QSAR Study for Carcinogenic Potency of Aromatic Amines Based on GEP and MLPs
Abstract
:1. Introduction
2. Methodology
2.1. Source of AA Data
2.2. Calculation of Molecular Descriptors
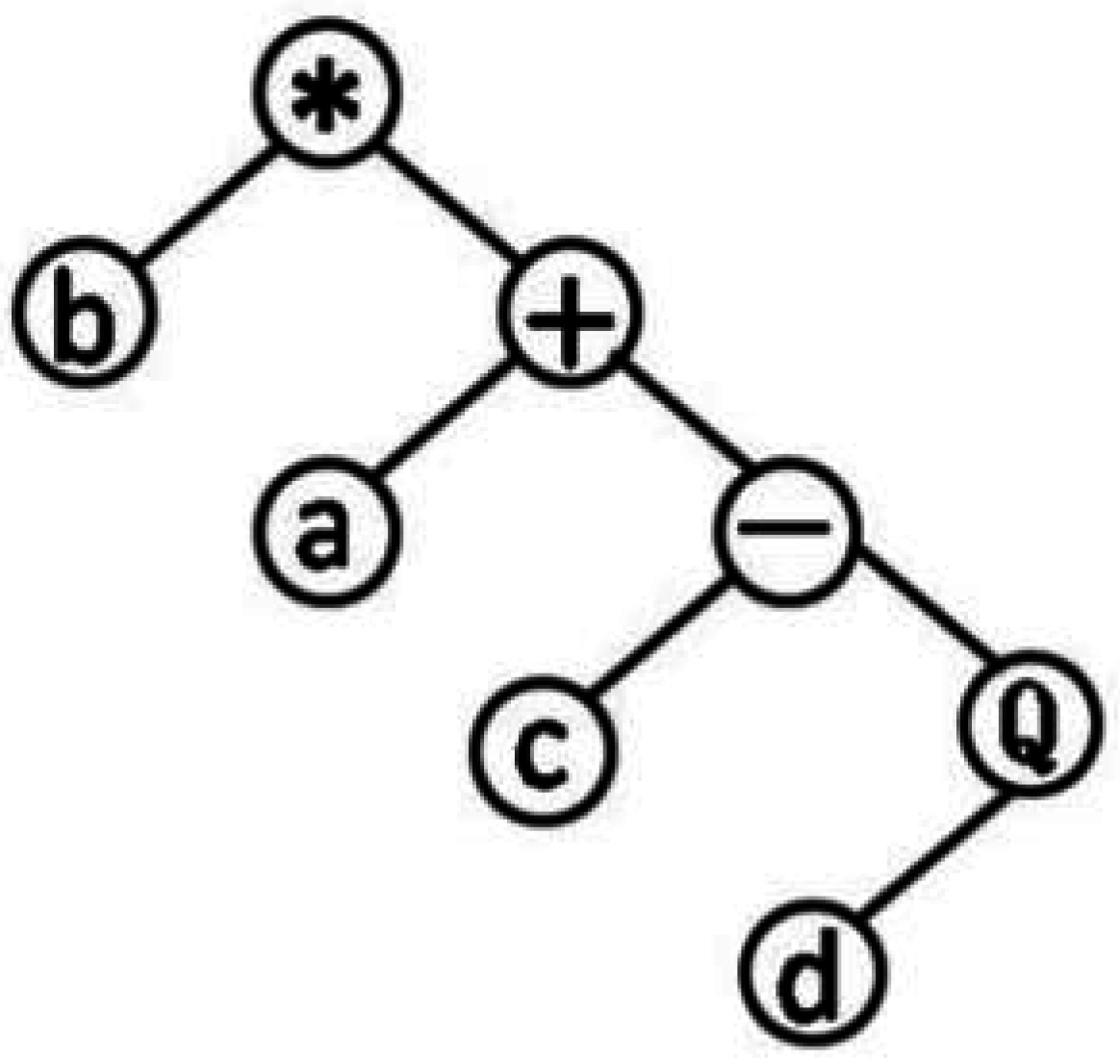
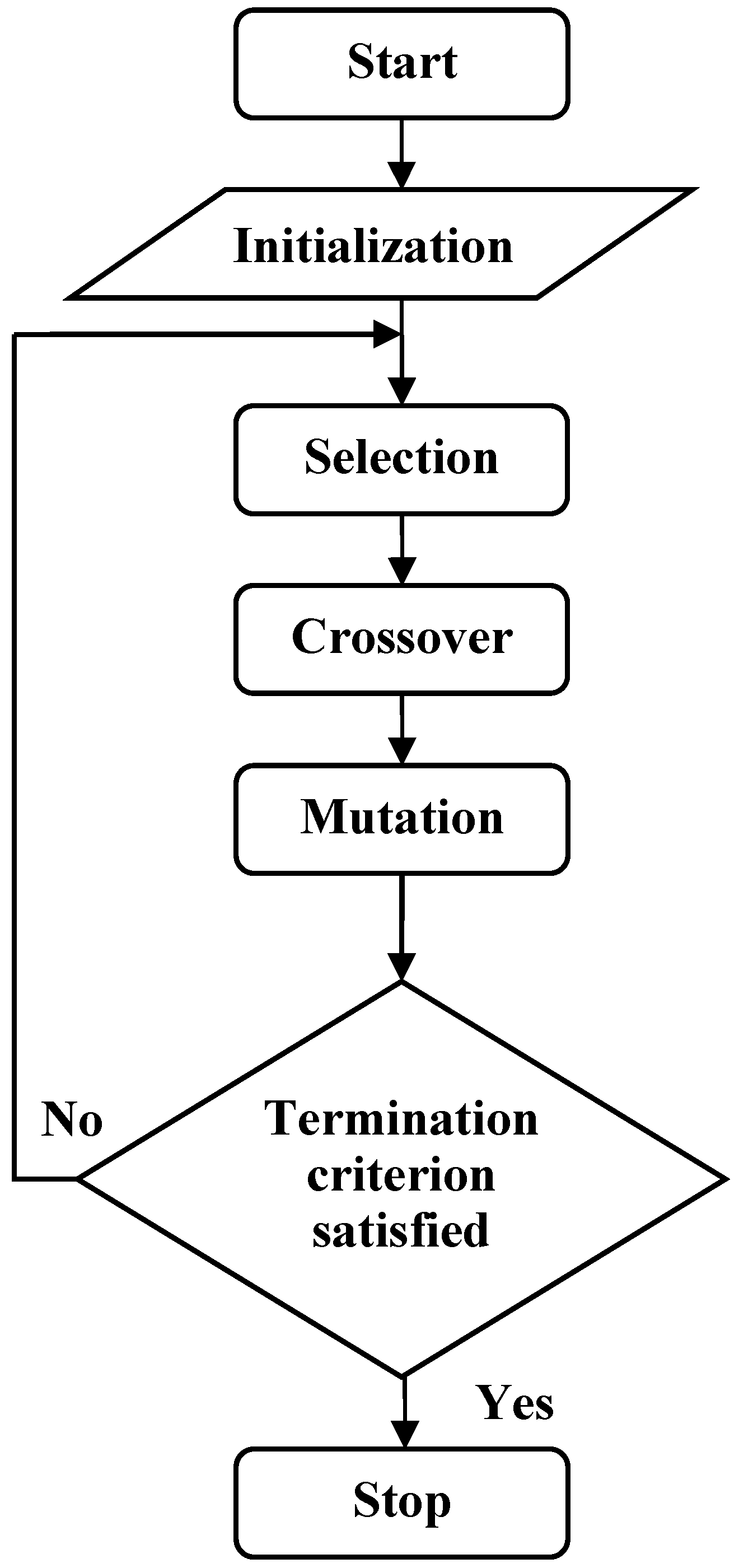
2.3. Theory of Gene Expression Programming
- If GEP_Rule (X) > 0 Then X ∈ class A
- ELSE X ∈ class B
- X stands for properties of instance.
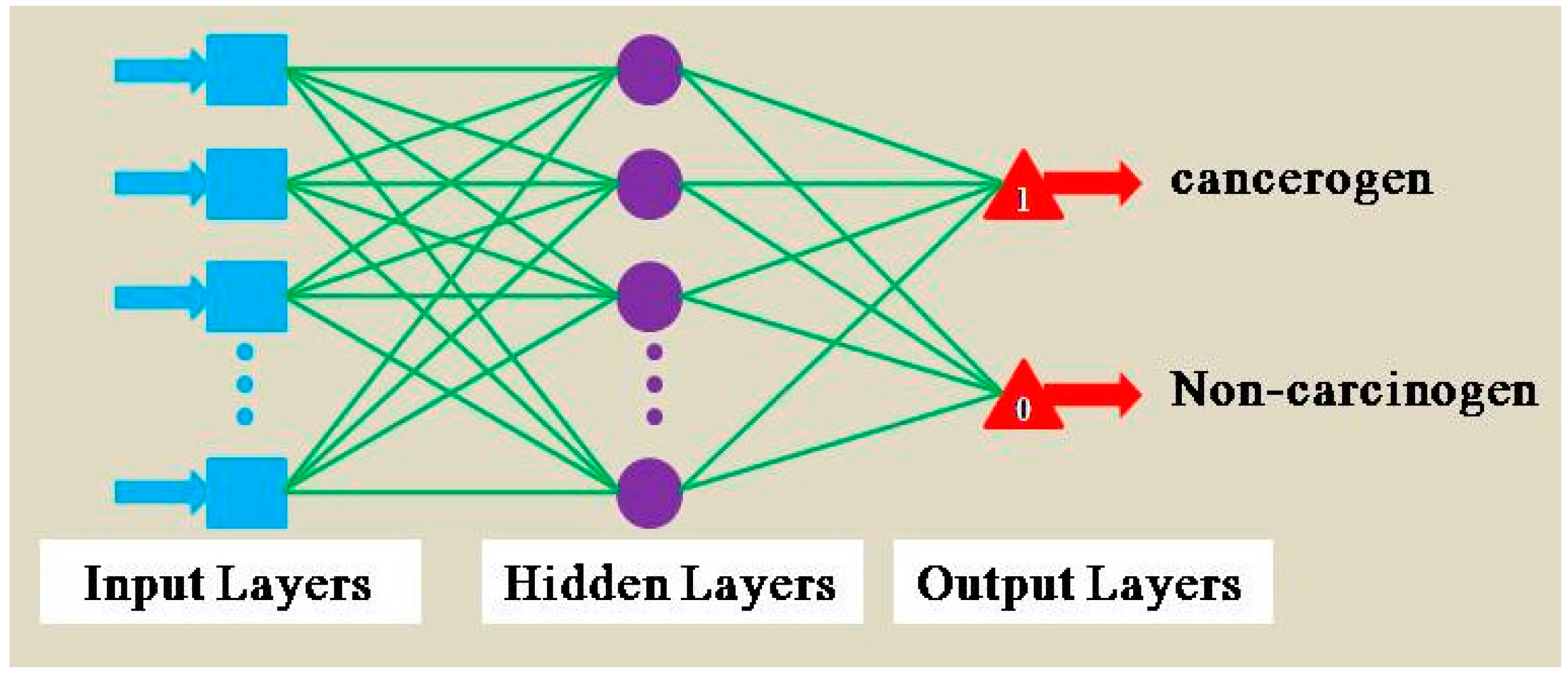
2.4. Multilayer Perceptrons (MLPs)
2.5. Platform of Weka
3. Results and Discussion
3.1. Significance of the Descriptors
3.2. Results of GEP
3.3. The Results of MLPs
3.4. Comparison between GEP and MLPs
4. Conclusions
Author Contributions
Conflicts of Interest
References
- Harding, A.P.; Popelier, P.L.A.; Harvey, J.; Giddings, A.; Foster, G.; Kranz, M. Evaluation of aromatic amines with different purities and different solvent vehicles in the Ames test. Regul. Toxicol. Pharm. 2015, 71, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Garrigós, M.C.; Reche, F.; Marín, M.L.; Pernías, K.; Jiménez, A. Optimization of the extraction of azo colorants used in toy products. J. Chromatogr. A 2002, 963, 427–433. [Google Scholar] [CrossRef]
- Sanchis, Y.; Coscollà, C.; Roca, M.; Yusà, V. Target analysis of primary aromatic amines combined with a comprehensive screening of migrating substances in kitchen utensils by liquid chromatography-high resolution mass spectrometry. Talanta 2015, 138, 290–297. [Google Scholar] [CrossRef] [PubMed]
- Ewald, R.; Thomas, M.; Joerg, D.; Lynda, C.; Regina, S. Heterocyclic aromatic amines and their contribution to the bacterial mutagenicity of the particulate phase of cigarette smoke. Toxicol. Lett. 2016, 243, 40–47. [Google Scholar]
- Wellner, T.; Lüersen, L.; Schaller, K.H.; Angerer, J.; Xler, H.; Korinth, G. Percutaneous absorption of aromatic amines—A contribution for human health risk assessment. Food Chem. Toxicol. 2008, 46, 1960–1968. [Google Scholar] [CrossRef] [PubMed]
- Inami, K.; Okazawa, M.; Mochizuki, M. Mutagenicity of aromatic amines and amides with chemical models for cytochrome p450 in Ames assay. Toxicol. In Vitro 2009, 23, 986–991. [Google Scholar] [CrossRef] [PubMed]
- Akceylan, E.; Bahadir, M.; Yılmaz, M. Removal efficiency of a calix[4]arene-based polymer for water-soluble carcinogenic direct azo dyes and aromatic amines. J. Hazard. Mater. 2009, 162, 960–966. [Google Scholar] [CrossRef] [PubMed]
- Bratberg, M.; Olsvik, P.A.; Edvardsen, R.B.; Brekken, H.K.; Vadla, R.; Meier, S. Effects of oil pollution and persistent organic pollutants (POPs) on glycerophospholipids in liver and brain of male Atlantic cod (Gadus morhua). Chemosphere 2013, 90, 2157–2171. [Google Scholar] [CrossRef] [PubMed]
- Merwe, J.P.V.D.; Hodge, M.; Olszowy, H.A.; Whittier, J.M.; Lee, S.Y. Using blood samples to estimate persistent organic pollutants and metals in green sea turtles (Chelonia mydas). Mar. Pollut. Bull. 2010, 60, 579–588. [Google Scholar] [CrossRef] [PubMed]
- Sondra, S.T.; Corrella, S.D. The biomechanisms of metal and metal-oxide nanoparticles’ interactions with cells. Int. J. Environ. Res. Public Health 2015, 12, 1112–1134. [Google Scholar]
- Sama, A.; Ayoub, K.J. Verifying the performance of artificial neural network and multiple linear regression in predicting the mean seasonal municipal solid waste generation rate: A case study of Fars province, Iran. Waste Manag. 2016, 48, 14–23. [Google Scholar]
- Li, Y.W.; Shi, X.L.; Zhang, Q.Z.; Hu, J.T.; Chen, J.M.; Wang, W.X. Computational evidence for the detoxifying mechanism of epsilon class glutathione transferase toward the insecticide DDT. Environ. Sci. Technol. 2014, 48, 5008–5016. [Google Scholar] [CrossRef] [PubMed]
- Uysala, M.; Tanyildizi, H. Estimation of compressive strength of self compacting concrete containing polypropylene fiber and mineral additives exposed to high temperature using artificial neural network. Constr. Build. Mater. 2012, 27, 404–414. [Google Scholar] [CrossRef]
- Ferreira, C. Gene expression programming: A new adaptive algorithm for solving problems. Complex Syst. 2001, 1, 87–129. [Google Scholar]
- Shaw, A.K.; Majumder, S.; Sarkar, S.; Sarkar, S.K. A novel EMD based watermarking of fingerprint biometric using GEP. Procedia Technol. 2013, 10, 172–183. [Google Scholar] [CrossRef]
- Prasenjit, D.; Ajoy, K.D. A utilization of GEP (gene expression programming) meta model and PSO (particle swarm optimization) tool to predict and optimize the forced convection around a cylinder. Energy 2016, 95, 447–458. [Google Scholar]
- Jędrzejowicz, J.; Jędrzejowicz, P. Experimental evaluation of two new GEP-based ensemble classifiers. Expert Syst. Appl. 2011, 38, 10932–10939. [Google Scholar] [CrossRef]
- Silva, A.A.; Lima Neto, I.A.; Misságia, R.M.; Ceia, M.A.; Carrasquilla, A.G.; Archilha, N.L. Artificial neural networks to support petrographic classification of carbonate-siliciclastic rocks using well logs and textural information. J. Appl. Geophys. 2015, 117, 118–125. [Google Scholar] [CrossRef]
- Zhu, Y.P.; Yu, Y.N.; Chen, X.R. Fisher discriminant analysis for carcinogenic potency of aromatic amines. Chin. J. Prev. Med. 1999, 1, 1–11. [Google Scholar]
- Si, H.Z.; Wang, T.; Zhang, K.J.; Hu, Z.D.; Fan, B.T. QSAR study of 1,4-dihydropyridine calcium channel antagonists based on gene expression programming. Bioorganic Med. Chem. 2006, 14, 4834–4841. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Y.; Luan, F.; Si, H.Z.; Hu, Z.D.; Liu, M.C. Prediction of retention times for a large set of pesticides or toxicants based on support vector machine and the heuristic method. Toxicol. Lett. 2007, 175, 136–144. [Google Scholar] [CrossRef] [PubMed]
- Servien, R.; Mamy, L.; Li, Z.; Rossard, V.; Latrille, E.; Bessac, F.; Patureau, D.; Benoit, P. Typol—A new methodology for organic compounds clustering based on their molecular characteristics and environmental behavior. Chemosphere 2014, 111, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Xiao, W.; Tirpak, T.M.; Nelson, P.C. Evolving Accurate and Compact Classification Rules with Gene Expression Programming. IEEE Trans. Evol. Comput. 2003, 7, 519–531. [Google Scholar] [CrossRef]
- Eriksson, L.; Andersson, P.L.; Johansson, E.; Tysklind, M. Megavariate analysis of environmental QSAR data. Part I—A basic framework founded on principal component analysis (PCA), partial least squares (PLS), and statistical molecular design (SMD). Mol. Divers. 2006, 10, 169–186. [Google Scholar] [CrossRef] [PubMed]
- Duan, L.; Tang, C.; Zhang, T.; Wei, D.; Zhang, H. Distance guided classification with gene expression programming. Adv. Data Min. Appl. 2006, 4093, 239–246. [Google Scholar]
- Teodorescu, L.; Sherwood, D. High energy physics event selection with gene expression programming. Comput. Phys. Commun. 2008, 178, 409–419. [Google Scholar] [CrossRef]
- Yadav, A.K.; Malik, H.; Chandel, S.S. Selection of most relevant input parameters using WEKA for artificial neural network based solar radiation prediction models. Renew. Sust. Energy Rev. 2014, 31, 509–519. [Google Scholar] [CrossRef]
- Mohammad, M.N.; Sulaiman, N.; Muhsin, O.A. A novel intrusion detection system by using intelligent data mining in WEKA environment. Procedia Comput. Sci. 2011, 3, 1237–1242. [Google Scholar] [CrossRef]
- Lievens, S.; Baets, B.D. Supervised ranking in the WEKA environment. Inf. Sci. 2010, 180, 4763–4771. [Google Scholar] [CrossRef]
- Singh, K.P.; Gupta, S.; Rai, P. Identifying pollution sources and predicting urban air quality using ensemble learning methods. Atmos. Environ. 2013, 80, 426–437. [Google Scholar] [CrossRef]
- Kar, S.; Roy, K. First report on development of quantitative interspecies structure-carcinogenicity relationship models and exploring discriminatory features for rodent carcinogenicity of diverse organic chemicals using OECD guidelines. Chemosphere 2012, 87, 339–355. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.C.; Zhang, Q.Z.; Wang, H.; Hu, J.T. Predicting carcinogenicity of organic compounds based on CPDB. Chemosphere 2015, 139, 81–90. [Google Scholar] [CrossRef] [PubMed]
- Helguera, A.M.; Pérez, M.A.C.; González, M.P.; Ruiz, R.M.; Díaz, H.G. A topological substructural approach applied to the computational prediction of rodent carcinogenicity. Bioorganic Med. Chem. 2005, 13, 2477–2488. [Google Scholar] [CrossRef] [PubMed]
- Basavaraja, J.; Inamdar, S.R.; Kumar, H.M.S. Solvents effect on the absorption and fluorescence spectra of 7-diethylamino-3-thenoylcoumarin: Evaluation and correlation between solvatochromism and solvent polarity parameters. Spectrochim. Acta A 2015, 137, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Sambathkumar, K.; Jeyavijayan, S.; Arivazhagan, M. Electronic structure investigations of 4-aminophthal hydrazide by UV-visible, NMR spectral studies and HOMO-LUMO analysis byab initioand DFT calculations. Spectrochim. Acta A 2015, 147, 124–138. [Google Scholar] [CrossRef] [PubMed]
- Lin, I.S.; Fan, P.L.; Chen, H.I.; Loh, C.H.; Shih, T.S.; Liou, S.H. Rapid and intermediate N-acetylators are less susceptible to oxidative damage among 4,4-methylenebis(2-chloroaniline) (MBOCA)-exposed workers. Int. J. Hydrogen Energy 2013, 216, 515–520. [Google Scholar] [CrossRef] [PubMed]
- Szczuka, M.; Ślęzak, D. Feedforward neural networks for compound signals. Theor. Comput. Sci. 2011, 412, 5960–5973. [Google Scholar] [CrossRef]
| No. | Aromatic Amines | Carcinogenicity (exp) | Carcinogenicity (GEP) | Carcinogenicity (MLPs) |
|---|---|---|---|---|
| 1 | N-Acetoxy-4-biphenylacetamide | 0 | 0 | 0 |
| 2 | N-Acetoxy-2-fluorenylacetamide | 0 | 0 | 0 |
| 3 | N-Acetoxy-4-phenanthrylacetamide | 0 | 0 | 0 |
| 4 | N-Acetoxy-N-(4-stilbenyl)acetamide | 0 | 0 | 0 |
| 5 | 3-Amino-s-triazole | 1 | 1 | 1 |
| 6 | 1-Anthramine | 0 | 0 | 0 |
| 7 | 9-Anthramine | 0 | 0 | 0 |
| 8 | 2-Anthranilacetamide | 0 | 0 | 0 |
| 9 | Benzidine | 1 | 0 | 1 |
| 10 | N-(Benzoyloxy)-fluorenylacetamide | 0 | 1 | 0 |
| 11 | 4-Biphenyldimethylamine | 0 | 0 | 0 |
| 12 | 3,6-Bis(dimethylamino)acridine | 1 | 0 | 1 |
| 13 | 2-Chloro-4-phenylaniline | 0 | 1 | 0 |
| 14 | 4′-Chloro-4-stilbenyl-N,N-dimethylamine | 0 | 0 | 0 |
| 15 | 2-Cyano-4-stilbenamine | 1 | 0 | 0 |
| 16 | 4,6-Diamino-2-(5-nitro-2-furyl)-s-triazine | 0 | 1 | 1 |
| 17 | 0,0′-Dianisidine | 0 | 0 | 0 |
| 18 | 3-Dibenzofuranylacetamide | 0 | 0 | 0 |
| 19 | 3-Dibenzothiophenylacetamide | 0 | 0 | 0 |
| 20 | 2,2′-Dichloro-4,4′-diaminostilbene | 1 | 0 | 1 |
| 21 | 3,3′-Dichloro-4,4′-diaminostilbene | 0 | 1 | 0 |
| 22 | 9,10-Dihydro-2-phenanthramine | 0 | 0 | 0 |
| 23 | 3,3′-Dihydroxybenzidine | 0 | 0 | 0 |
| 24 | 2-(4-(N,N-Dimethylamino)styryl) quinoline | 0 | 0 | 0 |
| 25 | 3,2′-Dimethyl-4-biphenylamine | 0 | 0 | 0 |
| 26 | 3,3′-Dimethyl-4-biphenylamine | 0 | 0 | 0 |
| 27 | 2-Fluorenylacetamide | 1 | 0 | 0 |
| 28 | 3-Fluorenylacetamide | 0 | 0 | 0 |
| 29 | 1-Fluorenylaceto hydroxamic acid | 0 | 0 | 0 |
| 30 | 2-Fluorenylaceto hydroxanic acid | 1 | 0 | 0 |
| 31 | N-Fluorenyl-2-benzamide | 0 | 1 | 0 |
| 32 | N-Fluorenyl-2-benzohydroxamic acid | 0 | 0 | 0 |
| 33 | 2-Fluorenyldiacetamide | 1 | 0 | 1 |
| 34 | 2-Fluorenyldimethylamine | 1 | 1 | 1 |
| 35 | 2,5-Fluorenylenediacetamide | 0 | 0 | 0 |
| 36 | 2-Fluorenylhydroxylamine | 0 | 0 | 0 |
| 37 | N-(2-Fluorenyl)-2,2,2-trifluoroacetamide | 1 | 0 | 1 |
| 38 | 4′-Fluoro-4-biphenylamine | 1 | 0 | 1 |
| 39 | 1-Fluoro-2-fluorenylacetamide | 0 | 0 | 1 |
| 40 | 3-Fluoro-2-fluorenylacetamide | 1 | 0 | 0 |
| 41 | 4-Fluoro-2-fluorenylacetamide | 0 | 0 | 0 |
| 42 | 5-Fluoro-2-fluorenylacetamide | 0 | 1 | 0 |
| 43 | 6-Fluoro-2-fluorenylacetamide | 1 | 0 | 0 |
| 44 | 7-Fluoro-2-fluorenylacetamide | 1 | 0 | 0 |
| 45 | 7-Fluoro-2-N-fluorenylacetohydroxamic acid | 1 | 1 | 0 |
| 46 | 4′-Fluoro-p-phenylaniline | 0 | 1 | 0 |
| 47 | 4′-Fluoro-4-stilbenamine | 1 | 1 | 0 |
| 48 | 4′-Fluoro-4-stilbenyl-N,N-dimethylamine | 1 | 0 | 0 |
| 49 | 2-Hydrazino-4-phenylthiazole | 0 | 1 | 0 |
| 50 | N-Hydroxy-N-(4-stilbenyl) acetamide | 0 | 1 | 0 |
| 51 | 3-Iodo-2-fluorenylacetamide | 0 | 0 | 0 |
| 52 | 7-Iodo-2-fluroenylacetamide | 0 | 0 | 0 |
| 53 | 2-Methoxy-3-benzofuranylamine | 0 | 0 | 0 |
| 54 | 7-Methoxy-2-fluorenylacetamide | 1 | 0 | 1 |
| 55 | 1-Methoxy-2-fluorenylamine | 1 | 0 | 1 |
| 56 | 3-Methoxy-2-fluorenylamine | 0 | 1 | 0 |
| 57 | 4-((p-Methoxyphenyl)azo)-o-anisidine | 1 | 0 | 1 |
| 58 | 2-Methyldiacetylbenzidine | 0 | 0 | 1 |
| 59 | 4,4′-Methylenebis(2-chloroaniline) | 1 | 1 | 1 |
| 60 | 4′-Methyl-4-phenylacetanilide | 0 | 0 | 0 |
| 61 | 3-Methyl-4-phenylaniline | 0 | 1 | 0 |
| 62 | 3-Methyl-4-stilbenamine | 0 | 0 | 0 |
| 63 | 1-Naphthylacetohydroxamic acid | 0 | 0 | 0 |
| 64 | 2-Naphthylhydroxylamine | 0 | 0 | 0 |
| 65 | 9-Oxo-2-fluorenylacetamide | 1 | 0 | 0 |
| 66 | 1-Phenanthrylacetamide | 0 | 0 | 0 |
| 67 | 2-Phenanthrylacetamide | 0 | 1 | 0 |
| 68 | 1-Phenanthrylamine | 0 | 0 | 0 |
| 69 | 3-Phenanthrylamine | 0 | 0 | 0 |
| 70 | 9-Phenanthrylamine | 0 | 0 | 0 |
| 71 | 4-(Phenylazo) acetanilide | 0 | 0 | 0 |
| 72 | 4-(Phenylazo) aniline | 0 | 0 | 0 |
| 73 | 4-(Phenylazo) diacetanilide | 0 | 0 | 0 |
| 74 | 4-(Phenylazo)-N-phenylacetohydroxamic acid | 0 | 0 | 0 |
| 75 | 4-Stilbenamine | 0 | 0 | 0 |
| 76 | N-(4-Stilbenyl) acetamide | 0 | 0 | 0 |
| 77 | 4-Stilbenyl-N,N-diethylamine | 0 | 0 | 0 |
| 78 | 4-Stilbenyl-N,N-dimethylamine | 0 | 0 | 0 |
| 79 | N-(4-Styrylphenyl) hydroxylamine | 0 | 0 | 0 |
| 80 | 3,2′,4′,6′-Tetramethyl-4-biphenylamine | 1 | 0 | 0 |
| 81 | o,o′-Tolidine | 0 | 1 | 0 |
| 82 | 4-(m-Tolylazo) acetanilide | 0 | 0 | 0 |
| 83 | 4-(m-Tolylazo) aniline | 0 | 0 | 0 |
| 84 | 2-(o-Tolylazo)-p-toluidine | 1 | 0 | 1 |
| 85 | 2-(p-Tolylazo)-p-toluidine | 0 | 0 | 0 |
| 86 | 4-(o-Tolylazo)-o-toluidine | 1 | 1 | 0 |
| 87 | 4-(o-Tolylazo)-m-toluidine | 0 | 0 | 0 |
| 88 | 4-(m-Tolylazo)-m-toluidine | 0 | 0 | 0 |
| 89 | 4-(p-Tolylazo)-o-toluidine | 0 | 0 | 0 |
| 90 | 4-(p-Tolylazo)-m-toluidine | 0 | 0 | 0 |
| 91 | N,N,2′-Trimethyl-4-stilbenamine | 0 | 0 | 0 |
| 92 | N,N,3′-Trimethyl-4-stilbenamine | 0 | 0 | 0 |
| 93 | N,N,4′-Trimethyl-4-stilbenamine | 0 | 0 | 0 |
| No. | Aromatic Amines | Carcinogenicity (exp) | Carcinogenicity (GEP) | Carcinogenicity (MLPs) |
|---|---|---|---|---|
| 1 | 2-Anthramine | 0 | 0 | 0 |
| 2 | 4-Biphenylacetamide | 0 | 0 | 0 |
| 3 | 4-Biphenylacetohydroxamic acid | 0 | 1 | 0 |
| 4 | 3-Carbazolylacetamide | 0 | 0 | 1 |
| 5 | 2,7-Diaminofluorene | 0 | 0 | 1 |
| 6 | 4,4′-Diaminostilbene | 1 | 1 | 0 |
| 7 | 2-Dibenzothiophenylacetamide | 0 | 0 | 0 |
| 8 | 3,3′-Dichlorobenzidine | 0 | 0 | 0 |
| 9 | 2-Fluorenamine | 1 | 1 | 0 |
| 10 | 1-Fluorenylacetamide | 0 | 0 | 0 |
| 11 | 3-Fluorenylaceto hydroxanic acid | 0 | 0 | 0 |
| 12 | 2,7-Fluorenyldiacetamide | 1 | 1 | 0 |
| 13 | 2-Fluorenyldiethylamine | 0 | 0 | 0 |
| 14 | N,2-Fluorenylformamide | 0 | 1 | 0 |
| 15 | 2-Fluorenylmethylamine | 1 | 0 | 0 |
| 16 | N,2-Fluorenylsuccinamic acid | 1 | 0 | 0 |
| 17 | 8-Fluoro-2-fluorenylacetamide | 1 | 0 | 1 |
| 18 | 2-Fluoro-4-phenylaniline | 0 | 0 | 0 |
| 19 | 3′-Fluoro-4-phenylaniline | 0 | 0 | 0 |
| 20 | 3-Methoxy-4-biphenylamine | 0 | 1 | 1 |
| 21 | 3-Methoxy-2-fluorenylacetamide | 0 | 1 | 0 |
| 22 | 4,4′-Methylenebis(2-methylaniline) | 1 | 0 | 1 |
| 23 | 3-Methyl-2-naphthylamine | 0 | 0 | 0 |
| 24 | 2-Methyl-4-phenylaniline | 0 | 0 | 0 |
| 25 | 2′-Methyl-4-phenylaniline | 0 | 0 | 0 |
| 26 | 2-Methyl-4-stilbenamine | 0 | 1 | 0 |
| 27 | 2-Naphthylamine | 0 | 0 | 0 |
| 28 | 1-Naphthylhydroxylamine | 0 | 0 | 0 |
| 29 | 9-Phenanthrylacetamide | 0 | 0 | 0 |
| 30 | 2-Phenanthrylacetohydroxamic acid | 0 | 0 | 0 |
| 31 | 2-Phenanthrylamine | 0 | 1 | 0 |
| 32 | 4-(Phynylazo)-o-anisidine | 1 | 1 | 0 |
| 33 | 1-(Phenylazo)-2-naphthylamine | 0 | 0 | 0 |
| 34 | 4-(Phenylazo)-N-phenylhydroxylamine | 0 | 0 | 0 |
| 35 | 3,2′,5′-Trimethyl-4-diphenylamine | 1 | 0 | 1 |
| Correlation | NCOS | NNOS | KFBI | BBI | SICI | TEIA | PLPT | LUMO |
|---|---|---|---|---|---|---|---|---|
| NCOS | 1.000 | −0.227 | 0.649 | −0.708 | 0.667 | 0.234 | −0.034 | −0.374 |
| NNOS | 1.000 | 0.175 | −0.014 | 0.159 | 0.312 | −0.201 | −0.111 | |
| KFBI | 1.000 | −0.569 | 0.730 | 0.433 | 0.007 | −0.18 | ||
| BBI | 1.000 | −0.681 | −0.259 | −0.173 | 0.438 | |||
| SICI | 1.000 | 0.620 | 0.250 | −0.456 | ||||
| TEIA | 1.000 | 0.339 | −0.107 | |||||
| PLPT | 1.000 | −0.277 | ||||||
| LUMO | 1.000 |
| Accuracy | Sensitivity | Specificity | Youden’s Index | |
|---|---|---|---|---|
| Training set of GEP | 0.914 | 0.947 | 0.905 | 0.852 |
| Test set of GEP | 0.829 | 0.667 | 0.885 | 0.552 |
| Training set of MLPS | 0.838 | 0.844 | 0.813 | 0.657 |
| Test set of MLPS | 0.743 | 0.793 | 0.500 | 0.293 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, F.; Zhang, A.; Liang, H.; Cui, L.; Li, W.; Si, H.; Duan, Y.; Zhai, H. QSAR Study for Carcinogenic Potency of Aromatic Amines Based on GEP and MLPs. Int. J. Environ. Res. Public Health 2016, 13, 1141. https://doi.org/10.3390/ijerph13111141
Song F, Zhang A, Liang H, Cui L, Li W, Si H, Duan Y, Zhai H. QSAR Study for Carcinogenic Potency of Aromatic Amines Based on GEP and MLPs. International Journal of Environmental Research and Public Health. 2016; 13(11):1141. https://doi.org/10.3390/ijerph13111141
Chicago/Turabian StyleSong, Fucheng, Anling Zhang, Hui Liang, Lianhua Cui, Wenlian Li, Hongzong Si, Yunbo Duan, and Honglin Zhai. 2016. "QSAR Study for Carcinogenic Potency of Aromatic Amines Based on GEP and MLPs" International Journal of Environmental Research and Public Health 13, no. 11: 1141. https://doi.org/10.3390/ijerph13111141






