Defining the Enterovirus Diversity Landscape of a Fecal Sample: A Methodological Challenge?
Abstract
:1. Introduction
2. History of Enterovirus Classification
2.1. Initial Isolation and Classification Strategies
2.2. Identifying the Value of VP1 in Enterovirus Identification
2.3. Equating Serotypes and Genotypes
2.4. Identifying the “Untypables”
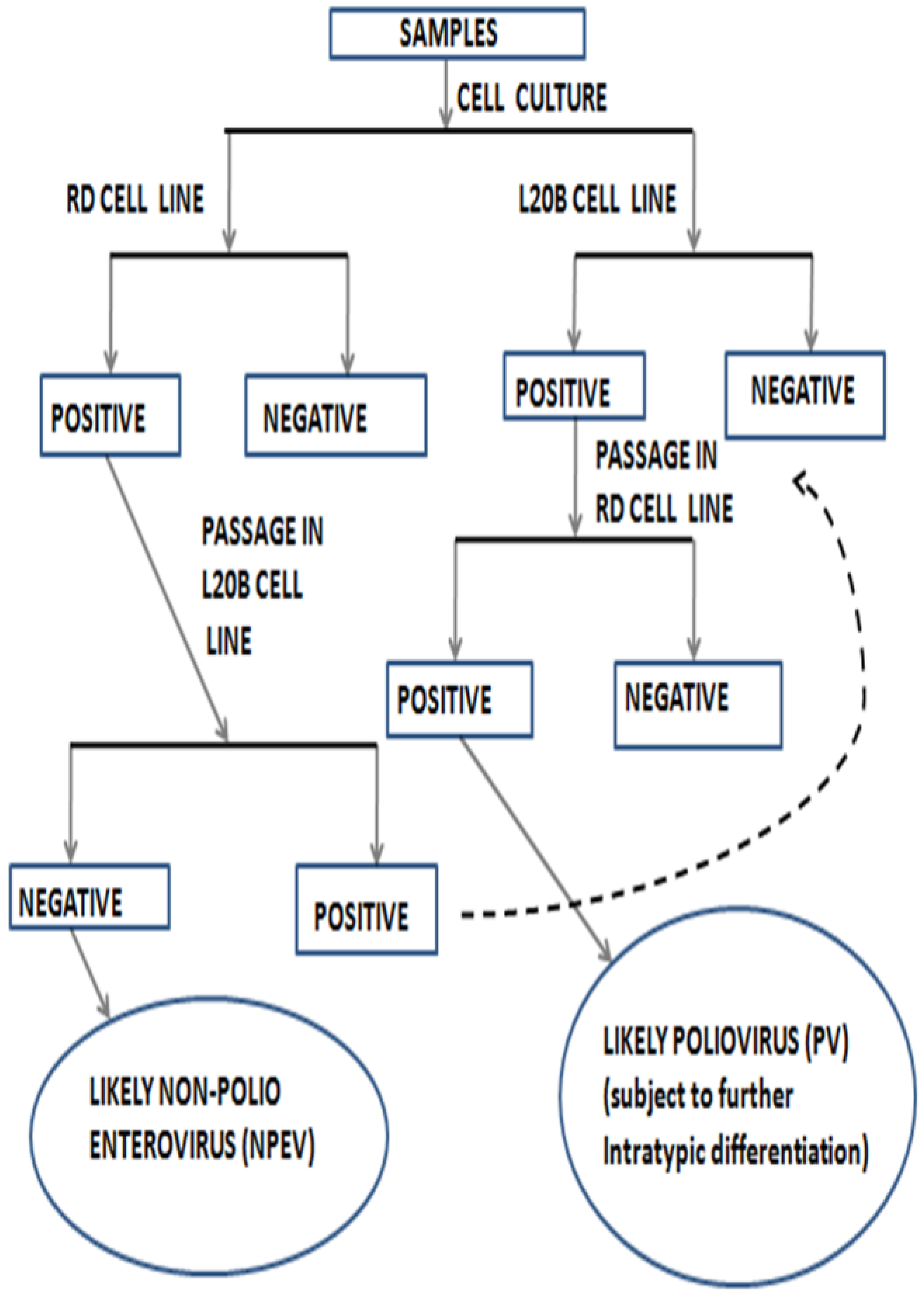
3. Impact of Cell Line Bias on the Enterovirus Diversity Landscape
4. Molecular Identification without Cell Culture
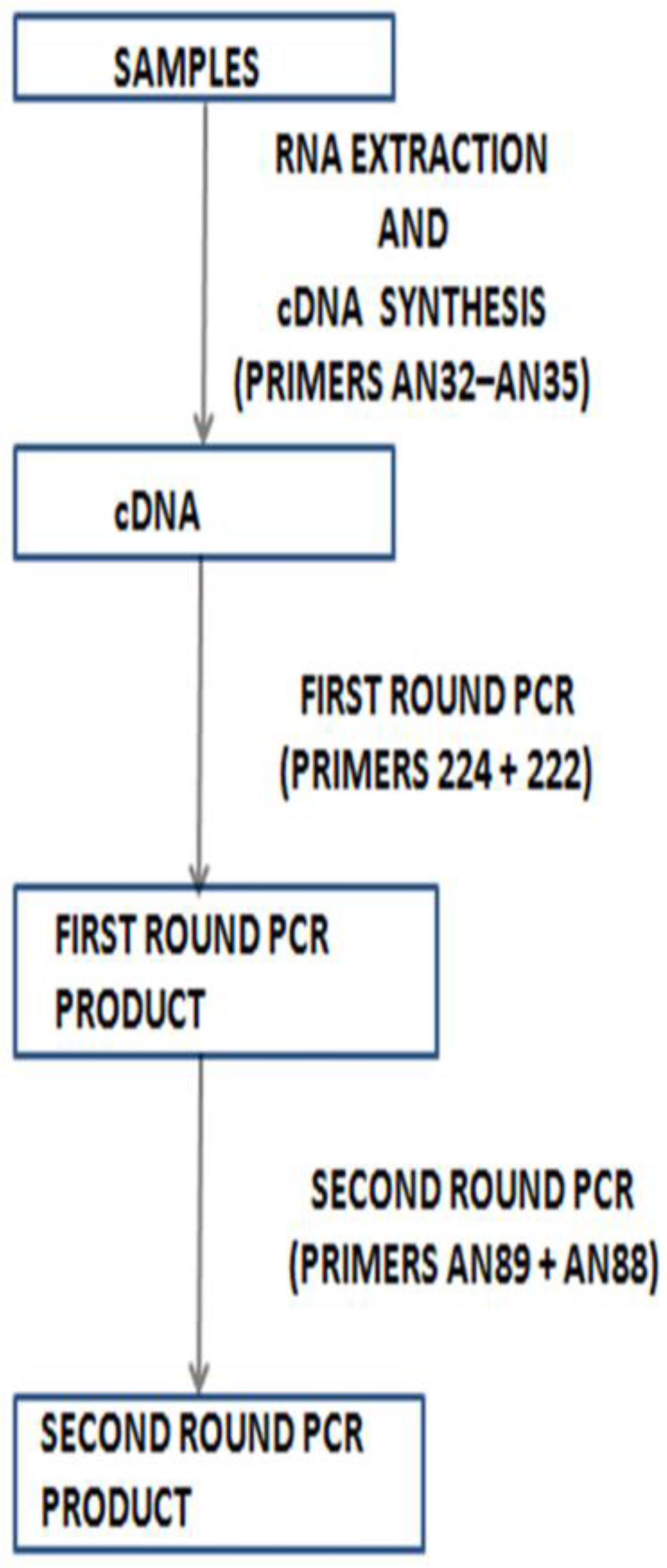
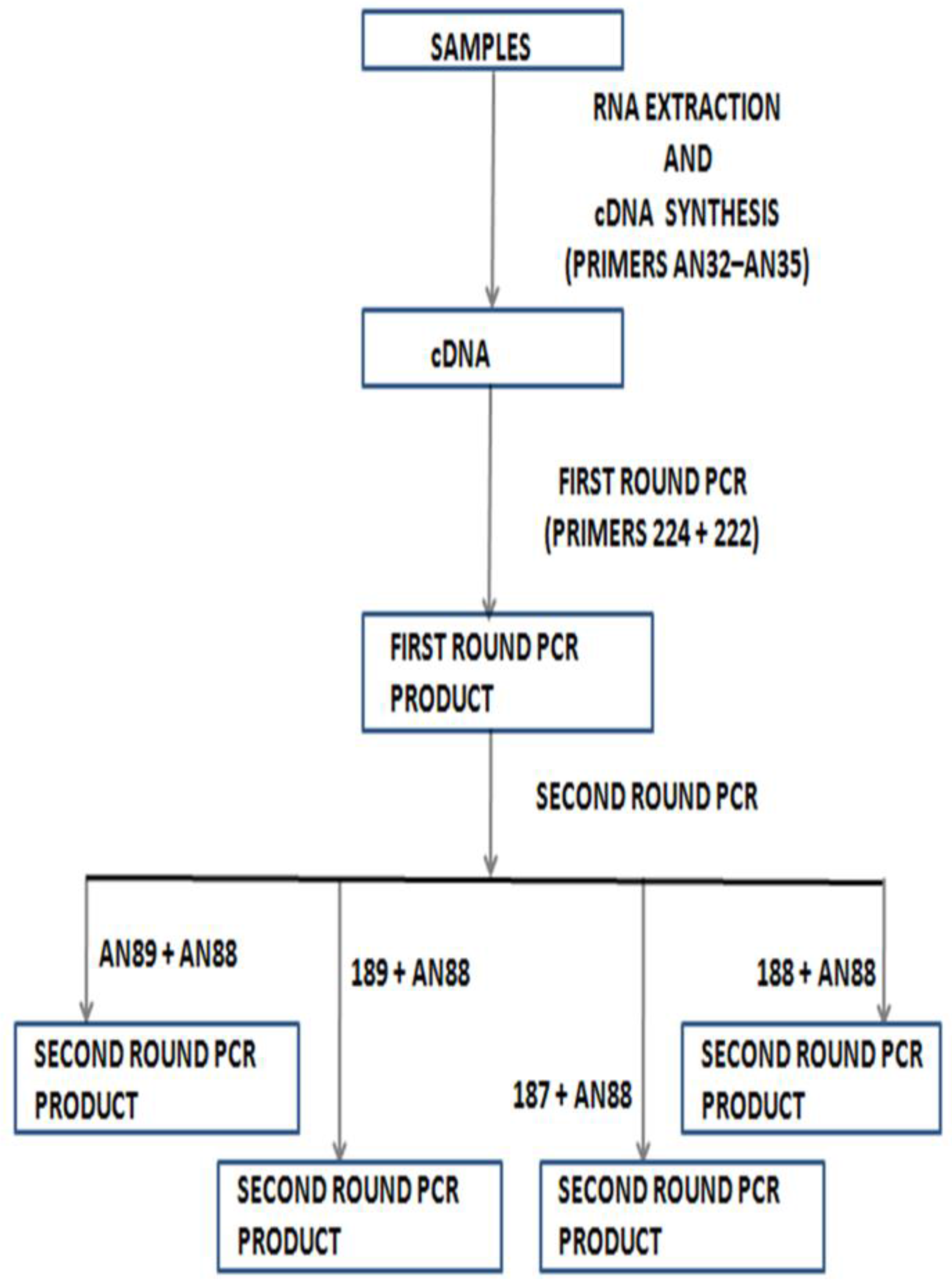
5. Shortcomings of Molecular Identification
6. Serotype is Not the Full Picture: Same Serotype, Different Phenotypes/Biological Properties
7. Conclusion: Enterovirus Diversity Landscape, What Is Really out There?
Acknowledgments
Author Contributions
Conflicts of Interest
References
- The Picornavirus Pages. Available online: http://www.picornaviridae.com (accessed on 12 January 2016).
- Tassin, M.; Martinovic, J.; Mirand, A.; Peigue-Lafeuille, H.; Picone, O.; Benachi, A.; Fellous, C.V. A case of congenital Echovirus11 infection acquired early in pregnancy. J. Clin. Virol. 2013, 59, 71–73. [Google Scholar] [CrossRef] [PubMed]
- Tapparel, C.; Siegrist, F.; Petty, T.J.; Kaiser, L. Picornavirus and enterovirus diversity with associated human disease. Infect. Genet. Evol. 2013, 12, 505–521. [Google Scholar] [CrossRef] [PubMed]
- Nathanson, N.; Kew, O.M. From Emergenceto Eradication: The Epidemiology of Poliomyelitis Deconstructed. Am. J. Epidemiol. 2010, 172, 1213–1229. [Google Scholar] [CrossRef] [PubMed]
- Strikas, R.A.; Anderson, L.; Parker, R.A. Temporal and geographic patterns of isolates of non polioenteroviruses in the United States, 1970–1983. J. Infect. Dis. 1986, 153, 346–351. [Google Scholar] [CrossRef] [PubMed]
- Landsteiner, K.; Popper, E. Mikroscopische präparate von einem menschlichen und zwei affentückermarker. Wien. Klin. Wochenschr. 1908, 21, 1830. [Google Scholar]
- Kessel, J.F.; Part, C.F. Differentiation of three groups of Poliomyelitis virus. Proc. Soc. Exp. Biol. Med. 1949, 70, 315–316. [Google Scholar] [CrossRef] [PubMed]
- Dalldorf, G.; Sickles, G.M. A virus recovered from the feaces of poliomyelitis patients pathogenic for suckling mice. J. Exp. Med. 1949, 89, 567–582. [Google Scholar] [CrossRef] [PubMed]
- Melnik, J.L.; Clarke, N.A.; Kraft, L.M. Immunological reaction of the coxsackie Viruses. Cross-protection test in infant mice born of vaccinated mothers: transfer of immunity through the milk. J. Exp. Med. 1950, 92, 499–505. [Google Scholar] [CrossRef]
- Melnick, J.L.; Agren, K. Poliomyelitis and Coxsackievirus isolated from Normal infant in Egypt. J. Exp. Biol. Med. 1952, 81, 621–624. [Google Scholar] [CrossRef]
- Chow, M.; Baltimore, D. Isolated poliovirus capsid protein VP1 induces a neutralizing response in rats. Proc. Natl. Acad. Sci. USA 1982, 79, 7518–7521. [Google Scholar] [CrossRef] [PubMed]
- Wychowski, C.; van der Werf, S.; Siffert, O.; Crainic, R.; Bruneau, P.; Girard, M. A poliovirus type 1 neutralization epitope is located within amino acid residues 93 to 104 of viral capsid polypeptide VP1. EMBO J. 1983, 2, 2019–2024. [Google Scholar] [PubMed]
- Minor, P.D.; Schild, G.C.; Bootman, J.; Evans, D.M.; Ferguson, M.; Reeve, P.; Spitz, M.; Stanway, G.; Cann, A.J.; Hauptmann, R.; et al. Location and primary structure of a major antigenic site for poliovirus neutralization. Nature 1983, 301, 674–679. [Google Scholar] [CrossRef] [PubMed]
- Emini, E.A.; Dorner, A.J.; Dorner, L.F.; Jameson, B.A.; Wimmer, E. Identification of a poliovirus neutralization epitope through use of neutralizing antiserum raised against a purified viral structural protein. Virology 1983, 124, 144–151. [Google Scholar] [CrossRef]
- Evans, D.M.; Minor, P.D.; Schild, G.S.; Almond, J.W. Critical role of an eight-amino acid sequence of VP1 in neutralization of poliovirus type 3. Nature 1983, 304, 459–462. [Google Scholar] [CrossRef] [PubMed]
- Chow, M.; Yabrov, R.; Bittle, J.; Hogle, J.; Baltimore, D. Synthetic peptides from four separate regions of the poliovirus type 1 capsid protein VP1 induce neutralizing antibodies. Proc. Natl. Acad. Sci. USA 1985, 82, 910–914. [Google Scholar] [CrossRef] [PubMed]
- Oberste, M.S.; Maher, K.; Kilpatrick, D.R.; Pallansch, M.A. Molecular evolution of the human enteroviruses: correlation of serotype with VP1 sequence and application to picornavirus classification. J. Virol. 1999, 73, 1941–1948. [Google Scholar] [PubMed]
- Oberste, M.S.; Maher, K.; Kilpatrick, D.R.; Flemister, M.R.; Brown, B.A.; Pallansch, M.A. Typing of human enteroviruses by partial sequencing of VP1. J. Clin. Microbiol. 1999, 37, 1288–1293. [Google Scholar] [PubMed]
- Oberste, M.S.; Schnurr, D.; Maher, K.; al-Busaidy, S.; Pallansch, M.A. Molecular identificationof new picornaviruses and characterization of a proposed enterovirus 73 serotype. J. Gen. Virol. 2001, 82, 409–416. [Google Scholar] [CrossRef] [PubMed]
- Oberste, M.S.; Nix, W.A.; Maher, K.; Pallansch, M.A. Improved molecular identification of enteroviruses by RT–PCR and amplicon sequencing. J. Clin. Virol. 2003, 26, 375–377. [Google Scholar] [CrossRef]
- Casas, I.; Palacios, G.F.; Trallero, G.; Cisterna, D.; Freire, M.C.; Tenorio, A. Molecular characterization of human enteroviruses in clinical samples: comparison between VP2, VP1, and RNA polymerase regions using RT nested PCR assays and direct sequencing of products. J. Med. Virol. 2001, 65, 138–148. [Google Scholar] [CrossRef] [PubMed]
- Caro, V.; Guillot, S.; Delpeyroux, F.; Crainic, R. Molecular strategy for “serotyping”of human enteroviruses. J. Gen. Virol. 2001, 82, 79–91. [Google Scholar] [CrossRef] [PubMed]
- Norder, H.; Bjerregaard, L.; Magnius, L.O. Homotypic echoviruses share amino terminal VP1 sequence homology applicable for typing. J. Med. Virol. 2001, 63, 35–44. [Google Scholar] [CrossRef]
- Thoelen, I.; Lemey, P.; van der Donck, I.; Beuselink, K.; Lindberg, A.M.; van Ranst, M. Molecular typing and epidemiology of enteroviruses identified from an outbreak of aseptic meningitis in Belgium during the summer of 2000. J. Med. Virol. 2003, 70, 420–429. [Google Scholar] [CrossRef] [PubMed]
- Blomqvist, S.; Paananen, A.; Savolainen-Kopra, C.; Hovi, T.; Roivainen, M. Eight years’ experience of molecular identification of human enteroviruses. J. Clin. Microbiol. 2008, 46, 2410–2413. [Google Scholar] [CrossRef] [PubMed]
- Brown, B.; Oberste, M.S.; Maher, K.; Pallansch, M.A. Complete genome sequencing shows that polioviruses and members of human enterovirus species C are closely related in the noncapsid coding region. J. Virol. 2003, 77, 8973–8984. [Google Scholar] [CrossRef] [PubMed]
- Oberste, M.S.; Maher, K.; Flemister, M.R.; Marchetti, G.; Kilpatrick, D.R.; Pallansch, M.A. Comparison of classic and molecular approaches for the identification of untypeable enteroviruses. J. Clin. Microbiol. 2000, 38, 1170–1174. [Google Scholar] [PubMed]
- Delpeyroux, F.; Colbère-Garapin, F. Editorial commentary: Emerging problems impeding the elimination of the last polioviruses: Silent circulation of wild strains in a well-immunized population. Clin. Infect. Dis. 2015, 60, 1065–1067. [Google Scholar] [PubMed]
- Faleye, T.O.C.; Adeniji, J.A. Enterovirus Species B Bias of RD Cell Line and Its Influence on Enterovirus Diversity Landscape. Food Environ. Virol. 2015, 7, 390–402. [Google Scholar] [CrossRef] [PubMed]
- Faleye, T.O.C.; Adeniji, J.A. Nonpolio Enterovirus-C (NPEV-C) strains circulating in South-Western Nigeria and their contribution to the emergence of recombinant cVDPV2 lineages. Br. J. Virol. under review.
- Sadeuh-Mba, S.A.; Bessaud, M.; Massenet, D.; Joffret, M.L.; Endegue, M.C.; Njouom, R.; Reynes, J.M.; Rousset, D.; Delpeyroux, F. High frequency and diversity of species C enteroviruses in Cameroon and neighboring countries. J. Clin. Microbiol. 2013, 51, 759–770. [Google Scholar] [CrossRef] [PubMed]
- Adeniji, J.A.; Faleye, T.O.C. Impact of Cell Lines Included in Enterovirus Isolation Protocol on Perception of Nonpolio Enterovirus Species C Diversity. J. Virol. Methods 2014, 207, 238–247. [Google Scholar] [CrossRef] [PubMed]
- Nix, W.A.; Oberste, M.S.; Pallansch, M.A. Sensitive, Seminested PCR Amplification of VP1 Sequences for Direct Identification of All Enterovirus Serotypes from Original Clinical Specimens. J. Clin. Microbiol. 2006, 44, 2698–2704. [Google Scholar] [CrossRef] [PubMed]
- World Health Organisation. Enterovirus Surveillance Guidelines: Guidelines for Enterovirus Surveillance in Support of the Polio Eradication Initiative; World Health Organisation: Geneva, Switzerland, 2015. [Google Scholar]
- World Health Organisation. Guidelines for Environmental Surveillance of Poliovirus Circulation; World Health Organisation: Geneva, Switzerland, 2003. [Google Scholar]
- World Health Organisation. Polio Laboratory Manual, 4th ed.; World Health Organisation: Geneva, Switzerland, 2004. [Google Scholar]
- Arita, M.; Kilpatrick, D.R.; Nakamura, T.; Burns, C.C.; Bukbuk, D.; Oderinde, S.B.; Oberste, M.S.; Kew, O.M.; Pallansch, M.A.; Shimizu, H. Development of an efficient entire-capsid-coding-region amplification method for direct detection of poliovirus from stool extracts. J. Clin. Microbiol. 2015, 53, 73–78. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Adeniji, J.A.; University of Ibadan, Ibadan, Nigeria. Personal Communication, 2015.
- Faleye, T.O.C.; Adewumi, M.O.; Kareem, S.A.; Adesuyan, Y.O.; Fapohunda, F.A.; Fasanya, S.T.; Jimeto, T.; Lawrence, O.E.; Obembe, A.A.; Adeniji, J.A. The impact of a panenterovirus VP1 assay on our perception of the enterovirus diversity landscape of a sample. 2015; under peer-review. [Google Scholar]
- Rao, C.D.; Yergolkar, P.; Shankarappa, K.S. Antigenic Diversity of Enteroviruses Associated with Nonpolio Acute Flaccid Paralysis, India, 2007–2009. Emerg. Infect. Dis. 2012, 18, 1833–1840. [Google Scholar] [CrossRef] [PubMed]
- Tao, Z.; Wang, H.; Liu, Y.; Li, Y.; Jiang, P.; Liu, G.; Lin, X.; Li, M.; Wang, S.; Ji, F.; et al. Non-Polio Enteroviruses from Acute Flaccid Paralysis Surveillance in Shandong Province, China, 1988–2013. Sci. Rep. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Chevaliez, S.; Szendröi, A.; Caro, V.; Balanant, J.; Guillot, S.; Berencsi, G.; Delpeyroux, F. Molecular comparison of echovirus 11 strains circulating in Europe during an epidemic of multisystem hemorrhagic disease of infants indicates that evolution generally occurs by recombination. Virology 2004, 325, 56–70. [Google Scholar] [CrossRef] [PubMed]
- Paananen, A.; Savolainen-Kopra, C.; Kaijalainen, S.; Vaarala, O.; Hovi, T.; Roivainen, M. Genetic and phenotypic diversity of echovirus 30 strains and pathogenesis of type 1 diabetes. J. Med. Virol. 2007, 79, 945–955. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Grard, G.; Lukashev, A.N.; Kozlovskaya, L.I.; Böttcher, S.; Uslu, G.; Reimerink, J.; Gmyl, A.P.; Taty-Taty, R.; Lekana-Douki, S.E.; et al. Robustness against serum neutralization of a poliovirus type 1 from a lethal epidemic of poliomyelitis in the Republic of Congo in 2010. Proc. Natl. Acad. Sci. USA 2014, 11, 12889–12894. [Google Scholar] [CrossRef] [PubMed]
- Shulman, L.M.; Martin, J.; Sofer, D.; Burns, C.C.; Manor, Y.; Hindiyeh, M.; Gavrilin, E.; Wilton, T.; Moran-Gilad, J.; Gamzo, R.; et al. Genetic analysis and characterization of wild poliovirus type 1 during sustained transmission in a population with >95% vaccine coverage, Israel 2013. Clin. Infect Dis. 2015, 60, 1057–1064. [Google Scholar] [CrossRef] [PubMed]
- Al-Hello, H.; Paananen, A.; Eskelinen, M.; Ylipaasto, P.; Hovi, T.; Salmela, K.; Lukashev, A.N.; Bobegamage, S.; Roivainen, M. An enterovirus strain isolated from diabetic child belongs to a genetic subcluster of echovirus 11, but is also neutralised with monotypic antisera to coxsackievirus A9. J. Gen. Virol. 2008, 89, 1949–1959. [Google Scholar]
- Savolainen-Kopra, C.; Al-Hello, H.; Paananen, A.; Blomqvist, S.; Klemola, P.; Sobotova, Z.; Roivainen, M. Molecular epidemiology and dual serotype specificity detection of echovirus 11 strains in Finland. Virus Res. 2009, 139, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Adeniji, J.A.; Faleye, T.O.C. Isolation and identification of enteroviruses from sewage and sewage contaminated water in Lagos, Nigeria. Food Environ. Virol. 2014, 6, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Bessaud, M.; Jegouic, S.; Joffret, M.L.; Barge, C.; Balanant, J.; Gouandjika-Vasilache, I.; Delpeyroux, F. Characterization of the genome of human enteroviruses: Design of generic primers for amplification and sequencing of different regions of the viral genome. J. Virol. Methods 2008, 149, 277–284. [Google Scholar] [CrossRef] [PubMed]
- Lukashev, A.N. Recombination among picornaviruses. Rev. Med. Virol. 2010, 20, 327–337. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.F.; Naguib, T.; Yang, S.J.; Nasr, E.; Jorba, J.; Ahmed, N.; Campagnoli, R.; van der Avoort, H.; Shimizu, H.; Yoneyama, T.; et al. Circulation of endemic type 2 vaccine-derived poliovirus in egypt from 1983 to 1993. J. Virol. 2003, 77, 8366–8377. [Google Scholar] [CrossRef] [PubMed]
- Blomqvist, S.; Savolainen, C.; Laine, P.; Hirttiö, P.; Lamminsalo, E.; Penttilä, E.; Jöks, S.; Roivainen, M.; Hovi, T. Characterization of a highly evolved vaccine-derived poliovirus type 3 isolated from sewage in Estonia. J. Virol. 2004, 78, 4876–4883. [Google Scholar] [CrossRef] [PubMed]
- Arita, M.; Zhu, S.L.; Yoshida, H.; Yoneyama, T.; Miyamura, T.; Shimizu, H. A sabin 3-derived poliovirus recombinant contained a sequence homologous with indigenous human enterovirus species c in the viral polymerase coding region. J. Virol. 2005, 79, 12650–12657. [Google Scholar] [CrossRef] [PubMed]
- Adu, F.; Iber, J.; Bukbuk, D.; Gumede, N.; Yang, S.J.; Jorba, J.; Campagnoli, R.; Sule, W.F.; Yang, C.F.; Burns, C.C.; et al. Isolation of recombinant type 2 vaccine-derived poliovirus (vdpv) from a nigerian child. Virus Res. 2007, 127, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Rakoto-Andrianarivelo, M.; Guillot, S.; Iber, J.; Balanant, J.; Blondel, B.; Riquet, F.; Martin, J.; Kew, O.; Randriamanalina, B.; Razafinimpiasa, L.; et al. Co-circulation and evolution of polioviruses and species C enteroviruses in a district of Madagascar. PLoS Pathogen. 2007, 3, e191. [Google Scholar] [CrossRef] [PubMed]
- Rakoto-Andrianarivelo, M.; Gumede, N.; Jegouic, S.; Balanant, J.; Andriamamonjy, S.N.; Rabemanantsoa, S.; Birmingham, M.; Randriamanalina, B.; Nkolomoni, L.; Venter, M.; et al. Reemergence of recombinant vaccine-derived poliovirus outbreak in madagascar. J. Infect. Dis. 2008, 197, 1427–1435. [Google Scholar] [CrossRef] [PubMed]
- Combelas, N.; Holmblat, B.; Joffret, M.; Colbère-Garapin, F.; Delpeyroux, F. Recombination between Poliovirus and Coxsackie A Viruses of Species C: A Model of Viral Genetic Plasticity and Emergence. Viruses 2011, 3, 1460–1484. [Google Scholar] [CrossRef] [PubMed]
- Burns, C.C.; Shaw, J.; Jorba, J.; Bukbuk, D.; Adu, F.; Gumede, N.; Pate, M.A.; Abanida, E.A.; Gasasira, A.; Iber, J.; et al. Multiple independent emergences of type 2 vaccine-derived polioviruses during a large outbreak in Northern Nigeria. J. Virol. 2013, 87, 4907–4922. [Google Scholar] [CrossRef] [PubMed]
- Smura, T. The Evolution of New Enterovirus Types EV-94, EV-96 and EV-97. Doctoral Dissertation, University of Helsinki, Helsinki, Finland, 2012. [Google Scholar]
- Tatsuo, H.; Ono, N.; Tanaka, K.; Yanagi, Y. SLAM (CDw150) is a cellular receptor for measles virus. Nature 2000, 406, 893–897. [Google Scholar] [PubMed]
- Noyce, R.S.; Bondre, D.G.; Ha, M.N.; Lin, L.T.; Sisson, G.; Tsao, M.S.; Richardson, C.D. Tumor cell marker PVRL4 (nectin 4) is an epithelial cell receptor for measles virus. PLoS Pathog. 2011. [Google Scholar] [CrossRef] [PubMed]
- Mühlebach, M.D.; Mateo, M.; Sinn, P.L.; Prüfer, S.; Uhlig, K.M.; Leonard, V.H.; Navaratnarajah, C.K.; Frenzke, M.; Wong, X.X.; Sawatsky, B.; et al. Adherens junction protein nectin-4 is the epithelial receptor for measles virus. Nature 2011, 480, 530–533. [Google Scholar] [CrossRef] [PubMed]
- Newcombe, N.G.; Andersson, P.; Johansson, E.S.; Au, G.G.; Lindberg, A.M.; Barry, R.D.; Shafren, D.R. Cellular receptor interactions of C-cluster human group A coxsackieviruses. J. Gen. Virol. 2003, 84, 3041–3050. [Google Scholar] [CrossRef] [PubMed]
- Arita, M. Development of poliovirus extraction method from stool extracts by using magnetic nanoparticles sensitized with soluble poliovirus receptor. J. Clin. Microbiol. 2013, 51, 2717–2720. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.X.; Terasaki, T.; Shiroki, K.; Ohka, S.; Aoki, J.; Tanabe, S.; Nomura, T.; Terada, E.; Sugiyama, Y.; Nomoto, A. Efficient delivery of circulating poliovirus to the central nervous system independently of poliovirus receptor. Virology 1997, 229, 421–428. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Faleye, T.O.C.; Adewumi, M.O.; Adeniji, J.A. Defining the Enterovirus Diversity Landscape of a Fecal Sample: A Methodological Challenge? Viruses 2016, 8, 18. https://doi.org/10.3390/v8010018
Faleye TOC, Adewumi MO, Adeniji JA. Defining the Enterovirus Diversity Landscape of a Fecal Sample: A Methodological Challenge? Viruses. 2016; 8(1):18. https://doi.org/10.3390/v8010018
Chicago/Turabian StyleFaleye, Temitope Oluwasegun Cephas, Moses Olubusuyi Adewumi, and Johnson Adekunle Adeniji. 2016. "Defining the Enterovirus Diversity Landscape of a Fecal Sample: A Methodological Challenge?" Viruses 8, no. 1: 18. https://doi.org/10.3390/v8010018
APA StyleFaleye, T. O. C., Adewumi, M. O., & Adeniji, J. A. (2016). Defining the Enterovirus Diversity Landscape of a Fecal Sample: A Methodological Challenge? Viruses, 8(1), 18. https://doi.org/10.3390/v8010018






