Antimicrobial Resistance in Enterococcus spp. Isolated from Environmental Samples in an Area of Intensive Poultry Production
Abstract
:1. Introduction
2. Experimental Section
2.1. Surface and Ground Water Sampling
2.2. Isolation and Identification of Enterococci
2.3. Antimicrobial Resistance Testing
2.4. Data Analysis
2.5. MAR Index
3. Results and Discussion
3.1. Bacterial Recovery
| No. (%) of samples containing: | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Sample | n | E. faecalis | E. faecium | E. casseliflavus | E. durans | E. gallinarum | E. hirae | E. mundtii | E. raffinosus | All other species |
| Surface Water | ||||||||||
| S1 | 8 | 0 | 4 (50) | 0 | 1 (12.5) | 0 | 0 | 3 (37.5) | 0 | 0 |
| S2 | 5 | 0 | 3 (60) | 2 (40) | 0 | 0 | 0 | 0 | 0 | 0 |
| S3 | 8 | 0 | 4 (50) | 1 (12.5) | 0 | 0 | 1 (12.5) | 0 | 1 (12.5) | 1 (12.5) |
| S4 | 10 | 5 (50) | 1 (10) | 0 | 0 | 0 | 1 (10) | 3 (30) | 0 | 0 |
| S5 | 2 | 0 | 1 (50) | 0 | 1 (50) | 0 | 0 | 0 | 0 | 0 |
| S6 | 10 | 4 (40) | 4 (40) | 0 | 1 (10) | 1 (10) | 0 | 0 | 0 | 0 |
| S7 | 7 | 1 (14.3) | 2 (28.6) | 1 (14.3) | 2 (28.6) | 1 (14.3) | 0 | 0 | 0 | 0 |
| S8 | 10 | 10 (100) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| S9 | 9 | 0 | 1 (11.1) | 0 | 0 | 0 | 0 | 0 | 0 | 8 (88.9) |
| S10 | 2 | 1 (50) | 1 (50) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| S11 | 7 | 2 (28.6) | 1 (14.3) | 2 (28.6) | 0 | 0 | 1 (14.3) | 1 (14.3) | 0 | 0 |
| S12 | 7 | 0 | 0 | 2 (28.6) | 1 (14.3) | 0 | 0 | 4 (57.1) | 0 | 0 |
| Total | 85 | 23 (27.1) | 22 (25.9) | 8 (9.4) | 6 (7.1) | 2 (2.4) | 3 (3.5) | 11 (12.9) | 1 (1.2) | 9 |
| Ground Water | ||||||||||
| BC-008 | 5 | 0 | 1 (20) | 0 | 3 (60) | 0 | 1 (20) | 0 | 0 | 0 |
| 91-11 | 1 | 1 (100) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| US-02 | 1 | 1 (100) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Total | 7 | 2 (28.6) | 1 (14.3) | 0 | 3 (42.9) | 0 | 1 (14.3) | 0 | 0 | 0 |
| Total Environment | 92 | 25 (27.2) | 23 (25) | 8 (8.7) | 9 (9.8) | 2 (2.2) | 4 (4.3) | 11 (12) | 1 (1.1) | 9 (9.8) |
| Poultry Litter | ||||||||||
| Layers | 29 | 0 | 29 (100) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Broilers (day 3) | 105 | 30 (28.6) | 16 (15.2) | 1 (0.95) | 1 (0.95) | 28 (26.7) | 27 (25.7) | 0 | 0 | 2 (1.9) |
| Broilers (day 35) | 29 | 6 (20.7) | 21 (72.4) | 0 | 1 (3.4) | 0 | 1 (3.4) | 0 | 0 | 0 |
| Total | 163 | 36 (22.1) | 66 (40.5) | 1 (0.6) | 2 (1.2) | 28 (17.2) | 28 (17.2) | 0 | 0 | 2 (1.2) |
3.2. Antimicrobial Resistance
3.2.1. Litter Samples
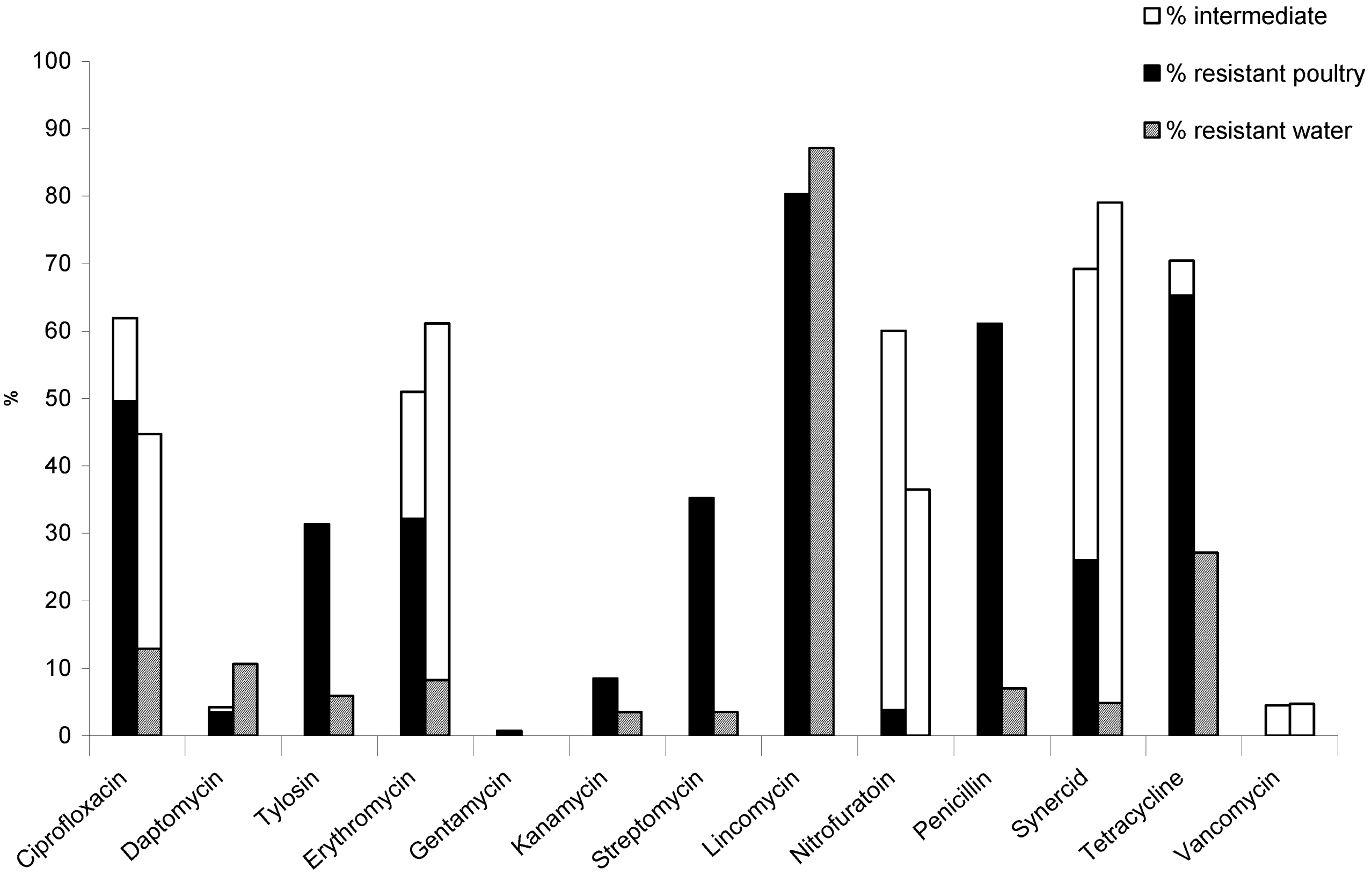
3.2.2. Water Samples
| Antimicrobial | Break-point (µg/mL) | Source | No. (%) of isolates resistant | |||||
|---|---|---|---|---|---|---|---|---|
| E. faecalis (n = 57) | E. faecium (n = 88) | E. gallinarum (n = 30) | E. hirae (n = 32) | E. durans (n = 11) | All other species (n = 32) | |||
| Chloramphenicol | ≥32 | Water | 1 (1.8) | 0 | 0 | 0 | 0 | 0 |
| Layers | 0 | |||||||
| Broilers | 0 | 0 | 0 | 0 | 0 | 0 | ||
| Ciprofloxacin | ≥4 | Water | 1 (1.8) | 11 (12.5) | 0 | 0 | 0 | 1 |
| Layers | 22 (25) | |||||||
| Broilers | 0 | 25 (28.4) | 2 (6.7) | 1 (3.1) | 0 | 0 | ||
| Daptomycin | ≥8 | Water | 0 | 0 | 0 | 1 (3.1) | 0 | 9 |
| Layers | 2 (2.3) | |||||||
| Broilers | 0 | 0 | 0 | 0 | 0 | 0 | ||
| Tylosin | ≥32 | Water | 2 (3.5) | 0 | 0 | 1 (3.1) | 3 (27.2) | 0 |
| Layers | 6 (6.8) | |||||||
| Broilers | 17 (29.8) | 6 (6.8) | 25 (83.3) | 2 (6.3) | 2 (18.2) | 3 | ||
| Erythromycin | ≥8 | Water | 2 (3.5) | 1 (1.1) | 0 | 1 (3.1) | 3 (27.2) | 1 |
| Layers | 6 (6.8) | |||||||
| Broilers | 17 (29.8) | 8 (9.1) | 25 (83.3) | 2 (6.3) | 2 (18.2) | 3 | ||
| Kanamycin | ≥1,024 | Water | 1 (1.8) | 1 (1.1) | 0 | 1 (3.1) | 0 | 0 |
| Layers | 1 (1.1) | |||||||
| Broilers | 4 (7.0) | 6 (6.8) | 5 (16.7) | 2 (6.3) | 0 | 0 | ||
| Streptomycin | >1,000 | Water | 1 (1.8) | 1 (1.1) | 0 | 0 | 3 (27.2) | 0 |
| Layers | 8 (9.1) | |||||||
| Broilers | 8 (14.0) | 22 (25) | 20 (66.7) | 1 (3.1) | 2 (18.2) | 1 | ||
| Lincomycin | >1,000 | Water | 24 (42.1) | 13 (14.8) | 2 (6.7) | 4 (12.5) | 9 (81.8) | 29 |
| Layers | 18 (20.5) | |||||||
| Broilers | 32 (56.1) | 34 (38.6) | 28 (87.5) | 2 (18.2) | 3 | |||
| Nitrofurantoin | ≥128 | Water | 0 | 2 (2.3) | 0 | 1 (3.1) | 4 (36.4) | 0 |
| Layers | 0 | |||||||
| Broilers | 0 | 4 (4.5) | 0 | 4 (12.5) | 1 (0.9) | 0 | ||
| Penicillin | ≥16 | Water | 0 | 5 (5.7) | 0 | 1 (3.1) | 0 | 0 |
| Layers | 23 (26.1) | |||||||
| Broilers | 0 | 31 (35.2) | 0 | 24 (75) | 0 | 0 | ||
| Synercid® | ≥4 | Water | 24 (42.1) | 1 (1.1) | 0 | 1 (3.1) | 0 | 1 |
| Layers | 2 (2.3) | |||||||
| Broilers | 32 (56.1) | 19 (21.6) | 3 (10) | 1 (3.1) | 2 (18.2) | 1 | ||
| Tetracycline | ≥16 | Water | 13 (22.8) | 7 (8.0) | 0 | 1 (3.1) | 5 (45.5) | 1 |
| Layers | 13 (14.8) | |||||||
| Broilers | 22 (38.6) | 31 (35.2) | 27 (90) | 27 (84.4) | 2 (18.2) | 2 | ||
| Gentamicin | ≥500 | Water | 0 | 0 | 0 | 0 | 0 | 0 |
| Layers | 0 | |||||||
| Broilers | 0 | 0 | 0 | 2 (6.3) | 0 | 0 | ||
3.2.3. Resistance Patterns
| No. antimicrobials | Resistance pattern a | Species (No. isolates) | Source | |
|---|---|---|---|---|
| Litter | Environment | |||
| 9 | Lin Tet Pen Tyl Cip Str Syn Kan Nit | E. faecium (1) | 1 | |
| Lin Tet Pen Tyl Ery Cip Str Syn Kan Ni | E. faecium (4) | 4 | ||
| 8 | Lin Tet Pen Tyl Ery Str Cip Syn | E. faecium (1) | 1 | |
| Lin Tet Tyl Ery Str Kan Chl | E. faecalis (1) | 1 | ||
| 7 | Lin Tet Pen Ery Str Syn Cip | E. faecium (1) | 1 | |
| Lin Tet Tyl Ery Str Syn Kan | E. faecalis (4) | 4 | ||
| E. gallinarum (3) | 3 | |||
| Lin Tet Pen Cip Str Syn Nit | E. hirae (1) | 1 | ||
| Lin Tet Tyl Ery Str Syn Nit | E. durans (1) | 1 | ||
| Lin Pen Tet Tyl Ery Cip Str | E. faecium (3) | 3 | ||
| Lin Pen Tet Tyl Ery Str Syn | E. faecium (1) | 1 | ||
| Lin Pet Tet Tyl Ery Str Kan | E. faecium (1) | 1 | ||
| Lin Pen Tyl Cip Ery Str Syn | E. faecium (1) | 1 | ||
| 6 | Lin Tet Pen Cip Syn Lin | E. faecium (9) | 9 | |
| Lin Tet Tyl Str Syn Ery | E. faecalis (1) | 1 | ||
| Lin Tet Tyl Ery Str Kan | E. gallinarum (2) | 2 | ||
| E. durans (1) | 1 | |||
| 5 | Lin Tet Pen Cip Str | E. faecium (4) | 4 | |
| Lin Tet Tyl Ery Syn | E. faecalis (13) | 12 | 1 | |
| Lin Tet Tyl Ery Str | E. gallinarum (15) | 15 | ||
| E. durans (2) | 2 | |||
| Lin Tet Pen Gen Kan | E. hirae (2) | 2 | ||
| Lin Tet Pen Cip Dap | E. faecium (2) | 2 | ||
| 4 | Lin Tet Pen Str | E. faecium (2) | 2 | |
| Lin Tet Pen Syn | E. faecium (2) | 2 | ||
| Lin Tet Pen Cip | E. faecium (3) | 3 | ||
| Lin Tet Ery Tyl | E. gallinarum (5) | 5 | ||
| E. hirae (2) | 2 | |||
| E. durans (1) | 1 | |||
| E. species (1) | 1 | |||
| Lin Tet Str Tet | E. faecium (1) | 1 | ||
| Lin Tet Ery Kan | E. faecium (1) | 1 | ||
| Lin Tet Pen Nit | E. hirae (3) | 3 | ||
| Lin Pen Str Cip | E. faecium (1) | 1 | ||
| 3 | Pen Cip Nit | E. faecium (1) | 1 | |
| Tet Pen Cip | E. faecium (4) | 3 | 1 | |
| Lin Tet Ery | E. faecium (1) | 1 | ||
| E. raffinosus (1) | 1 | |||
| Lin Tet Syn | E. faecalis (17) | 5 | 12 | |
| Lin Tet Str | E. durans (1) | 1 | ||
| Lin Str Syn | E. faecalis (3) | 3 | ||
| Lin Tet Cip | E. faecium (1) | 1 | ||
| E. gallinarum (1) | 1 | |||
| Lin Tet Pen | E. faecium (2) | 1 | 1 | |
| E. hirae (18) | 18 | |||
| Lin Tyl Ery | E. species (1) | 1 | ||
| Lin Syn Cip | E. faecalis (1) | 1 | ||
| Lin Pen Cip | E. faecium (6) | 6 | ||
| Tet Cip Str | E. faecium (1) | 1 | ||
| 2 | Lin Tet | E. faecium (4) | 4 | |
| E. gallinarum (1) | 1 | |||
| E. hirae (1) | 1 | |||
| E. durans (1) | 1 | |||
| Lin Nit | E. faecium (1) | 1 | ||
| Lin Syn | E. faecalis (16) | 7 | 9 | |
| E. hirae (1) | 1 | |||
| E. species (1) | 1 | |||
| Pen Tet | E. faecium (2) | 2 | ||
| E. gallinarum (1) | 1 | |||
| Pen Cip | E. faecium (6) | 3 | 3 | |
| E. casseliflavus (1) | 1 | |||
| Lin Dap | E. hirae (1) | 1 | ||
| E. mundtii (11) | 11 | |||
| 1 | Pen | E. faecium (2) | 2 | |
| Lin | E. faecium (5) | 5 | ||
| E. faecalis (1) | 1 | |||
| E. gallinarum (2) | 2 | |||
| E. hirae (2) | 1 | 1 | ||
| E. durans (4) | 4 | |||
| E. casseliflavus (7) | 7 | |||
| E. species (8) | 8 | |||
| E. mundtii (1) | 1 | |||
| Cip | E. faecium (10) | 3 | 7 | |
| Tet | E. faecium (1) | 1 | ||
| Total | 157 | 93 | ||
3.3. MAR Indices
| AR | MAR | MAR index | |||
|---|---|---|---|---|---|
| ≥1 (%) | ≥2 (%) | ≥5 (%) | |||
| Surface water | |||||
| S1 | 100 | 63 | 13 | 0.141 | |
| S2 | 100 | 40 | 0 | 0.071 | |
| S3 | 100 | 75 | 0 | 0.133 | |
| S4 | 100 | 80 | 10 | 0.138 | |
| S6 | 90 | 40 | 10 | 0.131 | |
| S7 | 100 | 29 | 29 | 0.125 | |
| S8 | 0 | 100 | 0 | 0.188 | |
| S9 | 100 | 0 | 0 | 0.063 | |
| S11 | 100 | 71 | 0 | 0.107 | |
| S12 | 100 | 71 | 0 | 0.107 | |
| Groundwater | |||||
| 0 | 5 | 0 | 0.188 | ||
| Poultry farms | |||||
| Layers | 97 | 83 | 28 | 0.218 | |
| Broilers (day3) | 100 | 98 | 44 | 0.248 | |
| Broilers (day 35) | 100 | 100 | 83 | 0.358 | |
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Araujo, C.; Torres, C.; Silva, N.; Carneiro, C.; Goncalves, A.; Radhouani, H.; Correia, S.; da Costa, P.M.; Paccheco, R.; Zarazaga, M.; Ruiz-Larrea, F.; Poeta, P.; Igrejas, G. Vancomycin-resistant enterococci from Portuguese wastewater treatment plants. J. Basic Microbiol. 2010, 50, 605–609. [Google Scholar]
- Lanthier, M.; Scott, A.; Zhang, Y.; Cloutier, M.; Durie, D.; Henderson, V.C.; Wilkes, G.; Lapen, D.R.; Topp, E. Distribution of selected virulence genes and antibiotic resistance in Enterococcus species isolated from the South Nation River drainage basin, Ontario, Canada. J. Appl. Microbiol. 2011, 110, 407–421. [Google Scholar]
- Novo, A.; Manaia, C.M. Factors influencing antibiotic resistance burden in municipal wastewater treatment plants. Appl. Microbiol. Biotechnol. 2010, 87, 1157–1166. [Google Scholar]
- Kotzamanidis, C.; Zdragas, A.; Kourelis, A.; Moraitou, E.; Papa, A.; Yiantzi, V.; Pantelidou, C.; Yiangou, M. Characterization of vanA-type Enterococcus faecium isolates from urban and hospital wastewater and pigs. J. Appl. Microbiol. 2009, 107, 997–1005. [Google Scholar]
- Jackson, C.R.; Furtula, V.; Farrell, E.G.; Barrett, J.B.; Hiott, L.M.; Chambers, P. A comparison of BOX-PCR and pulsed-field gel electrophoresis to determine genetic relatedness of enterococci from different environments. Microb. Ecol. 2012, 64, 378–387. [Google Scholar]
- Facklam, R.R.; Sahm, D.F.; Teixeiram, L.M. Enterococcus. In Manual of Clinical Microbiology, 7th; Murray, P.R., Baron, E.J., Pfaller, M.A., Tenover, F.C., Yolken, R.H., Eds.; ASM Press: Washington, DC, USA, 1999; pp. 297–305. [Google Scholar]
- Malani, P.N.; Kauffman, C.A.; Zervos, M.J. Enterococcal disease, epidemiology, and treatment. In The Enterococci: Pathogenesis, Molecular Biology, and Antibiotic Resistance; Gilmore, M.S., Clewell, D.B., Courvalin, P., Dunny, G.M., Murray, B.E., Rice, L.B., Eds.; ASM Press: Washington, DC, USA, 2002; pp. 385–408. [Google Scholar]
- Cetinkaya, Y.; Falk, P.; Mayhall, C.G. Vancomycin-resistant enterococci. Clin. Microbiol. Rev. 2000, 13, 686–707. [Google Scholar]
- Macedo, A.S.; Freitas, A.R.; Abreu, C.; Machado, E.; Peixe, L.; Sousa, J.C.; Novais, C. Characterization of antibiotic resistant enterococci isolated from untreated waters for human consumption in Portugal. Int. J. Food Microbiol. 2011, 145, 315–319. [Google Scholar]
- McGowan, L.L.; Jackson, C.R.; Barrett, J.B.; Hiott, L.M.; Fedorka-Cray, P.J. Prevalence and antimicrobial resistance of enterococci isolated from retail fruits, vegetables, and meats. J. Food Prot. 2006, 69, 2976–2982. [Google Scholar]
- Panda, S.K.; Patra, A.K.; Kar, R.N. Monitoring of multiple drug-resistant pathogens in a selected stretch of Bay of Bengal, India. Environ. Monit. Assess. 2011, 5, 83–98. [Google Scholar]
- Murray, B.E. Diversity among multidrug-resistant enterococci. Emerg. Infect. Dis. 1998, 4, 37–47. [Google Scholar]
- Brooks, J.P.; Adeli, A.; Read, J.J.; McLaughlin, M.R. Rainfall simulation in greenhouse microcosms to assess bacterial-associated runoff from land-applied poultry litter. J. Environ. Qual. 2009, 38, 218–229. [Google Scholar]
- Diarrassouba, F.; Diarra, M.S.; Bach, S.; Delaquis, P.; Pritchard, J.; Topp, E.; Skura, B.J. Antibiotic resistance and virulence genes in commensal Escherichia coli and Salmonella isolates from commercial broiler chicken farms. J. Food Prot. 2007, 70, 1316–1327. [Google Scholar]
- Furtula, V.; Farrell, E.G.; Diarrassouba, F.; Rempel, H.; Pritchard, J.; Diarra, M.S. Veterinary pharmaceuticals and antibiotic resistance of Escherichia coli isolates in poultry litter from commercial farms and controlled feeding trials. Poult. Sci. 2010, 89, 180–188. [Google Scholar]
- Kelley, T.R.; Pancorbo, O.C.; Merka, W.C.; Thompson, S.A.; Cabrera, M.L.; Barnhart, H.M. Bacterial pathogens and indicators in poultry litter during re-utilization. J. Appl. Poultry Res. 1995, 4, 366–373. [Google Scholar]
- Graham, J.P.; Evans, S.L.; Price, L.B.; Silbergeld, E.K. Fate of antimicrobial-resistant enterococci and staphylococci and resistance determinants in stored poultry litter. Environ. Res. 2009, 109, 682–689. [Google Scholar]
- Jenkins, M.B.; Truman, C.C.; Siragusa, G.; Line, E.; Bailey, J.S.; Frye, J.; Endale, D.M.; Franklin, D.H.; Schomberg, H.H.; Fisher, D.S.; Sharpe, R.R. Rainfall and tillage effects on transport of fecal bacteria and sex hormones 17beta-estradiol and testosterone from broiler litter applications to a Georgia Piedmont Ultisol. Sci. Total Environ. 2008, 403, 154–163. [Google Scholar]
- Servais, P.; Passerat, J. Antimicrobial resistance of fecal bacteria in waters of the Seine river watershed (France). Sci. Total Environ. 2009, 408, 365–372. [Google Scholar]
- Lata, P.; Ram, S.; Agrawal, M.; Shanker, R. Enterococci in river Ganga surface waters: Propensity of species distribution, dissemination of antimicrobial-resistance and virulence-markers among species along landscape. BMC Microbiol. 2009, 9, 140–149. [Google Scholar]
- Moore, D.F.; Guzman, J.A.; McGee, C. Species distribution and antimicrobial resistance of enterococci isolated from surface and ocean water. J. Appl. Microbiol. 2008, 105, 1017–1025. [Google Scholar]
- Furtula, V.; Jackson, C.R.; Osman, R.; Chambers, P.A. Use of Enterococcus, BST and sterols for poultry pollution source tracking in surface and groundwater. In Environmental Health-Emerging Issues and Practice; Oosthuizen, J., Ed.; InTech Open Access: Rijeka, Croatia, 2012; pp. 57–78. [Google Scholar]
- Improved Enumeration Methods for the Recreational Water Quality Indicators: Enterococci and Escherichia coli. USEPA: Washington, DC, USA, 2000; pp. 1–49.
- Jackson, C.R.; Fedorka-Cray, P.J.; Barrett, J.B. Use of a genus- and species-specific multiplex PCR for identification of enterococci. J. Clin. Microbiol. 2004, 42, 3558–3565. [Google Scholar]
- Bacterial Epidemiology and Antimicrobial Resistance: NARMS—National Antimicrobial Resistance Monitoring System Animal Isolates. Available online: http://www.ars.usda.gov/Main/docs.htm? docid=6750&page=3 (accessed on 7 March 2013).
- Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically, 7th ed; Clinical and Laboratory Standards Institute (CLSI): Villanova, PA, USA, 2006.
- Performance Standards for Antimicrobial Susceptibility Testing, Seventh Informational Supplement; Clinical and Laboratory Standards Institute (CLSI): Wayne, PA, USA, 2007.
- Krumperman, P.H. Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Appl. Environ. Microbiol. 1983, 46, 165–170. [Google Scholar]
- The European Parliament and the Council of the European Union Directive. Available online: http://eur-lex.europa.eu/LexUriServ/LexUriServ.do?uri=OJ:L:2006:064:0037:0051:EN:PDF (accessed on 29 October 2012).
- Luczkiewicz, A.; Jankowska, K.; Kurlenda, J.; Olanczuk-Neyman, K. Identification and antimicrobial resistance of Enterococcus spp.isolated from surface water. Water Sci. Technol. 2010, 62, 466–473. [Google Scholar] [CrossRef]
- Thal, L.A.; Chow, J.W.; Mahayni, R.; Bonilla, H.; Perri, M.B.; Donabedian, S.A.; Silverman, J.; Taber, S.; Zervos, M.J. Characterization of antimicrobial resistance in enterococci of animal origin. Antimicrob. Agents Chemother. 1995, 39, 2112–2115. [Google Scholar] [CrossRef]
- Simjee, S.; White, D.G.; McDermott, P.F.; Wagner, D.D.; Zervos, M.J.; Donabedian, S.M.; English, L.L.; Hayes, J.R.; Walker, R.D. Characterization of Tn1546 in vancomycin-resistant Enterococcus faecium isolated from canine urinary tract infections: Evidence of gene exchange between human and animal enterococci. J. Clin. Microbiol. 2002, 40, 4659–4665. [Google Scholar] [CrossRef]
- Fedorka-Cray, P.J.; Englen, M.D.; Gray, J.T.; Hudson, C.; Headrick, M.L. Programs for monitoring antimicrobial resistance. Animal Biotech. 2002, 13, 43–55. [Google Scholar]
- Fluit, A.C.; van der Bruggen, J.T.; Aarestrup, F.M.; Verhoef, J.; Jansen, W.T. Priorities for antibiotic resistance surveillance in Europe. Clin. Microbiol. Infect. 2006, 12, 410–417. [Google Scholar] [CrossRef]
- Butaye, P.; Devriese, L.A.; Haesebrouck, F. Differences in antibiotic resistance patterns of Enterococcus faecalis and Enterococcus faecium strains isolated from farm and pet animals. Antimicrob. Agents Chemother. 2001, 45, 1374–1378. [Google Scholar] [CrossRef]
- Petsaris, O.; Miszczak, F.; Gicquel-Bruneau, M.; Perrin-Guyomard, A.; Humbert, F.; Sanders, P.; Leclercq, R. Combined antimicrobial resistance in Enterococcus faecium isolated from chickens. Appl. Environ. Microbiol. 2005, 71, 2796–2799. [Google Scholar]
- Butaye, P.; Baele, M.; Devriese, L.A.; Haesebrouck, F. Comparison of susceptibility to antimicrobials of the enterococcal species isolated from pigeons (Columba livia). Microb. Drug Resist. 2002, 8, 215–218. [Google Scholar] [CrossRef]
- Yoshimura, H.; Ishimaru, M.; Endoh, Y.S.; Kojima, A. Antimicrobial susceptibilities of enterococci isolated from faeces of broiler and layer chickens. Lett. Appl. Microbiol. 2000, 31, 427–432. [Google Scholar] [CrossRef]
- Ruzauskas, M.; Siugzdiniene, R.; Spakauskas, V.; Povilonis, J.; Seputiene, V.; Suziedeliene, E.; Daugelavicius, R.; Pavilonis, A. Susceptibility of bacteria of the Enterococcus genus isolated from Lithuanian poultry farms. Vet. Med. 2009, 54, 583–588. [Google Scholar]
- Da Costa, P.M.; Belo, A.; Goncalves, J.; Bernardo, F. Field trial evaluating changes in prevalence and patterns of antimicrobial resistance among Escherichia coli and Enterococcus spp.isolated from growing broilers medicated with enrofloxacin, apramycin and amoxicillin. Vet. Microbiol. 2009, 139, 284–292. [Google Scholar]
- Aslam, M.; Diarra, M.S.; Service, C.; Rempel, H. Characterization of antimicrobial resistance in Enterococcus spp. recovered from a commercial beef processing plant. Foodborne. Pathog. Dis. 2010, 7, 235–241. [Google Scholar] [CrossRef]
- Jacob, M.E.; Fox, J.T.; Narayanan, S.K.; Drouillard, J.S.; Renter, D.G.; Nagaraja, T.G. Effects of feeding wet corn distillers grains with solubles with or without monensin and tylosin on the prevalence and antimicrobial susceptibilities of fecal foodborne pathogenic and commensal bacteria in feedlot cattle. J. Anim. Sci. 2008, 86, 1182–1190. [Google Scholar]
- Diarra, M.S.; Rempel, H.; Champagne, J.; Masson, L.; Pritchard, J.; Topp, E. Distribution of antimicrobial resistance and virulence genes in Enterococcus spp. and characterization of isolates from broiler chickens. Appl. Environ. Microbiol. 2010, 76, 8033–8043. [Google Scholar] [CrossRef]
- Hayes, J.R.; English, L.L.; Carr, L.E.; Wagner, D.D.; Joseph, S.W. Multiple-antibiotic resistance of Enterococcus spp. isolated from commercial poultry production environments. Appl. Environ. Microbiol. 2004, 70, 6005–6011. [Google Scholar]
- Aarestrup, F.M.; Agerso, Y.; Gerner-Smidt, P.; Madsen, M.; Jensen, L.B. Comparison of antimicrobial resistance phenotypes and resistance genes in Enterococcus faecalis and Enterococcus faecium from humans in the community, broilers, and pigs in Denmark. Diagn. Microbiol. Infect. Dis. 2000, 37, 127–137. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Furtula, V.; Jackson, C.R.; Farrell, E.G.; Barrett, J.B.; Hiott, L.M.; Chambers, P.A. Antimicrobial Resistance in Enterococcus spp. Isolated from Environmental Samples in an Area of Intensive Poultry Production. Int. J. Environ. Res. Public Health 2013, 10, 1020-1036. https://doi.org/10.3390/ijerph10031020
Furtula V, Jackson CR, Farrell EG, Barrett JB, Hiott LM, Chambers PA. Antimicrobial Resistance in Enterococcus spp. Isolated from Environmental Samples in an Area of Intensive Poultry Production. International Journal of Environmental Research and Public Health. 2013; 10(3):1020-1036. https://doi.org/10.3390/ijerph10031020
Chicago/Turabian StyleFurtula, Vesna, Charlene R. Jackson, Erin Gwenn Farrell, John B. Barrett, Lari M. Hiott, and Patricia A. Chambers. 2013. "Antimicrobial Resistance in Enterococcus spp. Isolated from Environmental Samples in an Area of Intensive Poultry Production" International Journal of Environmental Research and Public Health 10, no. 3: 1020-1036. https://doi.org/10.3390/ijerph10031020





