Microbial Community Response to Seasonal Temperature Variation in a Small-Scale Anaerobic Digester
Abstract
:1. Introduction
2. Results and Discussion
2.1. Digester Performance
| Month | OLR (kg VS/m3 day) | HRT (Days) | Gas Prod. (L/day) | CH4% | Air Temp. (°C) | Digester Temp. (°C) | pH | VFA (mg/L) | Alk. (mg/L) | VFA/Alk | Notes |
|---|---|---|---|---|---|---|---|---|---|---|---|
| April | 0.31 | 204 | 25 | 22 | 20.0 | 14.6 | 6.2 | - | - | - | - |
| May | - | - | 47 | 47 | 25.5 | 19.4 | 6.5 | - | - | - | re-inoculated |
| June | - | - | 411 | 54 | 26.5 | 24.8 | 7.2 | - | - | - | not loaded |
| July | 1.67 | 61 | 644 | 48 | 26.8 | 27.0 | 7.4 | 1,776 | 7,320 | 0.24 | - |
| August | 2.80 | 27 | 1016 | 51 | 26.0 | 26.9 | 7.4 | 2,263 | 8,184 | 0.28 | - |
| September | 2.95 | 26 | 1,172 | 56 | 24.5 | 25.1 | 7.3 | 2,477 | 9,238 | 0.27 | - |
| October | 3.28 | 27 | 814 | 55 | 20.4 | 23.2 | 7.4 | 2,249 | 10,550 | 0.21 | - |
| November | 2.87 | 28 | 650 | 50 | 11.9 | 18.9 | 7.4 | 2,174 | 9,390 | 0.23 | - |
| December | 1.46 | 49 | 194 | 35 | −1.8 | 11.9 | 7.3 | 4,498 | 8,188 | 0.55 | - |
| January | 1.38 | 51 | 79 | 29 | −3.0 | 8.5 | 6.9 | 6,561 | 6,800 | 0.99 | - |
| February | - | - | 5 | - | 2.7 | 5.4 | 6.8 | 8,814 | 5,550 | 1.59 | not loaded |
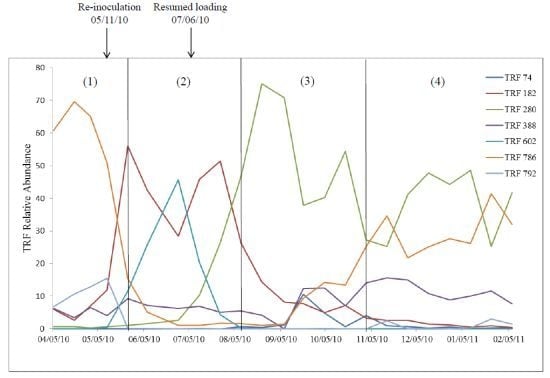
2.2. Relationship of Microbial Community Structure to Digester Phase
2.2.1. Sour Digester


2.2.2. Post Re-Inoculation
2.2.3. Peak Biogas Yield
2.2.4. Reduced Biogas Yield
2.3. Relationship between Biogas Yield and Microbial Community Structure
2.4. Archaea and Bacteria TRF Richness
2.5. Archaeal and Bacterial Community Rate of Change
3. Experimental Section
3.1. Anaerobic Digester Design
3.2. Feedstock Source and Digester Operation
3.3. Sample Collection
3.4. Microbial Community Analysis
3.4.1. Genomic DNA Extraction
3.4.2. Polymerase Chain Reaction
3.4.3. Restriction Digestions
3.4.4. T-RFLP Analysis
3.5. Clone Library Construction and Sequencing
3.6. Analysis of Biogas and pH
3.7. Alkalinity and Total Volatile Fatty Acids
3.8. Data Analyses
3.8.1. T-RFLP Data
3.8.2. DNA Sequence Data
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Gerardi, M.H. The Microbiology of Anaerobic Digesters; John Wiley & Sons: Hoboken, NJ, USA, 2003; pp. 3–10. [Google Scholar]
- Kashyap, D.R.; Dadhich, K.S.; Sharma, S.K. Biomethanation under psychrophilic conditions: A review. Bioresour. Technol. 2003, 87, 147–153. [Google Scholar] [PubMed]
- Huttunen, S.; Lampinen, A. Bioenergy Technology Evaluation and Potential in Costa Rica; University of Jyväskylä Printing House: Jyväskylä, Finland, 2005. [Google Scholar]
- Lansing, S.; Botero, R.B.; Martin, J.F. Waste treatment and biogas quality in small-scale agricultural digesters. Bioresour. Technol. 2008, 99, 5881–5890. [Google Scholar] [PubMed]
- Dhaked, R.K.; Singh, P.; Singh, L. Biomethanation under psychrophilic conditions. Waste Manag. 2010, 30, 2490–2496. [Google Scholar] [CrossRef] [PubMed]
- Massé, D.I.; Gilbert, Y.; Saady, N.M.C.; Liu, C. Low-temperature anaerobic digestion of swine manure in a plug-flow reactor. Environ. Technol. 2013. [Google Scholar] [CrossRef]
- Oz, N.A.; Ince, O.; Turker, G.; Ince, B.K. Effect of seed sludge microbial community and activity on the performance of anaerobic reactors during the start-up period. World J. Microbiol Biotechnol. 2012, 28, 637–647. [Google Scholar] [CrossRef] [PubMed]
- Marzorati, M.; Wittenbolle, L.; Boon, N.; Daffonchio, D.; Versraete, W. How to get more out of molecular fingerprints: Practical tools for microbial ecology. Environ. Microbiol. 2008, 10, 1571–1581. [Google Scholar] [CrossRef] [PubMed]
- Savant, D.V.; Shouche, Y.S.; Prakash, S.; Ranade, D.R. Methanobrevibacter acididurans sp. nov., a novel methoanogen from a sour anaerobic digester. Int. J. Syst. Evolut. Microbiol. 2002, 52, 1081–1087. [Google Scholar] [CrossRef]
- Pandey, J.; Ganesan, K.; Jain, R.K. Variations in T-RFLP profiles with differing chemistries of fluorescent dyes used for labeling the PCR primers. J. Microbiol. Methods 2006, 68, 633–638. [Google Scholar] [CrossRef] [PubMed]
- Chouari, R.; Paslier, D.L.; Daegelen, P.; Ginestet, P.; Weissenbach, J.; Sghir, A. Novel predominant archaeal and bacterial groups revealed by molecular analysis of an anaerobic sludge digester. Environ. Microbiol. 2005, 7, 1104–1115. [Google Scholar] [CrossRef] [PubMed]
- Ferry, J.G. Methanogenesis: Ecology, Physiology, Biochemistry and Genetics; Chapman and Hall: New York, NY, USA, 1993. [Google Scholar]
- Rastogi, G.; Ranade, D.R.; Yeole, T.Y.; Patole, M.S.; Shouche, Y.S. Investigation of methanogen population structure in biogas reactor by molecular characterization of methyl-co-enzyme M reductase A (mcrA) genes. Bioresour. Technol. 2008, 99, 5317–5326. [Google Scholar] [CrossRef] [PubMed]
- Demirel, B.; Scherer, P. The role of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: A review. Rev. Environ. Sci. Biotechnol. 2008, 7, 173–190. [Google Scholar] [CrossRef]
- Bialek, K.; Kumar, A.; Mahony, T.; Lens, P.N.; O’Flaherty, V. Microbial community structure and dynamics in anaerobic fluidized-bed and granular sludge-bed reactors: Influence of operational temperature and reactor configuration. Microb. Biotechnol. 2012, 5, 738–752. [Google Scholar] [CrossRef] [PubMed]
- Ziganshin, A.M.; Schmidt, T.; Scholwin, F.; Il’inskaya, O.N.; Harms, H.; Kleinsteuber, S. Bacteria and archaea involved in anaerobic digestion of distillers grains with solubles. Appl. Microbiol. Biotechnol. 2011, 89, 2039–2052. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Tovenen, K.; Lehtomäki, A.; Rintala, J. Microbial community structure in anaerobic co-digestion of grass silage and cow manure in a laboratory continuously stirred tank reactor. Biodegradation 2010, 21, 135–146. [Google Scholar]
- Liu, Y.; Whitman, W.B. Metabolic, phylogenetic and ecological diversity of the methonogenic archaea. Ann. N.Y. Acad. Sci. 2008, 1125, 171–189. [Google Scholar] [CrossRef] [PubMed]
- Carballa, M.; Smits, M.; Etchebeher, C.; Boon, N.; Verstraete, W. Correlations between molecular and operational parameters in continuous lab-scale anaerobic reactors. Appl. Microbiol. Biotechnol. 2011, 89, 303–314. [Google Scholar] [CrossRef] [PubMed]
- NIIR (National Institute of Industrial Research) Board. Handbook on Bio Gas and Its Applications; National Institute of Industrial Research: New Delhi, India, 2000. [Google Scholar]
- Kane, M.D.; Poulsen, L.K.; Stahl, D.A. Monitoring the enrichment and isolation of sulfate reducing bacteria by using oligonucleotide hybridization probes designed from environmentally derived 16S rRNA sequences. Appl. Environ. Microbiol. 1993, 59, 682–686. [Google Scholar] [PubMed]
- Lane, D.J.; Pace, B.; Olsen, G.J.; Stahl, D.A.; Sogin, M.L.; Pace, N.R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analysis. Proc. Natl. Acad. Sci. USA 1985, 82, 6955–6959. [Google Scholar] [CrossRef] [PubMed]
- Grosskopf, R.; Janssen, P.H.; Liesack, W. Diversity and structure of the methanogenic community in anoxic rice paddy soil microcosms as examined by cultivation and direct 16S rRNA gene sequence retrieval. Appl. Environ. Microbiol. 1998, 64, 960–969. [Google Scholar] [PubMed]
- Lueders, T.; Friedrich, M. Archaeal population dynamics during sequential reduction processes in rice field soil. Appl. Environ. Microbiol. 2000, 66, 2732–2742. [Google Scholar] [CrossRef] [PubMed]
- Padmasiri, S.I.; Zhang, J.; Fitch, M.; Norddahl, B.; Morgenroth, E.; Raskin, L. Methanogenic population dynamics and performance of an anaerobic membrane bioreactor (AnMBR) treating swine manure under high shear conditions. Water Res. 2007, 41, 134–144. [Google Scholar] [CrossRef] [PubMed]
- Microbial Community Analysis (MiCA) Web Page. Available online: http:/mica.ibest.uidaho.edu/ (accessed on 3 July 2012).
- New England Biolabs (NEB) Web Page. Available online: http://tools.neb.com/NEBcutter2/ (accessed on 20 August 2012).
- Lossie, U.; Pütz, P. Targeted Control of Biogas Plants with the Help of FOS/TAC; Practice Report; Hach Lange: Salford, UK, 2001. [Google Scholar]
- Krebs, C.J. Ecological Methodology, 2nd ed.; Benjamin Cummings: San Francisco, CA, USA, 1998; pp. 293–301. [Google Scholar]
- National Center for Biotechnology Information (NCBI) Web Page. Available online: http://www.ncbi.nlm.nih.gov/ (accessed on 13 June 2012).
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ciotola, R.J.; Martin, J.F.; Castańo, J.M.; Lee, J.; Michel, F. Microbial Community Response to Seasonal Temperature Variation in a Small-Scale Anaerobic Digester. Energies 2013, 6, 5182-5199. https://doi.org/10.3390/en6105182
Ciotola RJ, Martin JF, Castańo JM, Lee J, Michel F. Microbial Community Response to Seasonal Temperature Variation in a Small-Scale Anaerobic Digester. Energies. 2013; 6(10):5182-5199. https://doi.org/10.3390/en6105182
Chicago/Turabian StyleCiotola, Richard J., Jay F. Martin, Juan M. Castańo, Jiyoung Lee, and Frederick Michel. 2013. "Microbial Community Response to Seasonal Temperature Variation in a Small-Scale Anaerobic Digester" Energies 6, no. 10: 5182-5199. https://doi.org/10.3390/en6105182
APA StyleCiotola, R. J., Martin, J. F., Castańo, J. M., Lee, J., & Michel, F. (2013). Microbial Community Response to Seasonal Temperature Variation in a Small-Scale Anaerobic Digester. Energies, 6(10), 5182-5199. https://doi.org/10.3390/en6105182