3.2. Poly-l-Lactide-Based Nanoparticles
Model fitting to the data (
Table 1) was carried out and the quadratic model was found significant and was selected. To improve the model, insignificant terms were removed by backward elimination. Analysis of variance (ANOVA) of the selected model and terms (
Table 3) reveal that the selected model is significant (
p = 0.0020). Further, the model (Scheffe Polynomial) was also selected based on the estimation of several statistical parameters: multiple correlation coefficient (
R2), adjusted multiple correlation coefficient (adjusted
R2) and the predicted residual sum of squares (PRESS). Also, “lack of fit” was not statistically significant (
p = 0.4994) which is desirable. The resulting model (Scheffe polynomial) is shown in Equation (1) below:
where: A = Crosslinker (mmol); B = Initiators (mmol); C = Stabilizer (mmol); D = Macromonomer (mmol).
Table 3.
Analysis of variance table for particle size (Poly-l-lactide-based nanoparticles).
Table 3.
Analysis of variance table for particle size (Poly-l-lactide-based nanoparticles).
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value | |
|---|
| Model | 6011.26 | 7 | 858.75 | 6.83 | 0.0020 | s |
| Linear mixture | 1431.12 | 3 | 477.04 | 3.79 | 0.0401 | s |
| AB | 693.79 | 1 | 693.79 | 5.52 | 0.0368 | s |
| AC | 2723.37 | 1 | 2723.37 | 21.65 | 0.0006 | s |
| BC | 694.88 | 1 | 694.88 | 5.52 | 0.0367 | s |
| CD | 1247.85 | 1 | 1247.85 | 9.92 | 0.0084 | s |
| Residual | 1509.25 | 12 | 125.77 | | | |
| Lack of Fit | 895.66 | 7 | 127.95 | 1.04 | 0.4994 | ns |
| Pure Error | 613.59 | 5 | 122.72 | | | |
| Cor Total | 7520.52 | 19 | | | | |
Diagnostic plots (
Figure 2) show the validity of the model. The normal probability plot of the residuals (
Figure 2A) is the most important diagnostic and it checks for non-normality in the error term. A linear normal probability plot of the residuals, which indicates normality in the error term, was obtained.
Figure 2B shows a diagnostic plot that tests the assumption of constant variance. Both plots show no problem with our data. The Scheffe polynomial (Equation (1)) was used to generate the model graph (
Figure 3) which shows the design space and variation in particle size as a function of the mixture composition. (A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer) (Poly-
l-lactide-based nanoparticles). The predicted sizes for the four solutions are 276.7 nm, 283.6 nm, 291.6 nm and 302.7 nm while the experimentally obtained sizes are 244.2 nm ± 4.20 nm, 246.6 nm ± 0.87 nm, 250.4 nm ± 4.04 nm and 271 nm ± 4.62 nm respectively (
Table 4). The corresponding polydisperity index (PDI) values are 0.23, 0.28, 0.27 and 0.29, respectively.
Figure 2.
Diagnostic plots the particle size data of polylactide-based nanoparticles: (A) Normal Plot of Residuals; (B) Residuals vs. Predicted.
Figure 2.
Diagnostic plots the particle size data of polylactide-based nanoparticles: (A) Normal Plot of Residuals; (B) Residuals vs. Predicted.
Figure 3.
Model graph showing the design space and variation in particle size as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Polylactide-based nanoparticles).
Figure 3.
Model graph showing the design space and variation in particle size as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Polylactide-based nanoparticles).
Table 4.
Analysis of variance table for percent yield (Poly-l-lactide-based nanoparticles).
Table 4.
Analysis of variance table for percent yield (Poly-l-lactide-based nanoparticles).
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value (Prob > F) | |
|---|
| Model | 1163.07 | 5 | 232.61 | 2.98 | 0.0488 | s |
| Linear Mixture | 326.63 | 3 | 108.88 | 1.40 | 0.2854 | ns |
| AB | 508.70 | 1 | 508.70 | 6.52 | 0.0230 | s |
| AD | 822.41 | 1 | 822.41 | 10.54 | 0.0059 | s |
| Residual | 1092.30 | 14 | 78.02 | | | |
| Lack of Fit | 653.82 | 9 | 72.65 | 0.83 | 0.6211 | ns |
| Pure Error | 438.48 | 5 | 87.70 | | | |
| Cor Total | 2255.38 | 19 | | | | |
A similar model fitting was done for percent yield as shown in
Table 4. The resulting model (Scheffe polynomial) is shown in Equation (2) below:
where: A = Crosslinker (mmol); B = Initiators (mmol); C = Stabilizer (mmol); D = Macromonomer (mmol)
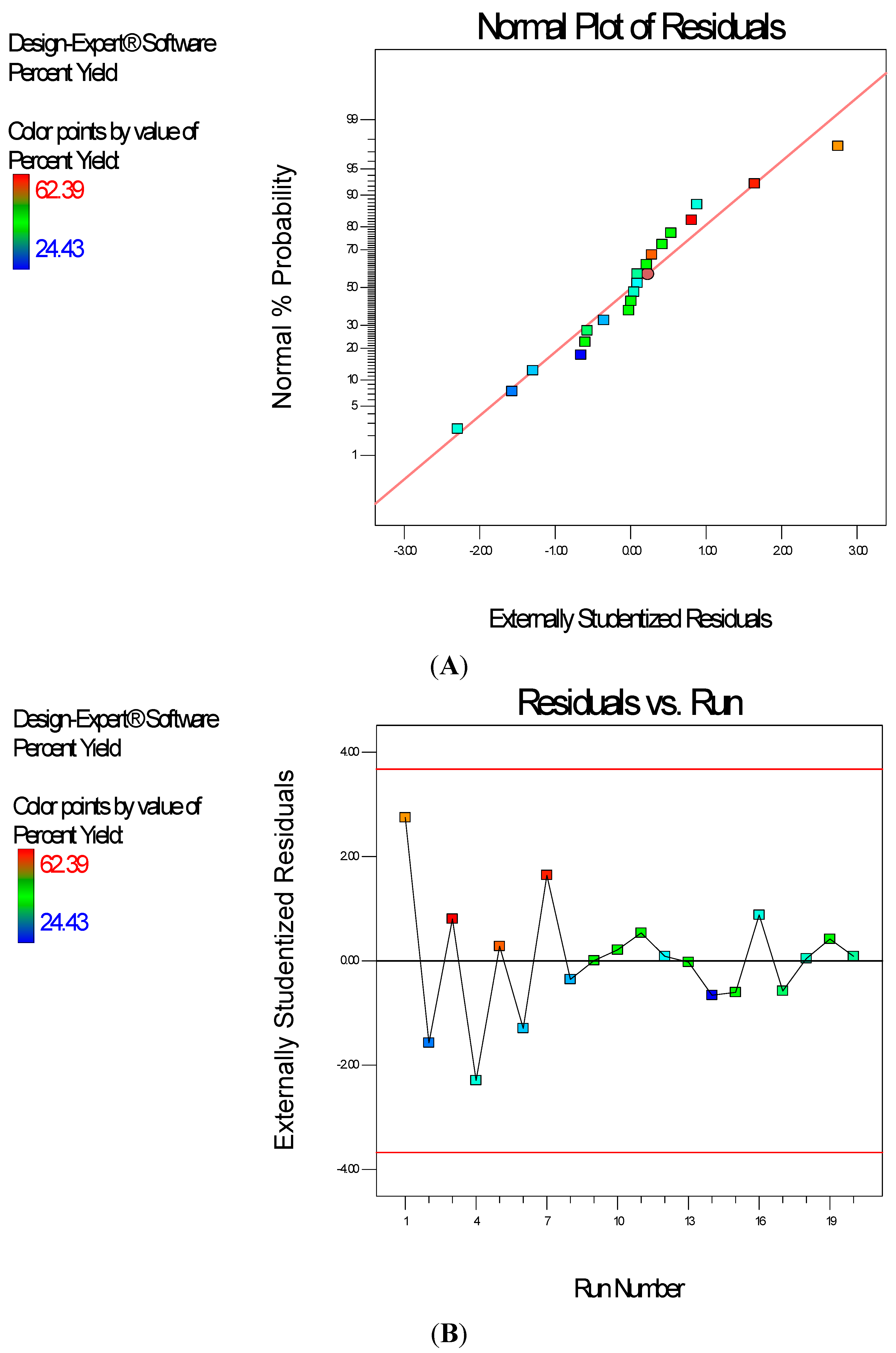
Diagnostic plots (
Figure 4) show the validity of the model. The Scheffe polynomial (Equation (2)) was used to generate the model graph (
Figure 5), which shows the design space and variation in percent yield as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Polylactide-based nanoparticles).
Figure 4.
Diagnostic plots the percent yield data of Polylactide-based nanoparticles: (A) Normal Plot of Residuals; (B) Residuals vs. Run.
Figure 4.
Diagnostic plots the percent yield data of Polylactide-based nanoparticles: (A) Normal Plot of Residuals; (B) Residuals vs. Run.
Figure 5.
Model graph showing the design space and variation in percent yield as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Polylactide-based nanoparticles).
Figure 5.
Model graph showing the design space and variation in percent yield as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Polylactide-based nanoparticles).
3.3. Poly-ɛ-Caprolactone-Based Nanoparticles
Logarithmic transformation was carried out before model fitting to particle size data (
Table 2). The quadratic model was found significant and was selected. To improve the model, insignificant terms were removed by backward elimination. Analysis of variance (ANOVA) of the selected model and terms (
Table 5) reveals that the selected model is significant (
p < 0.0001). The linear mixture (component linear terms) and the square of A term (crosslinker term) are significant:
p < 0.0001 and
p = 0.0005 respectively. Additionally, “lack of fit” is not significant (
p = 0.6921). Non-significant lack of fit is good as our desire is for the model to fit. The “Pred R-Squared” of 0.9326 is in reasonable agreement with the “Adj R-Squared” of 0.9455. Adequate Precision measures the signal to noise ratio. A ratio greater than 4 is desirable. The ratio of 27.332 obtained in this work indicates an adequate signal. Consequently, this model can be used to navigate the design space.
Table 5.
Analysis of variance table for particle size (Poly-ɛ-caprolactone-based nanoparticles.
Table 5.
Analysis of variance table for particle size (Poly-ɛ-caprolactone-based nanoparticles.
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value (Prob > F) | |
|---|
| Model | 1.98 | 4 | 0.49 | 126.80 | <0.0001 | s |
| Linear Mixture | 1.92 | 3 | 0.64 | 163.82 | <0.0001 | s |
| A2 | 0.061 | 1 | 0.061 | 15.71 | 0.0005 | s |
| Residual | 0.098 | 25 | 3.904e-003 | | | |
| Lack of Fit | 0.074 | 20 | 3.691e-003 | 0.78 | 0.6921 | ns |
| Pure Error | 0.024 | 5 | 4.756e-003 | | | |
| Cor Total | 2.08 | 29 | | | | |
The empirical model (Scheffe polynomial) is shown in Equation (3) below:
where: A = Crosslinker (mmol); B = Initiators (mmol); C = Stabilizer (mmol); D = Macromonomer (mmol).
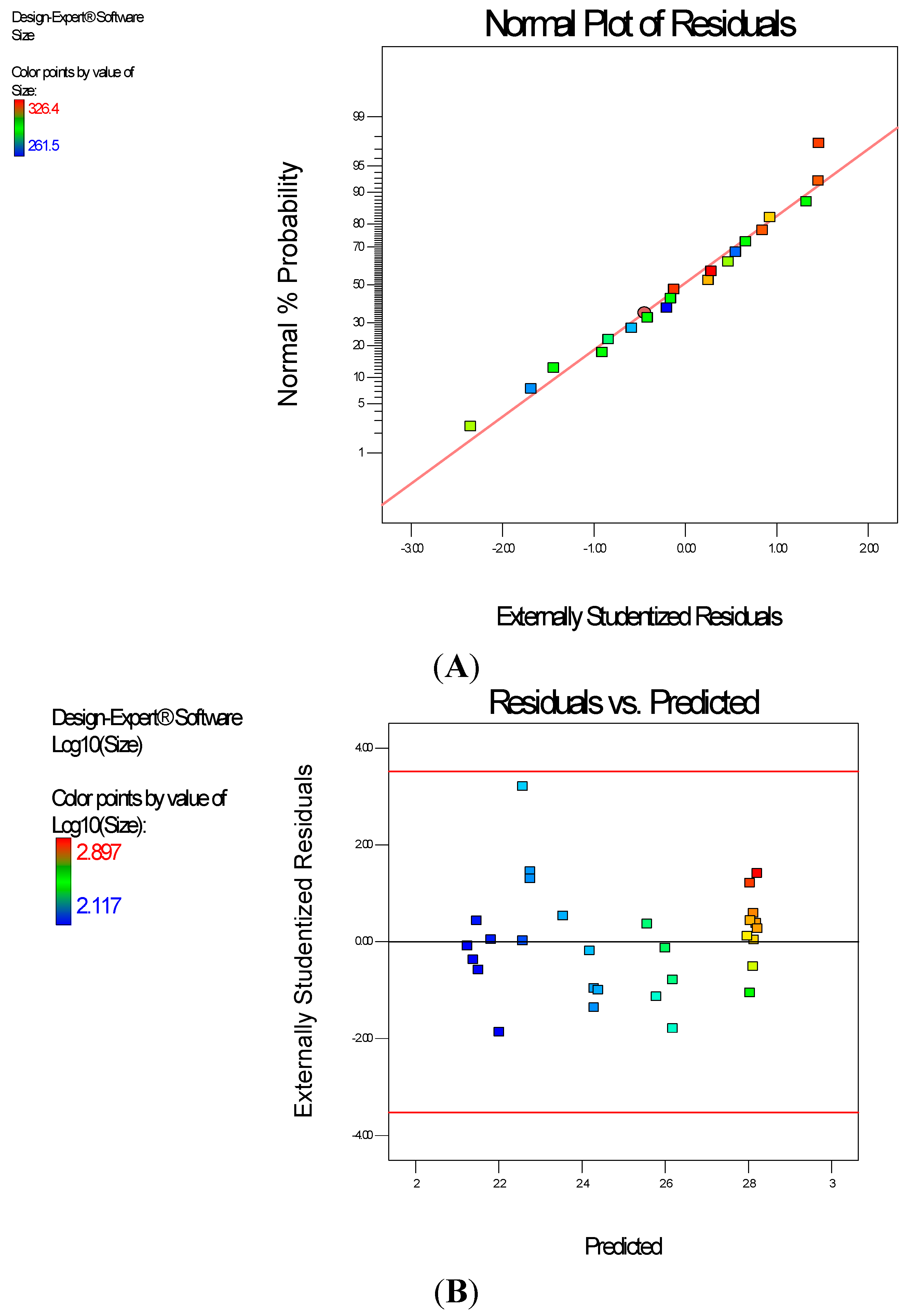
Diagnostic plots show the validity of the model (
Figure 6). The normal probability plot of the residuals is the most important diagnostic plot; it checks for non-normality in the error term. A linear normal probability plot of the residuals was obtained which indicates normality in the error term and therefore there is no problem with our data. Further, Residuals
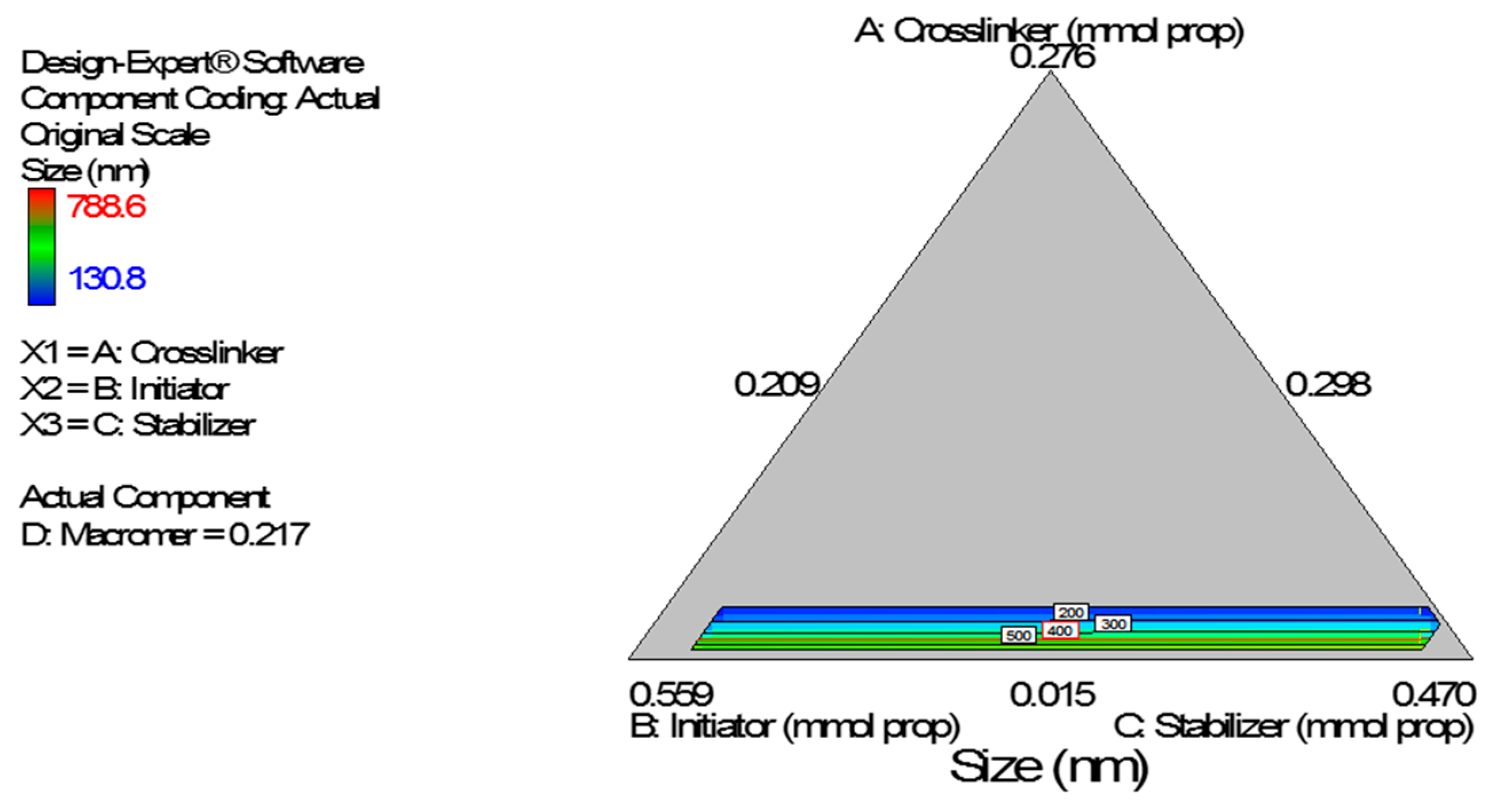
vs. Predicted tests the assumption of constant variance and it should be a random scatter within the upper and lower boundaries. The Scheffe polynomial (Equation (3)) was used to generate the model graph (
Figure 7) which shows the design space and variation in particle size as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Poly-ɛ-caprolactone based nanoparticles).
Figure 6.
Diagnostic plots the particle size data of poly-ɛ-aprolactone-based nanoparticles: (A) Normal Plot of Residuals (B) Residuals vs. Predicted.
Figure 6.
Diagnostic plots the particle size data of poly-ɛ-aprolactone-based nanoparticles: (A) Normal Plot of Residuals (B) Residuals vs. Predicted.
Figure 7.
Model graph showing the design space and variation in particle size as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (poly-ɛ-caprolactone-based nanoparticles).
Figure 7.
Model graph showing the design space and variation in particle size as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (poly-ɛ-caprolactone-based nanoparticles).
Following square root transformation, model fitting for zeta potential data was carried out. Quadratic model was found significant and the model was selected. To improve the model, insignificant terms were removed by backward elimination. Analysis of variance (ANOVA) (
Table 6) reveals that the selected model is significant (
p = 0.0437). The linear mixture terms (component linear terms) are not significant (
p = 0.7487); the quadratic term of A (crosslinker) by C (stabilizer) is significant
p = 0.0037. In addition, “lack of fit” is not significant (
p = 0.4389). Non-significant lack of fit is good; we want the model to fit. Adequate precision measures the signal to noise ratio. The ratio of 6.96 indicates an adequate signal. This model can be used to navigate the design space.
Table 6.
Analysis of variance table for zeta potential (Poly-ɛ-caprolactone-based nanoparticles.
Table 6.
Analysis of variance table for zeta potential (Poly-ɛ-caprolactone-based nanoparticles.
| Source | Sum of Squares | Df | Mean Square | F-Value | p-Value (Prob > F) | |
|---|
| Model | 43.01 | 4 | 10.75 | 2.87 | 0.0437 | s |
| Linear Mixture | 4.58 | 3 | 1.53 | 0.41 | 0.7487 | ns |
| AC | 38.43 | 1 | 38.43 | 10.27 | 0.0037 | s |
| Residual | 93.56 | 25 | 3.74 | | | |
| Lack of Fit | 77.93 | 20 | 3.90 | 1.25 | 0.4389 | ns |
| Pure Error | 15.63 | 5 | 3.13 | | | |
| Cor Total | 136.57 | 29 | | | | |
The empirical model (Scheffe polynomial) is shown in Equation (4) below:
where: A = Crosslinker (mmol); B = Initiators (mmol); C = Stabilizer (mmol); D = Macromonomer (mmol).
Diagnostic plots show the validity of the model (
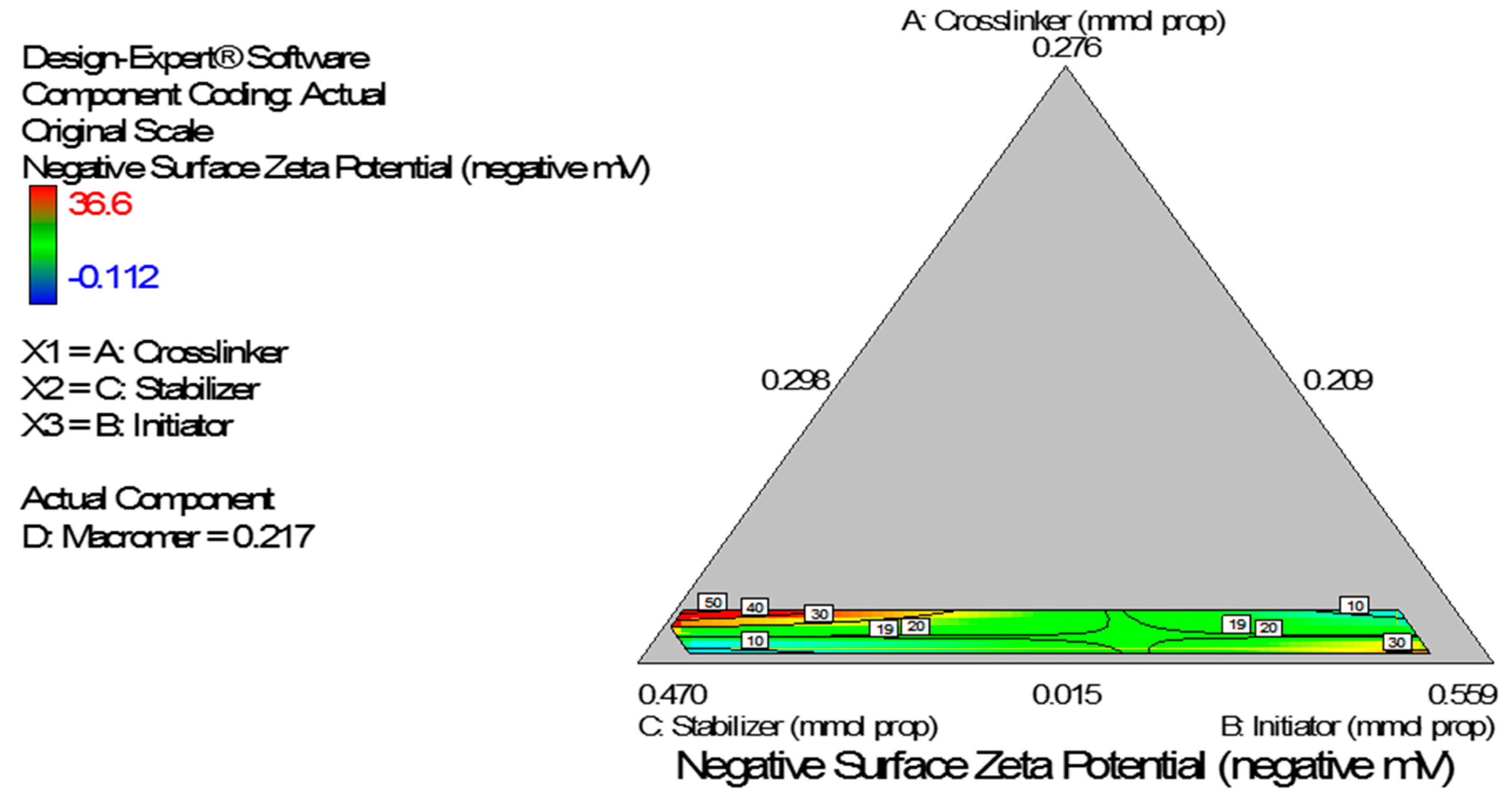
Figure 8). The Scheffe polynomial (Equation (4)) was used to generate the model graph (
Figure 9) which shows the design space and variation in zeta potential as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (Poly-ɛ-caprolactone based nanoparticles).
Figure 8.
Typical Diagnostic plots the zeta potential data of poly-ɛ-caprolactone-based nanoparticles (A) Residuals vs. Experimental Run (should show a random scatter); (B) Normal Plot of Residuals (should give a straight line).
Figure 8.
Typical Diagnostic plots the zeta potential data of poly-ɛ-caprolactone-based nanoparticles (A) Residuals vs. Experimental Run (should show a random scatter); (B) Normal Plot of Residuals (should give a straight line).
Figure 9.
Model graph showing the design space and variation in negative zeta potential as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (poly-ɛ-caprolactone-based nanoparticles).
Figure 9.
Model graph showing the design space and variation in negative zeta potential as a function of the mixture composition. A = Crosslinking agent; B = Initiators; C = Stabilizer and D = Macromonomer (poly-ɛ-caprolactone-based nanoparticles).