G.A.; Milani, C.; Mancabelli, L.; van Sinderen, D.; Ventura, M. MEGAnnotator: A user-friendly pipeline for microbial genomes assembly and annotation. FEMS Microbiol. Lett. 2016, 363, fnw049. [Google Scholar] [CrossRef] [PubMed]
- Bateman, A.; Coin, L.; Durbin, R.; Finn, R.D.; Hollich, V.; Griffiths-Jones, S.; Khanna, A.; Marshall, M.; Moxon, S.; Sonnhammer, E.L. The Pfam protein families database. Nucleic Acids Res. 2004, 32, D138–D141. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R. CDD: A Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef] [PubMed]
- Söding, J.; Biegert, A.; Lupas, A.N. The HHPred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 2005, 33, W244–W248. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. TRNAScan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Enright, A.J.; Van Dongen, S.; Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 2002, 30, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- Saeed, A.I.; Sharov, V.; White, J.; Li, J.; Liang, W.; Bhagabati, N.; Braisted, J.; Klapa, M.; Currier, T.; Thiagarajan, M.; et al. Tm4: A free, open-source system for microarray data management and analysis. Biotechniques 2003, 34, 374–378. [Google Scholar] [PubMed]
- Zhao, Y.; Wu, J.; Yang, J.; Sun, S.; Xiao, J.; Yu, J. PGAP: Pan-genomes analysis pipeline. Bioinformatics 2012, 28, 416–418. [Google Scholar] [CrossRef] [PubMed]
- Tettelin, H.; Masignani, V.; Cieslewicz, M.J.; Donati, C.; Medini, D.; Ward, N.L.; Angiuoli, S.V.; Crabtree, J.; Jones, A.L.; Durkin, A.S.; et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: Implications for the microbial “pan-genome”. Proc. Natl. Acad. Sci. USA 2005, 102, 13950–13955. [Google Scholar] [CrossRef] [PubMed]
- Murphy, J.; Bottacini, F.; Mahony, J.; Kelleher, P.; Neve, H.; Zomer, A.; Nauta, A.; van Sinderen, D. Comparative genomics and functional analysis of the 936 group of lactococcal Siphoviridae phages. Sci Rep. 2016, 6, 21345. [Google Scholar] [CrossRef] [PubMed]
- Di Grigoli, A.; Francesca, N.; Gaglio, R.; Guarrasi, V.; Moschetti, M.; Scatassa, M.L.; Settanni, L.; Bonanno, A. The influence of the wooden equipment employed for cheese manufacture on the characteristics of a traditional stretched cheese during ripening. Food Microbiol. 2015, 46, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Del Rio, B.; Binetti, A.G.; Martin, M.C.; Fernandez, M.; Magadan, A.H.; Alvarez, M.A. Multiplex PCR for the detection and identification of dairy bacteriophages in milk. Food Microbiol. 2007, 24, 75–81. [Google Scholar] [CrossRef] [PubMed]

| Phage | Lactococcus lactis Host (Reference) | Source (Sample Ref. No.) | Year |
|---|---|---|---|
| LW11 | 3107 [15] | Vastedda cheese whey (raw ewe’s milk) (W1.14) | 2014 |
| LW12 | 3107 | Vastedda cheese whey (W1.14) | 2014 |
| LW21 * | 3107 | Vastedda cheese whey (W2.14) | 2014 |
| LW22* | 3107 | Vastedda cheese whey (W2.14) | 2014 |
| LW31 | 3107 | Vastedda cheese whey (W3.14) | 2014 |
| LW32 | 3107 | Vastedda cheese whey (W3.14) | 2014 |
| LW33 | 3107 | Vastedda cheese whey (W3.14) | 2014 |
| LW34 | 3107 | Vastedda cheese whey (W3.14) | 2014 |
| LW4 | 3107 | Canestrato cheese whey (raw cow’s milk) (W4.14) | 2014 |
| LW71 * | 3107 | Caciocavallo cheese whey (raw cow’s milk) (W7.14) | 2014 |
| LW72 * | 3107 | Caciocavallo cheese whey (W7.14) | 2014 |
| LW81 | 3107 | Canestrato cheese whey (W8.14) | 2014 |
| LW82 | 3107 | Canestrato cheese whey (W8.14) | 2014 |
| LW91 | 3107 | Canestrato cheese whey (W9.14) | 2014 |
| LW92 * | 3107 | Canestrato cheese whey (W9.14) | 2014 |
| LW101 | 3107 | Canestrato cheese whey (W10.14) | 2014 |
| LW102 | 3107 | Canestrato cheese whey (W10.14) | 2014 |
| GW11 * | 3107 | Goat’s cheese whey (W11.14) | 2014 |
| CW12 * | 3107 | Cow’s cheese whey (W12.14) | 2014 |
| R1 | SMQ-86 [4] | Lamb rennet used in Caciocavallo cheese (R1.14) | 2014 |
| R2 | SMQ-86 | Lamb rennet used in Caciocavallo cheese (R2.14) | 2014 |
| R31 | SMQ-86 | Lamb rennet used in Vastedda cheese (R3.14) | 2014 |
| R32 | SMQ-86 | Lamb rennet used in Vastedda cheese (R3.14) | 2014 |
| R33 | SMQ-86 | Lamb rennet used in Vastedda cheese (R3.14) | 2014 |
| R3.4 | SMQ-86 | Lamb rennet used in Vastedda cheese (R3.14) | 2014 |
| R35 | SMQ-86 | Lamb rennet used in Vastedda cheese (R3.14) | 2014 |
| R4 | SMQ-86 | Lamb rennet used in Caciocavallo cheese (R4.14) | 2014 |
| AM1 | 3107 | Vastedda cheese whey (W4.16) | 2016 |
| AM2 | 3107 | Vastedda cheese whey (W4.16) | 2016 |
| AM3 | 3107 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM4 | 3107 | Caciocavallo cheese whey (W1.16) | 2016 |
| AM5 | 3107 | Caciocavallo cheese whey (W1.16) | 2016 |
| AM6 | SMQ-86 | Caciocavallo cheese whey (W10.16) | 2016 |
| AM7 | SMQ-86 | Caciocavallo cheese whey (W10.16) | 2016 |
| AM8 | SMQ-86 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM9 | SMQ-86 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM10 | 3107 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM11 | 3107 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM12 | IL1403 [28] | Caciocavallo cheese whey (W2.16) | 2016 |
| AM13 | C10 [29] | Vastedda cheese whey (W4.16) | 2016 |
| AM14 | C10 | Caciocavallo cheese whey (W5.16) | 2016 |
| AM15 | C10 | Caciocavallo cheese whey (W5.16) | 2016 |
| AM16 | C10 | Caciocavallo cheese whey (W5.16) | 2016 |
| AM17 | C10 | Caciocavallo cheese whey (W5.16) | 2016 |
| AM18 | C10 | Caciocavallo cheese whey (W5.16) | 2016 |
| AM19 | SMQ-86 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM20 | SMQ-86 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM21 | SMQ-86 | Caciocavallo cheese whey (W10.16) | 2016 |
| AM22 | SMQ-86 | Caciocavallo cheese whey (W10.16) | 2016 |
| AM23 | SMQ-86 | Caciocavallo cheese whey (W10.16) | 2016 |
| AM24 | 3107 | Caciocavallo cheese whey (W1.16) | 2016 |
| AM25 | 3107 | Caciocavallo cheese whey (W1.16) | 2016 |
| AM26 | 3107 | Caciocavallo cheese whey (W1.16) | 2016 |
| AM27 | 3107 | Caciocavallo cheese whey (W1.16) | 2016 |
| AM28 | 3107 | Caciocavallo cheese whey(W1.16) | 2016 |
| AM29 | 3107 | Caciocavallo cheese whey (W2.16) | 2016 |
| AM30 | 3107 | Vastedda cheese whey (W4.16) | 2016 |
| AM31 | 3107 | Vastedda cheese whey (W4.16) | 2016 |
| AM32 | 3107 | Vastedda cheese whey (W4.16) | 2016 |
| Phage | Phage Group | Genome Length (kb) | GC % | No. Predicted ORFs | tRNAs | Best Hit (% nt ID/Coverage) | Genbank Accession No. |
|---|---|---|---|---|---|---|---|
| LW31 | P087 | 60.551 | 34.3 | 85 | 3 (Cys, Asn, Thr) | P087 (97/91) | KY554762 |
| LW32 | P087 | 60.161 | 34.3 | 87 | 3 (Cys, Asn, Thr) | P087 (97/91) | KY554763 |
| LW33 | P087 | 59.899 | 34.4 | 86 | 3 (Cys, Asn, Thr) | P087 (97/91) | KY554764 |
| LW4 | P087 | 60.217 | 34.3 | 88 | 3 (Cys, Asn, Thr) | P087 (97/91) | KY554765 |
| LW81 | 949 | 128.179 | 32.6 | 178 | 4 (Trp, Asp) | WRP3 (95/84) | KY554777 |
| R31 | 936 | 27.203 | 35.4 | 52 | 2 (Trp, Pro) | Phi19 (90/82) | KY554761 |
| R3.4 | 936 | 27.704 | 35.4 | 51 | 2 (Trp, Pro) | Phi19 (96/82) | KY554760 |
| AM1 | 949 | 125.658 | 32.6 | 178 | 3 (Pro, Trp) | WRP3 (95/82) | KY554768 |
| AM2 | 949 | 125.656 | 32.5 | 177 | 3 (Pro, Trp) | WRP3 (95/82) | KY554769 |
| AM3 | 949 | 126.032 | 32.5 | 177 | 3 (Pro, Trp) | WRP3 (95/82) | KY554770 |
| AM4 | 949 | 132.949 | 32.5 | 194 | 3 (Arg, Pro, Met) | WRP3 (94/81) | KY554771 |
| AM5 | 949 | 128.178 | 32.6 | 182 | 3 (Arg, Pro, Met) | WRP3 (94/81) | KY554772 |
| AM6 | P087 | 62.054 | 34.3 | 90 | 4 (Pro, Thr, Asn, Cys) | P087 (97/96) | KY554766 |
| AM7 | P087 | 62.252 | 34.2 | 90 | 4 (Pro, Thr, Asn, Cys) | P087 (97/96) | KY554767 |
| AM8 | 949 | 126.177 | 32.6 | 178 | 3 (Pro, Trp) | WRP3 (95/82) | KY554773 |
| AM9 | 949 | 125.294 | 32.5 | 179 | 3 (Pro, Trp) | WRP3 (95/82) | KY554774 |
| AM11 | 949 | 126.161 | 32.6 | 178 | 3 (Pro, Trp) | WRP3 (95/82) | KY554775 |
| AM12 | 949 | 125.842 | 32.6 | 179 | 3 (Pro, Trp) | WRP3 (96/82) | KY554776 |
| Sample | Phage Titre on Lactococcal Strain (pfu·mL−1) | Plaque Morphology | |||
|---|---|---|---|---|---|
| 3107 | SMQ-86 | C10 | IL1403 | ||
| W1.14 | 6 × 103 | - | - | - | 1 mm clear |
| W2.14 | 8 × 102 | - | - | - | Pinpoint–0.5 mm |
| W3.14 | 4 × 102 | 2 × 103 | - | - | 1 mm |
| W4.14 | 1 × 102 | - | - | - | 1 mm |
| W7.14 | 1.3 × 104 | - | - | - | Pinpoint–1 mm |
| W8.14 | 4 × 103 | - | - | - | Pinpoint–1 mm |
| W9.14 | 2 × 103 | - | - | - | Pinpoint and 1.5 mm |
| W10.14 | 2.3 × 103 | - | - | - | Pinpoint and 1 mm |
| W11.14 | - | 8 × 102 | - | - | Pinpoint |
| W12.14 | - | 2.7 × 103 | - | - | Pinpoint |
| R1.14 | - | 3.3 × 103 | - | - | 1.5 mm |
| R2.14 | - | 5.9 × 103 | - | - | 1.5 mm |
| R3.14 | - | 1.4 × 103 | - | - | 1.5 mm |
| R4.14 | - | 7 × 102 | - | - | 1.5 mm |
| W1.16 | 8 × 102 | - | - | - | Pinpoint |
| W2.16 | 4 × 102 | 4 × 102 | - | 1 × 102 | Pinpoint |
| W4.16 | 5 × 102 | - | 1 × 102 | - | Pinpoint |
| W5.16 | - | - | 5 × 102 | - | Pinpoint |
| W10.16 | - | 5 × 102 | - | - | 1 mm |
| Phage | Phage Titre on Lactococcal Strains (pfu·mL−1) | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CWPS A Strains | CWPS B Strains | CWPS C Strains | ||||||||||||||
| SMQ-86 | SMQ-562 | WM1 | 275 | UC77 | IL1403 | 223 | 3107 | KH | NZ9000 | R1k10 | W34 | FD13 | JM3 | 1196 | 158 | |
| R31 a | 4 × 108 | 6 × 108 | ||||||||||||||
| R3.4 a | 3 × 107 | 2 × 108 | ||||||||||||||
| LW31 b | 2 × 107 | 3 × 106 | 2 × 102 | 4 × 107 | ||||||||||||
| LW32 b | 2 × 107 | 4 × 106 | 1 × 102 | 4 × 104 | 3 × 107 | |||||||||||
| LW33 b | 4 × 107 | 3 × 106 | 1 × 102 | 3 × 104 | 4 × 107 | |||||||||||
| LW4 b | 3 × 107 | 2 × 106 | 3 × 102 | 7 × 103 | 5 × 107 | |||||||||||
| AM6 b | 4 × 107 | |||||||||||||||
| AM7 b | 2 × 106 | 2 × 103 | ||||||||||||||
| LW81 c | 4 × 107 | 5 × 105 | 3 × 106 | 4 × 106 | 6 × 108 | 2 × 105 | ||||||||||
| AM1 c | 8 × 105 | 2 × 105 | 4 × 107 | 1 × 104 | 1 × 103 | 2 × 103 | ||||||||||
| AM2 c | 7 × 105 | 7 × 104 | 8 × 107 | 5 × 103 | 1 × 105 | |||||||||||
| AM3 c | 3 × 105 | 9 × 104 | 5 × 107 | 3 × 105 | 1 × 103 | 1 × 104 | 8 × 102 | |||||||||
| AM4 c | 2 × 107 | |||||||||||||||
| AM5 c | 3 × 107 | 3 × 104 | ||||||||||||||
| AM8 c | 5 × 107 | 3 × 105 | 1 × 103 | 4 × 106 | 1 × 105 | 4 × 104 | 9 × 104 | |||||||||
| AM9 c | 9 × 107 | 8 × 105 | 1 × 107 | 2 × 106 | 1 × 103 | 1 × 103 | 2 × 105 | 1 × 105 | 7 × 102 | 9 × 104 | ||||||
| AM11 c | 3 × 105 | 7 × 107 | 5 × 104 | |||||||||||||
| AM12 c | 5 × 105 | 9 × 107 | ||||||||||||||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mahony, J.; Moscarelli, A.; Kelleher, P.; Lugli, G.A.; Ventura, M.; Settanni, L.; Van Sinderen, D. Phage Biodiversity in Artisanal Cheese Wheys Reflects the Complexity of the Fermentation Process. Viruses 2017, 9, 45. https://doi.org/10.3390/v9030045
Mahony J, Moscarelli A, Kelleher P, Lugli GA, Ventura M, Settanni L, Van Sinderen D. Phage Biodiversity in Artisanal Cheese Wheys Reflects the Complexity of the Fermentation Process. Viruses. 2017; 9(3):45. https://doi.org/10.3390/v9030045
Chicago/Turabian StyleMahony, Jennifer, Angelo Moscarelli, Philip Kelleher, Gabriele A. Lugli, Marco Ventura, Luca Settanni, and Douwe Van Sinderen. 2017. "Phage Biodiversity in Artisanal Cheese Wheys Reflects the Complexity of the Fermentation Process" Viruses 9, no. 3: 45. https://doi.org/10.3390/v9030045
APA StyleMahony, J., Moscarelli, A., Kelleher, P., Lugli, G. A., Ventura, M., Settanni, L., & Van Sinderen, D. (2017). Phage Biodiversity in Artisanal Cheese Wheys Reflects the Complexity of the Fermentation Process. Viruses, 9(3), 45. https://doi.org/10.3390/v9030045







