Microbial Monitoring of Surface Water in South Africa: An Overview
Abstract
:1. Introduction
2. Legislative Framework for the Microbial Quality Monitoring of Surface Water in South Africa
3. National Microbial Monitoring Programme for Surface Water (NMMP)
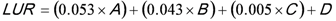
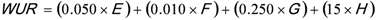
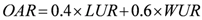
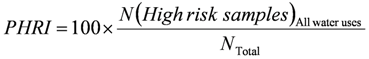
4. Advantages of the Existing Microbial Water Quality Monitoring Framework
5. Drawbacks of the Existing Framework for Microbial Water Quality Monitoring in South Africa
6. Faecal-Source Tracking and Proposed Changes to the Existing Framework of Microbial Water Quality Testing in South Africa
6.1. Viral Methods for Faecal-Source Tracking
6.2. Bacterial Methods for Faecal-Source Tracking
6.3. Microbial Water Quality Monitoring in Remote Areas
7. Conclusions
Acknowledgments
Conflict of Interest
References
- Schreiner, B.; van Zyl, F. Water Services: Yesterday, Today and Tomorrow—A Strategic Perspective. Durban, South Africa, 2006; Available online: http://www.dwa.gov.za/communications/DepartmentalSpeeches/2006/Wisa06Schreiner.doc (accessed on 4 December 2009). Presented at The WISA Biennial Conference and Exhibition 2006..
- Department of Water Affairs and Forestry. Water Supply and Sanitation Policy White Paper. 1994. Available online: http://www.dwaf.gov.za/Documents/Policies/WSSP.pdf (accessed on 21 February 2010).
- Statistics South Africa, General Household Survey 2010 (Revised Version); Statistics South Africa, Pretoria, South Africa, 2011; Statistical release P0318.
- Health Systems Trust. Diarrhoea Incidence in under 5 Year Old Children in South Africa. 2011. Available online: http://www.hst.org.za/recently-updated-indicators (accessed on 4 October 2011).
- Le Roux, A.; van Hyussteen, E. Socio-Economic Landscape: The South African Socio-Economic and Settlement Landscape. In South Africa Risk and Vulnerability Atlas; the South African National Department of Science and Technology, CPD Print: Pretoria, South Africa, 2010; pp. 15–21. [Google Scholar]
- Central Intelligence Agency. The World Factbook: South Africa. 2011. Available online: https://www.cia.gov/library/publications/the-world-factbook/fields/2212.html (accessed on 4 January 2011).
- Cohen, B. Urban growth in developing countries: A review of current trends and a caution regarding existing forecasts. World Dev. 2004, 32, 23–51. [Google Scholar] [CrossRef]
- Statistics South Africa, Mid-Year Population Estimates 2011; Statistics South Africa: Pretoria, South Africa, 2011; Statistical Release P0302.
- Van Hyussteen, E.; Oranje, M.C.; Robinson, S.; Makoni, E. South Africa’s city regions: A call for contemplation and action. Urban Forum 2009, 20, 175–195. [Google Scholar] [CrossRef]
- Statistics South Africa, GHS Series Volume III: Water and Sanitation 2002–2010 (In-Depth Analysis of the General Household Survey Data); Statistics South Africa: Pretoria, South Africa, 2011; Report No. 03-18-02 2002–2010.
- Statistics South Africa, Mid-Year Population Estimates 2007; Statistics South Africa: Pretoria, South Africa, 2007; Statistical release P0302.
- Statistics South Africa, South African Statistics; Statistics South Africa: Pretoria, South Africa, 2010.
- Luyt, C.D.; Muller, W.J.; Tandlich, R. Low-cost tools for microbial quality assessment of drinking water in South Africa. Int. J. Collab. Res. Int. Med. Public Health 2011, 3, 236. [Google Scholar]
- Lewin, S.; Norman, R.; Nannan, N.; Thomas, E.; Bradshaw, D. The South African comparative risk assessment collaborating group. Estimating the burden of disease attributable to unsafe water and lack of sanitation and hygiene in South Africa in 2000. S. Afr. Med. J. 2007, 97, 755–762. [Google Scholar]
- Parliamentary Monitoring Group. Water Cuts & Cholera Situation in City of Cape Town: City, Departmental & Civil Community Responses. Available online: http://www.pmg.org.za/report/20090204-department-health-department-water-affairs-forestry-responses-cholera (accessed on 11 February 2010).
- Haarhoff, J.; Rietveld, L.C.; Jagals, P. Rapid Technical Assessment and Troubleshooting of Rural Water Supply Systems. In Proceedings of the 10th Annual Water Distribution Systems Analysis Conference, Kruger National Park, South Africa, 17–20 August 2008.
- Momba, M.N.B.; Tyafa, Z..; Brouckaert, B.M.; Obi, C.L. Safe drinking water still a dream in rural areas of South Africa. Case study: The Eastern Cape Province. Water SA 2006, 32, 715–720. [Google Scholar]
- Tandlich, R.; Muller, W.J. Methods for Detection of Faecal Pollution and Identifying Its Origins: Linking the South African and the International Context; Water Research Commission: Pretoria, South Africa, 2008; Progress Report No. K8/806. [Google Scholar]
- Department of Water Affairs and Forestry of South Africa. South African Water Quality Guidelines, Volume 1. (Domestic Use). 1996. Available online: http://www.dwa.gov.za/iwqs/wq_guide/Pol_saWQguideFRESH_vol1_Domesticuse.PDF (accessed on 19 February 2010).
- South African Bureau of Standards, South African Standard Specifications for Water for Domestic Supplies; South African Bureau of Standards: Pretoria, South Africa, 2006; Standard No. 241/2006.
- Murray, K.; du Preez, M.; Kuhn, A.L.; van Niekerk, H. A Pilot Study to Demonstrate Implementation of the National Microbial Monitoring Programme; Water Research Commission: Pretoria, South Africa, 2004; WRC Report No. 1118/1/04. [Google Scholar]
- Monyai, P. Health-Related Water Quality and Surveillance Model for the Peddie District in the Eastern Cape; Water Research Commission: Pretoria, South Africa, 2004; WRC Report No.: 727/1/04. [Google Scholar]
- Grabow, W.O.K.; Favorov, M.O.; Khudyakova, N.S.; Taylor, M.B.; Fields, H.A. Hepatitis E seroprevalence in selected individuals in South Africa. J. Virol. Methods 1994, 44, 384–388. [Google Scholar] [CrossRef]
- Dalsgaard, A.; Forslund, A.; Sandvang, D.; Arntzen, L.; Keddy, K. Vibrio Cholerae O1 outbreak isolates in Mozambique and South Africa in 1998 are multiple-drug resistant, contain the SXT element and the aadA2 gene located on class 1 integrons. J. Antimicrob. Chemother. 2001, 48, 827–838. [Google Scholar] [CrossRef]
- Paulse, A.N.; Jackson, V.A.; Khan, W. Comparison of microbial contamination at various sites along the Plankenburg and Diep Rivers, Western Cape, South Africa. Water SA 2009, 35, 469–478. [Google Scholar]
- National Water Act No. 36 of 1998. South African Government Gazette No. 20706. 1998. Available online: http://www.dwaf.gov.za/Documents/Legislature/nw_act/NWA.pdf (accessed on 10 December 2009).
- National Heath Act No.61 of 2003 as amended 2008 (NHA; 2009). South African Government Gazette No. 31187. 2009. Available online: http://www.mangaung.co.za/docs/National%20Health%20Act.pdf (accessed on 10 December 2009).
- Du Preez, M.; Murray, K.; van Niekerk, H. A Pilot Study to Demonstrate Implementation of the National Microbial Monitoring Programme; Research Report; Water Research Commission: Pretoria, South Africa, 2002. [Google Scholar]
- Du Preez, M.; Murray, K.; van Niekerk, H. A Pilot Study to Demonstrate Implementation of the National Microbial Monitoring Programme; Progress Report; Water Research Commission: Pretoria, South Africa, 2001. [Google Scholar]
- Du Preez, M.; Venter, S.N.; van Ginkel, C.; Harris, J.; Kuhn, A.; Zingitwa, L.; Silberbauer, M. Research on the Selection of Procedures for Faecal Pollution Monitoring to Describe Health Risks; WRC Report No. K5/824/0/1; Water Research Commission: Pretoria, South Africa, 1999. [Google Scholar]
- Department of Water Affairs of South Africa. NMMP Details of Implementation; 2010. Available online: http://www.dwa.gov.za/iwqs/microbio/nmmp.asp (accessed on 11 February 2010).
- Venter, S.N.; Kühn, A.L.; Harris, J. A method for the prioritization areas experiencing microbial pollution of surface water. Water Sci. Technol. 1998, 38, 23–27. [Google Scholar]
- Department of Water Affairs and Forestry of South Africa. Prioritisation of Faecally Contaminated Areas. 2000. Available online: http://www.dwaf.gov.za/iwqs/microbio/Document/prioritisefin2.htm (accessed on 19 February 2010).
- Parliamentary Monitoring Group, Sanitation Programme Progress and Beneficiary List Management Update: Briefing by Department of Human Settlements. Meeting minutes held on 10 August 2010. Parliament of South Africa: Cape Town, South Africa, 2010.
- Department of Water Affairs and Forestry of South Africa, Middle Vaal Water Management Area: Water Resources Situation Assessment; Directorate of Water Resource Planning in the Department of Water Affairs and Forestry: Directorate of Water Resource Planning in the Department of Water Affairs and Forestry, 2002; 1.
- Department of Water Affairs of South Africa. NMMP Bimonthly Reports 2011. 2011. Available online: http://www.dwa.gov.za/iwqs/microbio/report/NMMPkey_2011.htm (accessed on 5 January 2012).
- Genthe, B.; Franck, M. A Tool for Assessing Microbial Quality in Small Community Water Supplies: An H2S Strip Test; WRC Report No. 961/1/99; Water Research Commission: Pretoria, South Africa, 1999. [Google Scholar]
- Medema, G.J.; Payment, P.; Dufour, A.; Robertson, W.; Waite, M.; Hunter, P.; Kirby, R.; Andersson, Y. Safe Drinking Water: An Ongoing Challenge. In Assessing Microbial Safety of Drinking Water—Improving Approaches and Methods; Medema, G.J., Payment, P., Dufour, A., Robertson, W., Waite, M., Hunter, P., Kirby, R., Andersson, Y., Eds.; WHO Press: Geneva, Switzerland, 2003. [Google Scholar]
- Pruss, A. Review of epidemiological studies on health effects from exposure to recreational water. Int. J. Epidemiol. 1998, 27, 1–9. [Google Scholar] [CrossRef]
- Hardina, C.M.; Fujioka, R.S. Soil—The environmental source of Escherichia coli and Enterococci in Hawaii streams. Environ. Toxicol. Water Contam. 1991, 6, 185–195. [Google Scholar]
- Desmarais, T.R.; Solo-Gabriele, H.M.; Palmer, C.J. Influence of soil on fecal indicator organisms in a tidally influenced subtropical environment. Appl. Environ. Microbiol. 2002, 68, 1165–1172. [Google Scholar]
- Hamilton, W.P.; Kim, M.; Thackston, E.L. Comparison of commercially available Escherichia coli enumeration tests: Implications for attaining water quality standards. Water Res. 2005, 39, 4869–4878. [Google Scholar]
- IDEXX. Validation of Colilert®-18/Quanti-Tray® for the Enumeration of E. coli and Coliform Bacteria from Water. 2008. Available online: http://www.idexx.fr/pdf/fr_fr/water/7537-01-colilert-18-report-eng2.pdf (accessed 30th June 2012).
- Luyt, C.D.; Muller, W.J.; Wilhelmi, B.S.; Tandlich, R. Health Implications of Flood Disaster Management in South Africa. In Proceedings of the 18th Annual Conference of the International Emergency Management of Society, Bucharest, Romania, 7–10 June 2011; pp. 376–385.
- Munnik, V.; Molose, V.; Moore, B.; Tempelhoff, J.; Gouws, I.; Motloung, S.; Sibiya, Z.; van Zyl, A.; Malapela, P.; Buang, B.; et al. Final Consolidated Report: The Potential of Civil Society Organisations in Monitoring and Improving Water Quality; WRC Report No.: K8/968/1; Water Research Commission: Pretoria, South Africa, 2011. [Google Scholar]
- Du Preez, M.; Genthe, B. Evaluation of Rapid Methods for the Detection of Indicator Organisms in Drinking Water. 1996. Available online: http://www.ewisa.co.za/literature/files/1996%20-%2074.pdf (accessed on 8 June 2012).
- Sundram, A.; Bailey, I.W.; Green, E. The Application of New Technology Used for Identifying Bacteria from Water Samples. In Proceedings of the WISA 2000 Biennial Conference, Sun City, South Africa, 28 May–1 June 2000; Available online: http://www.ewisa.co.za/literature/files/134sundram.pdf (accessed on 13 June 2012).
- Jackson, S.A.F.; Bailey, I.; Britz, R.; de Wet, C.; Fortune, A.; Johnston-Robertson, A.J.; Naidoo, K.M.; Sundram, A. Report on the South African Colilert™ Ring Trial. In Proceedings of the 2002 Biennial Conference of the Water Institute of Southern Africa (WISA), Durban, South Africa, 19–23 May 2002.
- Chao, K.K.; Chao, C.C.; Chao, W.L. Evaluation of Colilert-18 for detection of coliforms and Escherichia coli in subtropical freshwater. Appl. Environ. Microbiol. 2004, 70, 1242–1244. [Google Scholar] [CrossRef]
- Chao, W.L. Evaluation of Colilert-18 for the detection of coliforms and Escherichia coli in tropical fresh water. Lett. Appl. Microbiol. 2006, 42, 115–120. [Google Scholar] [CrossRef]
- Pisciotta, J.M.; Rath, D.F.; Stanek, P.A.; Flanery, D.M.; Harwood, V.J. Marine bacteria cause false-positive results in the Colilert-18 rapid identification test for Escherichia coli in Florida waters. Appl. Environ. Microbiol. 2002, 68, 539–544. [Google Scholar]
- Maheux, A.F.; Huppe, V.; Boissinot, M.; Picard, F.J.; Bissonnette, L.; Bernier, J.L.T.; Bergeron, M.G. Analytical limits of four beta-glucuronidase and beta-galactosidase-based commercial culture Methods used to detect Escherichia coli and total Coliforms. J. Microbiol. Methods 2008, 75, 506–514. [Google Scholar] [CrossRef]
- Martins, M.T.; Rivera, I.G.; Clark, D.L.; Stewart, M.H.; Wolfe, R.L.; Olson, B.H. Distribution of uidA gene sequences in Escherichia coli isolates in water sources and comparison with the expression of beta-glucuronidase activity in 4-methylumbelliferyl-beta-D-glucuronide media. Appl. Environ. Microbiol. 1993, 59, 2271–2276. [Google Scholar]
- Gronewold, N.D.; Wolpert, R.L. Modeling the relationship between most probable number (MPN) and colony-forming unit (CFU) estimates of fecal coliform concentration. Water Res. 2008, 42, 3327–3334. [Google Scholar] [CrossRef]
- Cho, K.H.; Han, D.; Park, Y.; Lee, S.W.; Cha, S.M.; Kang, J.-H.; Kim, J.H. Evaluation of the relationship between two different methods for enumeration fecal indicator bacteria: Colony-forming unit and most probable number. J. Environ. Sci. 2010, 22, 846–850. [Google Scholar]
- Rivett, U.; Loudon, M.; Wright, J. Introduction the Aquatest Project: Increasing the Ability to Monitor Water Quality at Supply Level. 2009. Available online: http://spatialdatamanagement.uct.ac.za/assets/RivettEtAl2009.pdf (accessed on 21 February 2010).
- Moraka, W. Water Quality Presentation to the Portfolio Committee on Water Affairs and Forestry: Local Government Perspective. Water Quality and Security: Public Hearings. In Proceedings of the Portfolio Committee on Water Affairs and Forestry, Cape Town, South Africa, 20 June 2006.
- Portfolio Committee on Water Affairs and Forestry, Water Quality and Water Challenges: Public Hearings; Meeting Minutes Held on 4 June 2008; Parliament of South Africa: Cape Town, South Africa, 2008.
- Grabow, W.O.K.; Vrey, A.; Uys, M.; de Villiers, J.C. Evaluation of the Application of Bacteriophages as Indicators of Water Quality; WRC Report No. 540/1/98; Water Research Commission: Pretoria, South Africa, 1998. [Google Scholar]
- Sundram, A.; Jumanlal, N.; Ehlers, M.M. Genotyping F-RNA coliphages isolated from wastewater and river water samples. Water SA 2006, 32, 65–70. [Google Scholar]
- Momba, M.N.B.; Sibewu, M.; Mandeya, A. Survival of somatic and F-RNA coliphages in treated wastewater effluents and their impact on viral quality of the receiving water bodies in the Eastern Cape Province-South Africa. J. Biol. Sci. 2009, 9, 648–654. [Google Scholar]
- Grabow, W.O.K. Bacteriophages: Update on application as models for viruses in water. Water SA 2001, 27, 251–268. [Google Scholar]
- Tartera, C.; Jofre, J. Bacteriophages active against Bacteroides fragilis in sewage-polluted waters. Appl. Environ. Microbiol. 1987, 53, 1632–1637. [Google Scholar]
- Tartera, C.; Bosch, A.; Jofre, J. The inactivation of bacteriophages infecting Bacteroides fragilis by chlorine treatment and UV-irradiation. FEMS Microbiol. Lett. 1988, 56, 313–316. [Google Scholar] [CrossRef]
- Schaper, M.; Jofre, J. Comparison of Methods for detecting genotypes of F-specific bacteriophages and fingerprinting the origin of faecal pollution in water samples. J. Virol. Methods 2000, 89, 1–10. [Google Scholar] [CrossRef]
- International Standardisation Organisation, Water Quality—Detection and Enumeration of Bacteriophages—Part 1: Enumeration of F-Specific RNA Bacteriophages. ISO 10705-1; International Standardisation Organisation: Geneva, Switzerland, 1995.
- Schaper, M.; Jofre, J.; Uys, M.; Grabow, W.O.K. Distribution of genotypes of F-specific RNA bacteriophages in human and non-human sources of faecal pollution in South Africa and Spain. J. Appl. Microbiol. 2002, 92, 657–667. [Google Scholar] [CrossRef]
- Grabow, W.O.K.; Neubrech, T.E.; Holtzhausen, C.S.; Jofre, J. Bacteroides fragilis and Escherichia coli bacteriophages: Excretion by humans and animals. Water Sci. Technol. 1995, 31, 223–230. [Google Scholar]
- Hsu, F.C.; Shieh, Y.S.C.; van Duin, J.; Beekwilder, M.J.; Sobsey, M.D. Genotyping male-specific bacteriophages by hybridization with oligonucleotide probes. Appl. Environ. Microbiol. 1995, 61, 3960–3966. [Google Scholar]
- Puig, A.; Jofre, J.; Araujo, R. Use of oxyrase enzyme (Oxyrase®) for the detection of bacteriophages of Bacteroides fragilis in aerobic incubating conditions (short communication). J. Microbiol. Methods 1998, 31, 205–207. [Google Scholar] [CrossRef]
- Puig, A.; Queralt, N.; Jofre, J.; Araujo, R. Diversity of Bacteroides fragilis strains in their capacity to recover phages from human and animal wastes and from fecally polluted wastewater. Appl. Environ. Microbiol. 1999, 65, 1772–1776. [Google Scholar]
- Goyal, S.M.; Gerba, C.P.; Bitton, G. Phage Ecology, 1st ed; John Wiley and Sons: New York, NY, USA, 1987. [Google Scholar]
- Kai, S.; Watanabe, S.; Furuse, K.; Osawa, A. Bacteroides bacteriophages isolated from human faeces. Microbiol. Immunol. 1985, 29, 895–899. [Google Scholar]
- Murray, K.; du Preez, M.; Taylor, M.B.; Meyer, R.; Parsons, R.; van Wyk, E.; Kuhn, A.; van Niekerk, H.; Ehlers, M.M. National Microbial Monitoring Programme for Groundwater Research Report; WRC Report No. 1277/1/04; Water Research Commission: Pretoria, South Africa, 2007. [Google Scholar]
- International Standardisation Organisation, Water Quality—Detection and Enumeration of Bacteriophages, Part 4: Enumeration of Bacteriophages Infecting Bacteroides fragilis; International Standardisation Organisation: Geneva, Switzerland, 1998; ISO/CD 10705-4.
- Stoeckel, D.M.; Harwood, V.J. Performance, design and analysis of microbial source tracking. Appl. Environ. Microbiol. 2007, 73, 2405–2415. [Google Scholar] [CrossRef]
- Harwood, V.J.; Wiggins, B.; Hagedorn, C.; Ellender, R.D.; Gooch, J.; Kern, J.; Samadpour, M.; Chapman, A.C.H.; Robinson, B.J.; Thompson, B.C. Phenotypic library-based microbial source tracking methods: Efficacy in the California collaborative study. J. Water Health 2003, 1, 153–166. [Google Scholar]
- Lin, J.; Biyela, P.T.; Puckree, T. Antibiotic resistance profiles of environmental isolates from Mhlathuze River, KwaZulu-Natal (RSA). Water SA 2004, 30, 23–28. [Google Scholar]
- Said, M.; le Roux, W.J.; Burke, L.; Said, H.; Paulsen, L.; Venter, S.N.; Potgieter, N.; Masoabi, D.; de Wet, C.M.E. Origin, Fate and Clinical Relevance of Water-borne Pathogens Present in Surface Waters; WRC Report No. 1398/1/05; Water Research Commission: Pretoria, South Africa, 2005. [Google Scholar]
- Doughari, H.J.; Ndakidemi, P.A.; Human, I.S.; Benade, S. Virulence factors and antibiotic susceptibility among verotoxic non O157: H7 Escherichia coli isolates obtained from water and wastewater samples in Cape Town, South Africa. Afr. J. Biotechnol. 2011, 10, 14160–14168. [Google Scholar]
- Obi, C.L.; Ramalivhana, J.; Momba, M.N.B.; Onabolu, B.; Igumbor, J.O.; Lukoto, M.; Mulaudzi, T.B.; Bessong, P.O.; Janse van Rensburg, E.L.; Green, E.; et al. Antibiotic resistance profiles and relatedness of enteric bacterial pathogens isolated from HIV/AIDS patients with and without diarrhoea and their household drinking water in rural communities in Limpopo Province South Africa. Afr. J. Biotechnol. 2007, 6, 1035–1047. [Google Scholar]
- Blanch, A.R.; Belanche-Muňoz, L.; Bonjoch, X.; Ebdon, J.; Gantzer, C.; Lucena, F.; Ottoson, J.; Kourtis, C.; Iversen, A.; Kühn, I.; et al. Integrated analysis of established and novel microbial and chemical methods for microbial source tracking. Appl. Environ. Microbiol. 2006, 72, 5915–5926. [Google Scholar] [CrossRef]
- Mitsuoka, T. Bifidobacteria and their role in human health. J. Ind. Microbiol. 1990, 6, 263–268. [Google Scholar] [CrossRef]
- Bonjoch, X.; Ballesté, E.; Blanch, A.R. Enumeration of bifidobacterial populations with selective media to determine the source of waterborne fecal pollution. Water Res. 2005, 39, 1621–1625. [Google Scholar] [CrossRef]
- Tannock, G.W. Probiotic properties of lactic-acid Bacteria: Plenty of scope for fundamental R & D. Trends Biotechnol. 1997, 15, 270–274. [Google Scholar] [CrossRef]
- Carillo, M.; Estrada, E.; Hazen, T.C. Survival and enumeration of the fecal indicators Bifidobacterium adolescentis and Escherichia coli in a tropical rain forest watershed. Appl. Environ. Microbiol. 1985, 50, 468–476. [Google Scholar]
- Ijabadeniyi, O.A.; Debusho, L.K.; van der Linde, M.; Buys, E.M. Irrigation water as a potential preharvest source of bacterial contamination of vegetables. J. Food Saf. 2011, 31, 452–461. [Google Scholar] [CrossRef]
- Resnick, I.G.; Levin, M.A. Quantitative procedure for enumeration of bifidobacteria. Appl. Environ. Microbiol. 1981, 42, 433–438. [Google Scholar]
- Resnick, I.G.; Levin, M.A. Assessment of bifidobacteria as indicators of human fecal pollution. Appl. Environ. Microbiol. 1981, 42, 427–432. [Google Scholar]
- Mara, D.D.; Oragui, J.I. Sorbitol-fermenting bifidobacteria as specific indicators of human faecal pollution. J. Appl. Bacteriol. 1983, 55, 349–357. [Google Scholar] [CrossRef]
- Bonjoch, X.; Lucena, F.; Blanch, A.R. The persistence of bifidobacteria populations in a river measured by molecular and culture techniques. J. Appl. Microbiol. 2009, 107, 1178–1185. [Google Scholar] [CrossRef]
- Ottoson, J.R. Bifidobacterial survival in surface water and implications for microbial source tracking. Can. J. Microbiol. 2009, 55, 642–647. [Google Scholar] [CrossRef]
- Jagals, P.; Grabow, W.O.K. An evaluation of sorbitol-fermenting bifidobacteria as specific indicators of human faecal pollution of environmental water. Water SA 1996, 22, 235–238. [Google Scholar]
- Jagals, P. Stormwater runoff from typical developed and developing South African urban developments: Definitely not for swimming. Water Sci. Technol. 1997, 35, 133–140. [Google Scholar] [CrossRef]
- American Public Health AssociationAmerican Water Works AssociationWater Environmental Federation (APHA)Standard Methods for the Examination of Water and Wastewater, 20th ed; American Public Health Association: Washington, DC, USA.
- Kromoredjo, P.; Fujioka, R.S. Evaluating three simple methods to assess the microbial quality of drinking water in Indonesia. Environ. Toxicol. Water Q. 1991, 6, 259–270. [Google Scholar] [CrossRef]
- Genthe, B.; Jagals, P. Application of an H2S Strip as a Field Test Kit for Microbial Water Quality Assessment; WRC Report No. 1105/1/03; Water Research Commission: Pretoria, South Africa, 2003. [Google Scholar]
- Mosley, L.M.; Sharp, D.S. The Hydrogen Sulphide (H2S) Paper-Strip Test: A Simple Test for Monitoring Drinking Water Quality in the Pacific Islands; 2005; South Pacific Applied Geoscience Commission and WHO Technical Report 373. [Google Scholar]
- Sobsey, M.D.; Pfaender, F.K. Evaluation of the H2S Method for Detection of Fecal Contamination of Drinking Water; WHO Report WHO/SDE/WSH/02.08; Water, Sanitation and Health, Department of Protection and the Human Environment, World Health Organization: Geneva, Switzerland, 2002. [Google Scholar]
- Ratto, A.; Dutka, B.J.; Vega, C.; Lopez, C.; El-Shaarawi, A. Potable water safety assessed by coliphage and bacterial tests. Water Res. 1989, 23, 253–255. [Google Scholar]
- Manja, K.S.; Maurya, M.S.; Rao, K.M. A simple field test for the detection of faecal pollution in drinking water. Bull. WHO 1982, 60, 797–801. [Google Scholar]
- Nagaraju, D.; Sastri, J.C.V. Confirmed faecal pollution to bore well waters of Mysore city. Environ. Geol. 1999, 38, 322–326. [Google Scholar]
- Proteus Vulgaris and Proteus Mirabilis. 2009. Available online: http://www.sunysccc.edu/academic/mst/microbes/17pvulg.htm (accessed on 2 March 2009).
- Shandera, W.X.; Tacket, C.O.; Blake, P.A. Food poisoning due to Clostridium perfringens in the United States. J. Infect. Dis. 1983, 147, 167–170. [Google Scholar] [CrossRef]
- Popoff, M.Y.; le Minor, L. Antigenic Formulas of the Salmonella Serovars, 7th ed; WHO Collaborating centre for reference research on Salmonella. Institut Pasteur: Paris, France, 1997. [Google Scholar]
- Abbott, S. Klebsiella, Enterobacter, Citrobacter, and Serratia. In Manual of Clinical Microbiology; Murray, P.R., Baron, E.J., Pfaller, M.A., Tenover, F.C., Yolken, R.H., Eds.; American Society for Microbiology: Washington, DC, USA, 1997; pp. 475–483. [Google Scholar]
- Sawant, A.A.; Hegde, N.V.; Straley, B.A.; Donaldson, S.C. Antimicrobial-resistant enteric bacteria from dairy cattle. Appl. Environ. Microbiol. 2007, 73, 156–163. [Google Scholar] [CrossRef]
- Duncan, D.W.; Razzell, W.E. Klebsiella biotypes among coliforms isolated from forest environments and farm produce. Appl. Microbiol. 1972, 24, 933–938. [Google Scholar]
- Luyt, C.D.; Muller, W.J.; Tandlich, R. Factors influencing the results of microbial surface water testing in South Africa. Int. J. Med. Microbiol. 2011, 301 (Suppl 1), 28–29. [Google Scholar]
- River Health Programme: SASS Proficiency Testing Procedure (SASS, 2010). The River Health Programme, 2010. Available online: http://www.dwa.gov.za/iwqs/rhp/quality/procedure.pdf (accessed on 9 April 2010).
- Chuang, P.; Trottier, S.; Murcott, S. Comparison and verification of four field-based microbiological tests: H2S test, Easygel®, Colilert®, PetrifilmTM. J. Water Sanit. Hyg. Dev. 2011, 1, 68–85. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Luyt, C.D.; Tandlich, R.; Muller, W.J.; Wilhelmi, B.S. Microbial Monitoring of Surface Water in South Africa: An Overview. Int. J. Environ. Res. Public Health 2012, 9, 2669-2693. https://doi.org/10.3390/ijerph9082669
Luyt CD, Tandlich R, Muller WJ, Wilhelmi BS. Microbial Monitoring of Surface Water in South Africa: An Overview. International Journal of Environmental Research and Public Health. 2012; 9(8):2669-2693. https://doi.org/10.3390/ijerph9082669
Chicago/Turabian StyleLuyt, Catherine D., Roman Tandlich, Wilhelmine J. Muller, and Brendan S. Wilhelmi. 2012. "Microbial Monitoring of Surface Water in South Africa: An Overview" International Journal of Environmental Research and Public Health 9, no. 8: 2669-2693. https://doi.org/10.3390/ijerph9082669
APA StyleLuyt, C. D., Tandlich, R., Muller, W. J., & Wilhelmi, B. S. (2012). Microbial Monitoring of Surface Water in South Africa: An Overview. International Journal of Environmental Research and Public Health, 9(8), 2669-2693. https://doi.org/10.3390/ijerph9082669




