Cytotoxicity of the Ascidian Cystodytes dellechiajei Against Tumor Cells and Study of the Involvement of Associated Microbiota in the Production of Cytotoxic Compounds
Abstract
:1. Introduction
2. Results and Discussion
2.1. In vitro antitumor activity from natural samples
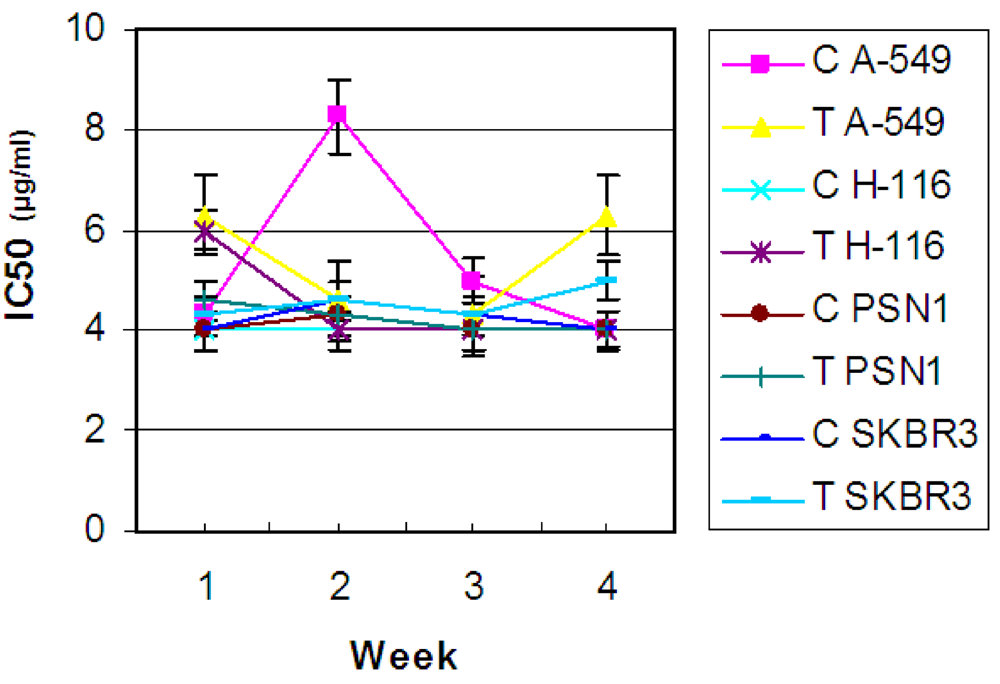
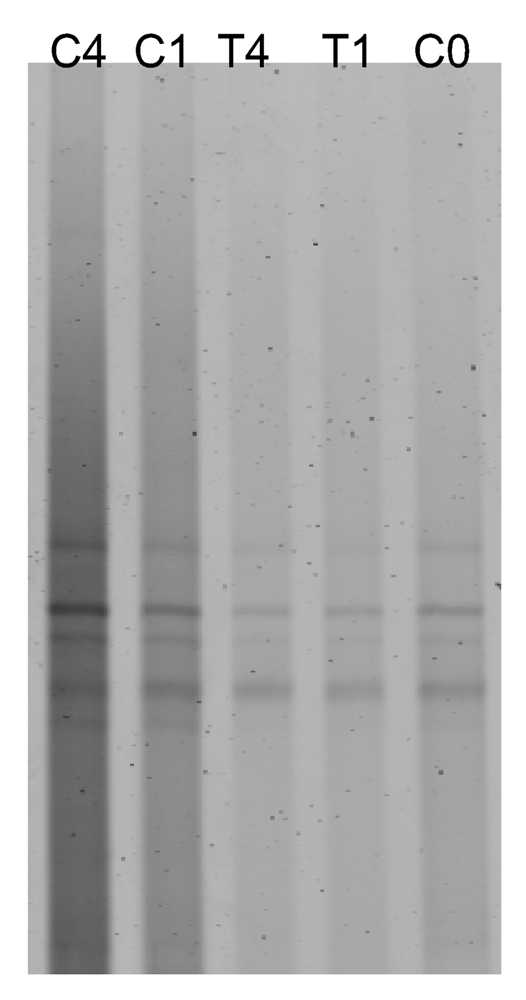
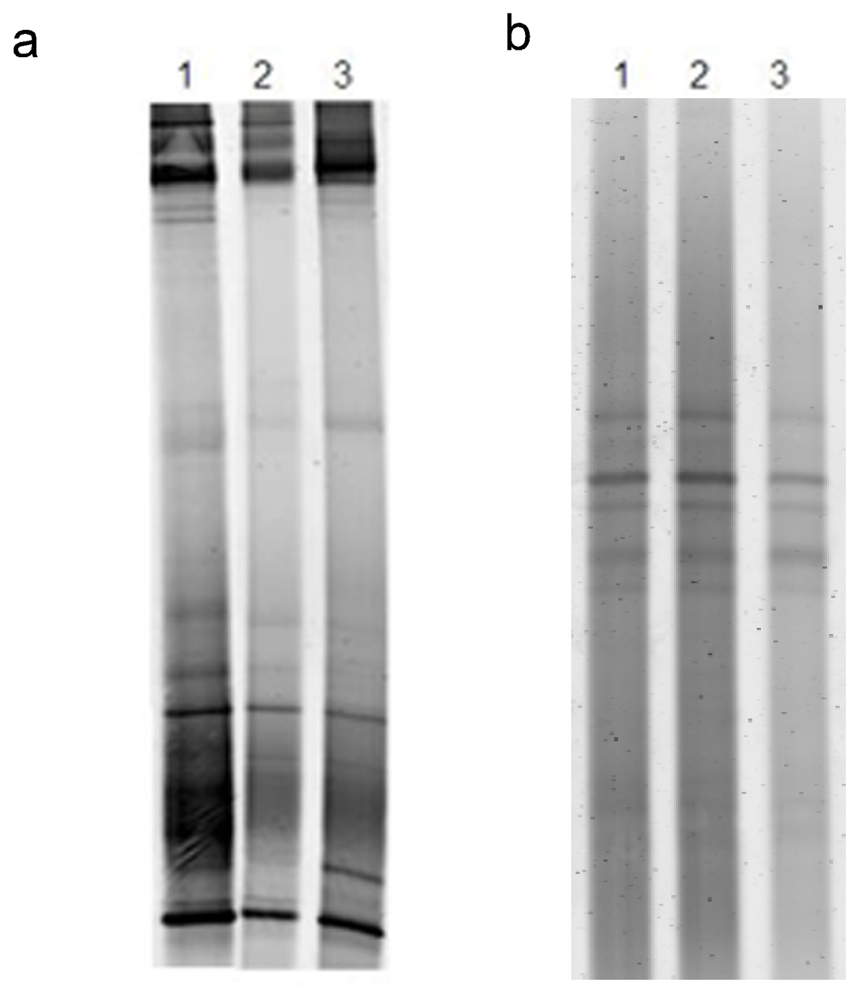
2.2. Antibiotic treatment of the C. dellechiajei colonies
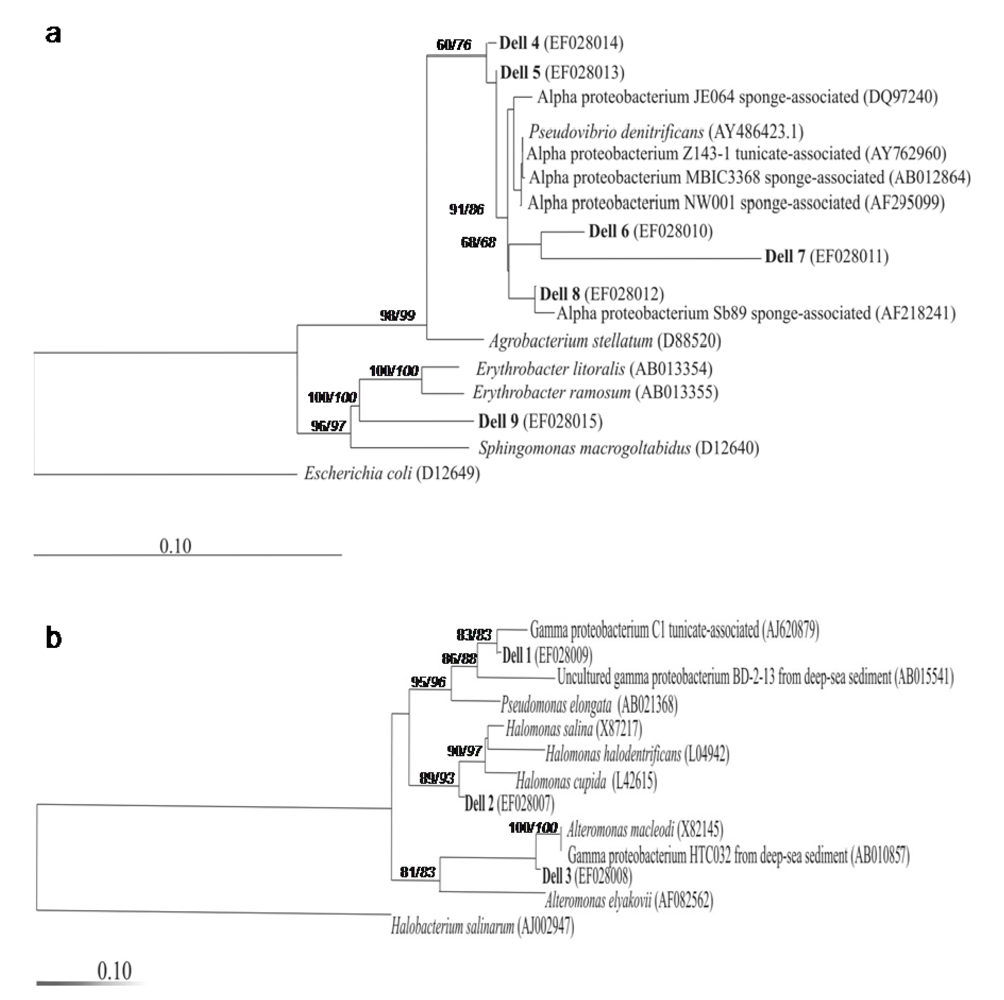
2.3. Culture of ascidia-associated bacteria and in vitro antitumor activity of the isolates
2.4. Is the microbial community involved in the synthesis of cytotoxic compounds?
3. Conclusions
4. Experimental Section
4.1. Sample collection
4.2. Maintenance of C. dellechiajei in aquarium and antibiotic treatment
4.3. DNA extraction, PCR and DGGE
4.4. DGGE data analysis
4.5. Bacterial isolation and 16S rRNA gene analysis
4.6. Phylogenetic analysis
4.7. Preparation of crude extracts from C. dellechiajei and microbial cultures for in vitro antitumor assays
4.8. Tumor cell cultures
4.9. In vitro antiproliferative assays
Acknowledgements
- Sample Availability: Available from the authors.
References and Notes
- Marinlit. Version september. A marine literature database produced and maintained by the Department of Chemistry, University of Canterbury, New Zealand, 2007; ( www.chem.canterbury.ac.nz/marinlit/marinlit.shtml)
- Faulkner, DJ. Marine natural products. Nat Prod Rep 2002, 19, 1–48. [Google Scholar]
- Proksch, P; Müller, WEG. Frontiers in Marine Biotechnology; Horizon Bioscience: Norfolk, U.K, 2006. [Google Scholar]
- Proksch, P; Edrada, RA; Ebel, R. Drugs from the sea-current status and microbiological implications. Appl Microbiol Biotechnol 2002, 59, 125–134. [Google Scholar]
- Harder, T; Lau, SCK; Dobretsov, S; Fang, TK; Qian, PY. A distinctive epibiotic bacterial community on the soft coral Dendronephtya sp. and antibacterial activity of coral tissue extracts suggest a chemical mechanism against bacterial epibiosis. FEMS Microbiol Ecol 2003, 43, 337–347. [Google Scholar]
- Harder, T; Lau, SCK; Tam, WY; Qian, PY. A bacterial culture-independient method to invesitigate chemically mediated control of bacterial epibiosis in marine invertebrates by TRFLP analysis and natural bacterial populations. FEMS Microbiol Ecol 2004, 47, 93–99. [Google Scholar]
- Mai-Pochnow, A; Evans, F; Dalisay-Saludes, D; Stelzer, S; Egan, S; James, S; Webb, JS; Kjelleberg, S. Biofilm development and cell death in the marine bacterium Pseudoalteromonas tunicata. Appl Environ Microbiol 2004, 70, 3232–3238. [Google Scholar]
- Kim, YM; Yasuda, M; Yamagishi, A; Oshima, A; Ohta, S. Characterization of the endo-symbiont of a deep-sea bivalve, Calyptogena soyoae. Appl Environ Microbiol 1995, 61, 823–827. [Google Scholar]
- Preston, CM; Wu, KY; Molinski, TF; DeLong, EF. A psycrophylic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov. Proc Natl Acad Sci USA 1996, 93, 6241–6246. [Google Scholar]
- Haygood, MG; Davidson, SK. Small subunit ribosomal RNA genes and in situ hybridation of bacterial symbionts in the larvae of the bryozoan Bugula neritina and proposal of “Candidatus Endobugula sertula”. Appl Env Microbiol 1997, 63, 4612–4616. [Google Scholar]
- Sipe, AR; Wilbur, AE; Cary, SC. Bacterial symbiont transmission in the wood-boring shipworm Bankia setacea (Bivalvia: Teredinidae). Appl Environ Microbiol 2000, 66, 1685–1691. [Google Scholar]
- Dubilier, N; Mulders, C; Ferdelman, T; de Beer, D; Pernthaler, A; Klein, M; Wagner, M; Erseus, C; Thiermann, F; Krieger, J; Giere, O; Amann, R. Endosymbiotic sulphate-reducing and sulphide-oxidizing bacteria in an oligochaete worm. Nature 2001, 411, 298–302. [Google Scholar]
- Kühl, M; Larkum, AW. Seckbach, J, Ed.; The microenvironment and photosynthetic performance of Prochloron sp. in symbiosis with Didemnid ascidians. In Symbiosis: Mechanisms and Models; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2002; pp. 273–290. [Google Scholar]
- Moss, C; Green, DH; Pérez, B; Velasco, A; Henríquez, R; McKenzie, JD. Intracellular bacteria associated with the ascidian Ecteinascidia turbinata: phylogenetic and in situ hybridisation analysis. Mar Biol 2003, 143, 99–110. [Google Scholar]
- Munn, CB. Watts, A, Ed.; Symbiotic associations. In Marine microbiology; Cromwell Press: Trowbridge, UK, 2004; pp. 167–181. [Google Scholar]
- Long, PF; Dunlap, WC; Batershill, CN; Jaspars, M. Shotgun cloning and heterologous expression of the patellamide gene clusters as strategy to achieving sustained metabolite production. Chembiochem 2005, 6, 1760–1765. [Google Scholar]
- Schmidt, EW; Nelson, JT; Rasko, DA; Sudek, S; Eisen, JA; Haygood, MG; Ravel, J. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc Natl Acad Sci USA 2005, 102, 7315–7320. [Google Scholar]
- Davidson, SK; Allen, SW; Lim, GE; Anderson, CM; Haygood, MG. Evidence for the biosynthesis of bryostatins by the bacterial symbiont "Candidatus Endobugula sertula" of the bryozoan Bugula neritina. Appl Environ Microbiol 2001, 67, 4531–4537. [Google Scholar]
- Hale, KJ; Hummersone, MG; Manaviazar, S; Frigerio, M. The chemistry and biology of the bryostatin antitumour macrolides. Nat Prod Rep 2002, 19, 413–453. [Google Scholar]
- Rottmayr, EM; Steffan, B; Wanner, G. Pigmentation and tunic cells in Cystodytes dellechiajei (Urochordata, Ascidiacea). Zoomorphology 2001, 120, 159–170. [Google Scholar]
- Turon, X; López-Legentil, S; Banaigs, B. Cell types, microsymbionts, and pyridoacridine distribution in the tunic of three color morphs of the genus Cystodytes (Ascidiacea, Polycitoridae). Invertebr Biol 2005, 124, 355–369. [Google Scholar]
- Steffan, B; Brix, K; Pütz, W. Biosynthesis of Shermilamine B. Tetrahedron Let 1993, 49, 6223–6228. [Google Scholar]
- Shimbo, K; Tsuda, M; Fukushi, E; Kawabata, J; Kobayashi, J. Dytesinins A and B, new clerodane-type diterpenes with a cyclopropane ring from the tunicate Cystodytes sp. Tethraedron 2000, 56, 7923–7926. [Google Scholar]
- Loukaci, A; Bultel-Poncé, V; Longeon, A; Guyot, M. New lipids from the tunicate Cystodytes cf. dellechiajei:as PLA2 inhibitors. J Nat Prod 2000, 63, 799–802. [Google Scholar]
- McDonald, LA; Eldredge, GS; Barrows, LR; Ireland, CM. Inhibition of topoisomerase II catalytic activity by pyridoacridine alkaloids from a Cystodytes sp. ascidian: A mechanism for the apparent intercalator-induced inhibition of topoisomerase II. J Med Chem 1994, 37, 3819–3827. [Google Scholar]
- Ciufolini, MA; Shen, YC; Bishop, MJ. A unified strategy for the synthesis of sulfur containing pyridoacridine alkaloids: antitumor agents of marine origin. J Am Chem Soc 1995, 117, 12460–12469. [Google Scholar]
- Dassonneville, L; Wattez, N; Baldeyrou, B; Mahieu, C; Lansiaux, A; Banaigs, B; Bonnard, I; Bailly, C. Inhibition of topoisomerase II by the marine alkaloid ascididemin and induction of apoptosis in leukemia cells. Biochem Pharmacol 2000, 60, 527–537. [Google Scholar]
- Torres, YR; Bugni, TS; Berlinck, RG; Ireland, CM; Magalhaes, A; Ferreira, AG; Rocha, RM. Sebastianines A and B, novel biologically active pyridoacridine alkaloids from the Brazilian ascidian Cystodytes dellechiajei. J Org Chem 2002, 67, 5429–5432. [Google Scholar]
- Prado, MP; Torres, YR; Berlinck, RGS; Desiderá, C; Sanchez, MA; Craveiro, MV; Hadju, H; da Rocha, RM; Machado-Santelli, GM. Effects of marine organisms extracts on microtubule integrity and cell cycle progression in cultured cells. J Exp Mar Biol Ecol 2004, 313, 125–137. [Google Scholar]
- Becerro, MA; Turon, X; Uriz, MJ. Multiple functions for secondary metabolites in encrusting marine invertebrates. J Chem Ecol 1997, 23, 1527–1547. [Google Scholar]
- Tarjuelo, I; López-Legentil, S; Codina, M; Turon, X. Defence mechanisms of adults and larvae of colonial ascidians: patterns of palatability and toxicity. Mar Ecol Prog Ser 2002, 235, 103–115. [Google Scholar]
- López-Legentil, S; Dieckmann, R; Bontemps-Subielos, N; Turon, X; Banaigs, B. Qualitative variation of alkaloids in colour morphs of Cystodytes (Ascidiacea). Bioch Syst Ecol 2005, 33, 1107–1119. [Google Scholar]
- Martínez-García, M; Diaz-Valdés, M; Wanner, G; Ramos-Esplá, A; Antón, J. Microbial community associated with the colonial ascidian Cystodytes dellechiajei. Environ Microbiol 2007, 9, 521–534. [Google Scholar]
- Schäfer, H; Bernard, L; Courtis, C; Lebaron, P; Servais, P; Pukall, P; Stackebrandt, E; Troussellier, M; Guindulain, T; Vives-Rego, J; Muyzer, G. Microbial community dynamics in Mediterranean nutrient-enriched seawater mesocosms: changes in the genetic diversity of bacterial populations. FEMS Microbiol Ecol 2001, 34, 243–253. [Google Scholar]
- Bonnard, I; Bontemps, N; Lahmy, S; Bonaigs, B; Combaut, G; Francisco, C; Colson, P; Houssie, C; Waring, MJ; Bailly, C. Binding to DNA and cytotoxic evaluation of ascididemin, the major alkaloid from the Mediterranean ascidian Cystodytes dellechiajei. Anti Cancer Drug Des 1995, 10, 333–346. [Google Scholar]
- Lindsay, B; Barrows, L; Copp, B. Structural requeriments for biological activity of the marine alkaloid ascididemin. Bioorg Med Chem Lett 1995, 5, 739–742. [Google Scholar]
- Lindquist, N; Hay, ME; Fenical, W. Defense of ascidians and their conspicuous larvae: adult vs. larval chemical defenses. Ecol Monogr 1992, 62, 547–568. [Google Scholar]
- Lindquist, N. Palatability of invertebrate larvae to corals and sea anemones. Mar Biol 1996, 126, 745–755. [Google Scholar]
- Lindquist, N; Hay, ME. Palatibility and chemical defense of marine invertebrate larvae. Ecol Monogr 1996, 66, 431–450. [Google Scholar]
- Hay, ME. Marine chemical ecology: what’s known and what’s next? J Exp Mar Biol Ecol 1996, 200, 103–134. [Google Scholar]
- Uriz, MJ; Turon, X; Becerro, MA; Galera, J. Feeding deterrence in sponges. The role of toxicity, physical defenses, energetic contents, and life-history stage. J Exp Mar Biol Ecol 1996, 205, 187–204. [Google Scholar]
- Roughgarden, J. Feldman, MW, Ed.; The evolution of marine life cycles. In Mathematical evolutionary theory; Princeton University Press: Princeton, NJ, 1989; pp. 270–300. [Google Scholar]
- Kobayashi, J; Cheng, J; Wälchli, MR; Nakamura, H; Hirata, Y; Sasaki, T; Ohizumi, Y. Cystodytins A, B, and C, novel tetracyclic aromatic alkaloids with potents antineoplasic activity from the okinawan tunicate Cystodytes dellechiajei. J Org Chem 1988, 53, 1800–1804. [Google Scholar]
- Carroll, AR; Cooray, NM; Pioner, A; Scheuer, PJ. A second shermilamine alkaloid from a tunicate Trididemnum spec. J Org Chem 1989, 54, 4231–4232. [Google Scholar]
- Kobayashi, J; Tsuda, M; Tanabe, A; Ishibashi, M. Cystodytins D-I, new cytotoxic tetracyclic aromatic alkaloids from the okinawan marine tunicate Cystodytes dellechiajei. J Nat Prod 1991, 54, 1634–1638. [Google Scholar]
- López-Legentil, S; Bontemps-Subielos, N; Turon, X; Banaigs, B. Temporal variation in the production of fours secondary metabolites in a colonial ascidian. J Chem Ecol 2006, 32, 2079–2084. [Google Scholar]
- Webster, NS; Hill, RT. The culturable microbial community of the Great Barrier Reef sponge Rhopaloeides odorabile is dominated by an α-Proteobacterium. Mar Biol 2001, 138, 843–851. [Google Scholar]
- Hentschel, U; Schmid, M; Wagner, M; Fieseler, L; Gernert, C; Hacker, J. Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the Mediterranean sponges Aplysina aerophoba and Aplysina cavernicola. FEMS Microbiol Ecol 2001, 35, 305–312. [Google Scholar]
- Hamada, T. Use of gyrB gene analysis to investigate phylogeny of marine bacteria. Unpublished. 1998. [Google Scholar]
- Enticknap, JJ; Kelly, M; Peraud, O; Hill, RT. Characterization of a culturable alpha-proteobacterial symbiont common to many marine sponges and evidence for vertical transmission through the gametes. Unpublished. 2005. [Google Scholar]
- Guzman, AS; Predicala, RZ; Bernardo, EB; Neilan, B; Elardo, SP; Mangalindan, GC; Tasdemir, D; Ireland, CM; Barraquio, WL; Concepcion, GP. Marine alpha-proteobacterium from a Philippine tunicate and its anti-Staphylococcus aureus metabolite heptylprodigiosin. Unpublished. 2004. [Google Scholar]
- Koblízek, M; Béjà, O; Bidigare, RR; Christensen, S; Benitez-Nelson, B; Vetriani, C; Kolber, MK; Falkowski, PG; Kolber, ZS. Isolation and characterization of Erythrobacter sp. strains from the upper ocean. Arch Microbiol 2003, 180, 327–338. [Google Scholar]
- Rottmayr, EM. Licht- und elektronenmikroskopische Untersuchungen der Tunika von Cystodytes dellechiajei Della Valle (Urochordata, Ascidiacea). In PhD thesis; Fakultät für Biologie der Ludwig-Maximilians-Universität: München, Germany, 2001. [Google Scholar]
- Lehmann, CM. Characterization of bacterium isolates C1 from Cystodytes dellechiajei. Unpublished. 2004. [Google Scholar]
- Debard, H; Banaigs, B; Francisco, F; Commeyras, A. Use of ascididemin and derivates as antifouling agents. PCT Int Appl WO 98/21959, 1998. [Google Scholar]
- In, Y; Doi, M; Inoue, M; Ishida, T; Hamada, Y; Shioiri, T. Patellamide A, a cytotoxic cyclic peptide from the ascidian Lissoclinum patella. Acta Crystalogr C 1994, 15, 432–434. [Google Scholar]
- Fu, X; Do, T; Schmitz, FJ; Andrusevich, V; Engel, MH. New cyclic peptides from the ascidian Lissoclinum patella. J Nat Prod 1998, 61, 1547–1551. [Google Scholar]
- Salomon, CE; Faulkner, DJ. Localization studies of bioactive cyclic peptides in the ascidian Lissoclinum patella. J Nat Prod 2002, 65, 689–692. [Google Scholar]
- Clardy, J. Using genomics to deliver natural products from symbiotic bacteria. Gen Biol 2005, 6, 232. [Google Scholar]
- Mojica, FJM. Cienfuegos, GA, Ruiz-Bravo, A, Pablo, MA, Eds.; Antiarchaeal antibiotics. In New approaches in the use of antibiotics; Research Singpost: Kerala, India, 2003; pp. 83–108. [Google Scholar]
- Sambrook, E; Fritsch, F; Maniatis, T. Molecular cloning A laboratory manual; Cold Spring Harbor Press: Cold Spring Harbor, NY, 1989. [Google Scholar]
- Muyzer, G; de Waal, EC; Uitterrlinden, AG. Profiling in complex microbial population by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Micriobiol 1993, 59, 695–700. [Google Scholar]
- Weisburg, WE; Barns, SM; Pelletier, DA; Lane, DJ. 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 1991, 173, 697–703. [Google Scholar]
- Vaneechoutte, M; Rossau, R; Vos, PD; Gillis, M; Janssens, D; Paepe, N; De Rouck; Fiers, T; Claeys, G; Kersters, K. Rapid identification of bacteria of the Comamonadaceae with amplified ribosomal DNA-restriction analysis (ARDRA). FEMS Microbiol Lett 1992, 93, 227–234. [Google Scholar]
- Amann, R; Ludwig, W; Schleifer, KH. Phylogenetic identification of individual microbial cells without cultivation. Microbiol Rev 1995, 59, 143–169. [Google Scholar]
- Altschul, SF; Madden, TL; Schäffer, AA; Zhang, J; Zhang, Z; Miller, W; Lipman, DJ. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res 1997, 25, 3389–3402. [Google Scholar]
- Strimmer, K; von Haeseler, A. Quartet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol Biol Evol 1996, 13, 964–969. [Google Scholar]
- Strimmer, K; von Haeseler, A. Likelihood-mapping: a simple method to visualize phylogenetic content of a sequence alignement. Proc Natl Acad Sci USA 1997, 94, 6815–6819. [Google Scholar]
- Moore, GE; Woods, LK. Culture media for human cells: RPMI 1630, RPMI 1634, RPMI 1640 and GEM 1717. T C A Man 1976, 3, 503–508. [Google Scholar]
- Mosmann, T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods 1983, 65, 55–63. [Google Scholar]

| IC50 for cancer cell line (μg/ml) | |||||
|---|---|---|---|---|---|
| Samples | aA-549 | bH-116 | cPSN-1 | dSKBR3 | eT98G |
| Blue colony 1 | 5 | <5 | 5 | 10 | 25 |
| Blue colony 2 | 5 | <5 | 5 | 5 | 10 |
| Blue colony 3 | 5 | <5 | 5 | 5 | 25 |
| Blue colony 4 | 5 | <5 | <5 | 5 | 10 |
| Blue colony 5 | 5 | <5 | <5 | 5 | 25 |
| Blue colony 6 | 5 | <5 | 5 | 5 | 25 |
| Blue colony 7 | 5 | <5 | 5 | 5 | 25 |
| Blue colony 8 | 5 | <5 | <5 | 5 | 25 |
| Blue colony 9 | 5 | <5 | 5 | 5 | 25 |
| Blue colony 10 | 5 | <5 | 5 | 5 | 25 |
| Blue colony 11 | <2.5 | 2.5 | 5 | 5 | 10 |
| Green colony 1 | 25 | 5 | - | - | - |
| Green colony 2 | 25 | 25 | - | - | - |
| Green colony 3 | 25 | 25 | - | - | - |
| Blue colony tunic | 2.5 | 2.5 | - | - | - |
| Blue colony zooids | 12.5 | 12.5 | - | - | - |
| Larva | 12.5 | 25 | - | - | - |
| DGGE bands | Accession no. (GenBank) | Closest taxon (% similarity BLASTn) | Division |
|---|---|---|---|
| 1 | EF028016 | 98% Janthinobacterium agaricidamnosus (AY167838) | Beta-proteobacteria |
| 2 | EF028017 | 99% Uncultured alpha-proteobacterium NJ1-1-1 (AY626827) | Alpha-proteobacteria |
| 3 | EF028018 | 97% Uncultured alpha-proteobacterium NJ1-1-1 (AY626827) | Alpha-proteobacteria |
| 4 | EF028019 | 98% Mesorhizobium sp. (AY690680) | Alpha-proteobacteria |
| 5 | EF028020 | 99% Mesorhizobium sp. (AY690680) | Alpha-proteobacteria |
| 6 | EF028021 | 94% Stappia sp. (AY307927) | Alpha-proteobacteria |
| 7 | EF028022 | 94% Stappia sp. (AY307927) | Alpha-proteobacteria |
| Strain | aMorphotype | No. of isolates | bCulture media | Closest taxon (% similarity BLASTn) |
|---|---|---|---|---|
| Dell4 | 1 | 7 | 2,3 | 99.3% Alpha-proteobacterium NW001 (AF295099) |
| Dell5 | 1 | 6 | 2,3 | 99.3% Alpha-proteobacterium NW001 (AF295099) |
| Dell6 | 1 | 4 | 2,3 | 96.1% Alpha-proteobacterium NW001 (AF295099) |
| Dell7 | 1 | 5 | 2,3 | 91% Alpha-proteobacterium NW001 (AF295099) |
| Dell8 | 1 | 1 | 2,3 | 98.2% Alpha-proteobacterium Sb89 (AF218241) |
| Dell9 | 2 | 1 | 1 | 93.5% Erythrobacter luteolus (AY5739662) |
| Dell1 | 3 | 1 | 2 | 97% Gamma-proteobacterium C1 (AJ620879) |
| Dell2 | 4 | 1 | 3 | 97.6% Halomonas salina (X87217) |
| Dell3 | 5 | 1 | 3 | 98.9% Alteromonas macleodi (X82145) |
Share and Cite
Martínez-García, M.; Díaz-Valdés, M.; Ramos-Esplá, A.; Salvador, N.; Lopez, P.; Larriba, E.; Antón, J. Cytotoxicity of the Ascidian Cystodytes dellechiajei Against Tumor Cells and Study of the Involvement of Associated Microbiota in the Production of Cytotoxic Compounds. Mar. Drugs 2007, 5, 52-70. https://doi.org/10.3390/md503052
Martínez-García M, Díaz-Valdés M, Ramos-Esplá A, Salvador N, Lopez P, Larriba E, Antón J. Cytotoxicity of the Ascidian Cystodytes dellechiajei Against Tumor Cells and Study of the Involvement of Associated Microbiota in the Production of Cytotoxic Compounds. Marine Drugs. 2007; 5(3):52-70. https://doi.org/10.3390/md503052
Chicago/Turabian StyleMartínez-García, Manuel, Marta Díaz-Valdés, Alfonso Ramos-Esplá, Nélida Salvador, Patricia Lopez, Eduardo Larriba, and Josefa Antón. 2007. "Cytotoxicity of the Ascidian Cystodytes dellechiajei Against Tumor Cells and Study of the Involvement of Associated Microbiota in the Production of Cytotoxic Compounds" Marine Drugs 5, no. 3: 52-70. https://doi.org/10.3390/md503052





