Characteristics of Antibiotic Resistance of Airborne Staphylococcus Isolated from Metro Stations
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection and Identification
2.2. Antimicrobial Susceptibility Tests
2.3. Detection of Methicillin and Disinfectant Resistance Genes and Sequence Analysis
2.4. Biofilm Detection
2.5. Statistical Analysis
3. Results
3.1. Strain Identification and Separation of CNS Strains
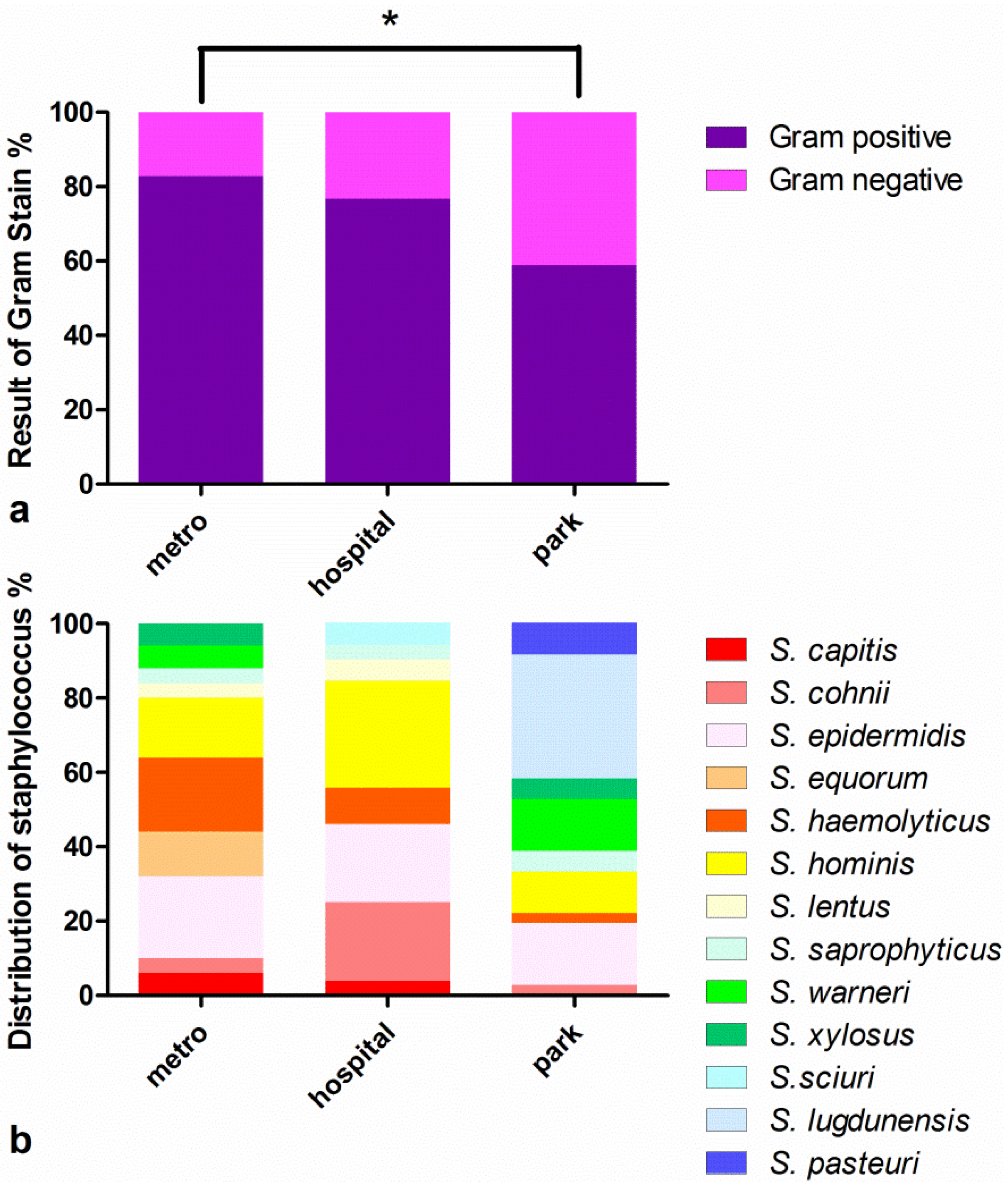
3.2. Antimicrobial Susceptibility Tests
| Antibiotics | Metro | Hospital | Park | p value | Average antibiotic resistance |
|---|---|---|---|---|---|
| n = 50 | n = 52 | n = 36 | |||
| Resistance rate | Resistance rate | Resistance rate | |||
| Cell wall synthesis | |||||
| Oxacillin | 22.00% | 21.15% | 36.11% | p > 0.05 | 33.33% |
| Penicillin (G) | 28.00% | 38.46% | 25.00% | p > 0.05 | 32.61% |
| Ampicillin | 20.00% | 36.54% | 13.89% | p < 0.05 * | 24.64% |
| Cefepime | 0.00% | 1.92% | 0.00% | p > 0.05 | 0.72% |
| Vancomycin | 0.00% | 0.00% | 0.00% | p > 0.05 | 0.00% |
| Amoxicillin/clavulanic acid | 8.00% | 19.23% | 0.00% | p > 0.05 | 10.14% |
| Synthesis of protein | |||||
| Gentamicin | 0.00% | 0.00% | 2.78% | p > 0.05 | 0.72% |
| Netilmicin | 0.00% | 0.00% | 0.00% | p > 0.05 | 0.00% |
| Erythromycin | 30.00% | 30.77% | 11.11% | p < 0.05 * | 25.36% |
| Tetracycline | 6.00% | 5.77% | 2.78% | p > 0.05 | 5.07% |
| Clindamycin | 24.00% | 23.08% | 19.44% | p > 0.05 | 22.46% |
| Synthesis of DNA | |||||
| Ciprofloxacin | 2.00% | 1.92% | 8.33% | p > 0.05 | 3.62% |
| Pefloxacin | 28.00% | 9.62% | 27.78% | p < 0.05 * | 21.01% |
| Synthesis of folacin | |||||
| Bactrim | 44.00% | 42.31% | 30.56% | p > 0.05 | 39.86% |
| Synthesis of RNA | |||||
| Rifampicin | 0.00% | 3.85% | 2.78% | p > 0.05 | 2.17% |
| Others | |||||
| Nitrofurantoin | 52.00% | 34.62% | 22.22% | p < 0.05 * | 37.68% |
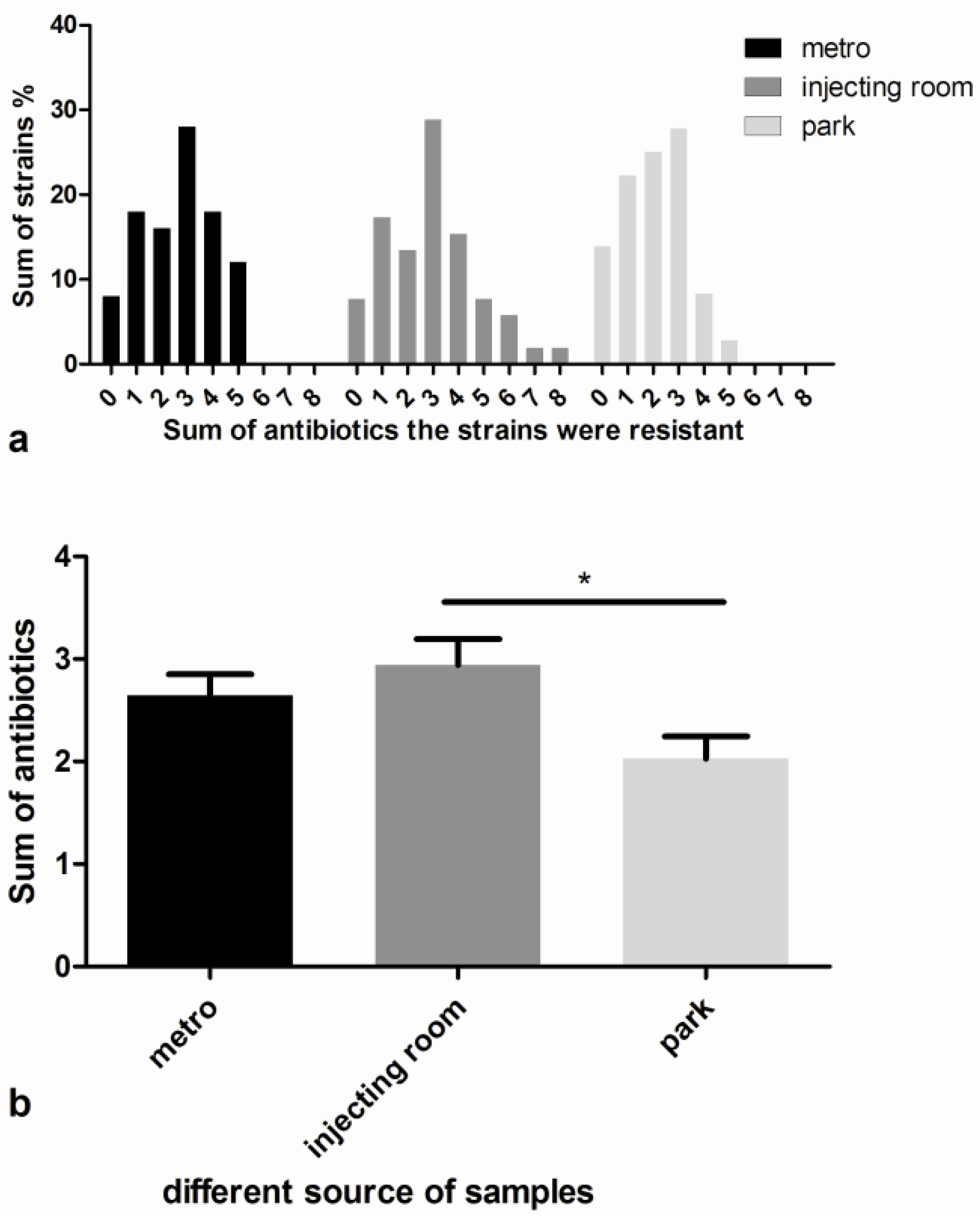
3.3. Detection of Methicillin and Disinfectant Resistance Genes
| Gene/Group | Metro station | Treatment room | Park | p value | |||
|---|---|---|---|---|---|---|---|
| n = 50 | n = 52 | n = 36 | |||||
| mecA | 14 | 28.00% | 10 | 19.23% | 2 | 5.56% | p < 0.05 |
| qac | 20 | 40.00% | 18 | 34.62% | 2 | 5.56% | p < 0.05 |
3.4. Sequence Analysis of the Resistance Genes
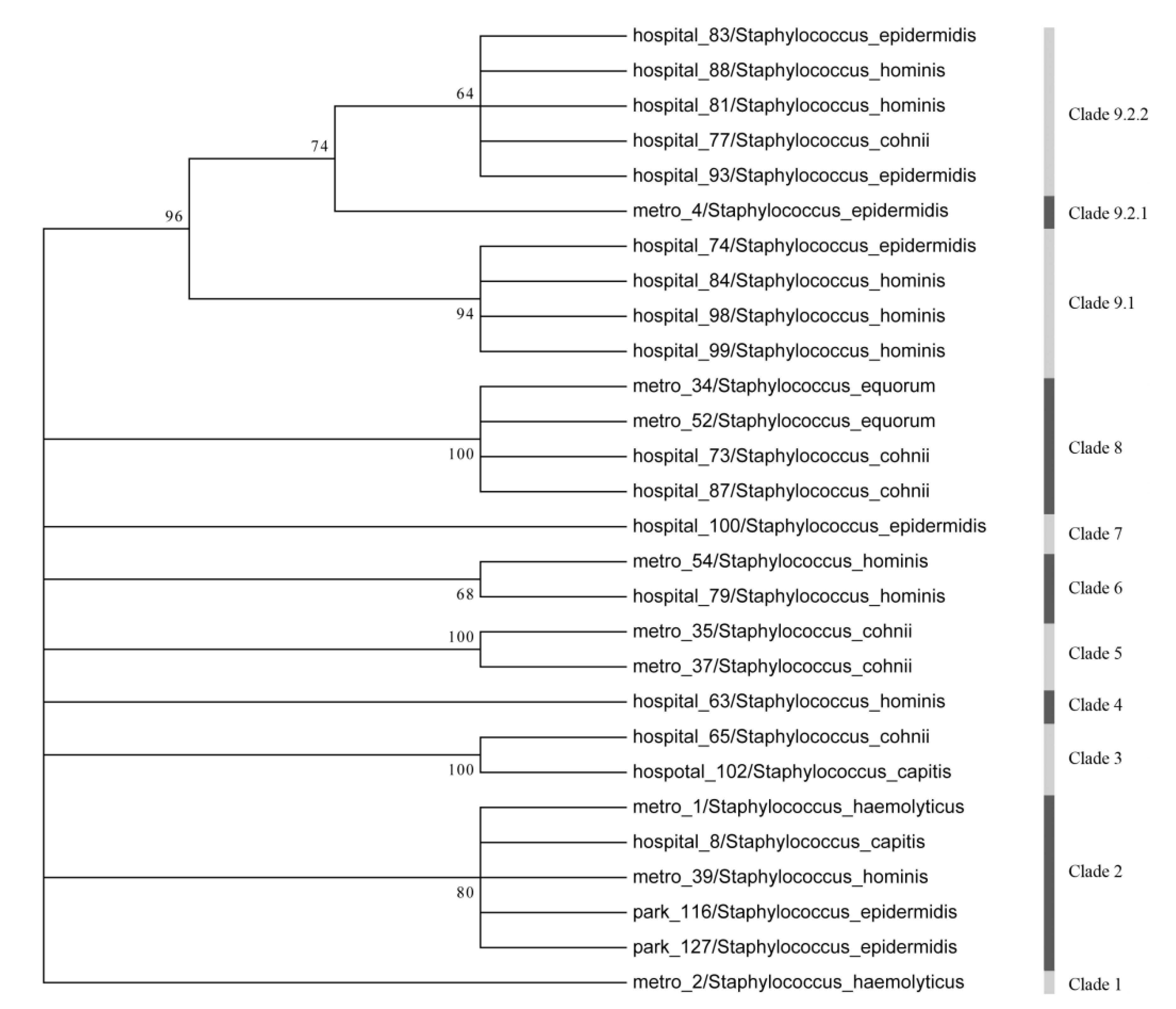

3.5. Biofilm Detection
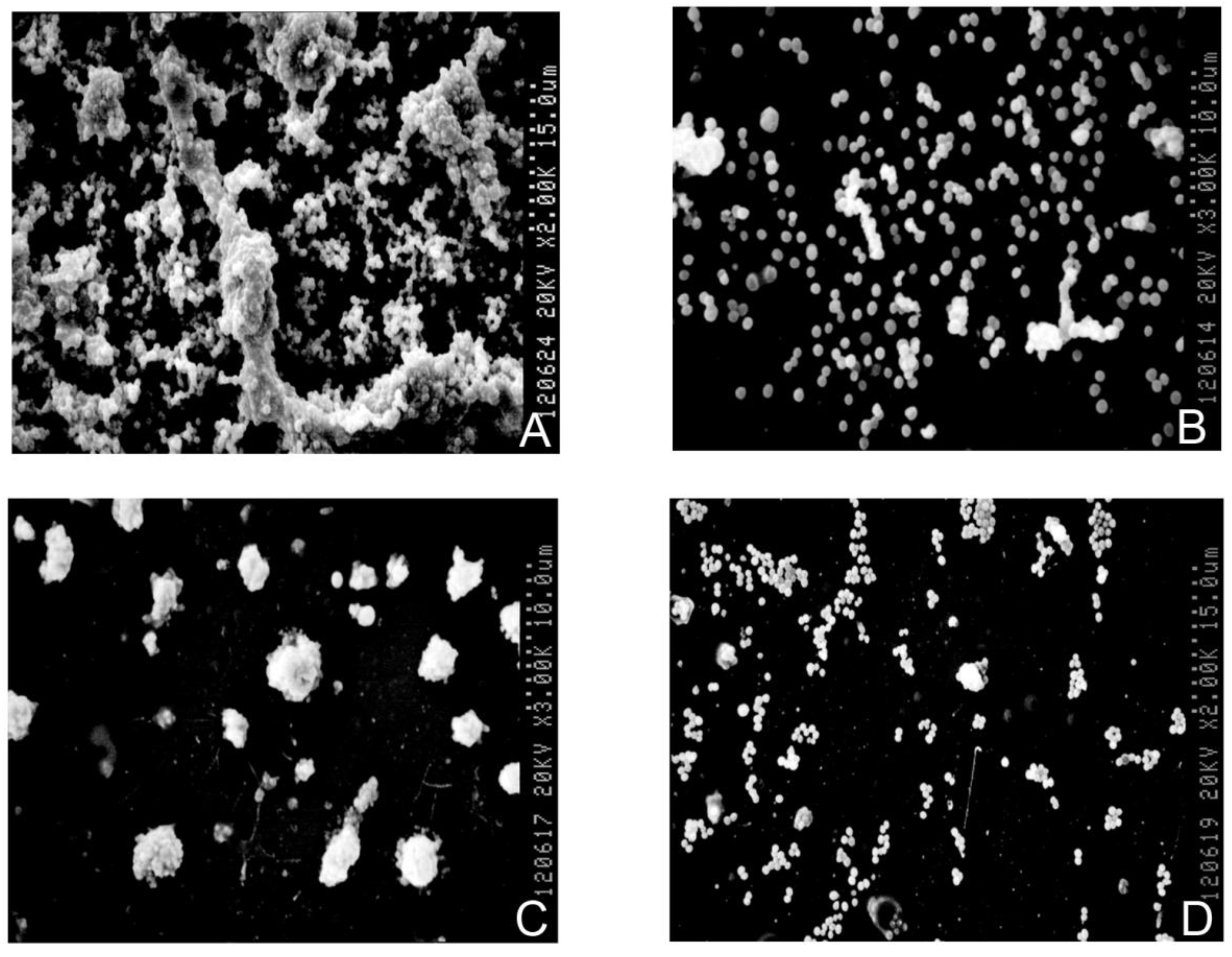
4. Discussion and Conclusions
Acknowledgments
Conflict of Interest
References
- Baba, T.; Takeuchi, F.; Kuroda, M.; Yuzawa, H.; Aoki, K.; Oguchi, A.; Nagai, Y.; Iwama, N.; Asano, K.; Naimi, T.; et al. Genome and virulence determinants of high virulence community-acquired MRSA. Lancet 2002, 359, 1819–1827. [Google Scholar] [CrossRef]
- Sobhy, N.; Aly, F.; Abd El Kader, O.; Ghazal, A.; Elbaradei, A. Community-acquired methicillin-resistant Staphylococcus aureus from skin and soft tissue infections (in a sample of Egyptian population): Analysis of mec gene and staphylococcal cassette chromosome. Braz. J. Infect. Dis. 2012, 16, 426–431. [Google Scholar] [CrossRef]
- Nobile, C.; Costantino, R.; Bianco, A.; Pileggi, C.; Pavia, M. Prevalence and pattern of antibiotic resistance of Campylobacter spp. in poultry meat in Southern Italy. Food Control 2013, 32, 715–718. [Google Scholar] [CrossRef]
- Reynolds, K.A.; Watt, P.M.; Boone, S.A.; Gerba, C.P. Occurrence of bacteria and biochemical markers on public surfaces. Int. J. Environ. Health Res. 2005, 15, 225–234. [Google Scholar] [CrossRef]
- Dybwad, M.; Granum, P.E.; Bruheim, P.; Blatny, J.M. Characterization of airborne bacteria at an underground subway station. Appl. Environ. Microbiol. 2012, 78, 1917–1929. [Google Scholar] [CrossRef]
- Gilleberg, S.B.; Faull, J.L.; Graeme-Cook, K.A. A preliminary survey of aerial biocontaminants at six London underground stations. Int. Biodeterior. Biodegrad. 1998, 41, 149–152. [Google Scholar] [CrossRef]
- Awad, A.H.A. Environmental study in subway metro stations in Cairo, Egypt. J. Occup. Health 2002, 44, 112–118. [Google Scholar] [CrossRef]
- Hwang, S.H.; Yoon, C.S.; Ryu, K.N.; Paik, S.Y.; Cho, J.H. Assessment of airborne environmental bacteria and related factors in 25 underground railway stations in Seoul, Korea. Atmos. Environ. 2010, 44, 1658–1662. [Google Scholar] [CrossRef]
- Kawasaki, T.; Kyotani, T.; Ushiogi, T.; Izumi, Y.; Lee, H.; Hayakawa, T. Distribution and identification of airborne fungi in railway stations in Tokyo, Japan. J. Occup. Health 2010, 52, 186–193. [Google Scholar] [CrossRef]
- Dong, S.; Yao, M. Exposure assessment in Beijing, China: Biological agents, ultrafine particles, and lead. Environ. Monit. Assess. 2010, 170, 331–343. [Google Scholar] [CrossRef]
- Yeh, P.; Simon, D.; Millar, J.; Alexander, F.; Franklin, D. A diversity of antibiotic-resistant Staphylococcus spp. in a public transportation system. Osong Public Health Res. Perspect. 2011, 2, 202–209. [Google Scholar] [CrossRef]
- Mandell, G.L.; Bennett, J.E.; Dolin, R. Mandell, Dauglas and Bennett’s Principles and Practice of Infectious Diseases, 5th ed.; Churchill Livingstone Inc: Philadelphia, PA, USA, 2000; p. 2049. [Google Scholar]
- Schreckenberger, P.C.; Ilendo, E.; Ristow, K.L. Incidence of constitutive and inducible clindamycin resistance in Staphylococcus aureus and coagulase-negative staphylococci in a community and a tertiary care hospital. J. Clin. Microbiol. 2004, 42, 2777–2779. [Google Scholar] [CrossRef]
- O’Toole, G.; Kaplan, H.B.; Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol. 2000, 54, 49–79. [Google Scholar] [CrossRef]
- Enright, M.C.; Robinson, D.A.; Randle, G.; Feil, E.J.; Grundmann, H.; Spratt, B.G. The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA). Proc. Natl. Acad. Sci. USA 2002, 99, 7687–7692. [Google Scholar]
- Koksal, F.; Yasar, H.; Samasti, M. Antibiotic resistance patterns of coagulase-negative Staphylococcus strains isolated from blood cultures of septicemic patients in Turkey. Microbiol. Res. 2009, 164, 404–410. [Google Scholar] [CrossRef]
- Poston, S.M.; Li Saw Hee, F.L. Genetic characterisation of resistance to metal ions in methicillin-resistant Staphylococcus aureus: Elimination of resistance to cadmium, mercury and tetracycline with loss of methicillin resistance. J. Med. Microbiol. 1991, 34, 193–201. [Google Scholar] [CrossRef]
- Mitchell, B.A.; Brown, M.H.; Skurray, R.A. QacA Multidrug Efflux Pump from Staphylococcus aureus: Comparative analysis of resistance to diamidines, biguanidines, and guanylhydrazones. Antimicrob. Agents Chemother. 1998, 42, 475–477. [Google Scholar] [CrossRef]
- Ministry of Health Law and Oversight Division, Disinfection Technical Specifications; Ministry of Health: Beijing, China, 2002.
- Onni, T.; Sanna, G.; Cubeddu, G.P.; Marogna, G.; Lollai, S.; Leori, G.; Tola, S. Identification of coagulase-negative staphylococci isolated from ovine milk samples by PCR-RFLP of 16S rRNA and gap genes. Vet. Microbiol. 2010, 144, 347–352. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. ClustalW and ClustalX version 2. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Li, M.; Wang, X.; Gao, Q.; Lu, Y. Molecular characterization of Staphylococcus epidermidis strains isolated from a teaching hospital in Shanghai, China. J. Med. Microbiol. 2009, 58, 456–461. [Google Scholar] [CrossRef]
- Wang, L.; Li, M.; Dong, D.; Bach, T.H.; Sturdevant, D.E.; Vuong, C.; Otto, M.; Gao, Q. SarZ is a key regulator of biofilm formation and virulence in Staphylococcus epidermidis. J. Infect. Dis. 2008, 197, 1254–1262. [Google Scholar] [CrossRef]
- Koskela, A.; Nilsdotter-Augustinsson, Å.; Persson, L.; Söderquist, B. Prevalence of the ica operon and insertion sequence IS256 among Staphylococcus epidermidis prosthetic joint infection isolates. Eur. J. Clin. Microbiol. Infect. Dis. 2009, 28, 655–660. [Google Scholar] [CrossRef]
- The World Health Report; World Health Organization: Geneva, Switzerland, 1996.
- Cosgrove, S.E.; Sakoulas, G.; Perencevich, E.N.; Schwaber, M.J.; Karchmer, A.W.; Carmeli, Y. Comparison of mortality associated with methicillin-resistant and methicillin-susceptible Staphylococcus aureus bacteremia: A meta-analysis. Clin. Infect. Dis. 2003, 36, 53–59. [Google Scholar] [CrossRef]
- Harbarth, S.; Rutschmann, O.; Sudre, P.; Pittet, D. Impact of methicillin resistance on the outcome of patients with bacteremia caused by Staphylococcus aureus. Arch. Intern. Med. 1998, 158, 182–189. [Google Scholar] [CrossRef]
- Gillespie, B.E.; Headrick, S.I.; Boonyayatra, S.; Oliver, S.P. Prevalence and persistence of coagulase-negative Staphylococcus species in three dairy research herds. Vet. Microbiol. 2009, 134, 65–72. [Google Scholar] [CrossRef]
- Lorenz, M.G.; Wackernagel, W. Bacterial gene transfer by natural genetic transformation in the environment. Microbiol. Rev. 1994, 58, 563–602. [Google Scholar]
- Zhu, B. Degradation of plasmid and plant DNA in water microcosms monitored by natural transformation and real-time polymerase chain reaction (PCR). Water Res. 2006, 40, 3231–3238. [Google Scholar] [CrossRef]
- Hyder, S.L.; Streitfeld, M.M. Transfer of erythromycin resistance from clinically isolated lysogenic strains of Streptococcus pyogenes via their endogenous phage. J. Infect. Dis. 1978, 138, 281–286. [Google Scholar] [CrossRef]
- Blahova, J.; Hupkova, M.; Babalova, M.; Krcmery, V.; Schafer, V. Transduction of resistance to imipenem, aztreonam and ceftazidime in nosocomial strains of Pseudomonas aeruginosa by wild-type phages. Acta Virol. 1993, 37, 429–436. [Google Scholar]
- Banks, D.J.; Porcella, S.F.; Barbian, K.D.; Beres, S.B.; Philips, L.E.; Voyich, J.M.; DeLeo, F.R.; Martin, J.M.; Somerville, G.A.; Musser, J.M. Progress toward characterization of the group A streptococcus metagenome: Complete genome sequence of a macrolide-resistant serotype M6 strain. J. Infect. Dis. 2004, 190, 727–738. [Google Scholar] [CrossRef]
- Kuenne, C.; Voget, S.; Pischimarov, J.; Oehm, S.; Goesmann, A.; Daniel, R.; Hain, T.; Chakraborty, T. Comparative analysis of plasmids in the genus Listeria. PLoS ONE 2010, 5, e12511. [Google Scholar] [CrossRef]
- Luo, M.; Fei, Y.E.; Wang, D.M.; He, J.L.; Zhong, H.Y. Analysis of coagulase-negative Staphylococcus from sputum. J. Modern Clin. Med. 2010, 5, 83–84. [Google Scholar]
- Cosgrove, S.E.; Sakoulas, G.; Perencevich, E.N.; Schwaber, M.J.; Karchmer, A.W.; Carmeli, Y. Comparison of mortality associated with methicillin-resistant and methicillin-susceptible Staphylococcus aureus bacteremia: A meta-analysis. Clin. Infect. Dis. 2003, 36, 53–59. [Google Scholar] [CrossRef]
- Harbarth, S.; Rutschmann, O.; Sudre, P.; Pittet, D. Impact of methicillin resistance on the outcome of patients with bacteremia caused by Staphylococcus aureus. Arch. Intern. Med. 1998, 158, 182–189. [Google Scholar] [CrossRef]
- Hisata, K.; Ito, T.; Matsunaga, N.; Komatsu, M.; Jin, J.; Li, S.; Watanabe, S.; Shimizu, T.; Hiramatsu, K. Dissemination of multiple MRSA clones among community-associated methicillin-resistant Staphylococcus aureus infections from Japanese children with impetigo. J. Infect. Chemother. 2011, 17, 609–621. [Google Scholar] [CrossRef]
- Barbier, F.; Ruppé, E.; Hernandez, D.; Lebeaux, D.; Francois, P.; Felix, B.; Desprez, A.; Maiga, A.; Woerther, P.L.; Gaillard, K.; et al. Methicillin-resistant coagulase-negative staphylococci in the community: High homology of SCCmec IVa between Staphylococcus epidermidis and major clones of methicillin-resistant Staphylococcus aureus. J. Infect. Dis. 2010, 202, 270–281. [Google Scholar]
- Schnellmann, C.; Gerber, V.; Rossano, A.; Jaquier, V.; Panchaud, Y.; Doherr, M.G.; Thomann, A.; Straub, R.; Perreten, V. Presence of new MecA and mph(C) variants conferring antibiotic resistance in Staphylococcus spp. isolated from the skin of horses before and after clinic admission. J. Clin. Microbiol. 2006, 44, 4444–4454. [Google Scholar] [CrossRef]
- Soge, O.O.; Meschke, J.S.; No, D.B.; Roberts, M.C. Characterization of methicillin-resistant Staphylococcus aureus and methicillin-resistant coagulase-negative Staphylococcus spp. Isolated from US West Coast public marine beaches. J. Antimicrob. Chemother. 2009, 64, 1148–1155. [Google Scholar] [CrossRef]
- Wei, W.P. Distribution and drug-resistance of 408 strains of coagulase negative Staphylococcus. China Prac. Med. 2010, 5, 67–69. [Google Scholar]
- Heir, E.; Sundheim, G.; Holck, A.L. The Staphylococcus qacH gene product: A new member of the SMR family encoding multidrug resistance. FEMS Microbiol. Lett. 1998, 163, 49–56. [Google Scholar] [CrossRef]
- Luthje, P.; von Kockritz-Blickwede, M.; Schwarz, S. Identification and characterization of nine novel types of small staphylococcal plasmids carrying the lincosamide. J. Antimicrob. Chemother. 2007, 59, 600–606. [Google Scholar] [CrossRef]
- Bjorland, J.; Steinum, T.; Sunde, M.; Waage, S.; Heir, E. Novel plasmid-borne gene qacJ mediates resistance to quaternary ammonium compounds in equine Staphylococcus aureus, Staphylococcus simulans, and Staphylococcus intermedius. Antimicrob. Agents Chemother. 2003, 47, 3046–3052. [Google Scholar] [CrossRef]
- O’Toole, G.; Kaplan, H.B.; Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol. 2000, 54, 49–79. [Google Scholar] [CrossRef]
- Siddiqui, A.R.; Bernstein, J.M. Chronic wound infection: Facts and controversies. Clin. Dermatol. 2010, 28, 519–526. [Google Scholar] [CrossRef]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilm formation: A clinically relevant microbiological process. Clin. Infect. Dis. 2001, 33, 1387–1392. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhou, F.; Wang, Y. Characteristics of Antibiotic Resistance of Airborne Staphylococcus Isolated from Metro Stations. Int. J. Environ. Res. Public Health 2013, 10, 2412-2426. https://doi.org/10.3390/ijerph10062412
Zhou F, Wang Y. Characteristics of Antibiotic Resistance of Airborne Staphylococcus Isolated from Metro Stations. International Journal of Environmental Research and Public Health. 2013; 10(6):2412-2426. https://doi.org/10.3390/ijerph10062412
Chicago/Turabian StyleZhou, Feng, and Yuyan Wang. 2013. "Characteristics of Antibiotic Resistance of Airborne Staphylococcus Isolated from Metro Stations" International Journal of Environmental Research and Public Health 10, no. 6: 2412-2426. https://doi.org/10.3390/ijerph10062412
APA StyleZhou, F., & Wang, Y. (2013). Characteristics of Antibiotic Resistance of Airborne Staphylococcus Isolated from Metro Stations. International Journal of Environmental Research and Public Health, 10(6), 2412-2426. https://doi.org/10.3390/ijerph10062412




