Abstract
Retroviruses exploit nuclear trafficking machinery at several distinct stages in their replication cycles. In this review, we will focus primarily on nucleocytoplasmic trafficking events that occur after the completion of reverse transcription and proviral integration. First, we will discuss nuclear export of unspliced viral RNA transcripts, which serves two essential roles: as the mRNA template for the translation of viral structural proteins and as the genome for encapsidation into virions. These full-length viral RNAs must overcome the cell’s quality control measures to leave the nucleus by co-opting host factors or encoding viral proteins to mediate nuclear export of unspliced viral RNAs. Next, we will summarize the most recent findings on the mechanisms of Gag nuclear trafficking and discuss potential roles for nuclear localization of Gag proteins in retrovirus replication.
1. Introduction: Nuclear Trafficking Events in Retrovirus Replication
Retroviruses interact with nuclear trafficking machinery during several different phases of their replication cycles (Figure 1). Retrovirus replication has been divided into “early” and “late” stages, with early events extending from virus entry through integration and late stages encompassing expression of viral RNA from the provirus through virus assembly and budding of immature virus particles from the host cell. During early infection, all retroviruses must gain access to the host chromatin for the provirus to integrate. Most retroviruses depend on mitosis and breakdown of the nuclear envelope to undergo integration. However, lentiviruses like human immunodeficiency virus type 1 (HIV-1) infect non-dividing cells, having developed strategies to enter the nucleus by passing through intact nuclear pores (reviewed in [1,2,3]). Although several viral factors, including HIV-1 MA (matrix), IN (integrase), Vpr, and the reverse-transcribed proviral DNA have been implicated in nuclear entry of the pre-integration complex (PIC), more recent data indicate that the CA (capsid) protein, nuclear import factor transportin-3 (TNPO3), and the nucleoporin Nup358 are important determinants [4,5,6,7,8,9,10,11,12,13,14,15]. Thus, retroviruses have evolved different strategies to promote proviral integration through complex interactions between host and viral factors. Although these early events are essential to establish retroviral infection, this review will focus primarily on interactions of retroviruses with nuclear transport machinery following integration.
Once integration is completed, viral RNA is synthesized by the cellular RNA polymerase II. Retroviral transcripts are co-transcriptionally modified with the addition of a 5’ cap and 3’ poly(A) tail, like cellular mRNAs. A fraction of the viral RNA is spliced and exported out of the nucleus to serve as mRNA for translation into viral proteins. The remainder of the viral RNA remains unspliced and must be transported from the nucleus into the cytoplasm where it serves two functions: (i) as the template for translation of the viral structural proteins, Gag and Gag-Pol; and (ii) as the viral genome, which is packaged into virus particles. Because export of unspliced and incompletely spliced cellular RNAs is prevented by host machinery to prevent translation of abnormal proteins (reviewed in [17]), retroviruses must circumvent this cellular blockade to export their full-length RNA molecules (reviewed in [16,18]). Prior translation of the unspliced viral RNA is not a prerequisite for genome encapsidation, as genomic RNA can be packaged in trans (reviewed in [16]). The genomic RNA forms a noncovalent dimer and is encapsidated through an interaction between the psi (ψ) packaging sequence near the 5’ end of the genome and the NC (nucleocapsid) domain of the Gag protein.
The Gag protein, which directs the assembly and budding of virus particles from the plasma membrane, is localized primarily in the cytoplasm of infected cells. However, the Gag proteins of Rous sarcoma virus (RSV), feline immunodeficiency virus (FIV), mouse mammary tumor virus (MMTV), prototype foamy virus (PFV), murine leukemia virus (MLV), Mason-Pfizer monkey virus (MPMV), HIV-1, and several retrotransposons undergo nuclear localization under certain conditions (Table 1). The RSV, FIV, and PFV Gag proteins utilize the cellular CRM1 protein for nuclear export [19,20,21], but the host importins involved in nuclear import of Gag have only been defined for RSV. In this review, we will focus on nuclear transport events associated with the nucleocytoplasmic trafficking of unspliced retroviral RNAs and Gag proteins and their roles in virion assembly.

Figure 1.
Model of retrovirus replication. Retroviral infection is initiated with binding of the viral Env protein to a cell surface receptor and fusion of the viral envelope with the cellular membrane (step 1). The viral RNA genome is reverse transcribed from RNA to DNA (step 2) to form the provirus, which is stably integrated into the genome of the host cell (step 3). Viral RNA is transcribed by the host polymerase II, and a portion of the RNA is spliced to direct the translation of the Env glycoprotein and other viral proteins (step 4). A portion of the viral RNA remains unspliced and is exported from the nucleus by the host factor TAP/NXF1 (e.g., simple retroviruses MPMV and RSV) or virally encoded Rev-like proteins (e.g., complex retroviruses like HIV-1 and HTLV-I) (step 5) to serve as a template for Gag and Gag-Pol translation (step 6). The Gag proteins of some retroviruses traffic through the nucleus during assembly (step 7). It has been postulated that the nuclear population of Gag (denoted by “?”) might select genomic RNA (gRNA) and transport it into the cytoplasm for packaging. Alternatively, selection of genomic RNA may occur in the cytoplasm (step 8). In either case, the Gag-gRNA complex is transported to the plasma membrane (step 9) where additional Gag molecules bind the viral RNP to complete assembly of the immature virus particle, which buds from the plasma membrane (step 10). The steps in replication that are covered in this review are indicated in bold. Figure modified from [22] and used with the author’s permission.

Table 1.
Retroviral proteins that undergo nuclear trafficking*
| Retroviral Protein | Localization of the Population Associated with the Nuclear Compartment | Nuclear Localization Mechanism | Nuclear Export Mechanism |
|---|---|---|---|
| Alpharetrovirus | |||
| RSV Gag | Nucleus [
20,23] Punctate Nuclear Foci [23,24] Nucleoli [24] | Imp11 (MA domain) [
25,26] TNPO3 (MA Domain) [26] Importin α/β (NC Domain) [25,26] | CRM1 [ 20,25] |
| RSV NC | Nucleoli [24] | Importin α/β [25,26] | None |
| RSV RT, β subunit | Nucleus [27] | Unknown | None |
| RSV IN | Nucleus [28] | Shares Import Pathway with Histone H1 [29] | None |
| Betaretrovirus | |||
| MMTV Gag | Nucleoli [30] | Unknown [30]; Nucleolar Localization Increased with RPL9 Overexpression [30] | No Identified NES |
| MMTV NC | Nucleoli [24,30] | Unknown; | None |
| MMTV Rem | Nucleoli [31] | Retrotranslocation from Endoplasmic Reticulum to Nucleus [32] | CRM1 [31] |
| MPMV Gag | Nuclear Pore Complex; Low Levels in Nucleus [33,34] | Unknown; Nuclear localization Increased with Ubc9 Overexpression [34] | CRM1 [35] |
| JSRV Rej | Nucleus and Nucleoli [36,37] | Unknown | Rej Function Depends on CRM1 [37] |
| Gammaretrovirus | |||
| MLV Gag | Nucleus [38] | Unknown | No Identified NES |
| MLV NC | Nucleus [24,39] Nucleoli [24,39] | Unknown | None |
| MLV p12 | Mitotic Chromatin [40,41] | Unknown | None |
| MLV IN | Nucleus and Nucleoli [39] | Unknown; Interacts withBrd4 in Nucleus [42] | None |
| Deltaretrovirus | |||
| HTLV Rex | Nucleus and Nucleoli [43,44,45] Punctate Nuclear and Nucleolar Foci [43] | Importin β [46] | CRM1 [47,48] |
| BLV Rex | Punctate Nuclear Foci [49] Nuclear Pore Complex [49] | Unknown | CRM1 [49] |
| Lentivirus | |||
| HIV-1 Gag | Nucleolar [24] | Unknown | Not CRM1 [19,35,50] |
| HIV-1 IN | Nucleus [14,51,52,53,54,55] | Importin α3 [51] Importin α [14,52] Importin 7 [56] | None |
| HIV-1 NC | Nucleoli [24] Nucleus [57,58] | Unknown | None |
| HIV-1 Rev | Nucleus andNnucleoli [44,59] Nuclear Foci [59] HIV-1 Transcription Sites [60] | Importin β [61,62] Transportin, Importin 5, and Importin 7 [63] | CRM1 [47,64] |
| HIV-1 Vpr | Nucleus [65,66] Nuclear Pore Complex [6,66] | Importin α [66] Interacts Directly with Nuclear Pore Complex [67,68] | CRM1 [65] |
| HIV-2 Rev | Nucleoli [69] | Unknown | Unknown |
| HIV-2 Vpx | Nucleoplasm [70,71] | Unknown | No Identified NES |
| FIV Gag | Nucleus [19], Nucleoli [19], and Nuclear Envelope [72] | Unknown | CRM1 [19] |
| FIV Rev | Nucleolus [73] | Unknown | CRM1 [74] |
| EAIV Rev | Nucleus [74] | Unknown | CRM1 [74] |
| BIV Rev | Nucleus and Nucleoli [75,76] | Importin α [77] | CRM1 [77] |
| MVV Rev | Nucleoli [78] | Unknown | Unknown |
| CAEV Rev | Nucleoli [79,80] | Unknown | Unknown |
| Spumaretrovirus | |||
| PFV Gag | Nucleus [21,81] Punctate Nuclear Foci [82,83] Mitotic Chromatin [82] | Binds to H2A/H2B on Mitotic Chromatin [82,83] | CRM1 [21] |
* Note: Transcriptional activators related to HIV-1 Tat were not included in the table.
2. Nuclear Export of Unspliced and Incompletely Spliced RNAs of Complex Retroviruses
Productive retroviral infection requires unspliced viral transcripts to be transported into the cytoplasm where they are translated into the essential viral proteins Gag and Gag-Pol. To circumvent intrinsic cellular blockades that prevent the export of incompletely spliced RNAs from the nucleus, complex retroviruses encode trans-acting viral proteins that export their intron-containing viral RNAs from the nucleus. HIV-1 Rev was the first member of this family to be discovered; however, Rev-like proteins have been described in the Lentivirus [e.g., Rev proteins of human immunodeficiency virus type-2 (HIV-2), simian immunodeficiency virus (SIV), FIV, equine infectious anemia virus (EIAV), bovine immunodeficiency virus (BIV), Maedi-visna virus (MVV) and caprine encephalitis-anemia virus, CAEV)] [73,84,85,86,87,88,89,90,91,92,93], Deltaretrovirus [(e.g., Rex proteins of human T cell leukemia virus type-I (HTLV-I) and bovine leukemia virus (BLV)], and Betaretrovirus [e.g., Rem protein of MMTV and Rej protein of Jaagsiekte sheep retrovirus (JSRV)] genera [31,36,37,77,94,95,96]. Rev-like proteins localize to the nucleus and nucleolus through interactions with a variety of import factors (see (1), and they contain CRM1-dependent nuclear export signals (NESs) [45,47,77,97,98,99,100].
HIV-1 Rev recognizes and binds to the highly structured cis-acting Rev-responsive element (RRE) in HIV-1 RNAs [101,102,103,104] and undergoes multimerization, which is important for its export function (reviewed in [105]). Multimerization of Rev was demonstrated within the nucleolus in living cells, suggesting that the nucleolus may be the site of Rev multimer formation [106]. However, the relevance of these experiments is somewhat limited because they were not performed in the context of HIV-1 infection, nor was the RRE sequence expressed in the cells. Attempts to define whether nucleolar localization of Rev is important for binding to RRE-containing RNAs have been difficult because the Rev RNA binding domain overlaps with the nuclear/nucleolar localization signal. However, HIV-1 unspliced RNA appears to undergo nucleolar trafficking based on the finding that the RNA is cleaved by ribozymes artificially targeted to the nucleolus [107] and small nucleolar RNAs engineered to contain the RRE are exported into the cytoplasm by Rev [108,109]. Together, these data provide evidence that Rev and the RRE-containing viral transcripts both traffic through the nucleolus, but there is no definitive evidence that nucleolar trafficking of the Rev-RRE complex is essential for nuclear export of HIV-1 RNA during natural virus infection.
In the nucleoplasm, Rev co-localizes with the SR-domain splicing factor SC-35 in nuclear speckles [59,110], intrachromatin granule clusters enriched in mRNA splicing factors and snRNPs located adjacent to sites of active transcription [111,112,113,114,115]. The association of Rev with splicing factors in speckles suggests that Rev binds to the RRE co-transcriptionally, just as splicing factors bind to cellular transcripts as they are synthesized [116]. In support of this idea, previous studies demonstrated that the Rev-RRE interaction is abrogated by transcription inhibitors [117]. More recently, it was shown that Rev co-localizes with HIV-1 RNA at transcription sites, providing strong evidence that Rev binds RRE-containing transcripts co-transcriptionally [60].
HIV-1 Rev mediates export of unspliced viral RNAs through an interaction between the NES of Rev and the CRM1/RanGTP export complex. However, many other cellular proteins also interact with Rev, indicating that nuclear export of unspliced HIV-1 RNA depends upon a complex network of interactions [60,62,105,118,119,120,121,122,123,124,125,126,127]. For example, several RNA helicases interact with Rev, including DDX1 and DDX3, which help to maintain Rev localization in the nucleolus and nucleus [124,126,127,128]. In addition, the nuclear matrix-associated protein Matrin 3 binds HIV-1 RNA co-transcriptionally and facilitates Rev-mediated nuclear export of unspliced RNA [60,129]. Although the precise mechanism by which Matrin 3 facilitates HIV-1 RNA export has not been elucidated, Matrin 3 increases the stability and expression of cellular mRNAs, suggesting that it may have a similar effect on HIV-1 RNA [130]. The finding that Matrin 3 interacts with HIV-1 RNA also creates an intriguing connection between HIV-1 RNA export and the nuclear matrix, which not only provides structural support to the nucleus [131] but also associates with areas of euchromatin involved in ongoing transcription [132]. Recent evidence also suggests that components of the nuclear matrix called nuclear regulatory networks bind genomic DNA and form a tubular pathway leading to nuclear pore complexes for nuclear export of transcripts and proteins [133,134]. Therefore, it is intriguing to postulate that Matrin 3 bridges the interaction between Rev and active HIV-1 RNA transcription sites [60,129,135,136], recruiting the CRM1 nuclear export machinery associated with nuclear regulatory networks to transport viral ribonucleoprotein complexes (RNPs) through the nuclear pore and into the cytoplasm.
3. The Role of cis-Acting Sequences in RNA Export of Simple Retroviruses
In contrast to complex retroviruses that encode trans-acting factors to facilitate nuclear export of unspliced RNA, simple retroviruses have evolved cis-elements to circumvent the blockade to export of unspliced transcripts from the nucleus. MPMV and other type D retroviruses, including simian retrovirus-1 and 2 (SRV-1 and SRV-2), contain a small cis element, the constitutive transport element (CTE), which is required for nuclear export of unspliced viral RNA [137,138]. When inserted into unspliced or incompletely spliced HIV-1 transcripts, the MPMV CTE sequence replaces the function of the Rev/RRE complex, leading to expression of Gag and Env followed by the production of infectious virus particles [137]. Thus, Rev/RRE and the CTE provide similar roles in the nuclear export of unspliced RNA in complex and simple retroviruses.
Insight into the mechanism by which CTE-containing RNAs are exported from the nucleus was provided by proteomic studies that identified the host nuclear export protein Tip-associating protein/Nuclear RNA export factor 1 (TAP/NXF1) as a binding partner of CTE complexes [139,140]. Microinjection of Xenopus oocyte nuclei expressing TAP/NXF1 and an intron containing the CTE resulted in nuclear export of the RNA in the absence of splicing [141,142]. The TAP/NXF1 protein, homologous to the mRNA export protein Mex67p in yeast, forms a heterodimer with NXT1 to transport mRNAs out of the nucleus [139,140,143,144,145]. The N-terminal domain of TAP/NXF1 contains an RNA recognition motif that binds to a structured stem-loop in the CTE, inducing structural changes in both TAP/NXF1 and the CTE-containing RNA to promote nuclear export of the viral RNP [146]. Mutations in the RNA or in the coding region of TAP/NXF1 that disrupt CTE-TAP/NXF1 complex formation prevent expression of CTE-containing reporters in vivo [146].
A putative cis-acting unspliced RNA transport element was also identified in RSV, which lies within 115 nucleotide direct repeat (DR) sequences flanking the v-src oncogene [147]. DR elements are highly conserved in avian retroviruses [148], and strains missing the src sequence maintain at least a single DR element to remain replication-competent [149,150]. The biological role of the DR elements is complex; pleotropic, contradictory effects on virus replication have been reported, including differences in levels of cytoplasmic accumulation of RSV RNA, viral RNA stability, expression of the Gag polyprotein, viral RNA packaging and virus assembly [148,151,152,153]. These conflicting results may be explained by differences in cell types or the use of subviral reporter constructs in some studies and full-length, replication-competent viruses in others.
RSV RNAs containing the DR elements are exported by the cellular mRNA export factor TAP/NXF1 and the RNA helicase Dbp5 [139,154,155]. An additional host factor may bridge the interaction because neither TAP/NXF1 nor Dbp5 bind the DR element directly. Because the RSV Gag protein was reported to traffic through the nucleus [20], LeBlanc et al. used a subviral reporter construct containing either the DR sequence or the ψ sequence to examine whether Gag could enhance translation by promoting nuclear export of unspliced RNA [155]. Gag did not enhance translation of the reporter; however, nucleocytoplasmic fractionation of the RNA was not performed, so it is unclear whether Gag had an effect on cytoplasmic levels of the DR- or ψ-containing RNAs. Thus, these experiments suggest that DR elements mediate nuclear export through TAP/NXF1 and Dbp5 to stimulate translation of RSV unspliced RNA, but Gag is not likely to be involved in DR-mediated RNA transport.
Taking the available data into account, we postulate that there is a temporal switch in RSV replication, such that viral transcripts produced early after integration are exported using DR-mediated interactions with Tap/NXF1 and Dbp5 to initiate the synthesis of Gag and GagPol proteins. We hypothesize that as the levels of these viral structural proteins increase, the Gag protein enters the nucleus where it may bind unspliced viral RNA and export it into the cytoplasm for encapsidation into virions [20,156]. It is possible that other simple retroviruses may use a similar mechanism. We speculate that MPMV RNA export could be similarly regulated, since a subpopulation of MPMV Gag localizes to the nucleus and nuclear envelope, and the pp24 domain NLS has been linked to genomic RNA incorporation [33,34,35]. Complex retroviruses regulate this temporal switch between early and late gene expression differently; the Rev protein is synthesized from a fully spliced mRNA and then traffics into the nucleus to promote nucleocytoplasmic transport of unspliced RNAs for structural gene expression and genome packaging.
4. Foamy Viruses Use a Unique Pathway among Retroviruses for Nuclear Export of Viral RNAs
FVs, members of the genus Spumavirus, share similarities with both simple and complex retroviruses, yet they have several unique characteristics (reviewed in [157,158]). Unlike other retroviruses, Gag is the only protein translated from an unspliced transcript [159,160,161]. Additionally, instead of being expressed as a fusion protein with Gag using frameshifting or termination codon suppression, Pol is expressed from a separate spliced viral mRNA [162]. Similar to complex retroviruses however, FVs encode the transcriptional transactivator protein Tas, which functions analogously to HIV-1 Tat [163]. FVs do not encode an accessory protein with Rev-like functionality [164], therefore, FVs rely entirely on host factors to mediate the export of unspliced RNA from the nucleus, similar to the simple retroviruses MPMV and RSV.
Among retroviruses, PFV utilizes a unique set of cellular factors that bind to its viral RNA for nucleocytoplasmic transport. Rather than using TAP/NXF1, like the cis-acting RSV and MPMV RNA transport elements [139,155], spliced and unspliced PFV RNAs interact with host proteins HuR and ANP32A/B to exit the nucleus through the CRM1 pathway [165]. HuR associates with PFV RNAs and the adaptor proteins ANP32A or ANP32B, which bridge the association with CRM1 [165,166,167]. The HuR-interacting RNA sequence likely resides in the 3’ region of the genome, which is shared among spliced and unspliced PFV RNAs [165], although the PFV 3’ end does not share homology with previously-characterized RNA sequences that bind HuR [165]. Although both the spliced and unspliced PFV transcripts appear to be exported by the same pathway, subsequent interaction with the cytoplasmic mRNA processing factor DDX6 may distinguish the genomic RNA from the viral transcripts earmarked for translation on polysomes [168]. It is not yet known whether all PFV RNPs exported by HuR-ANP32A/B-CRM1 traffic to the same subcellular location where DDX6 co-localizes with genomic RNA in association with Gag. However, having spliced and unspliced RNAs targeted to the same cytoplasmic localization may not pose a problem for selective genomic RNA selection by PFV Gag because both segments of the bipartite cis packaging signal are present only on unspliced viral RNA [169].
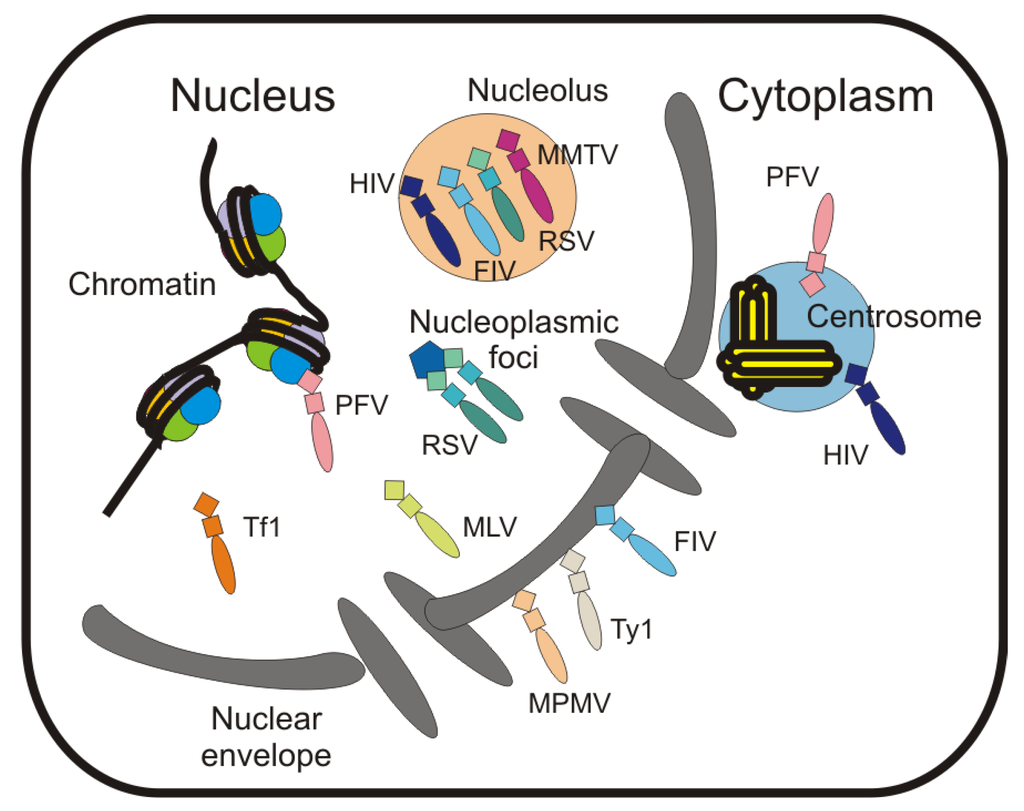
7. Localization of Gag to Subnuclear Sites
The Gag proteins of RSV and HIV-1 localize to the nucleolus under certain conditions [24] (Figure 2), and nucleolar trafficking of RSV Gag is dependent on basic residues in the NC domain. For RSV NC NoLS activity resides in residues 36–39 (KKRK), 61–63 (RKR) and 70R/73K. By contrast, the HIV-1 NC domain contains two independent NoLSs (R10/K11 and R32/K33/K34), each of which is sufficient for nucleolar localization of NC [24]. HIV-1 NC localizes to the nucleus during infection [57,58], but it is not clear whether nucleolar localization of the mature NC protein plays an important role during early infection. It is intriguing that many capsid and nucleic acid binding proteins from diverse viruses localize to the nucleolus, prompting the nucleolus to be referred to as “the gateway to viral infection" [19,24,26,39,57,58,207].
Due to the transient nature of its trafficking through the nucleus, detecting RSV Gag in nucleoli requires disruption of the nuclear export activity of Gag, either by mutating the p10 NES or treating infected cells with LMB [24]. The retention of Gag in nucleoli is decreased in the presence of the RSV Ψ packaging signal, suggesting that Gag binding to the packaging signal may either induce the Gag-RNA complex to leave the nucleolus or may prevent trafficking of the RSV RNP through the nucleolus. Whether there is a host protein or RNA that binds to RSV Gag-RNA complexes in the nucleolus is unknown. A subset of the HIV-1 Gag protein also localizes to nucleoli when the provirus is expressed at high levels from an inducible, Rev-dependent, integrated provirus. HIV-1 Gag accumulates in nucleoli when co-expressed with Rev or NC, and positive FRET (fluorescence resonance energy transfer) was observed between Gag and Rev, indicating an intimate association between these proteins within the nucleolus [24]. Whether the nucleolar localization of HIV-1 Gag or the interaction between Gag and Rev has any significant role in genomic RNA encapsidation or virus replication remains to be determined.
Other retroviral Gag proteins have been reported to localize to the nucleolus under certain conditions. For example, LMB treatment of transfected cells causes FIV Gag to accumulate in nucleoli when expressed alone and nucleolar localization was dependent on NC [19] (Figure 2). However, the importance of Gag or NC nucleolar localization was not further investigated. In addition, MMTV Gag is partially nucleolar-localized in a murine mammary cell line chronically infected with a highly tumorigenic strain of the virus [30]. Furthermore, a subset of MMTV Gag can be induced to accumulate in the nucleolus with overexpression of ribosomal protein L9, which interacts with MMTV Gag in an extraribosomal context (Figure 2). A functional role for this interaction was demonstrated by knockdown of L9, which causes a decrease in MMTV particle production [30]. Thus, it is possible that nucleolar trafficking of MMTV Gag is an important step in virus assembly pathway, although further experiments will be needed to test this intriguing idea.
In addition to localizing to the nucleolus, the RSV Gag protein also accumulates in discrete nucleoplasmic foci when restricted to the nucleus by mutating the NES or inhibiting CRM1-mediated nuclear export [20,23] (Figure 2). It will be interesting to determine whether RSV Gag is associated with a particular subnuclear body or tethered to a specific cellular protein or RNA at these sites. The appearance of these subnuclear foci of RSV Gag is similar to the localization of Spumaretrovirus PFV Gag in nucleoplasmic foci during prophase [82]. In the case of PFV, it was recently discovered that the Gag protein binds to the host chromosome in mitotic cells using a chromatin-tethering domain encoded in the glycine-arginine box II domain, which binds to core histones H2A and H2B [82]. The PFV Gag-chromatin interaction facilitates proviral integration, although the precise mechanism has not been defined. In addition, a CRM1-dependent nuclear export pathway was reported for PFV Gag, although this finding is controversial [21,83]. An intriguing idea not yet experimentally tested is whether the association PFV Gag with the proviral integration site [82] could position Gag at the site of PFV RNA synthesis to facilitate selection of genomic RNA. Additional studies are required to determine whether PFV Gag nuclear localization plays a role in genomic RNA encapsidation or whether its function is restricted to early events, when it facilitates integration of the provirus.
8. Remaining Questions about Nuclear Trafficking during Retrovirus Assembly
In the past decade, great strides have been made in understanding the mechanisms that guide nuclear import, nuclear export, and the intranuclear activities of retroviral and retrotransposon Gag proteins. The use of a comparative approach to identify common and unique features of different Gag proteins in the nucleus has been very illuminating. Although it is not yet clear whether all Gag proteins transit through the nucleus at some low level, examination of the roles of nucleocytoplasmic trafficking in those Gag proteins that do enter the nucleus or associate with the nuclear envelope (RSV, MPMV, MLV, HIV-1, FIV, PFV, Ty1 and Tf1) are important to pursue.
Is Gag nuclear trafficking involved in genomic RNA packaging? To date, studies of RSV, MPMV, and Ty1 support this idea. PFV Gag associates with chromatin and plays a role in integration, although it might also function in genome encapsidation of nuclear transcripts [21,83]. In simple retroviruses, there could be a temporal switch with unspliced viral RNA export mediated by Tap/NXF1 for translation of structural proteins early after integration. As Gag proteins accumulate, they could enter the nucleus and export viral RNA for packaging. Thus, could there be two pathways of nuclear export of unspliced viral RNAs, or two distinct viral RNPs, one primed for packaging and the other for translation? If so, it is logical to suggest that the cytoplasmic fates of retroviral RNAs may be pre-determined in the nucleus according to the composition of the RNP.
Finding that several retroviral Gag proteins undergo nucleolar localization raises intriguing questions about whether there is a unifying function of the nucleolus in virus replication. One possibility is that retroviral Gag proteins interact with a host protein or RNA in the nucleolus involved in genome packaging or virus assembly. It is intriguing that several RNA pol III transcripts are enriched in retroviral particles, including RNAs that are processed in the nucleolus such as tRNA, U6 small nuclear RNA, 7SL [208,209], and 5S rRNA, [187,210,211,212,213,214,215,216]. Whether these RNAs are recruited into virus particles as passengers encountered along the subcellular trafficking route of Gag or whether they play a facilitating role in virus replication remains to be seen. As an interesting connection with endogenous retroelements and the nucleolus, there is strong genetic evidence that the nucleolus may be the site of RNP maturation for the LINE (long-interspersed nuclear element) retrotransposons, with U6 potentially playing a central role [217]. Thus, further investigation into the role of nuclear machinery in retrovirus and retrotransposon RNA processing, nuclear export, and RNP trafficking may generate new paradigms about genomic RNA encapsidation and virus particle assembly.
Acknowledgments
This work was supported by grants from the National Institutes of Health R01CA76534 (LJP), P50GM103297 (LJP), F30 CA165774 (DVB), F31 CA171862 (RJK) and CURE Funding from the Pennsylvania Department of Health (LJP). The Pennsylvania Department of Health specifically disclaims responsibility for any analyses, interpretations, or conclusions of this project.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Jayappa, K.D.; Ao, Z.; Yao, X. The HIV–1 passage from cytoplasm to nucleus: the process involving a complex exchange between the components of HIV–1 and cellular machinery to access nucleus and successful integration. IJBMB 2012, 3, 70–85. [Google Scholar]
- Sherman, M.P.; Greene, W.C. Slipping through the door: HIV entry into the nucleus. Microbes Infect. 2002, 4, 67–73. [Google Scholar] [CrossRef]
- Suzuki, Y.; Craigie, R. The road to chromatin – nuclear entry of retroviruses. Nat. Rev. Microbiol. 2007, 5, 187–96. [Google Scholar] [CrossRef]
- Stetor, S.R.; Rausch, J.W.; Guo, M.J.; Burnham, J.P.; Boone, L.R.; Waring, M.J.; Le Grice, S.F. Characterization of (+) strand initiation and termination sequences located at the center of the equine infectious anemia virus genome. Biochem. 1999, 38, 3656–3667. [Google Scholar] [CrossRef]
- Charneau, P.; Mirambeau, G.; Roux, P.; Paulous, S.; Buc, H.; Clavel, F. HIV–1 reverse transcription. A termination step at the center of the genome. J. Mol. Biol. 1994, 241, 651–662. [Google Scholar] [CrossRef]
- Fouchier, R.A.; Meyer, B.E.; Simon, J.H.; Fischer, U.; Albright, A.V.; Gonzalez–Scarano, F.; Malim, M.H. Interaction of the human immunodeficiency virus type 1 Vpr protein with the nuclear pore complex. J. Virol. 1998, 72, 6004–6013. [Google Scholar]
- Haffar, O.K.; Popov, S.; Dubrovsky, L.; Agostini, I.; Tang, H.; Pushkarsky, T.; Nadler, S.G.; Bukrinsky, M. Two nuclear localization signals in the HIV–1 matrix protein regulate nuclear import of the HIV–1 pre–integration complex. J. Mol. Biol. 2000, 299, 359–368. [Google Scholar] [CrossRef]
- Krishnan, L.; Matreyek, K.A.; Oztop, I.; Lee, K.; Tipper, C.H.; Li, X.; Dar, M.J.; Kewalramani, V.N.; Engelman, A. The requirement for cellular transportin 3 (TNPO3 or TRN–SR2) during infection maps to human immunodeficiency virus type 1 capsid and not integrase. J. Virol. 2010, 84, 397–406. [Google Scholar] [CrossRef]
- Lee, K.; Ambrose, Z.; Martin, T.D.; Oztop, I.; Mulky, A.; Julias, J.G.; Vandegraaff, N.; Baumann, J.G.; Wang, R.; Yuen, W.; Takemura, T.; Shelton, K.; Taniuchi, I.; Li, Y.; Sodroski, J.; Littman, D.R.; Coffin, J.M.; Hughes, S.H.; Unutmaz, D.; Engelman, A.; KewalRamani, V.N. Flexible use of nuclear import pathways by HIV–1. J. Chom. 2010, 7, 221–333. [Google Scholar]
- De Iaco, A.; Luban, J. Inhibition of HIV–1 infection by TNPO3 depletion is determined by capsid and detectable after viral cDNA enters the nucleus. Retrovirol. 2011, 8, 98. [Google Scholar] [CrossRef]
- Yamashita, M.; Perez, O.; Hope, T.J.; Emerman, M. Evidence for direct involvement of the capsid protein in HIV infection of nondividing cells. PLoS Pathog. 2007, 3, 1502–1510. [Google Scholar]
- Yamashita, M.; Emerman, M. Capsid is a dominant determinant of retrovirus infectivity in nondividing cells. J. Virol. 2004, 78, 5670–5678. [Google Scholar] [CrossRef]
- Zennou, V.; Petit, C.; Guetard, D.; Nerhbass, U.; Montagnier, L.; Charneau, P. HIV–1 genome nuclear import is mediated by a central DNA flap. Cell 2000, 101, 173–185. [Google Scholar] [CrossRef]
- Gallay, P.; Hope, T.; Chin, D.; Trono, D. HIV–1 infection of nondividing cells through the recognition of integrase by the importin/karyopherin pathway. Proc. Natl. Acad. Sci. U.S.A. 1997, 94, 9825–9830. [Google Scholar] [CrossRef]
- Schaller, T.; Ocwieja, K.E.; Rasaiyaah, J.; Price, A.J.; Brady, T.L.; Roth, S.L.; Hue, S.; Fletcher, A.J.; Lee, K.; KewalRamani, V.N.; Noursadeghi, M.; Jenner, R.G.; James, L.C.; Bushman, F.D.; Towers, G.J. HIV–1 capsid–cyclophilin interactions determine nuclear import pathway, integration targeting and replication efficiency. PLoS Pathog. 2011, 7, e1002439. [Google Scholar] [CrossRef]
- Butsch, M.; Boris–Lawrie, K. Destiny of unspliced retroviral RNA: ribosome and/or virion? J. Virol. 2002, 76, 3089–3094. [Google Scholar] [CrossRef]
- Schmid, M.; Jensen, T.H. Quality control of mRNP in the nucleus. Chromosoma 2008, 117, 419–429. [Google Scholar] [CrossRef]
- D'Souza, V.; Summers, M.F. How retroviruses select their genomes. Nat. Rev. Microbiol. 2005, 3, 643–655. [Google Scholar] [CrossRef]
- Kemler, I.; Saenz, D.; Poeschla, E. Feline immunodeficiency virus Gag is a nuclear shuttling protein. J. Virol. 2012, 86, 8402–8411. [Google Scholar] [CrossRef]
- Scheifele, L.Z.; Garbitt, R.; Rhoads, J.D.; Parent, L.J. Nuclear entry and CRM1–dependent nuclear export of the Rous sarcoma virus Gag polyprotein. Proc. Natl. Acad. Sci. U.S.A. 2002, 99, 3944–3949. [Google Scholar]
- Renault, N.; Tobaly–Tapiero, J.; Paris, J.; Giron, M.L.; Coiffic, A.; Roingeard, P.; Saib, A. A nuclear export signal within the structural Gag protein is required for prototype foamy virus replication. Retrovirol. 2011, 8, 6. [Google Scholar] [CrossRef]
- Bann, D.V. Intracellular Trafficking of Retroviral Gag and RNA During Late Replication. Doctoral Dissertation, The Pennsylvania State University College of Medicine, Hershey, PA, USA, 2013. [Google Scholar]
- Kenney, S.P.; Lochmann, T.L.; Schmid, C.L.; Parent, L.J. Intermolecular Interactions between Retroviral Gag Proteins in the Nucleus. J. Virol. 2007, 82, 683–691. [Google Scholar]
- Lochmann, T.L.; Bann, D.V.; Ryan, E.P.; Beyer, A.R.; Mao, A.; Cochrane, A.; Parent, L.J. NC–mediated nucleolar localization of retroviral gag proteins. Virus Res. 2013, 171, 304–318. [Google Scholar] [CrossRef]
- Gudleski, N.; Flanagan, J.M.; Ryan, E.P.; Bewly, M.C.; Parent, L.J. Directionality of nucleocytoplasmic transport of the retroviral gag protein depends on sequential binding of karyopherins and viral RNA. Proc. Natl. Acad. Sci. U.S.A. 2010, 107, 9358–9363. [Google Scholar]
- Butterfield–Gerson, K.L.; Scheifele, L.Z.; Ryan, E.P.; Hopper, A.K.; Parent, L.J. Importin–beta Family Members Mediate Alpharetrovirus Gag Nuclear Entry via Interactions with Matrix and Nucleocapsid. J. Virol. 2006, 80, 1798–1806. [Google Scholar] [CrossRef]
- Werner, S.; Hindmarsh, P.; Napirei, M.; Vogel–Bachmayr, K.; Wohrl, B.M. Subcellular localization and integration activities of rous sarcoma virus reverse transcriptase. J. Virol. 2002, 76, 6205–6212. [Google Scholar] [CrossRef]
- Kukolj, G.; Jones, K.S.; Skalka, A.M. Subcellular localization of avian sarcoma virus and human immunodeficiency virus type 1 integrases. J. Virol. 1997, 71, 843–847. [Google Scholar]
- Andrake, M.D.; Sauter, M.M.; Boland, K.; Goldstein, A.D.; Hussein, M.; Skalka, A.M. Nuclear import of Avian Sarcoma Virus integrase is facilitated by host cell factors. Retrovirol. 2008, 5, 73. [Google Scholar] [CrossRef]
- Beyer, A.R.; Bann, D.V.; Rice, B.; Pultz, I.S.; Kane, M.; Goff, S.P.; Golovkina, T.V.; Parent, L.J. Nucleolar trafficking of the mouse mammary tumor virus gag protein induced by interaction with ribosomal protein L9. J. Virol. 2013, 87, 1069–1082. [Google Scholar] [CrossRef]
- Indik, S.; Gunzburg, W.H.; Salmons, B.; Rouault, F. A novel, mouse mammary tumor virus encoded protein with Rev–like properties. Virology 2005, 337, 1–6. [Google Scholar] [CrossRef]
- Byun, H.; Halani, N.; Mertz, J.A.; Ali, A.F.; Lozano, M.M.; Dudley, J.P. Retroviral Rem protein requires processing by signal peptidase and retrotranslocation for nuclear function. Proc. Natl. Acad. Sci. U.S.A. 2010, 107, 12287–12292. [Google Scholar]
- Bohl, C.; Brown, S.; Weldon, R. The pp24 phosphoprotein of Mason–Pfizer monkey virus contributes to viral genome packaging. Retrovirol. 2005, 2, 68. [Google Scholar] [CrossRef]
- Weldon Jr, R.A.; Sarkar, P.; Brown, S.M.; Weldon, S.K. Mason–Pfizer monkey virus Gag proteins interact with the human sumo conjugating enzyme, hUbc9. Virology 2003, 314, 62–73. [Google Scholar] [CrossRef]
- Baluyot, M.F.; Grosse, S.A.; Lyddon, T.D.; Janaka, S.K.; Johnson, M.C. CRM1–dependent trafficking of retroviral Gag proteins revisited. J. Virol. 2012, 86, 4696–4700. [Google Scholar] [CrossRef]
- Hofacre, A.; Nitta, T.; Fan, H. Jaagsiekte sheep retrovirus encodes a regulatory factor, Rej, required for synthesis of Gag protein. J. Virol. 2009, 83, 12483–12498. [Google Scholar] [CrossRef]
- Caporale, M.; Arnaud, F.; Mura, M.; Golder, M.; Murgia, C.; Palmarini, M. The signal peptide of a simple retrovirus envelope functions as a posttranscriptional regulator of viral gene expression. J. Virol. 2009, 83, 4591–4604. [Google Scholar] [CrossRef]
- Nash, M.A.; Meyer, M.K.; Decker, G.L.; Arlinghaus, R.B. A Subset of Pr65gag Is Nucleus Associated in Murine Leukemia Virus–Infected Cells. J. Virol. 1993, 67, 1350–1356. [Google Scholar]
- Risco, C.; Menendez–Arias, L.; Copeland, T.D.; Pinto da Silva, P.; Oroszlan, S. Intracellular transport of the murine leukemia virus during acute infection of NIH 3T3 cells: nuclear import of nucleocapsid protein and integrase. J. Cell Sci. 1995, 108, 3039–3050. [Google Scholar]
- Elis, E.; Ehrlich, M.; Prizan–Ravid, A.; Laham–Karam, N.; Bacharach, E. p12 Tethers the Murine Leukemia Virus Pre–integration Complex to Mitotic Chromosomes. PLoS Pathog. 2012, 8, e1003103. [Google Scholar] [CrossRef]
- Schneider, W.M.; Brzezinski, J.D.; Aiyer, S.; Malani, N.; Gyuricza, M.; Bushman, F.D.; Roth, M.J. Viral DNA tethering domains complement replication–defective mutations in the p12 protein of MuLV Gag. Proc. Natl. Acad. Sci. U.S.A. 2013, 110, 9487–9492. [Google Scholar]
- Sharma, A.; Larue, R.C.; Plumb, M.R.; Malani, N.; Male, F.; Slaughter, A.; Kessl, J.J.; Shkriabai, N.; Coward, E.; Aiyer, S.S.; Green, P.L.; Wu, L.; Roth, M.J.; Bushman, F.D.; Kvaratskhelia, M. BET proteins promote efficient murine leukemia virus integration at transcription start sites. Proc. Natl. Acad. Sci. U.S.A. 2013, 110, 12036–12041. [Google Scholar] [CrossRef]
- Nosaka, T.; Miyazaki, Y.; Takamatsu, T.; Sano, K.; Nakai, M.; Fujita, S.; Martin, T.E.; Hatanaka, M. The Post–transcriptional Regulator Rex of the Human T–Cell Leukemia Virus Type I Is Present as Nucleolar Speckles in Infected Cells. Exp. Cell Res. 1995, 219, 122–129. [Google Scholar] [CrossRef]
- Kubota, S.; Siomi, H.; Satoh, T.; Endo, S.; Maki, M.; Hatanaka, M. Functional similarity of HIV–I rev and HTLV–I rex proteins: identification of a new nucleolar–targeting signal in rev protein. BBRC 1989, 162, 963–970. [Google Scholar]
- Siomi, H.; Shida, H.; Nam, S.H.; Nosaka, T.; Maki, M.; Hatanaka, M. Sequence requirements for nucleolar localization of human T cell leukemia virus type I pX protein, which regulates viral RNA processing. Cell 1988, 55, 197–209. [Google Scholar] [CrossRef]
- Palmeri, D.; Malim, M.H. Importin β Can Mediate the Nuclear Import of an Arginine–Rich Nuclear Localization Signal in the Absence of Importin α. Mol. Cell. Biol. 1999, 19, 1218–1225. [Google Scholar]
- Neville, M.; Stutz, F.; Lee, L.; Davis, L.I.; Rosbash, M. The importin–beta family member Crm1p bridges the interaction between Rev and the nuclear pore complex during nuclear export. Curr. Biol. 1997, 7, 767–775. [Google Scholar]
- Hakata, Y.; Umemoto, T.; Matsushita, S.; Shida, H. Involvement of human CRM1 (exportin 1) in the export and multimerization of the Rex protein of human T–cell leukemia virus type 1. J. Virol. 1998, 72, 6602–6607. [Google Scholar]
- Choi, E.A.; Hope, T.J. Mutational analysis of bovine leukemia virus Rex: Identification of a dominant–negative inhibitor. J. Virol. 2005, 79, 7172–7181. [Google Scholar] [CrossRef]
- Grewe, B.; Hoffmann, B.; Ohs, I.; Blissenbach, M.; Brandt, S.; Tippler, B.; Grunwald, T.; Uberla, K. Cytoplasmic utilization of human immunodeficiency virus type 1 genomic RNA is not dependent on a nuclear interaction with gag. J. Virol. 2012, 86, 2990–3002. [Google Scholar] [CrossRef]
- Ao, Z.; Danappa Jayappa, K.; Wang, B.; Zheng, Y.; Kung, S.; Rassart, E.; Depping, R.; Kohler, M.; Cohen, E.A.; Yao, X. Importin alpha3 interacts with HIV–1 integrase and contributes to HIV–1 nuclear import and replication. J. Virol. 2010, 84, 8650–8663. [Google Scholar] [CrossRef]
- Hearps, A.C.; Jans, D.A. HIV–1 integrase is capable of targeting DNA to the nucleus via an importin alpha/beta–dependent mechanism. Biochem. J. 2006, 398, 475–484. [Google Scholar] [CrossRef]
- Bouyac–Bertoia, M.; Dvorin, J.D.; Fouchier, R.A.; Jenkins, Y.; Meyer, B.E.; Wu, L.I.; Emerman, M.; Malim, M.H. HIV–1 infection requires a functional integrase NLS. Mol. Cell. 2001, 7, 1025–1035. [Google Scholar] [CrossRef]
- Depienne, C.; Mousnier, A.; Leh, H.; Le Rouzic, E.; Dormont, D.; Benichou, S.; Dargemont, C. Characterization of the nuclear import pathway for HIV–1 integrase. J. Biol. Chem. 2001, 276, 18102–18107. [Google Scholar]
- Petit, C.; Schwartz, O.; Mammano, F. The karyophilic properties of human immunodeficiency virus type 1 integrase are not required for nuclear import of proviral DNA. J. Virol. 2000, 74, 7119–7126. [Google Scholar] [CrossRef]
- Ao, Z.; Huang, G.; Yao, H.; Xu, Z.; Labine, M.; Cochrane, A.W.; Yao, X. Interaction of human immunodeficiency virus type 1 integrase with cellular nuclear import receptor importin 7 and its impact on viral replication. J. Biol. Chem. 2007, 282, 13456–13467. [Google Scholar] [CrossRef]
- Zhang, J.; Crumpacker, C.S. Human immunodeficiency virus type 1 nucleocapsid protein nuclear localization mediates early viral mRNA expression. J. Virol. 2002, 76, 10444–10454. [Google Scholar] [CrossRef]
- Gallay, P.; Swingler, S.; Song, J.; Bushman, F.; Trono, D. HIV nuclear import is governed by the phosphotyrosine–mediated binding of matrix to the core domain of integrase. Cell 1995, 83, 569–576. [Google Scholar] [CrossRef]
- Kalland, K.H.; Szilvay, A.M.; Langhoff, E.; Haukenes, G. Subcellular distribution of human immunodeficiency virus type 1 Rev and colocalization of Rev with RNA splicing factors in a speckled pattern in the nucleoplasm. J. Virol. 1994, 68, 1475–1485. [Google Scholar]
- Kula, A.; Gharu, L.; Marcello, A. HIV–1 pre–mRNA commitment to Rev mediated export through PSF and Matrin 3. Virology 2013, 435, 329–340. [Google Scholar] [CrossRef]
- Truant, R.; Cullen, B.R. The arginine–rich domains present in human immunodeficiency virus type 1 Tat and Rev function as direct importin beta–dependent nuclear localization signals. Mol. Cell. Biol. 1999, 19, 1210–1217. [Google Scholar]
- Henderson, B.R.; Percipalle, P. Interactions between HIV Rev and nuclear import and export factors: the Rev nuclear localisation signal mediates specific binding to human importin–beta. J. Mol. Biol. 1997, 274, 693–707. [Google Scholar] [CrossRef]
- Arnold, M.; Nath, A.; Hauber, J.; Kehlenbach, R.H. Multiple importins function as nuclear transport receptors for the Rev protein of human immunodeficiency virus type 1. J. Biol. Chem. 2006, 281, 20883–20890. [Google Scholar] [CrossRef]
- Fornerod, M.; Ohno, M.; Yoshida, M.; Mattaj, I.W. CRM1 is an export receptor for leucine–rich nuclear export signals. Cell 1997, 90, 1051–1060. [Google Scholar] [CrossRef]
- Sherman, M.P.; de Noronha, C.M.; Heusch, M.I.; Greene, S.; Greene, W.C. Nucleocytoplasmic shuttling by human immunodeficiency virus type 1 Vpr. J. Virol. 2001, 75, 1522–1532. [Google Scholar] [CrossRef]
- Vodicka, M.A.; Koepp, D.M.; Silver, P.A.; Emerman, M. HIV–1 Vpr interacts with the nuclear transport pathway to promote macrophage infection. Gene Dev. 1998, 12, 175–185. [Google Scholar] [CrossRef]
- Jenkins, Y.; McEntee, M.; Weis, K.; Greene, W.C. Characterization of HIV–1 vpr nuclear import: analysis of signals and pathways. J. Cell Biol. 1998, 143, 875–885. [Google Scholar] [CrossRef]
- Le Rouzic, E.; Mousnier, A.; Rustum, C.; Stutz, F.; Hallberg, E.; Dargemont, C.; Benichou, S. Docking of HIV–1 Vpr to the nuclear envelope is mediated by the interaction with the nucleoporin hCG1. J. Biol. Chem. 2002, 277, 45091–45098. [Google Scholar]
- Dillon, P.J.; Nelbock, P.; Perkins, A.; Rosen, C.A. Structural and functional analysis of the human immunodeficiency virus type 2 Rev protein. J. Virol. 1991, 65, 445–449. [Google Scholar]
- Belshan, M.; Ratner, L. Identification of the nuclear localization signal of human immunodeficiency virus type 2 Vpx. Virology 2003, 311, 7–15. [Google Scholar] [CrossRef]
- Pancio, H.A.; Vander Heyden, N.; Ratner, L. The C–terminal proline–rich tail of human immunodeficiency virus type 2 Vpx is necessary for nuclear localization of the viral preintegration complex in nondividing cells. J. Virol. 2000, 74, 6162–6167. [Google Scholar] [CrossRef]
- Kemler, I.; Meehan, A.; Poeschla, E.M. Live–Cell Coimaging of the Genomic RNAs and Gag Proteins of Two Lentiviruses. J. Virol. 2010, 84, 6352–6366. [Google Scholar] [CrossRef]
- Phillips, T.R.; Lamont, C.; Konings, D.A.; Shacklett, B.L.; Hamson, C.A.; Luciw, P.A.; Elder, J.H. Identification of the Rev transactivation and Rev–responsive elements of feline immunodeficiency virus. J. Virol. 1992, 66, 5464–5471. [Google Scholar]
- Otero, G.C.; Harris, M.E.; Donello, J.E.; Hope, T.J. Leptomycin B Inhibits Equine Infectious Anemia Virus Rev and Feline Immunodeficiency Virus Rev Function but Not the Function of the Hepatitis B Virus Posttranscriptional Regulatory Element. J. Virol. 1998, 72, 7593–7597. [Google Scholar]
- Oberste, M.S.; Williamson, J.C.; Greenwood, J.D.; Nagashima, K.; Copeland, T.D.; Gonda, M.A. Characterization of Bovine Immunodeficiency Virus Rev Cdnas and Identification and Subcellular–Localization of the Rev Protein. J. Virol. 1993, 67, 6395–6405. [Google Scholar]
- Gomez Corredor, A.; Archambault, D. The bovine immunodeficiency virus Rev protein: identification of a novel nuclear import pathway and nuclear export signal among retroviral Rev/Rev–like proteins. J. Virol. 2012, 86, 4892–4905. [Google Scholar]
- Corredor, A.G.; Archambault, D. The bovine immunodeficiency virus Rev protein: identification of a novel nuclear import pathway and nuclear export signal among retroviral Rev/Rev–like proteins. J. Virol. 2012, 86, 4892–4905. [Google Scholar] [CrossRef]
- Tiley, L.S.; Brown, P.H.; Le, S.Y.; Maizel, J.V.; Clements, J.E.; Cullen, B.R. Visna virus encodes a post–transcriptional regulator of viral structural gene expression. Proc. Natl. Acad. Sci. U.S.A. 1990, 87, 7497–7501. [Google Scholar]
- Schoborg, R.V.; Saltarelli, M.J.; Clements, J.E. A Rev Protein Is Expressed in Caprine Arthritis–Encephalitis Virus (Caev)–Infected Cells and Is Required for Efficient Viral Replication. Virology 1994, 202, 1–15. [Google Scholar] [CrossRef]
- Saltarelli, M.J.; Schoborg, R.; Pavlakis, G.N.; Clements, J.E. Identification of the caprine arthritis encephalitis virus Rev protein and its cis–acting Rev–responsive element. Virology 1994, 199, 47–55. [Google Scholar] [CrossRef]
- Schliephake, A.W.; Rethwilm, A. Nuclear localization of foamy virus Gag precursor protein. J. Virol. 1994, 68, 4946–4954. [Google Scholar]
- Tobaly–Tapiero, J.; Bittoun, P.; Lehmann–Che, J.; Delelis, O.; Giron, M.L.; de The, H.; Saib, A. Chromatin tethering of incoming foamy virus by the structural Gag protein. Traffic 2008, 9, 1717–1727. [Google Scholar] [CrossRef]
- Müllers, E.; Stirnnagel, K.; Kaulfuss, S.; Lindemann, D. Prototype Foamy Virus Gag Nuclear Localization: a Novel Pathway among Retroviruses. J. Virol. 2011, 85, 9276–9285. [Google Scholar]
- Kiyomasu, T.; Miyazawa, T.; Furuya, T.; Shibata, R.; Sakai, H.; Sakuragi, J.; Fukasawa, M.; Maki, N.; Hasegawa, A.; Mikami, T.; et al. Identification of feline immunodeficiency virus rev gene activity. J. Virol. 1991, 65, 4539–4542. [Google Scholar]
- Hirsch, V.M.; Olmsted, R.A.; Murphey–Corb, M.; Purcell, R.H.; Johnson, P.R. An African primate lentivirus (SIVsm) closely related to HIV–2. Nature 1989, 339, 389–392. [Google Scholar] [CrossRef]
- Cheng, S.M.; Blume, M.; Lee, S.G.; Hung, P.P.; Hirsch, V.M.; Johnson, P.R. Coexpression of biologically active simian immunodeficiency virus (SIV) Rev and Env in an SV40 system: the SIV rev gene regulates env expression. Virology 1990, 177, 816–819. [Google Scholar] [CrossRef]
- Gomez Corredor, A.; Archambault, D. The bovine immunodeficiency virus rev protein: identification of a novel lentiviral bipartite nuclear localization signal harboring an atypical spacer sequence. J. Virol. 2009, 83, 12842–12853. [Google Scholar] [CrossRef]
- Stephens, R.M.; Derse, D.; Rice, N.R. Cloning and characterization of cDNAs encoding equine infectious anemia virus tat and putative Rev proteins. J. Virol. 1990, 64, 3716–3725. [Google Scholar]
- Fridell, R.A.; Partin, K.M.; Carpenter, S.; Cullen, B.R. Identification of the activation domain of equine infectious anemia virus rev. J. Virol. 1993, 67, 7317–7323. [Google Scholar]
- Fischer, U.; Meyer, S.; Teufel, M.; Heckel, C.; Luhrmann, R.; Rautmann, G. Evidence that HIV–1 Rev directly promotes the nuclear export of unspliced RNA. EMBO J. 1994, 13, 4105–4112. [Google Scholar]
- Guyader, M.; Emerman, M.; Sonigo, P.; Clavel, F.; Montagnier, L.; Alizon, M. Genome organization and transactivation of the human immunodeficiency virus type 2. Nature 1987, 326, 662–669. [Google Scholar] [CrossRef]
- Malim, M.H.; Hauber, J.; Le, S.Y.; Maizel, J.V.; Cullen, B.R. The HIV–1 rev trans–activator acts through a structured target sequence to activate nuclear export of unspliced viral mRNA. Nature 1989, 338, 254–257. [Google Scholar] [CrossRef]
- Garvey, K.J.; Oberste, M.S.; Elser, J.E.; Braun, M.J.; Gonda, M.A. Nucleotide sequence and genome organization of biologically active proviruses of the bovine immunodeficiency–like virus. Virology 1990, 175, 391–409. [Google Scholar] [CrossRef]
- Mertz, J.A.; Simper, M.S.; Lozano, M.M.; Payne, S.M.; Dudley, J.P. Mouse mammary tumor virus encodes a self–regulatory RNA export protein and is a complex retrovirus. J. Virol. 2005, 79, 14737–14747. [Google Scholar] [CrossRef]
- Hidaka, M.; Inoue, J.; Yoshida, M.; Seiki, M. Post–transcriptional regulator (rex) of HTLV–1 initiates expression of viral structural proteins but suppresses expression of regulatory proteins. EMBO J. 1988, 7, 519–523. [Google Scholar]
- Sagata, N.; Yasunaga, T.; Tsuzuku–Kawamura, J.; Ohishi, K.; Ogawa, Y.; Ikawa, Y. Complete nucleotide sequence of the genome of bovine leukemia virus: its evolutionary relationship to other retroviruses. Proc. Natl. Acad. Sci. U.S.A. 1985, 82, 677–681. [Google Scholar]
- Dultz, E.; Hildenbeutel, M.; Martoglio, B.; Hochman, J.; Dobberstein, B.; Kapp, K. The signal peptide of the mouse mammary tumor virus Rem protein is released from the endoplasmic reticulum membrane and accumulates in nucleoli. J. Biol. Chem. 2008, 283, 9966–9976. [Google Scholar] [CrossRef]
- Nosaka, T.; Siomi, H.; Adachi, Y.; Ishibashi, M.; Kubota, S.; Maki, M.; Hatanaka, M. Nucleolar targeting signal of human T–cell leukemia virus type I rex–encoded protein is essential for cytoplasmic accumulation of unspliced viral mRNA. Proc. Natl. Acad. Sci. U.S.A. 1989, 86, 9798–9802. [Google Scholar] [CrossRef]
- Nitta, T.; Hofacre, A.; Hull, S.; Fan, H. Identification and mutational analysis of a Rej response element in Jaagsiekte sheep retrovirus RNA. J. Virol. 2009, 83, 12499–12511. [Google Scholar] [CrossRef]
- Cullen, B.R.; Hauber, J.; Campbell, K.; Sodroski, J.G.; Haseltine, W.A.; Rosen, C.A. Subcellular localization of the human immunodeficiency virus trans–acting art gene product. J. Virol. 1988, 62, 2498–2501. [Google Scholar]
- Kjems, J.; Brown, M.; Chang, D.D.; Sharp, P.A. Structural analysis of the interaction between the human immunodeficiency virus Rev protein and the Rev response element. Proc. Natl. Acad. Sci. U.S.A. 1991, 88, 683–687. [Google Scholar] [CrossRef]
- Daugherty, M.D.; Booth, D.S.; Jayaraman, B.; Cheng, Y.; Frankel, A.D. HIV Rev response element (RRE) directs assembly of the Rev homooligomer into discrete asymmetric complexes. Proc. Natl. Acad. Sci. U.S.A. 2010, 107, 12481–12486. [Google Scholar]
- Tiley, L.S.; Malim, M.H.; Tewary, H.K.; Stockley, P.G.; Cullen, B.R. Identification of a high–affinity RNA–binding site for the human immunodeficiency virus type 1 Rev protein. Proc. Natl. Acad. Sci. U.S.A. 1992, 89, 758–762. [Google Scholar]
- Cook, K.S.; Fisk, G.J.; Hauber, J.; Usman, N.; Daly, T.J.; Rusche, J.R. Characterization of HIV–1 REV protein: binding stoichiometry and minimal RNA substrate. Nucleic. Acids. Res. 1991, 19, 1577–1583. [Google Scholar] [CrossRef]
- Pollard, V.W.; Malim, M.H. The HIV–1 Rev protein. Annu. Rev. Microbiol. 1998, 52, 491–532. [Google Scholar] [CrossRef]
- Daelemans, D.; Costes, S.V.; Cho, E.H.; Erwin–Cohen, R.A.; Lockett, S.; Pavlakis, G.N. In vivo HIV–1 Rev multimerization in the nucleolus and cytoplasm identified by fluorescence resonance energy transfer. J. Biol. Chem. 2004, 279, 50167–50175. [Google Scholar]
- Michienzi, A.; Cagnon, L.; Bahner, I.; Rossi, J.J. Ribozyme–mediated inhibition of HIV 1 suggests nucleolar trafficking of HIV–1 RNA. Proc. Natl. Acad. Sci. U.S.A. 2000, 97, 8955–8960. [Google Scholar] [CrossRef]
- Buonomo, S.B.C.; Michienzi, A.; De Angelis, F.G.; Bozzoni, I. The Rev protein is able to transport to the cytoplasm small nucleolar RNAs containing a Rev binding element. Rna 1999, 5, 993–1002. [Google Scholar] [CrossRef]
- Fischer, U.; Pollard, V.W.; Luhrmann, R.; Teufel, M.; Michael, M.W.; Dreyfuss, G.; Malim, M.H. Rev–mediated nuclear export of RNA is dominant over nuclear retention and is coupled to the Ran–GTPase cycle. Nucleic. Acids. Res. 1999, 27, 4128–4134. [Google Scholar] [CrossRef]
- Favaro, J.P.; Borg, K.T.; Arrigo, S.J.; Schmidt, M.G. Effect of Rev on the intranuclear localization of HIV–1 unspliced RNA. Virology 1998, 249, 286–296. [Google Scholar] [CrossRef]
- Shopland, L.S.; Johnson, C.V.; Byron, M.; McNeil, J.; Lawrence, J.B. Clustering of multiple specific genes and gene–rich R–bands around SC–35 domains: evidence for local euchromatic neighborhoods. J. Cell. Biol. 2003, 162, 981–990. [Google Scholar] [CrossRef]
- Moen, P.T.; Johnson, C.V.; Byron, M.; Shopland, L.S.; de la Serna, I.L.; Imbalzano, A.N.; Lawrence, J.B. Repositioning of muscle–specific genes relative to the periphery of SC–35 domains during skeletal myogenesis. Mol. Biol. Cell. 2004, 15, 197–206. [Google Scholar]
- Szczerbal, I.; Bridger, J.M. Association of adipogenic genes with SC–35 domains during porcine adipogenesis. Chromosome Res. 2010, 18, 887–895. [Google Scholar] [CrossRef]
- Spector, D.L.; Lamond, A.I. Nuclear Speckles. In Csh Perspect Biol; 2011; Vol. 3. [Google Scholar]
- Sutherland, H.; Bickmore, W.A. Transcription factories: gene expression in unions? Nat. Rev. Genet. 2009, 10, 457–466. [Google Scholar] [CrossRef]
- Listerman, I.; Sapra, A.K.; Neugebauer, K.M. Cotranscriptional coupling of splicing factor recruitment and precursor messenger RNA splicing in mammalian cells. Nat. Struct. Mol. Biol. 2006, 13, 815–822. [Google Scholar] [CrossRef]
- Iacampo, S.; Cochrane, A. Human immunodeficiency virus type 1 Rev function requires continued synthesis of its target mRNA. J. Virol. 1996, 70, 8332–8339. [Google Scholar]
- Naji, S.; Ambrus, G.; Cimermancic, P.; Reyes, J.R.; Johnson, J.R.; Filbrandt, R.; Huber, M.D.; Vesely, P.; Krogan, N.J.; Yates, J.R.; Saphire, A.C.; Gerace, L. Host Cell Interactome of HIV–1 Rev Includes RNA Helicases Involved in Multiple Facets of Virus Production. In Mol Cell Proteomics; 2012; Vol. 11. [Google Scholar]
- Nekhai, S.; Jeang, K.T. Transcriptional and post–transcriptional regulation of HIV–1 gene expression: role of cellular factors for Tat and Rev. Future Microbiol. 2006, 1, 417–426. [Google Scholar] [CrossRef]
- Hadian, K.; Vincendeau, M.; Mausbacher, N.; Nagel, D.; Hauck, S.M.; Ueffing, M.; Loyter, A.; Werner, T.; Wolff, H.; Brack–Werner, R. Identification of a heterogeneous nuclear ribonucleoprotein–recognition region in the HIV Rev protein. J. Biol. Chem. 2009, 284, 33384–33391. [Google Scholar] [CrossRef]
- Ruhl, M.; Himmelspach, M.; Bahr, G.M.; Hammerschmid, F.; Jaksche, H.; Wolff, B.; Aschauer, H.; Farrington, G.K.; Probst, H.; Bevec, D.; et al. Eukaryotic initiation factor 5A is a cellular target of the human immunodeficiency virus type 1 Rev activation domain mediating trans–activation. J. Cell Biol. 1993, 123, 1309–1320. [Google Scholar] [CrossRef]
- Schatz, O.; Oft, M.; Dascher, C.; Schebesta, M.; Rosorius, O.; Jaksche, H.; Dobrovnik, M.; Bevec, D.; Hauber, J. Interaction of the HIV–1 rev cofactor eukaryotic initiation factor 5A with ribosomal protein L5. Proc. Natl. Acad. Sci. U.S.A. 1998, 95, 1607–1612. [Google Scholar] [CrossRef]
- Yasuda–Inoue, M.; Kuroki, M.; Ariumi, Y. Distinct DDX DEAD–box RNA helicases cooperate to modulate the HIV–1 Rev function. Biochem. Biophys. Res. Commun. 2013, 434, 803–808. [Google Scholar] [CrossRef]
- Fang, J.; Kubota, S.; Yang, B.; Zhou, N.; Zhang, H.; Godbout, R.; Pomerantz, R.J. A DEAD box protein facilitates HIV–1 replication as a cellular co–factor of Rev. Virology 2004, 330, 471–480. [Google Scholar] [CrossRef]
- Zhou, X.X.; Luo, J.; Mills, L.; Wu, S.X.; Pan, T.; Geng, G.N.; Zhang, J.; Luo, H.H.; Liu, C.; Zhang, H. DDX5 Facilitates HIV–1 Replication as a Cellular Co–Factor of Rev. Plos One 2013, 8, (5). [Google Scholar]
- Edgcomb, S.P.; Carmel, A.B.; Naji, S.; Ambrus–Aikelin, G.; Reyes, J.R.; Saphire, A.C.; Gerace, L.; Williamson, J.R. DDX1 is an RNA–dependent ATPase involved in HIV–1 Rev function and virus replication. J. Mol. Biol. 2012, 415, 61–74. [Google Scholar] [CrossRef]
- Robertson–Anderson, R.M.; Wang, J.; Edgcomb, S.P.; Carmel, A.B.; Williamson, J.R.; Millar, D.P. Single–molecule studies reveal that DEAD box protein DDX1 promotes oligomerization of HIV–1 Rev on the Rev response element. J. Mol. Biol. 2011, 410, 959–971. [Google Scholar] [CrossRef]
- Yedavalli, V.S.; Neuveut, C.; Chi, Y.H.; Kleiman, L.; Jeang, K.T. Requirement of DDX3 DEAD box RNA helicase for HIV–1 Rev–RRE export function. Cell 2004, 119, 381–392. [Google Scholar] [CrossRef]
- Yedavalli, V.S.R.K.; Jeang, K.T. Matrin 3 is a co–factor for HIV–1 Rev in regulating post–transcriptional viral gene expression. In Retrovirology; 2011; Vol. 8. [Google Scholar]
- Salton, M.; Elkon, R.; Borodina, T.; Davydov, A.; Yaspo, M.L.; Halperin, E.; Shiloh, Y. Matrin 3 Binds and Stabilizes mRNA. In Plos One; 2011; Vol. 6. [Google Scholar]
- Butin–Israeli, V.; Adam, S.A.; Goldman, A.E.; Goldman, R.D. Nuclear lamin functions and disease. Trends Genet. 2012, 28, 464–471. [Google Scholar] [CrossRef]
- Ciejek, E.M.; Tsai, M.J.; Omalley, B.W. Actively Transcribed Genes Are Associated with the Nuclear Matrix. Nature 1983, 306, 607–609. [Google Scholar] [CrossRef]
- Malecki, M.; Malecki, B. Nuclear routing networks span between nuclear pore complexes and genomic DNA to guide nucleoplasmic trafficking of biomolecules. In J Fertili In Vitro, 2013/01/01 ed.; 2012; Vol. 2. [Google Scholar]
- Malecki, M.; Malecki, B. Routing of Biomolecules and Transgenes' Vectors in Nuclei of Oocytes. J. Fertili. In Vitro 2012, 2, 108–118. [Google Scholar]
- Zeitz, M.J.; Malyavantham, K.S.; Seifert, B.; Berezney, R. Matrin 3: chromosomal distribution and protein interactions. J. Cell. Biochem. 2009, 108, 125–33. [Google Scholar] [CrossRef]
- Malyavantham, K.S.; Bhattacharya, S.; Barbeitos, M.; Mukherjee, L.; Xu, J.; Fackelmayer, F.O.; Berezney, R. Identifying functional neighborhoods within the cell nucleus: proximity analysis of early S–phase replicating chromatin domains to sites of transcription, RNA polymerase II, HP1gamma, matrin 3 and SAF–A. J. Cell. Biochem. 2008, 105, 391–403. [Google Scholar] [CrossRef]
- Bray, M.; Prasad, S.; Dubay, J.W.; Hunter, E.; Jeang, K.–T.; Rekosh, D.; Hammarskjöld, M.L. A small element from the Mason–Pfizer monkey virus genome makes human immunodeficiency virus type 1 expression and replication Rev–independent. Proc. Natl. Acad. Sci. U.S.A. 1994, 91, 1256–1260. [Google Scholar]
- Ernst, R.K.; Bray, M.; Rekosh, D.; Hammarskjold, M.L. A structured retroviral RNA element that mediates nucleocytoplasmic export of intron–containing RNA. Mol. Cell. Biol. 1997, 17, 135–144. [Google Scholar]
- Grüter, P.; Tabernero, C.; von Kobbe, C.; Schmitt, C.; Saavedra, C.; Bachi, A.; Wilm, M.; Felber, B.K.; Izaurralde, E. TAP, the Human Homolog of Mex67p, Mediates CTE–Dependent RNA Export from the Nucleus. Mol. Cell. 1998, 1, 649–659. [Google Scholar] [CrossRef]
- Katahira, J.; Strasser, K.; Podtelejnikov, A.; Mann, M.; Jung, J.U.; Hurt, E. The Mex67p–mediated nuclear mRNA export pathway is conserved from yeast to human. EMBO J. 1999, 18, 2593–2609. [Google Scholar] [CrossRef]
- Saavedra, C.; Felber, B.; Izaurralde, E. The simian retrovirus–1 constitutive transport element, unlike the HIV–1 RRE, uses factors required for cellular mRNA export. Curr. Biol. 1997, 7, 619–628. [Google Scholar] [CrossRef]
- Pasquinelli, A.E.; Ernst, R.K.; Lund, E.; Grimm, C.; Zapp, M.L.; Rekosh, D.; Hammarskjold, M.L.; Dahlberg, J.E. The constitutive transport element (CTE) of Mason–Pfizer monkey virus (MPMV) accesses a cellular mRNA export pathway. EMBO J. 1997, 16, 7500–7510. [Google Scholar] [CrossRef]
- Sergef, A.; Sharma, K.; Doye, V.; Hellwig, A.; Huber, J.; Lührmann, R.; Hurt, E. Mex67p, a novel factor for nuclear mRNA export, binds to both poly(A)+ RNA and nuclear pores. EMBO J. 1997, 16, 3256–3271. [Google Scholar] [CrossRef]
- Kang, Y.; Cullen, B.R. The human Tap protein is a nuclear mRNA export factor that contains novel RNA–binding and nucleocytoplasmic transport sequences. Genes Dev. 1999, 13, 1126–1139. [Google Scholar] [CrossRef]
- Stutz, F.; Bachi, A.; Doerks, T.; Braun, I.C.; Séraphin, B.; Wilm, M.; Bork, P.; Izaurralde, E. REF, an evolutionarily conserved family of hnRNP–like proteins, interacts with TAP/Mex67p and participates in mRNA nuclear export. Rna 2000, 6, 638–650. [Google Scholar] [CrossRef]
- Teplova, M.; Wohlbold, L.; Khin, N.W.; Izaurralde, E.; Patel, D.J. Structure–function studies of nucleocytoplasmic transport of retroviral genomic RNA by mRNA export factor TAP. Nature 2011, 18, 990–998. [Google Scholar]
- Schwartz, D.E.; Tizard, R.; Gilbert, W. Nucleotide Sequence of Rous Sarcoma Virus. Cell 1983, 32, 853–869. [Google Scholar] [CrossRef]
- Ogert, R.A.; Lee, L.H.; Beemon, K. Avian Retroviral RNA Element Promotes Unspliced RNA Accumulation in the Cytoplasm. J. Virol. 1996, 70, 3834–3843. [Google Scholar]
- Omer, C.A.; Pogue–Geile, K.; Guntaka, R.; Staskus, K.A.; Faras, A.J. Involvement of Directly Repeated Sequences in the Generation of Deletions of the Avian Sarcoma Virus src Gene. J. Virol. 1983, 47, 380–382. [Google Scholar]
- Yamamoto, T.; Tyagi, J.S.; Fagan, J.B.; Jay, G.; de Crombrugghe, B.; Pastan, I. Molecular Mechanism for the Capture and Excision of the Transforming Gene of Avian Sarcoma Virus as Suggested by Analysis of Recombinant Clones. J. Virol. 1980, 35, 436–443. [Google Scholar]
- Aschoff, J.M.; Foster, D.; Coffin, J.M. Point mutations in the avian sarcoma/leukosis virus 3' untranslated region result in a packaging defect. J. Virol. 1999, 73, 7421–7429. [Google Scholar]
- Simpson, S.B.; Guo, W.; Winistorfer, S.C.; Craven, R.C.; Stoltzfus, C.M. The Upstream Direct Repeat Sequence of Prague A Rous Sarcoma Virus is Deficient in Mediating Efficiient Gag Assembly and Particle Release. Virology 1998, 247, 86–96. [Google Scholar] [CrossRef]
- Simpson, S.B.; Zhang, L.; Craven, R.C.; Stoltzfus, C.M. Rous sarcoma virus direct repeat cis elements exert effects at several points in the virus life cycle. J. Virol. 1997, 71, 9150–9156. [Google Scholar]
- Paca, R.E.; Ogert, R.A.; Hibbert, C.S.; Izaurralde, E.; Beemon, K. Rous Sarcoma Virus DR Postranscriptional Elements Use a Novel RNA Export Pathway. J. Virol. 2000, 74, 9507–9514. [Google Scholar] [CrossRef]
- LeBlanc, J.J.; Uddowla, S.; Abraham, B.; Clatterbuck, S.; Beemon, K.L. Tap and Dbp5, but not Gag, are involved in DR–mediated nuclear export of unspliced Rous sarcoma virus RNA. Virology 2007, 363, 376–386. [Google Scholar] [CrossRef]
- Garbitt–Hirst, R.; Kenney, S.P.; Parent, L.J. Genetic Evidence for a Connection between Rous Sarcoma Virus Gag Nuclear Trafficking and Genomic RNA Packaging. J. Virol. 2009, 83, 6790–6797. [Google Scholar] [CrossRef]
- Mullers, E. The foamy virus Gag proteins: what makes them different? Viruses 2013, 5, 1023–1041. [Google Scholar] [CrossRef]
- Linial, M.L. Foamy Viruses Are Unconventional Retroviruses. J. Virol. 1999, 73, 1747–1755. [Google Scholar]
- Enssle, J.; Jordan, I.; Mauer, B.; Rethwilm, A. Foamy virus reverse transcriptase is expressed independently from the Gag protein. Proc. Natl. Acad. Sci. U.S.A. 1996, 93, 4137–4141. [Google Scholar] [CrossRef]
- Lochelt, M.; Flugel, R.M. The human foamy virus pol gene is expressed as a Pro–Pol polyprotein and not as a gag–pol fusion protein. J. Virol. 1996, 70, 1033–1040. [Google Scholar]
- Yu, S.F.; Edelmann, K.; Strong, R.K.; Moebes, A.; Rethwilm, A.; Linial, M.L. The carboxyl terminus of the human foamy virus Gag protein contains separable nucleic acid binding and nuclear transport domains. J. Virol. 1996, 70, 8255–8262. [Google Scholar]
- Jordan, I.; Enssle, J.; Güttler, E.; Mauer, B.; Rethwilm, A. Expression of Human Foamy Virus Reverse Transcriptase Involves a Spliced pol mRNA. Virology 1996, 224, 314–319. [Google Scholar] [CrossRef]
- Keller, A.; Partin, K.M.; Löchelt, M.; Bannert, H.; Flügel, R.M.; Cullen, B.R. Characterization of the Transcriptional trans Activator of Human Foamy Retrovirus. J. Virol. 1991, 65, 2589–2594. [Google Scholar]
- Baunach, G.; Maurer, B.; Hahn, H.; Kranz, M.; Rethwilm, A. Functional Analysis of Human Foamy Virus Accessory Reading Frames. J. Virol. 1993, 67, 5411–5418. [Google Scholar]
- Bodem, J.; Schied, T.; Gabriel, R.; Rammling, M.; Rethwilm, A. Foamy virus nuclear RNA export is distinct from that of other retroviruses. J. Virol. 2011, 85, 2333–2341. [Google Scholar]
- Gallouzi, I.E.; Steitz, J.A. Delineation of mRNA export pathways by the use of cell–permeable peptides. Science 2001, 294, 1895–1901. [Google Scholar] [CrossRef]
- Brennan, C.M.; Gallouzi, I.–E.; Steitz, J.A. Protein Ligands to HuR Modulate Its Interaction with target mRNAs In Vivo. J. Cell Biol. 2000, 151, 1–13. [Google Scholar]
- Yu, S.F.; Lujan, P.; Jackson, D.L.; Emerman, M.; Linial, M.L. The DEAD–box RNA helicase DDX6 is required for efficient encapsidation of a retroviral genome. PLoS Pathog 2011, 7, e1002303. [Google Scholar] [CrossRef]
- Erlwein, O.; Bieniasz, P.D.; McClure, M.O. Sequences in pol are required for transfer of human foamy virus–based vectors. J. Virol. 1998, 72, 5510–5516. [Google Scholar]
- Scheifele, L.Z.; Ryan, E.P.; Parent, L.J. Detailed Mapping of the Nuclear Export Signal in the Rous Sarcoma Virus Gag Protein. J. Virol. 2005, 79, 8732–8741. [Google Scholar] [CrossRef]
- Brun, S.; Solignat, M.; Gay, B.; Bernard, E.; Chaloin, L.; Fenard, D.; Devaux, C.; Chazal, N.; Briant, L. VSV–G pseudotyping rescues HIV–1 CA mutations that impair core assembly or stability. Retrovirol. 2008, 5, 57. [Google Scholar] [CrossRef]
- Thys, W.; De Houwer, S.; Demeulemeester, J.; Taltynov, O.; Vancraenenbroeck, R.; Gerard, M.; De Rijck, J.; Gijsbers, R.; Christ, F.; Debyser, Z. Interplay between HIV entry and transportin–SR2 dependency. Retrovirol. 2011, 8, 7. [Google Scholar] [CrossRef]
- Garbitt, R.A.; Bone, K.R.; Parent, L.J. Insertion of a Classical Nuclear Import Signal into the Matrix Domain of the Rous Sarcoma Virus Gag Protein Interferes with Virus Replication. J. Virol. 2004, 78, 13534–13542. [Google Scholar] [CrossRef]
- Kataoka, N.; Bachorik, J.L.; Dreyfuss, G. Transportin–SR, a Nuclear Import Receptor for SR Proteins. J. Cell. Biol. 1999, 145, 1145–1152. [Google Scholar] [CrossRef]
- Lai, M.C.; Lin, R.I.; Tarn, W.Y. Transportin–SR2 mediates nuclear import of phosphorylated SR proteins. Proc. Natl. Acad. Sci. U.S.A. 2001, 98, 10154–10159. [Google Scholar] [CrossRef]
- Lai, M.C.; Lin, R.I.; Huang, S.Y.; Tsai, C.W.; Tarn, W.Y. A human importin–beta family protein, transportin–SR2, interacts with the phosphorylated RS domain of SR proteins. J. Biol. Chem. 2000, 275, 7950–7957. [Google Scholar]
- Flynn, J.A.; An, W.; King, S.R.; Telesnitsky, A. Nonrandom dimerization of murine leukemia virus genomic RNAs. J. Virol. 2004, 78, 12129–12139. [Google Scholar] [CrossRef]
- Flynn, J.A.; Telesnitsky, A. Two distinct Moloney murine leukemia virus RNAs produced from a single locus dimerize at random. Virology 2006, 344, 391–400. [Google Scholar] [CrossRef]
- Kharytonchyk, S.A.; Kireyeva, A.I.; Osipovich, A.B.; Fomin, I.K. Evidence for preferential copackaging of Moloney murine leukemia virus genomic RNAs transcribed in the same chromosomal site. Retrovirol. 2005, 2, 3. [Google Scholar] [CrossRef]
- Rasmussen, S.V.; Pedersen, F.S. Co–localization of gammaretroviral RNAs at their transcription site favours co–packaging. J. Gen. Virol. 2006, 87, 2279–2289. [Google Scholar] [CrossRef]
- Maurel, S.; Mougel, M. Murine leukemia virus RNA dimerization is coupled to transcription and splicing processes. Retrovirol. 2010, 7, 64. [Google Scholar] [CrossRef]
- Levin, J.G.; Rosenak, M. J. Synthesis of murine leukemia virus proteins associated with virions assembled in actinomycin D–treated cells: evidence for persistence of viral messenger RNA. Proc. Natl. Acad. Sci. U.S.A. 1976, 73, 1154–1158. [Google Scholar] [CrossRef]
- Levin, J.G.; Grimley, P.M.; Ramseur, J.M.; Berezesky, I.K. Deficiency of 60 to 70S RNA in murine leukemia virus particles assembled in cells treated with actinomycin D. J. Virol. 1974, 14, 152–161. [Google Scholar]
- Dorman, N.; Lever, A. Comparison of viral genomic RNA sorting mechanisms in human immunodeficiency virus type 1 (HIV–1), HIV–2, and Moloney murine leukemia virus. J. Virol. 2000, 74, 11413–11417. [Google Scholar] [CrossRef]
- Messer, L.I.; Levin, J.G.; Chattopadhyay, S.K. Metabolism of viral RNA in murine leukemia virus–infected cells; evidence for differential stability of viral message and virion precursor RNA. J. Virol. 1981, 40, 683–690. [Google Scholar]
- D'Souza, V.; Summers, M.F. Structural basis for packaging the dimeric genome of Moloney murine leukaemia virus. Nature 2004, 431, 586–590. [Google Scholar] [CrossRef]
- Garcia, E.L.; Onafuwa–Nuga, A.; Sim, S.; King, S.R.; Wolin, S.L.; Telesnitsky, A. Packaging of host mY RNAs by murine leukemia virus may occur early in Y RNA biogenesis. J. Virol. 2009, 83, 12526–12534. [Google Scholar] [CrossRef]
- Poole, E.; Strappe, P.; Mok, H.P.; Hicks, R.; Lever, A.M. HIV–1 Gag–RNA interaction occurs at a perinuclear/centrosomal site; analysis by confocal microscopy and FRET. Traffic 2005, 6, 741–755. [Google Scholar] [CrossRef]
- Yu, S.F.; Eastman, S.W.; Linial, M.L. Foamy virus capsid assembly occurs at a pericentriolar region through a cytoplasmic targeting/retention signal in Gag. Traffic 2006, 7, 966–977. [Google Scholar] [CrossRef]
- Molle, D.; Segura–Morales, C.; Camus, G.; Berlioz–Torrent, C.; Kjems, J.; Basyuk, E.; Bertrand, E. Endosomal trafficking of HIV–1 gag and genomic RNAs regulates viral egress. J. Biol. Chem. 2009, 284, 19727–19743. [Google Scholar]
- Basyuk, E.; Galli, T.; Mougel, M.; Blanchard, J.M.; Sitbon, M.; Bertrand, E. Retroviral genomic RNAs are transported to the plasma membrane by endosomal vesicles. Dev. Cell. 2003, 5, 161–174. [Google Scholar] [CrossRef]
- Checkley, M.A.; Mitchell, J.A.; Eizenstat, L.D.; Lockett, S.J.; Garfinkel, D.J. Ty1 gag enhances the stability and nuclear export of Ty1 mRNA. Traffic 2013, 14, 57–69. [Google Scholar]
- Jouvenet, N.; Bieniasz, P.D.; Simon, S.M. Imaging the biogenesis of individual HIV–1 virions in live cells. Nature 2008, 454, 236–240. [Google Scholar] [CrossRef]
- Jouvenet, N.; Simon, S.M.; Bieniasz, P.D. Imaging the interaction of HIV–1 genomes and Gag during assembly of individual viral particles. Proc. Natl. Acad. Sci. U.S.A. 2009, 106, 19114–19119. [Google Scholar] [CrossRef]
- Jouvenet, N.; Simon, S.M.; Bieniasz, P.D. Visualizing HIV–1 assembly. J. Mol. Biol. 2011, 410, 501–511. [Google Scholar] [CrossRef]
- Kutluay, S.B.; Bieniasz, P.D. Analysis of the initiating events in HIV–1 particle assembly and genome packaging. PLoS Pathog. 2010, 6, e1001200. [Google Scholar] [CrossRef]
- Levesque, K.; Halvorsen, M.; Abrahamyan, L.; Chatel–Chaix, L.; Poupon, V.; Gordon, H.; DesGroseillers, L.; Gatignol, A.; Mouland, A.J. Trafficking of HIV–1 RNA is mediated by heterogeneous nuclear ribonucleoprotein A2 expression and impacts on viral assembly. Traffic 2006, 7, 1177–1193. [Google Scholar] [CrossRef]
- Lund, N.; Milev, M.P.; Wong, R.; Sanmuganantham, T.; Woolaway, K.; Chabot, B.; Abou Elela, S.; Mouland, A.J.; Cochrane, A. Differential effects of hnRNP D/AUF1 isoforms on HIV–1 gene expression. Nucleic. Acids. Res. 2012, 40, 3663–3675. [Google Scholar] [CrossRef]
- Stoltzfus, C.M.; Madsen, J.M. Role of viral splicing elements and cellular RNA binding proteins in regulation of HIV–1 alternative RNA splicing. Curr. HIV Res. 2006, 4, 43–55. [Google Scholar] [CrossRef]
- Lehmann, M.; Milev, M.P.; Abrahamyan, L.; Yao, X.J.; Pante, N.; Mouland, A.J. Intracellular transport of human immunodeficiency virus type 1 genomic RNA and viral production are dependent on dynein motor function and late endosome positioning. J. Biol. Chem. 2009, 284, 14572–14585. [Google Scholar] [CrossRef]
- Malagon, F.; Jensen, T.H. The T body, a new cytoplasmic RNA granule in Saccharomyces cerevisiae. Mol. Cell. Biol. 2008, 28, 6022–6032. [Google Scholar] [CrossRef]
- Checkley, M.A.; Nagashima, K.; Lockett, S.J.; Nyswaner, K.M.; Garfinkel, D.J. P–body components are required for Ty1 retrotransposition during assembly of retrotransposition–competent virus–like particles. Mol. Cell. Biol. 2010, 30, 382–398. [Google Scholar] [CrossRef]
- Dutko, J.A.; Kenny, A.E.; Gamache, E.R.; Curcio, M.J. 5' to 3' mRNA decay factors colocalize with Ty1 gag and human APOBEC3G and promote Ty1 retrotransposition. J. Virol. 2010, 84, 5052–5066. [Google Scholar] [CrossRef]
- Malagon, F.; Jensen, T.H. T–body formation precedes virus–like particle maturation in S. cerevisiae. RNA Biol. 2011, 8, 184–189. [Google Scholar] [CrossRef]
- Larsen, L.S.; Beliakova–Bethell, N.; Bilanchone, V.; Zhang, M.; Lamsa, A.; Dasilva, R.; Hatfield, G.W.; Nagashima, K.; Sandmeyer, S. Ty3 nucleocapsid controls localization of particle assembly. J. Virol. 2008, 82, 2501–2514. [Google Scholar] [CrossRef]
- Dang, V.D.; Levin, H.L. Nuclear import of the retrotransposon Tf1 is governed by a nuclear localization signal that possesses a unique requirement for the FXFG nuclear pore factor Nup124p. Mol. Cell. Biol. 2000, 20, 7798–7812. [Google Scholar] [CrossRef]
- Hiscox, J.A. The nucleolus – a gateway to viral infection? Arch. Virol. 2002, 147, 1077–1089. [Google Scholar] [CrossRef]
- Politz, J.C.; Yarovoi, S.; Kilroy, S.M.; Gowda, K.; Zwieb, C.; Pederson, T. Signal recognition particle components in the nucleolus. Proc. Natl. Acad. Sci. U.S.A. 2000, 97, 55–60. [Google Scholar] [CrossRef]
- Jacobson, M.R.; Pederson, T. Localization of signal recognition particle RNA in the nucleolus of mammalian cells. Proc. Natl. Acad. Sci. U.S.A. 1998, 95, 7981–7986. [Google Scholar] [CrossRef]
- Keene, S.E.; King, S.R.; Telesnitsky, A. 7SL RNA is retained in HIV–1 minimal virus–like particles as an S–domain fragment. J. Virol. 2010, 84, 9070–9077. [Google Scholar]
- Keene, S.E.; Telesnitsky, A. cis–Acting determinants of 7SL RNA packaging by HIV–1. J. Virol. 2012, 86, 7934–7942. [Google Scholar] [CrossRef]
- Onafuwa–Nuga, A.A.; Telesnitsky, A.; King, S.R. 7SL RNA, but not the 54–kd signal recognition particle protein, is an abundant component of both infectious HIV–1 and minimal virus–like particles. Rna 2006, 12, 542–546. [Google Scholar] [CrossRef]
- Rulli, S.J., Jr.; Hibbert, C.S.; Mirro, J.; Pederson, T.; Biswal, S.; Rein, A. Selective and nonselective packaging of cellular RNAs in retrovirus particles. J. Virol. 2007, 81, 6623–6631. [Google Scholar] [CrossRef]
- Giles, K.E.; Caputi, M.; Beemon, K.L. Packaging and reverse transcription of snRNAs by retroviruses may generate pseudogenes. Rna 2004, 10, 299–307. [Google Scholar] [CrossRef]
- Sawyer, R.C.; Hanafusa, H. Comparison of the Small RNAs of Polymerase–Deficient and Polymerase–Positive Rous Sarcoma Virus and Another Species of Avian Retrovirus. J. Virol. 1979, 29, 863–871. [Google Scholar]
- Faras, A.J.; Garapin, A.C.; Levinson, W.E.; Bishop, J.M.; Goodman, H.M. Characterization of low–molecular–weight RNAs associated with 70s RNA of Rous sarcoma virus. J. Virol. 1973, 12, 334–342. [Google Scholar]
- Buzdin, A.; Gogvadze, E.; Lebrun, M.H. Chimeric retrogenes suggest a role for the nucleolus in LINE amplification. FEBS Lett 2007, 581, 2877–2882. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).