Simultaneous Detection of Nine Key Bacterial Respiratory Pathogens Using Luminex xTAG® Technology
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethics
2.2. Bacterial Strains
2.3. Clinical Specimens
2.4. DNA Extraction
2.5. Primers for Pathogen Amplification
2.6. Multiplex PCR (mPCR) Amplification
2.7. PCR Product Identification
2.8. Luminex Assay
2.9. MPLT Data Analysis
2.10. The Evaluation of Sensitivity
2.11. The Evaluation of Specificity
2.12. Detection of DNA from Clinical BALF Samples
2.13. Real-Time PCR for Bacterial Detection
2.14. Statistical Analyses
3. Results
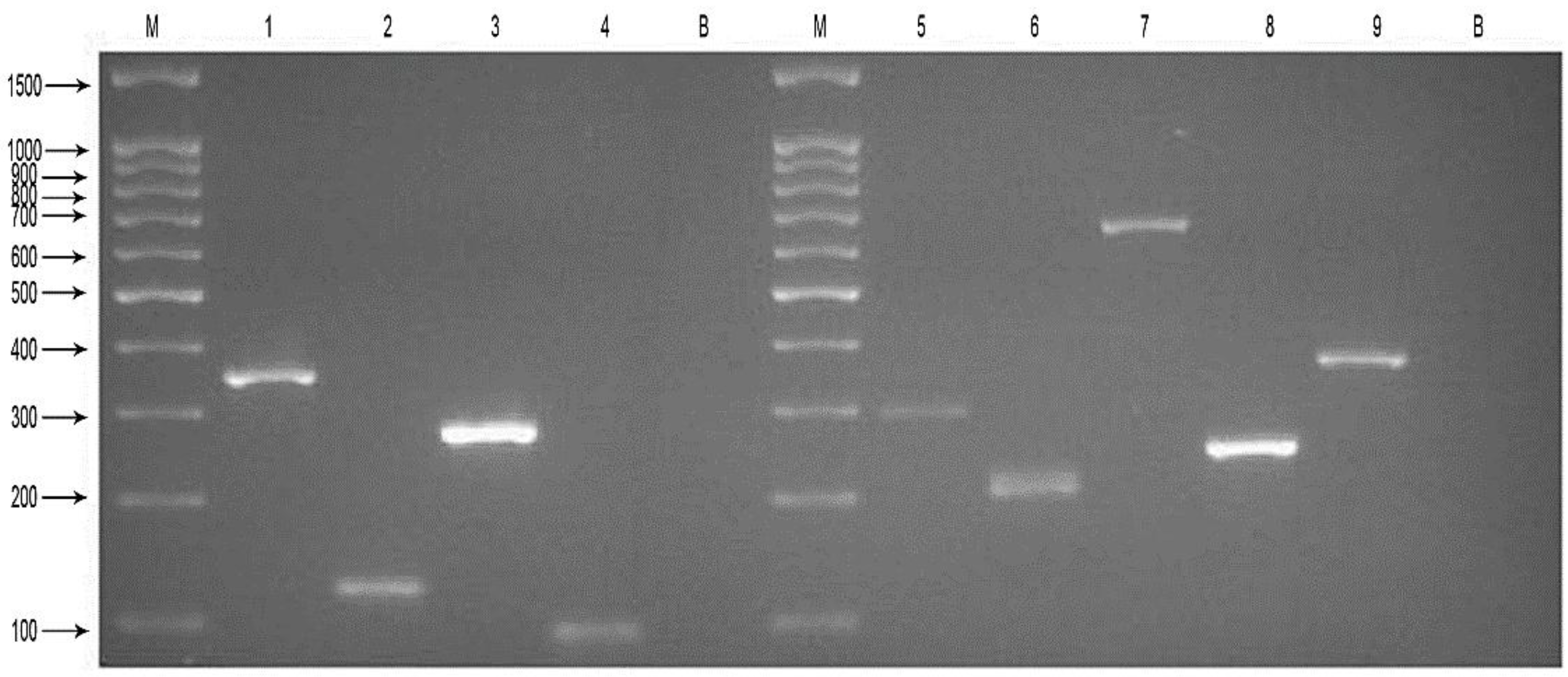
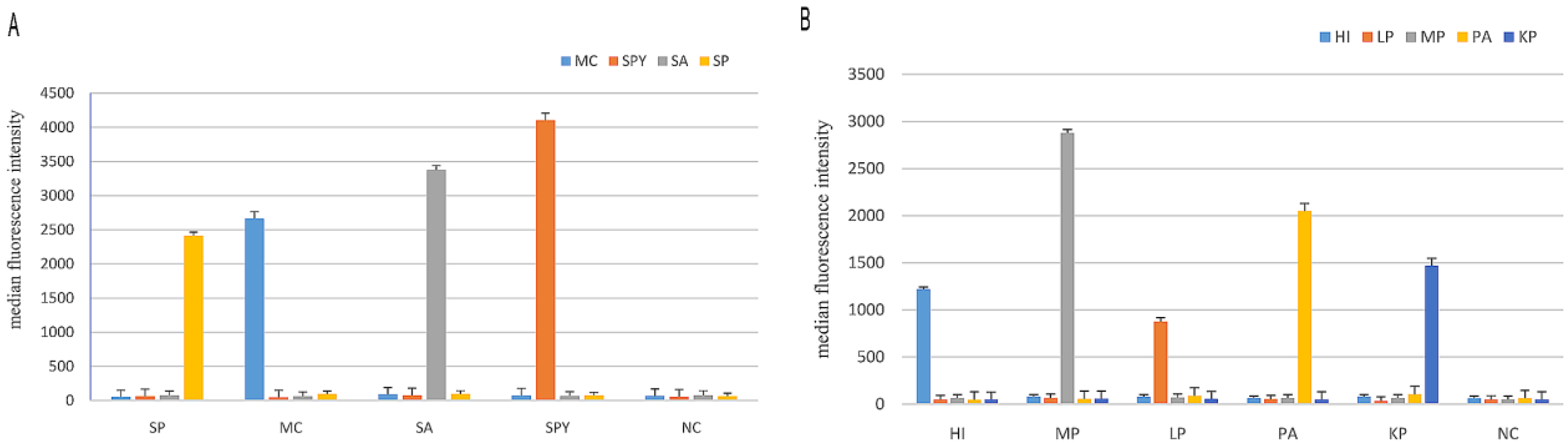
3.1. Specificity of the Two mPCR Assays and the MPLT Assay
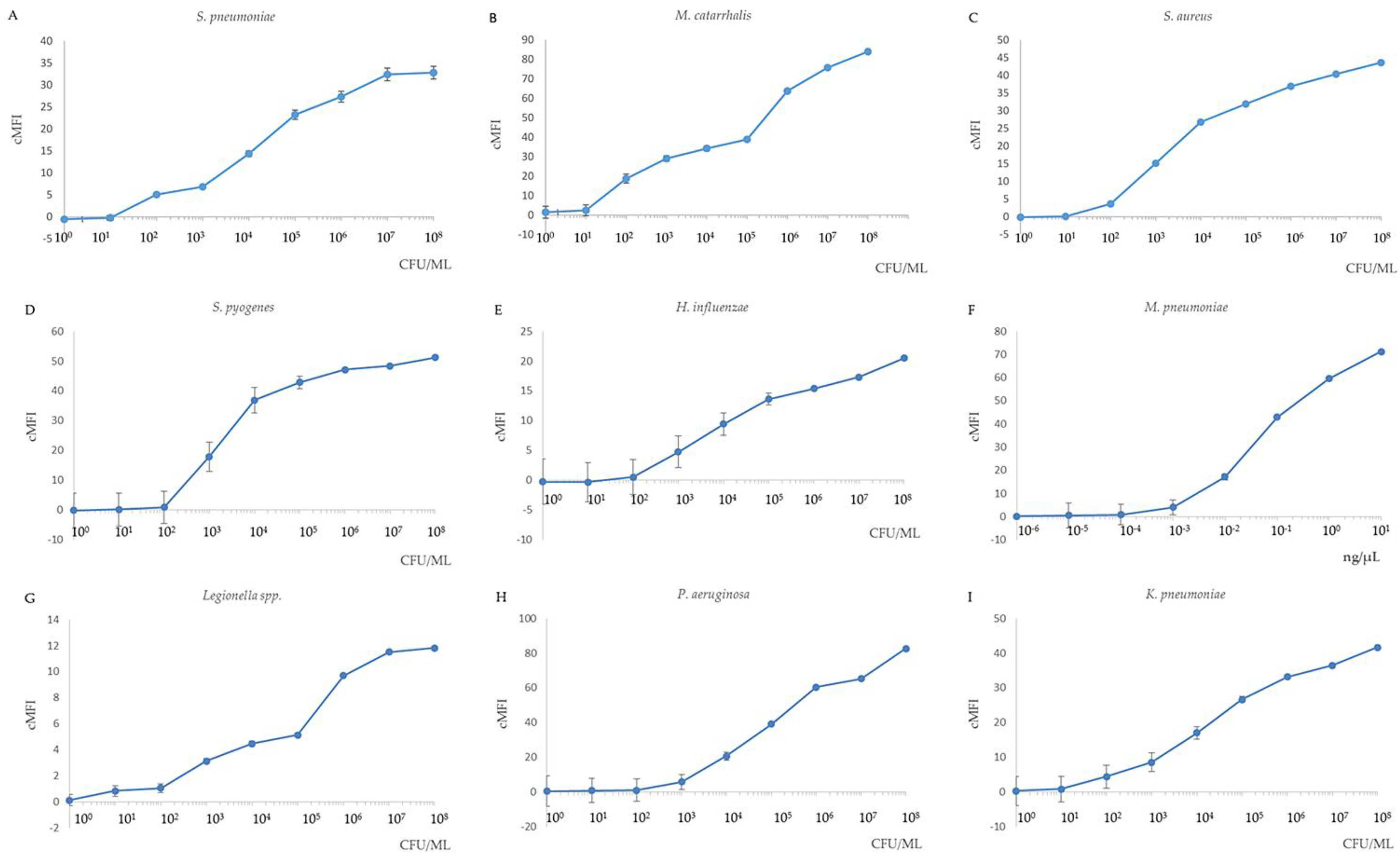
3.2. Sensitivity of the MPLT Assay
3.3. Application to Clinical Samples
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| LRTI | lower respiratory tract infection |
| LRTIs | lower respiratory tract infections |
| MPLT | multiplex PCR and Luminex technology |
| PCR | polymerase chain reaction |
| BALF | bronchoalveolar lavage fluid |
| S. pneumonia | Streptococcus pneumoniae |
| M. catarrhalis | Moraxella catarrhalis |
| S. aureus | Staphylococcus aureus |
| S. pyogenes | Streptococcus pyogenes |
| H. influenza | Haemophilus influenzae |
| M. pneumonia | Mycoplasma pneumoniae |
| P. aeruginosa | Pseudomonas aeruginosa |
| K. pneumonia | Klebsiella pneumoniae |
| mPCR | multiplex PCR; SAPE, Streptavidin, R-Phycoerythrin Conjugate |
| MFI | median fluorescence intensity |
| cMFI | corrected MFI |
| SEM | standard error of the mean |
| CFU | colony forming unit |
| Ct | threshold cycle |
References
- Woodhead, M.; Blasi, F.; Ewig, S.; Garau, J.; Huchon, G.; Ieven, M.; Ortqvist, A.; Schaberg, T.; Torres, A.; van der Heijden, G.; et al. Joint Taskforce of the European Respiratory Society and European Society for Clinical Microbiology and Infectious Diseases Guidelines for the management of adult lower respiratory tract infections—Full version. Clin. Microbiol. Infect. 2011, 17, E1–E59. [Google Scholar] [CrossRef] [PubMed]
- Prina, E.; Ranzani, O.T.; Torres, A. Community-acquired pneumonia. Lancet 2015, 386, 1097–1108. [Google Scholar] [CrossRef]
- Carroll, K.C. Laboratory diagnosis of lower respiratory tract infections: Controversy and conundrums. J. Clin. Microbiol. 2002, 40, 3115–3120. [Google Scholar] [CrossRef] [PubMed]
- Johansson, N.; Kalin, M.; Tiveljung-Lindell, A.; Giske, C.G.; Hedlund, J. Etiology of community-acquired pneumonia: Increased microbiological yield with new diagnostic methods. Clin. Infect. Dis. 2010, 50, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Lim, W.S.; Baudouin, S.V.; George, R.C.; Hill, A.T.; Jamieson, C.; Le Jeune, I.; Macfarlane, J.T.; Read, R.C.; Roberts, H.J.; Levy, M.L.; et al. Pneumonia Guidelines Committee of the BTS Standards of Care Committee BTS guidelines for the management of community acquired pneumonia in adults: Update 2009. Thorax 2009, 64, iii1–iii55. [Google Scholar] [CrossRef] [PubMed]
- Mandell, L.A.; Wunderink, R.G.; Anzueto, A.; Bartlett, J.G.; Campbell, G.D.; Dean, N.C.; Dowell, S.F.; File, T.M.; Musher, D.M.; Niederman, M.S.; et al. Infectious Diseases Society of America/American Thoracic Society Consensus Guidelines on the Management of Community-Acquired Pneumonia in Adults. Clin. Infect. Dis. 2007, 44, S27–S72. [Google Scholar] [CrossRef] [PubMed]
- Mandell, L.A. Community-acquired pneumonia: An overview. Postgrad. Med. 2015, 127, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Strålin, K. Usefulness of aetiological tests for guiding antibiotic therapy in community-acquired pneumonia. Int. J. Antimicrob. Agents 2008, 31, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Saranglao, A.; Smith, P.R. Diagnostic tests for CAP: Current approaches and future perspectives. Expert Rev. Mol. Diagn. 2002, 2, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Pai, R.; Gertz, R.E.; Beall, B. Sequential multiplex PCR approach for determining capsular serotypes of Streptococcus pneumoniae isolates. J. Clin. Microbiol. 2006, 44, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Mangold, K.A.; Santiano, K.; Broekman, R.; Krafft, C.A.; Voss, B.; Wang, V.; Hacek, D.M.; Usacheva, E.A.; Thomson, R.B.; Kaul, K.L.; et al. Real-time detection of blaKPC in clinical samples and surveillance specimens. J. Clin. Microbiol. 2011, 49, 3338–3339. [Google Scholar] [CrossRef] [PubMed]
- Thong, K.L.; Lai, M.Y.; Teh, C.S.J.; Chua, K.H. Simultaneous detection of methicillin-resistant Staphylococcus aureus, Acinetobacter baumannii, Escherichia coli, Klebsiella pneumoniae and Pseudomonas aeruginosa by multiplex PCR. Trop. Biomed. 2011, 28, 21–31. [Google Scholar] [PubMed]
- Tian, G.Z.; Zhang, L.J.; Wang, X.L.; Zhang, L.; Li, S.F.; Gu, C.M.; Sun, J.; Cui, B.Y. Rapid detection of Haemophilus influenzae and Haemophilus parainfluenzae in nasopharyngeal swabs by multiplex PCR. Biomed. Environ. Sci. 2012, 25, 367–371. [Google Scholar] [PubMed]
- Gok, U.; Bulut, Y.; Keles, E.; Yalcin, S.; Doymaz, M.Z. Bacteriological and PCR analysis of clinical material aspirated from otitis media with effusions. Int. J. Pediatr. Otorhinolaryngol. 2001, 60, 49–54. [Google Scholar] [CrossRef]
- Lynch, H.E.; Sanchez, A.M.; D’Souza, M.P.; Rountree, W.; Denny, T.N.; Kalos, M.; Sempowski, G.D. Development and implementation of a proficiency testing program for Luminex bead-based cytokine assays. J. Immunol. Methods 2014, 409, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.; Liu, Z.; Ji, S.; Gottschalk, M.; Zheng, H.; Xu, J. Simultaneous detection of 33 Streptococcus suis serotypes using the luminex xTAG® assayTM. J. Microbiol. Methods 2015, 117, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, S.A. Applications of Luminex xMAP technology for rapid, high-throughput multiplexed nucleic acid detection. Clin. Chim. Acta 2006, 363, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Abdeldaim, G.M.K.; Strålin, K.; Korsgaard, J.; Blomberg, J.; Welinder-Olsson, C.; Herrmann, B. Multiplex quantitative PCR for detection of lower respiratory tract infection and meningitis caused by Streptococcus pneumoniae, Haemophilus influenzae and Neisseria meningitidis. BMC Microbiol. 2010, 10, 310. [Google Scholar] [CrossRef] [PubMed]
- Park, H.K.; Lee, S.-J.; Yoon, J.W.; Shin, J.W.; Shin, H.-S.; Kook, J.-K.; Myung, S.C.; Kim, W. Identification of the cpsA gene as a specific marker for the discrimination of Streptococcus pneumoniae from viridans group streptococci. J. Med. Microbiol. 2010, 59, 1146–1152. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Hu, F.Z.; Gieseke, A.; Nistico, L.; Nguyen, D.; Hayes, J.; Forbes, M.; Greenberg, D.P.; Dice, B.; Burrows, A.; et al. Direct detection of bacterial biofilms on the middle-ear mucosa of children with chronic otitis media. JAMA 2006, 296, 202–211. [Google Scholar] [CrossRef] [PubMed]
- Brakstad, O.G.; Aasbakk, K.; Maeland, J.A. Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J. Clin. Microbiol. 1992, 30, 1654–1660. [Google Scholar] [PubMed]
- Karmakar, A.; Dua, P.; Ghosh, C. Biochemical and Molecular Analysis of Staphylococcus aureus Clinical Isolates from Hospitalized Patients. Can. J. Infect. Dis. Med. Microbiol. 2016, 2016, 9041636. [Google Scholar] [PubMed]
- Slinger, R.; Goldfarb, D.; Rajakumar, D.; Moldovan, I.; Barrowman, N.; Tam, R.; Chan, F. Rapid PCR detection of group A Streptococcus from flocked throat swabs: A retrospective clinical study. Ann. Clin. Microbiol. Antimicrob. 2011, 10, 33. [Google Scholar] [CrossRef] [PubMed]
- Strålin, K.; Bäckman, A.; Holmberg, H.; Fredlund, H.; Olcén, P. Design of a multiplex PCR for Streptococcus pneumoniae, Haemophilus influenzae, Mycoplasma pneumoniae and Chlamydophila pneumoniae to be used on sputum samples. APMIS 2005, 113, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Ursi, J.P.; Ursi, D.; Ieven, M.; Pattyn, S.R. Utility of an internal control for the polymerase chain reaction. Application to detection of Mycoplasma pneumoniae in clinical specimens. APMIS 1992, 100, 635–639. [Google Scholar] [CrossRef] [PubMed]
- Loens, K.; Ursi, D.; Goossens, H.; Ieven, M. Molecular diagnosis of Mycoplasma pneumoniae respiratory tract infections. J. Clin. Microbiol. 2003, 41, 4915–4923. [Google Scholar] [CrossRef] [PubMed]
- Rafiee, M.; Jahangiri-Rad, M.; Hajjaran, H.; Mesdaghinia, A.; Hajaghazadeh, M. Detection and identification of Legionella species in hospital water supplies through Polymerase Chain Reaction (16S rRNA). J. Environ. Health Sci. Eng. 2014, 12, 83. [Google Scholar] [CrossRef]
- De Vos, D.; Lim, A.; Pirnay, J.P.; Struelens, M.; Vandenvelde, C.; Duinslaeger, L.; Vanderkelen, A.; Cornelis, P. Direct detection and identification of Pseudomonas aeruginosa in clinical samples such as skin biopsy specimens and expectorations by multiplex PCR based on two outer membrane lipoprotein genes, oprI and oprL. J. Clin. Microbiol. 1997, 35, 1295–1299. [Google Scholar] [PubMed]
- Sun, Z.; Chen, Z.; Hou, X.; Li, S.; Zhu, H.; Qian, J.; Lu, D.; Liu, W. Locked nucleic acid pentamers as universal PCR primers for genomic DNA amplification. PLoS ONE 2008, 3. [Google Scholar] [CrossRef] [PubMed]
- Bøving, M.K.; Pedersen, L.N.; Møller, J.K. Eight-plex PCR and liquid-array detection of bacterial and viral pathogens in cerebrospinal fluid from patients with suspected meningitis. J. Clin. Microbiol. 2009, 47, 908–913. [Google Scholar] [CrossRef] [PubMed]
- Taniuchi, M.; Verweij, J.J.; Sethabutr, O.; Bodhidatta, L.; Garcia, L.; Maro, A.; Kumburu, H.; Gratz, J.; Kibiki, G.; Houpt, E.R. Multiplex polymerase chain reaction method to detect Cyclospora, Cystoisospora, and Microsporidia in stool samples. Diagn. Microbiol. Infect. Dis. 2011, 71, 386–390. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Wu, R.; Shi, X.; Feng, D.; Feng, G.; Yang, Y.; Dai, W.; Bian, T.; Liu, T.; He, Y.; et al. Simultaneous Detection of Five Pathogens from Cerebrospinal Fluid Specimens Using Luminex Technology. Int. J. Environ. Res. Public Health 2016, 13, 193. [Google Scholar] [CrossRef] [PubMed]
- Maria da Gloria, S.C.; Tondella, M.L.; McCaustland, K.; Weidlich, L.; McGee, L.; Mayer, L.W.; Steigerwalt, A.; Whaley, M.; Facklam, R.R.; Fields, B. Evaluation and improvement of real-time PCR assays targeting lytA, ply, and psaA genes for detection of pneumococcal DNA. J. Clin. Microbiol. 2007, 45, 2460–2466. [Google Scholar]
- Hirama, T.; Yamaguchi, T.; Miyazawa, H.; Tanaka, T.; Hashikita, G.; Kishi, E.; Tachi, Y.; Takahashi, S.; Kodama, K.; Egashira, H.; et al. Prediction of the pathogens that are the cause of pneumonia by the battlefield hypothesis. PLoS ONE 2011, 6, e24474. [Google Scholar] [CrossRef] [PubMed]
- Chiba, N.; Murayama, S.Y.; Morozumi, M.; Nakayama, E.; Okada, T.; Iwata, S.; Sunakawa, K.; Ubukata, K. Rapid detection of eight causative pathogens for the diagnosis of bacterial meningitis by real-time PCR. J. Infect. Chemother. 2009, 15, 92–98. [Google Scholar] [CrossRef] [PubMed]
- Gosiewski, T.; Szaał, L.; Pietrzyk, A.; Brzychczy-Wołch, M.; Heczko, P.B.; Bulanda, M. Comparison of methods for isolation of bacterial and fungal DNA from human blood. Curr. Microbiol. 2014, 68, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Morozumi, M.; Nakayama, E.; Iwata, S.; Aoki, Y.; Hasegawa, K.; Kobayashi, R.; Chiba, N.; Tajima, T.; Ubukata, K. Simultaneous detection of pathogens in clinical samples from patients with community-acquired pneumonia by real-time PCR with pathogen-specific molecular beacon probes. J. Clin. Microbiol. 2006, 44, 1440–1446. [Google Scholar] [CrossRef] [PubMed]
- Welti, M.; Jaton, K.; Altwegg, M.; Sahli, R.; Wenger, A.; Bille, J. Development of a multiplex real-time quantitative PCR assay to detect Chlamydia pneumoniae, Legionella pneumophila and Mycoplasma pneumoniae in respiratory tract secretions. Diagn. Microbiol. Infect. Dis. 2003, 45, 85–95. [Google Scholar] [CrossRef]
- Al-Marzooq, F.; Imad, M.A.M.; How, S.H.; Kuan, Y.C. Development of multiplex real-time PCR for the rapid detection of five bacterial causes of community acquired pneumonia. Trop. Biomed. 2011, 28, 545–556. [Google Scholar] [PubMed]
- Nolte, F.S. Molecular diagnostics for detection of bacterial and viral pathogens in community-acquired pneumonia. Clin. Infect. Dis. 2008, 47, S123–S126. [Google Scholar] [CrossRef] [PubMed]
- Chan, Y.R.; Morris, A. Molecular diagnostic methods in pneumonia. Curr. Opin. Infect. Dis. 2007, 20, 157–164. [Google Scholar] [CrossRef] [PubMed]
- Wessels, E.; Rusman, L.G.; Bussel, M.J.; Claas, E.C. Added value of multiplex Luminex Gastrointestinal Pathogen Panel (xTAG® GPP) testing in the diagnosis of infectious gastroenteritis. Clin. Microbiol. Infect. 2014, 20, O182–O187. [Google Scholar] [CrossRef] [PubMed]
- Stentzel, S.; Sundaramoorthy, N.; Michalik, S.; Nordengrün, M.; Schulz, S.; Kolata, J.; Kloppot, P.; Engelmann, S.; Steil, L.; Hecker, M.; et al. Specific serum IgG at diagnosis of Staphylococcus aureus bloodstream invasion is correlated with disease progression. J. Proteom. 2015, 128, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Jokela, P.; Piiparinen, H.; Mannonen, L.; Auvinen, E.; Lappalainen, M. Performance of the Luminex xTAG Respiratory Viral Panel Fast in a clinical laboratory setting. J. Virol. Methods 2012, 182, 82–86. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-S.; Li, H.-R.; Zhang, W.; Hua, Z.-D.; Lin, X.-H.; Lin, M.-Q.; Huang, W.-S.; Huang, L.-P.; Yu, X.-L.; Xu, N.-L.; et al. Development of a bead-based suspension array for the detection of pathogens in acute respiratory tract infections. Exp. Biol. Med. 2016, 241, 1551–1558. [Google Scholar] [CrossRef] [PubMed]
- Lengger, S.; Otto, J.; Elsässer, D.; Schneider, O.; Tiehm, A.; Fleischer, J.; Niessner, R.; Seidel, M. Oligonucleotide microarray chip for the quantification of MS2, ΦX174, and adenoviruses on the multiplex analysis platform MCR 3. Anal. Bioanal. Chem. 2014, 406, 3323–3334. [Google Scholar] [CrossRef] [PubMed]
- Donhauser, S.C.; Niessner, R.; Seidel, M. Sensitive quantification of Escherichia coli O157:H7, Salmonella enterica, and Campylobacter jejuni by combining stopped polymerase chain reaction with chemiluminescence flow-through DNA microarray analysis. Anal. Chem. 2011, 83, 3153–3160. [Google Scholar] [CrossRef] [PubMed]
- Booth, S.A.; Drebot, M.A.; Martin, I.E.; Ng, L.K. Design of oligonucleotide arrays to detect point mutations: Molecular typing of antibiotic resistant strains of Neisseria gonorrhoeae and hantavirus infected deer mice. Mol. Cell. Probes 2003, 17, 77–84. [Google Scholar] [CrossRef]
- Chou, C.-C.; Chen, C.-H.; Lee, T.-T.; Peck, K. Optimization of probe length and the number of probes per gene for optimal microarray analysis of gene expression. Nucleic Acids Res. 2004, 32, e99. [Google Scholar] [CrossRef] [PubMed]
- Binder, H.; Preibisch, S. Specific and nonspecific hybridization of oligonucleotide probes on microarrays. Biophys. J. 2005, 89, 337–352. [Google Scholar] [CrossRef] [PubMed]
- Steel, A.B.; Levicky, R.L.; Herne, T.M.; Tarlov, M.J. Immobilization of nucleic acids at solid surfaces: Effect of oligonucleotide length on layer assembly. Biophys. J. 2000, 79, 975–981. [Google Scholar] [CrossRef]
- Wilson, W.J.; Erler, A.M.; Nasarabadi, S.L.; Skowronski, E.W.; Imbro, P.M. A multiplexed PCR-coupled liquid bead array for the simultaneous detection of four biothreat agents. Mol. Cell. Probes 2005, 19, 137–144. [Google Scholar] [CrossRef] [PubMed]
| Number | Species | Strain ID and Description |
|---|---|---|
| 1 | Streptococcus pneumoniae | ATCC 49619 |
| 2 | Haemophilus influenzae | L3 (clinical isolates from China) |
| 3 | Moraxella catarrhalis | ATCC 25238 |
| 4 | Pseudomonas aeruginosa | ATCC 27853 |
| 5 | Klebsiella pneumoniae | A915 (clinical isolates from China) |
| 6 | Staphylococcus aureus | ATCC 25923 |
| 7 | Legionella pneumophila | ATCC 33152 |
| 8 | Streptococcus pyogenes | ATCC 19615 |
| 9 | Mycoplasma pneumonia * | ATCC 29342 |
| 10 | Escherichia coli | ATCC 25922 |
| 11 | Bordetella pertussis | ATCC 9797 |
| 12 | Corynebacterium diphtheriae | CMCC 38106 |
| 13 | Mycobacterium tuberculosis complex * | H37R |
| Organism | Target Gene | No. of Beads Coupled with “Anti-TAG” Sequences | Sequences (5′-3′) | Working Concentration (μM) | Length (bp) | Detection Limits | Source |
|---|---|---|---|---|---|---|---|
| S. pneumoniae | cpsA | 21 | F-TCAAACTCTCAATTCTTACTTAAT-12C-ACGCAACTGACGAGTGTGAC R-biotin-GATCGCGACACCGAACTAAT | 0.3 | 350 | 0.1 pg | [19] |
| M. catarrhalis | 16SrRNA | 8 | F-AAATAACTCACTATTTCACTTACA-12C-TTGGCTTGTGCTAAAATATC R-biotin-GTCATCGCTATCATTCACCT | 0.3 | 140 | 0.1 pg | [20] |
| S. aureus | nuc | 12 | F-CATAATCAATTTCAACTTTCTACT-12C-GCGATTGATGGTGATACGGTT R-biotin-AGCCAAGCCTTGACGAACTAAAGC | 0.3 | 270 | 0.1 pg | [21,22] |
| S. pyogenes | dnaseB | 9 | F-CACATCTAATACTTTATACAATTC-12C-TGATTCCAAGAGCTGTCGTG R-biotin-TGGTGTAGCCATTAGCTGTGTT | 0.3 | 93 | 0.1 pg | [23] |
| H. influenzae | p6 | 8 | F-AAATAACTCACTATTTCACTTACA-12C-TTGGCGGWTACTCTGTTGCT R-biotin-TGCAGGTTTTTCTTCACCGT | 0.3 | 296 | 0.1 pg | [24] |
| M. pneumoniae | P1 | 12 | F-CATAATCAATTTCAACTTTCTACT-12C-GCCACCCTCGGGGGCAGTCAG R-biotin-GAGTCGGGATTCCCCGCGGAGG | 0.3 | 209 | 1 pg | [25,26] |
| Legionella spp. | 16SrRNA | 9 | F-CACATCTAATACTTTATACAATTC-12C-AAGATTAGCCTGCGTCCGAT R-biotin-GTCAACTTATCGCGTTTGCT | 0.3 | 650 | 1 pg | [27] |
| P. aeruginosa | oprI | 14 | F-AATTTCTTCTCTTTCTTTCACAAT-12C-ATGAACAACGTTCTGAAATTCTCTGCT R-biotin-CTTGCGGCTGGCTTTTTCCAG | 0.3 | 249 | 1 pg | [28] |
| K. pneumoniae | Mdh | 15 | F-TACTTCTTTACTACAATTTACAAC-12C-GCGTGGCGGTAGATCTAAGTCATA R-biotin-TTCAGCTCCGCCACAAAGGTA | 0.3 | 364 | 0.1 pg | [12,29] |
| Organism | Gene Target | Oligonucleotide/Oligo Sequence | Reference |
|---|---|---|---|
| S. pneumoniae | lytA | Forward: ACGCAATCTAGCAGATGAAGCA
Reverse: TCGTGCGTTTTAATTCCAGCT Probe: FAM-GCCGAAAACGCTTGATACAGGGAG-BHQ1 | [33] |
| M. catarrhalis | copB | Forward: CGTGCGTGTTGACCGTTTTGACTTTA
Reverse: CACGCTGCCAAAAATAACTGCCAAAG Probe: FAM-CAGCGGTAACCTAATCTATGCCACTC-BHQ1 | [34] |
| S. aureus | spa | Forward: TACATGTCGTTAAACCTGGTG
Reverse: TACAGTTGTACCGATGAATGG Probe: FAM-CGCGATCCAAGAACTTGTTGTTGATAAGAAGCAACCGATCGCG-BHQ1 | [35,36] |
| S. pyogenes | 16SrRNA | Forward: GAGAGACTAACGCATGTTAGTA
Reverse: TAGTTACCGTCACTTGGTGG Probe: FAM-CGCGATCGCGACGATACATAGCCGACCTGGATCGCG-BHQ1 | [37] |
| H. influenzae | 16SrRNA | Forward: TTGACATCCTAAGAAGAGCTCAGAGA
Reverse: CTTCCCTCTGTATACGCCATTGTAGC Probe: FAM-ATGGCTGTCGTCAGCTCGTGTT-BHQ1 | [34] |
| M. pneumoniae | P1 | Forward: CCAACCAAACAACAACGTTCA
Reverse: ACCTTGACTGGAGGCCGTTA Probe: FAM-TCAATCCGAATAACGGTGACTTCTTACCACTG-BHQ1 | [38] |
| Legionella spp. | 16SrRNA | Forward: AGGCTAATCTTAAAGCGCCAGGCC
Reverse: GCATGCTTAACACATGCAAGTCGAAC Probe: FAM-CATATTCCTACGCGTTACTCACCCGT-BHQ1 | [34] |
| P. aeruginosa | 16SrRNA | Forward: GACGGGTGAGTAATGCCTAGGA
Reverse: CCACTGGTGTTCCTTCCTATATCT Probe: FAM-AGTGGGGGATCTTCGGACCTCA-BHQ1 | [34] |
| K. pneumoniae | gapA | Forward: TGAAGTATGACTCCACTCACGGT
Reverse: CTTCAGAAGCGGCTTTGATGGCTT Probe: FAM-CCGGTATCTTCCTGACCGACGA-BHQ1 | [34] |
| Pathogens | Real-Time PCR Assay | MPLT Assay | Performance of the MPLT Assay | Measures of Agreement Kappa Values | |||
|---|---|---|---|---|---|---|---|
| Positive | Negative | Consistency (%) | Sensitivity (%) | Specificity (%) | |||
| S. pneumoniae | Positive | 7 | 0 | 98.8 | 87.5 | 100 | 0.927 (p < 0.001) |
| Negative | 1 | 78 | |||||
| M. catarrhalis | Positive | 0 | 0 | - | - | - | * |
| Negative | 0 | 86 | |||||
| S. aureus | Positive | 11 | 0 | 100 | 100 | 100 | 1.000 (p < 0.001) |
| Negative | 0 | 75 | |||||
| S. pyogenes | Positive | 0 | 0 | - | - | - | * |
| Negative | 0 | 86 | |||||
| H. influenzae | Positive | 8 | 0 | 100 | 100 | 100 | 1.000 (p < 0.001) |
| Negative | 0 | 78 | |||||
| M. pneumoniae | Positive | 1 | 0 | 100 | 100 | 100 | 1.000 (p < 0.001) |
| Negative | 0 | 85 | |||||
| Legionella spp. | Positive | 0 | 0 | - | - | - | * |
| Negative | 0 | 86 | |||||
| P. aeruginosa | Positive | 10 | 0 | 100 | 100 | 100 | 1.000 (p < 0.001) |
| Negative | 0 | 76 | |||||
| K. pneumoniae | Positive | 5 | 0 | 100 | 100 | 100 | 1.000 (p < 0.001) |
| Negative | 0 | 81 | |||||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, L.; Ren, H.; Zhou, H.; Qin, T.; Chen, Y. Simultaneous Detection of Nine Key Bacterial Respiratory Pathogens Using Luminex xTAG® Technology. Int. J. Environ. Res. Public Health 2017, 14, 223. https://doi.org/10.3390/ijerph14030223
Jiang L, Ren H, Zhou H, Qin T, Chen Y. Simultaneous Detection of Nine Key Bacterial Respiratory Pathogens Using Luminex xTAG® Technology. International Journal of Environmental Research and Public Health. 2017; 14(3):223. https://doi.org/10.3390/ijerph14030223
Chicago/Turabian StyleJiang, Luxi, Hongyu Ren, Haijian Zhou, Tian Qin, and Yu Chen. 2017. "Simultaneous Detection of Nine Key Bacterial Respiratory Pathogens Using Luminex xTAG® Technology" International Journal of Environmental Research and Public Health 14, no. 3: 223. https://doi.org/10.3390/ijerph14030223





