Production of the Marine Carotenoid Astaxanthin by Metabolically Engineered Corynebacterium glutamicum
Abstract
:1. Introduction
2. Results
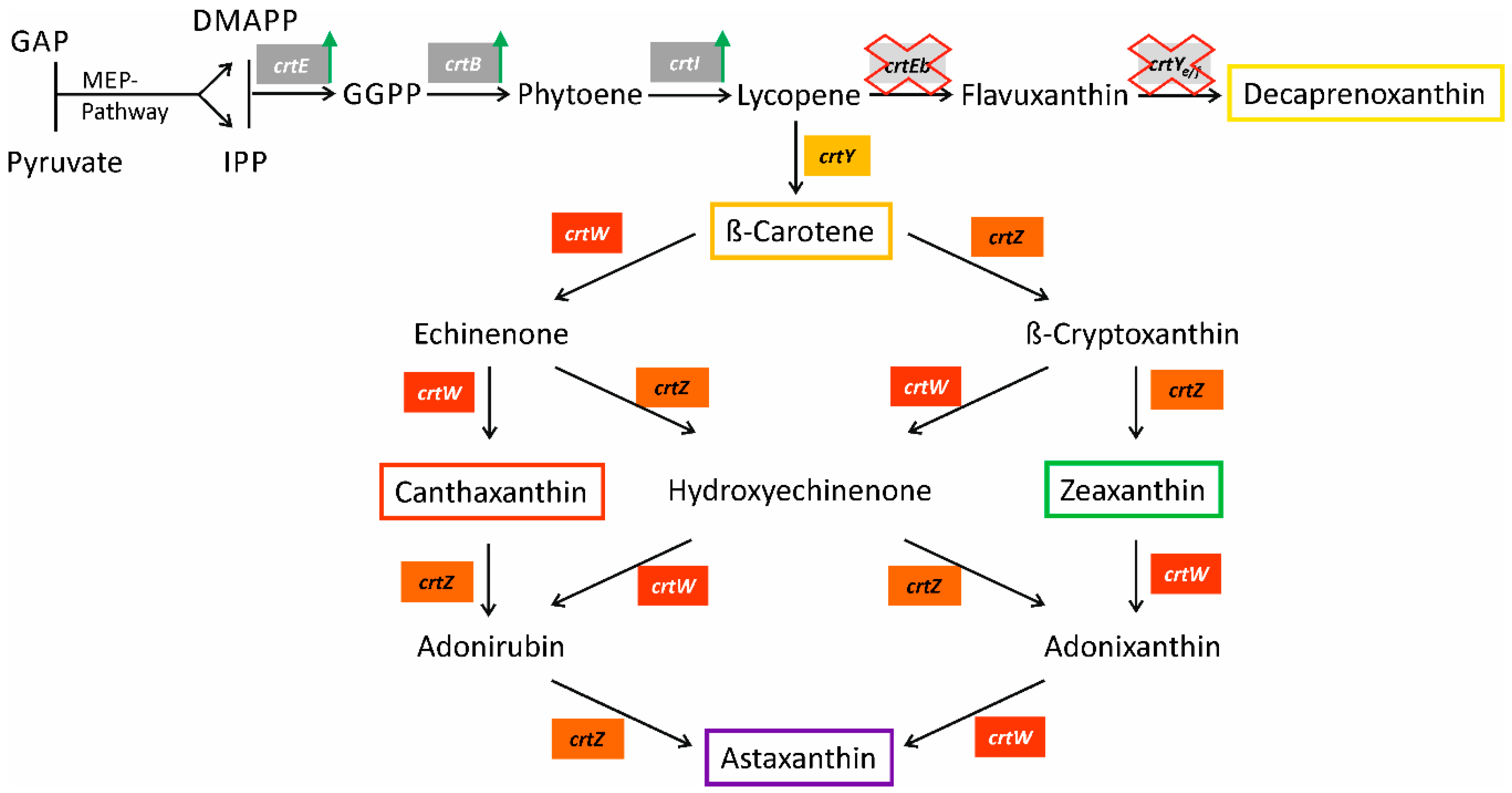
2.1. Construction of a β-Carotene Producing C. glutamicum Base Strain
2.2. Design of the Combinatorial Gene Assembly and Library Construction for Engineering Astaxanthin Production in C. glutamicum
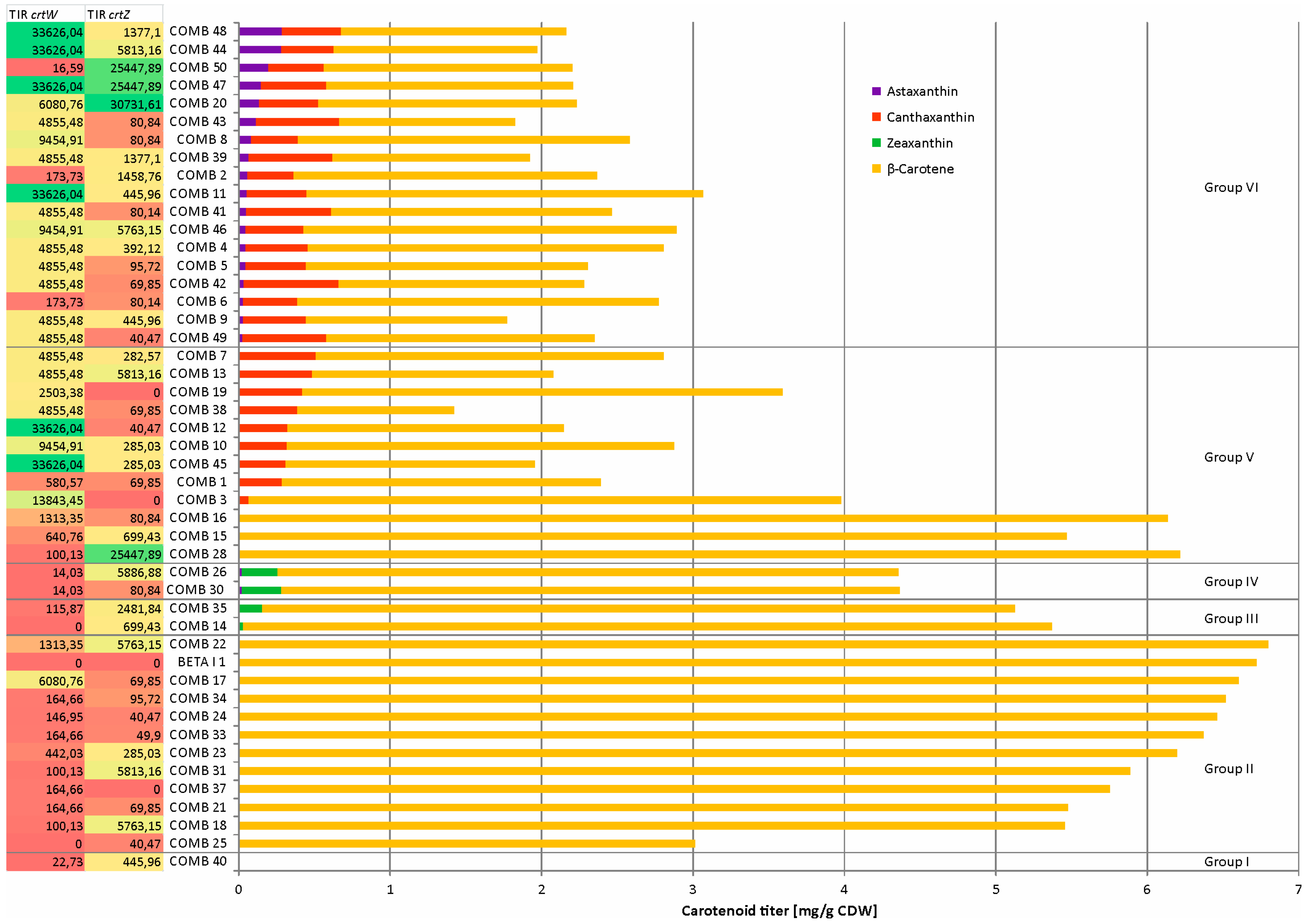
2.3. Combinatorial Engineering Covered Vastly Different Astaxanthin, β-Carotene, Zeaxanthin and Canthaxanthin Titers
2.4. In Silico Analysis of the Carotenogenic Genes crtZ and crtW from Marine and Non-Marine Bacteria for Heterologous Expression in C. glutamicum
2.5. High Astaxanthin Production by C. glutamicum Strains Expressing crtW and crtZ from F. pelagi
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Media and Growth Conditions
4.2. Recombinant DNA Work
4.3. Construction of Expression Vector pSH1
4.4. Deletion and Exchenge Mutagenesis in the Genome of C. glutamicum
4.5. Combinatorial Gene Assembly, Library Construction and Overexpression of Carotenogenic Genes
4.6. Extraction and Quantification of Carotenoids
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| TIR | Translation Initiation Rate |
| MDPI | Multidisciplinary Digital Publishing Institute |
| DOAJ | Directory of open access journals |
| TLA | Three letter acronym |
| LD | Linear dichroism |
References
- Cooper, D.A.; Eldridge, A.L.; Peters, J.C. Dietary carotenoids and certain cancers, heart disease, and age-related macular degeneration: A review of recent research. Nutr. Rev. 1999, 57, 201–214. [Google Scholar] [CrossRef] [PubMed]
- Krinsky, N.I.; Johnson, E.J. Carotenoid actions and their relation to health and disease. Mol. Asp. Med. 2005, 26, 459–516. [Google Scholar] [CrossRef] [PubMed]
- Ye, V.M.; Bhatia, S.K. Pathway engineering strategies for production of beneficial carotenoids in microbial hosts. Biotechnol. Lett. 2012, 34, 1405–1414. [Google Scholar] [CrossRef] [PubMed]
- BBC Research. The Global Market for Carotenoids. Available online: http://www.bccresearch.com/market-research/food-and-beverage/carotenoids-global-market-fod025d.html (accessed on 13 June 2016).
- Belviranli, M.; Okudan, N. Well-Known Antioxidants and Newcomers in Sport Nutrition: Coenzyme Q10, Quercetin, Resveratrol, Pterostilbene, Pycnogenol and Astaxanthin. In Antioxidants in Sport Nutrition; Lamprecht, M., Ed.; Taylor & Francis Group: Boca Raton, FL, USA, 2015. [Google Scholar]
- Gassel, S.; Schewe, H.; Schmidt, I.; Schrader, J.; Sandmann, G. Multiple improvement of astaxanthin biosynthesis in Xanthophyllomyces dendrorhous by a combination of conventional mutagenesis and metabolic pathway engineering. Biotechnol. Lett. 2013, 35, 565–569. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Osawa, T. Cis astaxanthin and especially 9-cis astaxanthin exhibits a higher antioxidant activity in vitro compared to the all-trans isomer. Biochem. Biophys. Res. Commun. 2007, 357, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Hussein, G.; Nakamura, M.; Zhao, Q.; Iguchi, T.; Goto, H.; Sankawa, U.; Watanabe, H. Antihypertensive and neuroprotective effects of astaxanthin in experimental animals. Biol. Pharm. Bull. 2005, 28, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Giuffrida, D.; Sutthiwong, N.; Dugo, P.; Donato, P.; Cacciola, F.; Girard-Valenciennes, E.; le Mao, Y.; Monnet, C.; Fouillaud, M.; Caro, Y.; et al. Characterisation of the C50 carotenoids produced by strains of the cheese-ripening bacterium Arthrobacter arilaitensis. Int. Dairy J. 2016, 55, 10–16. [Google Scholar] [CrossRef]
- Chen, C.W.; Hsu, S.H.; Lin, M.T.; Hsu, Y.H. Mass production of C50 carotenoids by Haloferax mediterranei in using extruded rice bran and starch under optimal conductivity of brined medium. Bioprocess Biosyst. Eng. 2015, 38, 2361–2367. [Google Scholar] [CrossRef] [PubMed]
- Furubayashi, M.; Ikezumi, M.; Takaichi, S.; Maoka, T.; Hemmi, H.; Ogawa, T.; Saito, K.; Tobias, A.V.; Umeno, D. A highly selective biosynthetic pathway to non-natural C50 carotenoids assembled from moderately selective enzymes. Nat. Commun. 2015, 6, 7534. [Google Scholar] [CrossRef] [PubMed]
- Umeno, D.; Arnold, F.H. A C35 carotenoid biosynthetic pathway. Appl. Environ. Microbiol. 2003, 69, 3573–3579. [Google Scholar] [CrossRef] [PubMed]
- Takaichi, S.; Sandmann, G.; Schnurr, G.; Satomi, Y.; Suzuki, A.; Misawa, N. The carotenoid 7,8-dihydro-y end group can be cyclized by the lycopene cyclases from the bacterium Erwinia uredovora and the higher plant Capsicum annuum. Eur. J. Biochem. 1996, 241, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Fassett, R.G.; Coombes, J.S. Astaxanthin in cardiovascular health and disease. Molecules 2012, 17, 2030–2048. [Google Scholar] [CrossRef] [PubMed]
- Ohgami, K.; Shiratori, K.; Kotake, S.; Nishida, T.; Mizuki, N.; Yazawa, K.; Ohno, S. Effects of astaxanthin on lipopolysaccharide-induced inflammation in vitro and in vivo. Investig. Ophthalmol. Vis. Sci. 2003, 44, 2694–2701. [Google Scholar] [CrossRef]
- Baralic, I.; Andjelkovic, M.; Djordjevic, B.; Dikic, N.; Radivojevic, N.; Suzin-Zivkovic, V.; Radojevic-Skodric, S.; Pejic, S. Effect of Astaxanthin Supplementation on Salivary IgA, Oxidative Stress, and Inflammation in Young Soccer Players. Evid. Based. Complement. Altern. Med. 2015, 2015, 783761. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhu, D.; Niu, J.; Shen, S.; Wang, G. An economic assessment of astaxanthin production by large scale cultivation of Haematococcus pluvialis. Biotechnol. Adv. 2011, 29, 568–574. [Google Scholar] [CrossRef] [PubMed]
- Zhu, F.; Zhong, X.; Hu, M.; Lu, L.; Deng, Z.; Liu, T. In vitro Reconstitution of Mevalonate Pathway and targeted engineering of farnesene overproduction in Escherichia coli. Biotechnol. Bioeng. 2014, 111, 1396–1405. [Google Scholar] [CrossRef] [PubMed]
- Capelli, B.; Bagchi, D.; Cysewski, G. Synthetic astaxanthin is significantly inferior to algal-based astaxanthin as an antioxidant and may not be suitable as a human nutraceutical supplement. Nutrafoods 2013, 12, 145–152. [Google Scholar] [CrossRef]
- Lee, P.C.; Schmidt-Dannert, C. Metabolic engineering towards biotechnological production of carotenoids in microorganisms. Appl. Microbiol. Biotechnol. 2002, 60, 1–11. [Google Scholar] [PubMed]
- George, B.; Synnove, L.J.; Hanspeter, P. Carotenoids Handbook; Birkhauser Verlag: Basel, Switzerland, 2004. [Google Scholar]
- Cutzu, R.; Coi, A.; Rosso, F.; Bardi, L.; Ciani, M.; Budroni, M.; Zara, G.; Zara, S.; Mannazzu, I. From crude glycerol to carotenoids by using a Rhodotorula glutinis mutant. World J. Microbiol. Biotechnol. 2013, 29, 1009–1017. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, S.; Udaka, S.; Shimono, M. Studies on the amino acid fermentation. Production of L-glutamic acid by various microorganisms. J. Gen. Appl. Microbiol. 1957, 3, 193–205. [Google Scholar] [CrossRef]
- Zahoor, A.; Otten, A.; Wendisch, V.F. Metabolic engineering of Corynebacterium glutamicum for glycolate production. J. Biotechnol. 2014, 192, 366–375. [Google Scholar] [CrossRef] [PubMed]
- Schneider, J.; Wendisch, V.F. Putrescine production by engineered Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2010, 88, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Blombach, B.; Eikmanns, B.J. Current knowledge on isobutanol production with Escherichia coli, Bacillus subtilis and Corynebacterium glutamicum. Bioeng. Bugs 2011, 2, 346–350. [Google Scholar] [CrossRef] [PubMed]
- Frohwitter, J.; Heider, S.A.; Peters-Wendisch, P.; Beekwilder, J.; Wendisch, V.F. Production of the sesquiterpene (+)-valencene by metabolically engineered Corynebacterium glutamicum. J. Biotechnol. 2014, 191, 205–213. [Google Scholar] [CrossRef] [PubMed]
- Heider, S.A.; Peters-Wendisch, P.; Wendisch, V.F.; Beekwilder, J.; Brautaset, T. Metabolic engineering for the microbial production of carotenoids and related products with a focus on the rare C50 carotenoids. Appl. Microbiol. Biotechnol. 2014, 98, 4355–4368. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, S.; Tanaka, K. Glutamic acid. In The Microbial Production of Amino Acids; Yamada, K., Kinoshita, S., Tsunoda, T., Aida, K., Eds.; Halsted Press: New York, NY, USA, 1972; pp. 263–324. [Google Scholar]
- Blombach, B.; Seibold, G.M. Carbohydrate metabolism in Corynebacterium glutamicum and applications for the metabolic engineering of L-lysine production strains. Appl. Microbiol. Biotechnol. 2010, 86, 1313–1322. [Google Scholar] [CrossRef] [PubMed]
- Meiswinkel, T.M.; Rittmann, D.; Lindner, S.N.; Wendisch, V.F. Crude glycerol-based production of amino acids and putrescine by Corynebacterium glutamicum. Bioresour. Technol. 2013, 145, 254–258. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, V.; Meiswinkel, T.M.; Wendisch, V.F.; Nampoothiri, K.M. Amino acid production from rice straw and wheat bran hydrolysates by recombinant pentose-utilizing Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2011, 92, 985–996. [Google Scholar] [CrossRef] [PubMed]
- Uhde, A.; Youn, J.W.; Maeda, T.; Clermont, L.; Matano, C.; Kramer, R.; Wendisch, V.F.; Seibold, G.M.; Marin, K. Glucosamine as carbon source for amino acid-producing Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2013, 97, 1679–1687. [Google Scholar] [CrossRef] [PubMed]
- Matano, C.; Uhde, A.; Youn, J.W.; Maeda, T.; Clermont, L.; Marin, K.; Kramer, R.; Wendisch, V.F.; Seibold, G.M. Engineering of Corynebacterium glutamicum for growth and L-lysine and lycopene production from N-acetyl-glucosamine. Appl. Microbiol. Biotechnol. 2014, 98, 5633–5643. [Google Scholar] [CrossRef] [PubMed]
- Tsuchidate, T.; Tateno, T.; Okai, N.; Tanaka, T.; Ogino, C.; Kondo, A. Glutamate production from beta-glucan using Endoglucanase-secreting Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2011, 90, 895–901. [Google Scholar] [CrossRef] [PubMed]
- Seibold, G.; Auchter, M.; Berens, S.; Kalinowski, J.; Eikmanns, B.J. Utilization of soluble starch by a recombinant Corynebacterium glutamicum strain: Growth and lysine production. J. Biotechnol. 2006, 124, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Heider, S.A.; Peters-Wendisch, P.; Wendisch, V.F. Carotenoid biosynthesis and overproduction in Corynebacterium glutamicum. BMC Microbiol. 2012, 12, 198. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heider, S.A.; Peters-Wendisch, P.; Netzer, R.; Stafnes, M.; Brautaset, T.; Wendisch, V.F. Production and glucosylation of C50 and C40 carotenoids by metabolically engineered Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2014, 98, 1223–1235. [Google Scholar] [CrossRef] [PubMed]
- Heider, S.A.; Wendisch, V.F. Engineering microbial cell factories: Metabolic engineering of Corynebacterium glutamicum with a focus on non-natural products. Biotechnol. J. 2015, 10, 1170–1184. [Google Scholar] [CrossRef] [PubMed]
- Krubasik, P.; Kobayashi, M.; Sandmann, G. Expression and functional analysis of a gene cluster involved in the synthesis of decaprenoxanthin reveals the mechanisms for C50 carotenoid formation. Eur. J. Biochem. 2001, 268, 3702–3708. [Google Scholar] [CrossRef] [PubMed]
- Heider, S.A.; Peters-Wendisch, P.; Beekwilder, J.; Wendisch, V.F. IdsA is the major geranylgeranyl pyrophosphate synthase involved in carotenogenesis in Corynebacterium glutamicum. FEBS J. 2014, 281, 4906–4920. [Google Scholar] [CrossRef] [PubMed]
- Heider, S.A.; Wolf, N.; Hofemeier, A.; Peters-Wendisch, P.; Wendisch, V.F. Optimization of the IPP precursor supply for the production of lycopene, decaprenoxanthin and astaxanthin by Corynebacterium glutamicum. Front. Bioeng. Biotechnol. 2014, 2, 28. [Google Scholar] [CrossRef] [PubMed]
- Farasat, I.; Kushwaha, M.; Collens, J.; Easterbrook, M.; Guido, M.; Salis, H.M. Efficient search, mapping, and optimization of multi-protein genetic systems in diverse bacteria. Mol. Syst. Biol. 2014, 10, 731. [Google Scholar] [CrossRef] [PubMed]
- Gibson, D.G.; Young, L.; Chuang, R.Y.; Venter, J.C.; Hutchison, C.A.; Smith, H.O. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 2009, 6, 343–345. [Google Scholar] [CrossRef] [PubMed]
- Takano, H.; Agari, Y.; Hagiwara, K.; Watanabe, R.; Yamazaki, R.; Beppu, T.; Shinkai, A.; Ueda, K. LdrP, a cAMP receptor protein/FNR family transcriptional regulator, serves as a positive regulator for the light-inducible gene cluster in the megaplasmid of Thermus thermophilus. Microbiology 2014, 160, 2650–2660. [Google Scholar] [CrossRef] [PubMed]
- Takano, H.; Mise, K.; Hagiwara, K.; Hirata, N.; Watanabe, S.; Toriyabe, M.; Shiratori-Takano, H.; Ueda, K. The role and function of LitR, an AdoB12-bound light-sensitive regulator of Bacillus megaterium QM B1551, in the regulation of carotenoid production. J. Bacteriol. 2015, 197, 2301–2315. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Rouviere, P.E.; Cheng, Q. A carotenoid synthesis gene cluster from a non-marine Brevundimonas that synthesizes hydroxylated astaxanthin. Gene 2006, 379, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Scaife, M.A.; Ma, C.A.; Ninlayarn, T.; Wright, P.C.; Armenta, R.E. Comparative analysis of beta-carotene hydroxylase genes for astaxanthin biosynthesis. J. Nat. Prod. 2012, 75, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Poindexter, J.S. Biological Properties and Classification of the Caulobacter Group. Bacteriol. Rev. 1964, 28, 231–295. [Google Scholar] [PubMed]
- Cho, J.C.; Giovannoni, S.J. Fulvimarina pelagi gen. nov., sp. nov., a marine bacterium that forms a deep evolutionary lineage of descent in the order “Rhizobiales”. Int. J. Syst. Evol. Microbiol. 2003, 53, 1853–1859. [Google Scholar] [PubMed]
- Asker, D.; Amano, S.; Morita, K.; Tamura, K.; Sakuda, S.; Kikuchi, N.; Furihata, K.; Matsufuji, H.; Beppu, T.; Ueda, K. Astaxanthin dirhamnoside, a new astaxanthin derivative produced by a radio-tolerant bacterium, Sphingomonas astaxanthinifaciens. J. Antibiot. 2009, 62, 397–399. [Google Scholar] [CrossRef] [PubMed]
- Asker, D.; Beppu, T.; Ueda, K. Sphingomonas astaxanthinifaciens sp. nov., a novel astaxanthin-producing bacterium of the family Sphingomonadaceae isolated from Misasa, Tottori, Japan. FEMS Microbiol. Lett. 2007, 273, 140–148. [Google Scholar] [CrossRef] [PubMed]
- Kang, I.; Oh, H.M.; Lim, S.I.; Ferriera, S.; Giovannoni, S.J.; Cho, J.C. Genome sequence of Fulvimarina pelagi HTCC2506T, a Mn(II)-oxidizing alphaproteobacterium possessing an aerobic anoxygenic photosynthetic gene cluster and Xanthorhodopsin. J. Bacteriol. 2010, 192, 4798–4799. [Google Scholar] [CrossRef] [PubMed]
- Knoll, A.; Bartsch, S.; Husemann, B.; Engel, P.; Schroer, K.; Ribeiro, B.; Stockmann, C.; Seletzky, J.; Buchs, J. High cell density cultivation of recombinant yeasts and bacteria under non-pressurized and pressurized conditions in stirred tank bioreactors. J. Biotechnol. 2007, 132, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Aflalo, C.; Meshulam, Y.; Zarka, A.; Boussiba, S. On the relative efficiency of two- vs. one-stage production of astaxanthin by the green alga Haematococcus pluvialis. Biotechnol. Bioeng. 2007, 98, 300–305. [Google Scholar] [PubMed]
- Landry, A.P.; Ding, H. Redox Control of Human Mitochondrial Outer Membrane Protein MitoNEET [2Fe-2S] Clusters by Biological Thiols and Hydrogen Peroxide. J. Biol. Chem. 2014, 289, 4307–4315. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Liu, M.; Lv, X.; Lu, W.; Gu, J.; Yu, H. Construction of a controllable beta-carotene biosynthetic pathway by decentralized assembly Strategy in Saccharomyces Cerevisiae. Biotechnol. Bioeng. 2014, 111, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Lemuth, K.; Steuer, K.; Albermann, C. Engineering of a plasmid-free Escherichia coli strain for improvedin vivo biosynthesis of astaxanthin. Microb. Cell Fact. 2011, 10, 29. [Google Scholar] [CrossRef] [PubMed]
- Steiger, S.; Sandmann, G. Cloning of two carotenoid ketolase genes from Nostoc punctiforme for the heterologous production of canthaxanthin and astaxanthin. Biotechnol. Lett. 2004, 26, 813–817. [Google Scholar] [CrossRef] [PubMed]
- Fraser, P.D.; Miura, Y.; Misawa, N. In vitro characterization of astaxanthin biosynthetic enzymes. J. Biol. Chem. 1997, 272, 6128–6135. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Huang, Y.; Zhang, R.; Wang, S.; Liu, Y. Four different methods comparison for extraction of astaxanthin from green alga Haematococcus pluvialis. Sci. World J. 2014, 2014, 694305. [Google Scholar] [CrossRef] [PubMed]
- Scaife, M.A.; Burja, A.M.; Wright, P.C. Characterization of cyanobacterial beta-carotene ketolase and hydroxylase genes in Escherichia coli, and their application for astaxanthin biosynthesis. Biotechnol. Bioeng. 2009, 103, 944–955. [Google Scholar] [CrossRef] [PubMed]
- Tejera, N.; Cejas, J.R.; Rodriguez, C.; Bjerkeng, B.; Jerez, S.; Bolanos, A.; Lorenzo, A. Pigmentation, carotenoids, lipid peroxides and lipid composition of skin of red porgy (Pagrus pagrus) fed diets supplemented with different astaxanthin sources. Aquaculture 2007, 270, 218–230. [Google Scholar] [CrossRef]
- Rodriguez-Saiz, M.; de la Fuente, J.L.; Barredo, J.L. Xanthophyllomyces dendrorhous for the industrial production of astaxanthin. Appl. Microbiol. Biotechnol. 2010, 88, 645–658. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, I.; Schewe, H.; Gassel, S.; Jin, C.; Buckingham, J.; Humbelin, M.; Sandmann, G.; Schrader, J. Biotechnological production of astaxanthin with Phaffia rhodozyma/Xanthophyllomyces dendrorhous. Appl. Microbiol. Biotechnol. 2011, 89, 555–571. [Google Scholar] [CrossRef] [PubMed]
- Ni, H.; Chen, Q.H.; He, G.Q.; Wu, G.B.; Yang, Y.F. Optimization of acidic extraction of astaxanthin from Phaffia rhodozyma. J. Zhejiang Univ. Sci. B 2008, 9, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Monnet, C.; Loux, V.; Gibrat, J.F.; Spinnler, E.; Barbe, V.; Vacherie, B.; Gavory, F.; Gourbeyre, E.; Siguier, P.; Chandler, M.; et al. The Arthrobacter arilaitensis Re117 genome sequence reveals its genetic adaptation to the surface of cheese. PLoS ONE 2010, 5, e15489. [Google Scholar] [CrossRef] [PubMed]
- Brown, N.L.; Stoyanov, J.V.; Kidd, S.P.; Hobman, J.L. The MerR family of transcriptional regulators. FEMS Microbiol. Rev. 2003, 27, 145–163. [Google Scholar] [CrossRef]
- Lorenz, R.T.; Cysewski, G.R. Commercial potential for Haematococcus microalgae as a natural source of astaxanthin. Trends Biotechnol. 2000, 18, 160–167. [Google Scholar] [CrossRef]
- Bumbak, F.; Cook, S.; Zachleder, V.; Hauser, S.; Kovar, K. Best practices in heterotrophic high-cell-density microalgal processes: Achievements, potential and possible limitations. Appl. Microbiol. Biotechnol. 2011, 91, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, G.; Jolly, S.; Sedmak, J.; Skatrud, T.; Wasileski, J. Astaxanthin over-Producing Strains of Phaffia Rhodozyma, Methods for their Cultivation, and their Use in Animal Feeds. U.S. Patent 6,015,684, 12 August 1999. [Google Scholar]
- Baumgart, M.; Unthan, S.; Ruckert, C.; Sivalingam, J.; Grunberger, A.; Kalinowski, J.; Bott, M.; Noack, S.; Frunzke, J. Construction of a prophage-free variant of Corynebacterium glutamicum ATCC 13032 for use as a platform strain for basic research and industrial biotechnology. Appl. Environ. Microbiol. 2013, 79, 6006–6015. [Google Scholar] [CrossRef] [PubMed]
- Eggeling, L.; Reyes, O. Experiments. In Handbook of Corynebacterium Glutamicum; Eggeling, L., Bott, M., Eds.; CRC Press: Boca Raton, FL, USA, 2005; pp. 3535–3566. [Google Scholar]
- Abe, S.; Takayarna, K.; Kinoshita, S. Taxonomical studies on glutamic acid producing bacteria. J. Gen. Appl. Microbiol. 1967, 13, 279–301. [Google Scholar] [CrossRef]
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Misawa, N.; Nakagawa, M.; Kobayashi, K.; Yamano, S.; Izawa, Y.; Nakamura, K.; Harashima, K. Elucidation of the Erwinia uredovora carotenoid biosynthetic pathway by functional analysis of gene products expressed in Escherichia coli. J. Bacteriol. 1990, 172, 6704–6712. [Google Scholar] [PubMed]
- Abraham, W.R.; Strömpl, C.; Meyer, H.; Lindholst, S.; Moore, E.R.; Christ, R.; Vancanneyt, M.; Tindall, B.J.; Bennasar, A.; Smit, J.; et al. Phylogeny and polyphasic taxonomy of Caulobacter species. Proposal of Maricaulis gen. nov. with Maricaulis maris (Poindexter) comb. nov. as the type species, and emended description of the genera Brevundimonas and Caulobacter. Int. J. Syst. Bacteriol. 1999, 49, 1053–1073. [Google Scholar] [CrossRef] [PubMed]
- Busing, K.H.; Doll, W.; Freytag, K. Bacterial flora of the medicinal leech. Arch. Mikrobiol. 1953, 19, 52–86. [Google Scholar] [PubMed]
- Kirchner, O.; Tauch, A. Tools for genetic engineering in the amino acid-producing bacterium Corynebacterium glutamicum. J. Biotechnol. 2003, 104, 287–299. [Google Scholar] [CrossRef]
- Stansen, C.; Uy, D.; Delaunay, S.; Eggeling, L.; Goergen, J.L.; Wendisch, V.F. Characterization of a Corynebacterium glutamicum lactate utilization operon induced during temperature-triggered glutamate production. Appl. Environ. Microbiol. 2005, 71, 5920–5928. [Google Scholar] [CrossRef] [PubMed]
- Peters-Wendisch, P.G.; Schiel, B.; Wendisch, V.F.; Katsoulidis, E.; Mockel, B.; Sahm, H.; Eikmanns, B.J. Pyruvate carboxylase is a major bottleneck for glutamate and lysine production by Corynebacterium glutamicum. J. Mol. Microbiol. Biotechnol. 2001, 3, 295–300. [Google Scholar] [PubMed]
- Schäfer, A.; Tauch, A.; Jäger, W.; Kalinowski, J.; Thierbach, G.; Puhler, A. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: Selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 1994, 145, 69–73. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratoy Press: New York, NY, USA, 2001. [Google Scholar]
- Van der Rest, M.E.; Lange, C.; Molenaar, D. A heat shock following electroporation induces highly efficient transformation of Corynebacterium glutamicum with xenogeneic plasmid DNA. Appl. Microbiol. Biotechnol. 1999, 52, 541–545. [Google Scholar] [CrossRef] [PubMed]
- Eggeling, L.; Bott, M. (Eds.) Handbook of Corynebacterium Glutamicum; CRC Press: Boca Raton, FL, USA, 2005.
- Eikmanns, B.J.; Rittmann, D.; Sahm, H. Cloning, sequence analysis, expression, and inactivation of the Corynebacterium glutamicum icd gene encoding isocitrate dehydrogenase and biochemical characterization of the enzyme. J. Bacteriol. 1995, 177, 774–782. [Google Scholar] [PubMed]


| Name | Strain | Lycopene (mg·(g·CDW)−1) |
|---|---|---|
| LYC3 | crtYeYfEb deletion mutant of C. glutamicum MB001 | 0.04 ± 0.01 |
| LYC3-Ptufdxs | LYC3::Ptufdxs | 0.09 ± 0.01 |
| LYC4 | LYC3::PtufcrtEBI | 0.32 ± 0.01 |
| LYC5 | LYC4::Ptufdxs | 0.43 ± 0.02 |
| Name | Strain | β-Carotene (mg·(g·CDW)−1) |
|---|---|---|
| BETA1 | LYC5 (pEXEx3_crtYPa) | 5.2 ± 1.0 |
| BETA2 | LYC5 (pSH1_crtYPa) | 5.9 ± 0.8 |
| BETA3 | LYC5::Ptuf_ crtYPa | 6.5 ± 1.3 |
| Strain Growth | Carotenoid Titer (mg·g−1·CDW) | Volumetric Productivity (mg·L−1·h−1) | |||||
|---|---|---|---|---|---|---|---|
| BETA4 Transformed with | Final OD600 nm | Astaxanthin | Canthaxanthin | β-Carotene | Astaxanthin | Canthaxanthin | β-Carotene |
| - | 28 ± 1 | <0.1 | <0.1 | 11.7 ± 2.0 | <0.1 | <0.1 | 3.4 ± 0.5 |
| (pSH1_crtWBb) (pEC-XT_crtZBb) | 21 ± 1 | <0.1 | <0.1 | 4.9 ± 0.4 | <0.1 | <0.1 | 1.1 ± 0.1 |
| (pSH1_crtWSa) (pEC-XT_crtZSa) | 22 ± 2 | < 0.1 | 0.3 ± 0.1 | 3.3 ± 0.5 | <0.1 | <0.1 | 0.8 ± 0.1 |
| (pSH1_crtWSa) (pEC-XT_crtZFp) | 24 ± 1 | 0.7 ± 0.3 | 0.2 ± 0.1 | 1.8 ± 0.1 | 0.2 ± 0.1 | <0.1 | 0.5 ± 0.1 |
| (pSH1_crtWBa) (pEC-XT_crtZFp) | 22 ± 1 | 1.7 ± 0.3 | 0.1 ± 0.1 | 2.0 ± 0.5 | 0.4 ± 0.1 | <0.1 | 0.4 ± 0.2 |
| (pSH1_crtWFp) (pEC-XT_crtZFp) = ASTA1 | 23 ± 1 | 1.6 ± 0.3 | 0.1 ± 0.1 | 0.3 ± 0.1 | 0.4 ± 0.1 | <0.1 | 0.1 ± 0.1 |
| Strain; Plasmid | Relevant Characteristics | Reference |
|---|---|---|
| C. glutamicum Strains | ||
| WT | Wild type, ATCC 13032 | [74] |
| MB001 | prophage cured, genome reduced ATCC 13032 | [72] |
| LYC3 | crtYeYfEb deletion mutant of C. glutamicum MB001 | [42] |
| LYC4 | LYC3 derivative with an artificial operon containing crtE, crtB, and crtI under control of the Ptuf promoter integrated into the chromosome | this work |
| LYC5 | LYC4 derivative with dxs under control of the Ptuf promoter integrated into the chromosome | this work |
| BETA1 | LYC5 derivative (pEKEx3_ crtYPa) | this work |
| BETA2 | LYC5 derivative (pSH1_ crtYPa) | this work |
| BETA3 | LYC5 derivative with crtYPa under control of the Ptuf promoter integrated into the chromosome | this work |
| BETA4 | cg0725 deletion mutant of C. glutamicum BETA3 | this work |
| ASTA1 | C. glutamicum BETA4 carrying pSH1_crtW1Fp and pEC-XT_crtZFp | this work |
| Other Strains | ||
| E. coli DH5α | F- thi-1 endA1 hsdr17(r-, m-) supE44 ΔlacU169 (Φ80lacZΔM15) recA1 gyrA96 | [75] |
| Pantoea ananatis | Wild type, ATCC 33244, DSM 17873, Z96081 | [76] |
| Brevundimonas aurantiaca | Wild type, ATCC 15266, DSM 4731, NR028889 | [77] |
| Brevundimonas bacteroides | Wild type, ATCC 15254, DSM 4726, AJ227782 | [49] |
| Brevundimonas vesicularis | Wild type, ATCC 11426, DSM 7226, LN681560 | [78] |
| Fulvimarina pelagi | Wild type, ATCC BAA-666, DSM 15513, AY178860 | [50] |
| Sphingomonas astaxanthinifaciens | Wild type, NBRC 102146, DSM 22298, AB277583 | [52] |
| Plasmids | ||
| pEC-XT99A (pEC-XT) | TetR, PtrclacIq, pGA1 oriVCg, C. glutamicum/E. coli expression shuttle vector | [79] |
| pEC-XT_crtZBb | pEC-XT derivative for IPTG-inducible expression of crtZ from B. bacteroides containing an artificial ribosome binding site | this work |
| pEC-XT_crtZBv | pEC-XT derivative for IPTG-inducible expression of crtZ from B. vesicularis containing an artificial ribosome binding site | this work |
| pEC-XT_crtZFp | pEC-XT derivative for IPTG-inducible expression of crtZ from F. pelagi containing an artificial ribosome binding site | this work |
| pEC-XT_crtZSa | pEC-XT derivative for IPTG-inducible expression of crtZ from S. astaxanthinifaciens containing an artificial ribosome binding site | this work |
| pEKEx3 | SpecR, PtaclacIq, pBL1 oriVCg, C. glutamicum/E. coli expression shuttle vector | [80] |
| pEKEx3_crtYPa | pEKEx3 derivative for IPTG-inducible expression of crtY from P. ananatis containing an artificial ribosome binding site | this work |
| pVWEx1 | KmR, PtaclacIq, pHM519 oriVCg, C. glutamicum/E. coli expression shuttle vector | [81] |
| pSH1 | KmR, Ptuf, pHM519 oriVCg, C. glutamicum/E. coli expression shuttle vector | this work |
| pSH1_crtYPa | pSH1 derivative for constitutive expression of crtY from P. ananatis containing an artificial ribosome binding site | this work |
| pSH1_crtWBa_crtZPa | pSH1 derivative for constitutive expression of crtW from B. aurantiaca and crtZ from P. ananatis containing artificial ribosome binding sites | this work |
| pSH1_crtWBa | pSH1 derivative for constitutive expression of crtW from B. aurantiaca containing an artificial ribosome binding site | this work |
| pSH1_crtWBb | pSH1 derivative for constitutive expression of crtW from B. bacteroides containing an artificial ribosome binding site | this work |
| pSH1_crtW1Bv | pSH1 derivative for constitutive expression of crtW from B. vesicularis containing an artificial ribosome binding site | this work |
| pSH1_crtW2Bv | pSH1 derivative for constitutive expression of crtW from B. vesicularis containing an artificial ribosome binding site | this work |
| pSH1_crtW1Fp | pSH1 derivative for constitutive expression of crtW from F. pelagi containing an artificial ribosome binding site | this work |
| pSH1_crtW2Fp | pSH1 derivative for constitutive expression of crtW from F. pelagi containing an artificial ribosome binding site | this work |
| pSH1_crtW3Fp | pSH1 derivative for constitutive expression of crtW from F. pelagi containing an artificial ribosome binding site | this work |
| pSH1_crtWSa | pSH1 derivative for constitutive expression of crtW from S. astaxanthinifaciens containing an artificial ribosome binding site | this work |
| pK19mobsacB | KmR; E. coli/C. glutamicum shuttle vector for construction of insertion and deletion mutants in C. glutamicum (pK18 oriVEc sacB lacZα) | [82] |
| pK19mobsacB-cg0725 | pK19mobsacB with a cg0725 deletion construct | - |
| pK19mobsacB-Ptuf-dxs | pK19mobsacB derivative with a tuf promoter region (200 bp upstream of the coding sequence of the tuf gene(cg0587) construct for the promoter exchange of dxs | [42] |
| pK19mobsacB-IntcrtEBI | pK19mobsacB derivative containing the artificial operon crtE_crtBI under the control of the Ptuf promoter with an addition ribosome binding site in front of crtB for integration in the cgp2 cured region of C. glutamicum MB001 | this work |
| pVWEx1-crtEBI | pVWEx1 derivative for IPTG-inducible expression of crtE, crtB and crtI from C. glutamicum containing artificial ribosome binding sites in front of crtE and crtBI | [38] |
| pK19mobsacB-IntcrtY | pK19mobsacB derivative containing crtY of Pantoea ananatis under the control of the Ptuf promoter for integration in the cgp1 cured region of C. glutamicum MB001 | this work |
| Oligonucleotide | Sequence (5'→3') |
|---|---|
| N1 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGACCGCCGCCGTCGCCGAG |
| N2 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGGTGACCGCCGCCGTCGCCGAG |
| N3 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCATGACCGCCGCCGTCGCCGAG |
| N4 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCGTGACCGCCGCCGTCGCCGAG |
| N5 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCATGACCGCCGCCGTCGCCGAG |
| N6 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCGTGACCGCCGCCGTCGCCGAG |
| N7 | CATGCCTGCAGGTCGACTCTAGAGGAAAGAAGGCCCTTCAGATGACCGCCGCCGTCGCCGAG |
| N8 | CATGCCTGCAGGTCGACTCTAGAGGAAAGAAGGCCCTTCAGGTGACCGCCGCCGTCGCCGAG |
| N9 | CATGCCTGCAGGTCGACTCTAGAGGAAAGAAGGCCCTTCATGACCGCCGCCGTCGCCGAG |
| N10 | CATGCCTGCAGGTCGACTCTAGAGGAAAGAAGGCCCTTCGTGACCGCCGCCGTCGCCGAG |
| N11 | CATGCCTGCAGGTCGACTCTAGAGGAAAGAAGGCCCATGACCGCCGCCGTCGCCGAG |
| N12 | CATGCCTGCAGGTCGACTCTAGAGGAAAGAAGGCCCGTGACCGCCGCCGTCGCCGAG |
| N13 | CATGCCTGCAGGTCGACTCTAGAGGAATGGAGGCCCTTCAGATGACCGCCGCCGTCGCCGAG |
| N14 | CATGCCTGCAGGTCGACTCTAGAGGAATGGAGGCCCTTCAGGTGACCGCCGCCGTCGCCGAG |
| N15 | CATGCCTGCAGGTCGACTCTAGAGGAATGGAGGCCCTTCATGACCGCCGCCGTCGCCGAG |
| N16 | CATGCCTGCAGGTCGACTCTAGAGGAATGGAGGCCCTTCGTGACCGCCGCCGTCGCCGAG |
| N17 | CATGCCTGCAGGTCGACTCTAGAGGAATGGAGGCCCATGACCGCCGCCGTCGCCGAG |
| N18 | CATGCCTGCAGGTCGACTCTAGAGGAATGGAGGCCCGTGACCGCCGCCGTCGCCGAG |
| N19 | CATGCCTGCAGGTCGACTCTAGAGGAATGAAGGCCCTTCAGATGACCGCCGCCGTCGCCGAG |
| N20 | CATGCCTGCAGGTCGACTCTAGAGGAATGAAGGCCCTTCAGGTGACCGCCGCCGTCGCCGAG |
| N21 | CATGCCTGCAGGTCGACTCTAGAGGAATGAAGGCCCTTCATGACCGCCGCCGTCGCCGAG |
| N22 | CATGCCTGCAGGTCGACTCTAGAGGAATGAAGGCCCTTCGTGACCGCCGCCGTCGCCGAG |
| N23 | CATGCCTGCAGGTCGACTCTAGAGGAATGAAGGCCCATGACCGCCGCCGTCGCCGAG |
| N24 | CATGCCTGCAGGTCGACTCTAGAGGAATGAAGGCCCGTGACCGCCGCCGTCGCCGAG |
| N25 | AACTGCCACACGAACGAAAGGAGGCCCTTCAGATGTTGTGGATTTGGAATGCCCTGATC |
| N26 | AACTGCCACACGAACGAAAGGAGGCCCTTCAGGTGTTGTGGATTTGGAATGCCCTGATC |
| N27 | AACTGCCACACGAACGAAAGGAGGCCCTTCATGTTGTGGATTTGGAATGCCCTGATC |
| N28 | AACTGCCACACGAACGAAAGGAGGCCCTTCGTGTTGTGGATTTGGAATGCCCTGATC |
| N29 | AACTGCCACACGAACGAAAGGAGGCCCATGTTGTGGATTTGGAATGCCCTGATC |
| N30 | AACTGCCACACGAACGAAAGGAGGCCCGTGTTGTGGATTTGGAATGCCCTGATC |
| N31 | AACTGCCACACGAACGAAAGAAGGCCCTTCAGATGTTGTGGATTTGGAATGCCCTGATC |
| N32 | AACTGCCACACGAACGAAAGAAGGCCCTTCAGGTGTTGTGGATTTGGAATGCCCTGATC |
| N33 | AACTGCCACACGAACGAAAGAAGGCCCTTCATGTTGTGGATTTGGAATGCCCTGATC |
| N34 | AACTGCCACACGAACGAAAGAAGGCCCTTCGTGTTGTGGATTTGGAATGCCCTGATC |
| N35 | AACTGCCACACGAACGAAAGAAGGCCCATGTTGTGGATTTGGAATGCCCTGATC |
| N36 | AACTGCCACACGAACGAAAGAAGGCCCGTGTTGTGGATTTGGAATGCCCTGATC |
| N37 | AACTGCCACACGAACGAATGGAGGCCCTTCAGATGTTGTGGATTTGGAATGCCCTGATC |
| N38 | AACTGCCACACGAACGAATGGAGGCCCTTCAGGTGTTGTGGATTTGGAATGCCCTGATC |
| N39 | AACTGCCACACGAACGAATGGAGGCCCTTCATGTTGTGGATTTGGAATGCCCTGATC |
| N40 | AACTGCCACACGAACGAATGGAGGCCCTTCGTGTTGTGGATTTGGAATGCCCTGATC |
| N41 | AACTGCCACACGAACGAATGGAGGCCCATGTTGTGGATTTGGAATGCCCTGATC |
| N42 | AACTGCCACACGAACGAATGGAGGCCCGTGTTGTGGATTTGGAATGCCCTGATC |
| N43 | AACTGCCACACGAACGAATGAAGGCCCTTCAGATGTTGTGGATTTGGAATGCCCTGATC |
| N44 | AACTGCCACACGAACGAATGAAGGCCCTTCAGGTGTTGTGGATTTGGAATGCCCTGATC |
| N45 | AACTGCCACACGAACGAATGAAGGCCCTTCATGTTGTGGATTTGGAATGCCCTGATC |
| N46 | AACTGCCACACGAACGAATGAAGGCCCTTCGTGTTGTGGATTTGGAATGCCCTGATC |
| N47 | AACTGCCACACGAACGAATGAAGGCCCATGTTGTGGATTTGGAATGCCCTGATC |
| N48 | AACTGCCACACGAACGAATGAAGGCCCGTGTTGTGGATTTGGAATGCCCTGATC |
| N49 | GTTCGTGTGGCAGTTTTAGTGGTGGTGGTGGTGGTGAGACTCGCCGCGCCACAGACG |
| N50 | ATTCGAGCTCGGTACCCGGGGATCTTACTTTTCGAACTGTGGGTGGGACCACTTCCCGGATGCGGGCTC |
| BaW1 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGACCGCCGCCGTCGCCGAG |
| BaW2 | CGGTACCCGGGGATCTCAAGACTCGCCGCGCCAC |
| BbW1 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGACGCGGGAACGCCA |
| BbW2 | ATTCGAGCTCGGTACCCGGGGATCTTAGAGACGTTCGCTACGC |
| BbZ1 | ATGGAATTCGAGCTCGGTACCCGGGGAAAGGAGGCCCTTCAGATGACGATCGTCTGGTTCAC |
| BbZ2 | GCATGCCTGCAGGTCGACTCTAGAGGATCTTACTCGGCCGGGATGTCC |
| BvW1 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGGGGCAAGCGAACAG |
| BvW2 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGCGGCAAGCGAACAG |
| BvW3 | ATTCGAGCTCGGTACCCGGGGATCCTAGCTGAACAAACTCCACCAG |
| BvZ1 | ATGGAATTCGAGCTCGGTACCCGGGGAAAGGAGGCCCTTCAGATGTCCTGGCCGACGATG |
| BvZ2 | GCATGCCTGCAGGTCGACTCTAGAGGATCTTAGGCGCCGTTGCTGGAT |
| FpW1 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGACCCTCAGCCCAACCTC |
| FpW2 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGAGACCCTACCAAACGACG |
| FpW3 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGCATGGTTCGCTGGC |
| FpW4 | ATTCGAGCTCGGTACCCGGGGATCTTAGGACTGGCGAGTATGCG |
| FpZ1 | ATGGAATTCGAGCTCGGTACCCGGGGAAAGGAGGCCCTTCAGATGACGATCTGGACTCTCTACTAC |
| FpZ2 | GCATGCCTGCAGGTCGACTCTAGAGGATCTTACCGAACCGGCGCGT |
| SaW1 | CATGCCTGCAGGTCGACTCTAGAGGAAAGGAGGCCCTTCAGATGGCCCCCATGCTCAGTG |
| SaW2 | ATTCGAGCTCGGTACCCGGGGATCTTAGGCGGGAAGCGCAAG |
| SaZ1 | ATGGAATTCGAGCTCGGTACCCGGGGAAAGGAGGCCCTTCAGATGTCCTGGCCTGCCG |
| SaZ2 | GCATGCCTGCAGGTCGACTCTAGAGGATCTTAGGCCCGCTCCTCGTG |
| pSH1 fw | ACCGGCTCCAGATTTATCAG |
| pVWEx/ pSH1 rv | ATCTTCTCTCATCCGCCA |
| pEC-XT fw | AATACGCAAACCGCCTCTCC |
| pEC-XT rv | TACTGCCGCCAGGCAAATTC |
| pV_Ptuf -fw | CGGAATCTTGCACGCCCTTGGCCGTTACCCTGCGAATG |
| pV_Ptuf -rv | CTGCAGGCATGCAAGCTTTGTATGTCCTCCTGGACTTC |
| pV1-fw | GAAGTCCAGGAGGACATACAAAGCTTGCATGCCTGCAG |
| pV6962-rv | CATTCGCAGGGTAACGGCCAAGGGCGTGCAAGATTCCG |
| cg0725-A | GCAGGTCGACTCTAGAGGATCCCCGCGCGAAGATTTGATGGG |
| cg0725-B | CCCATCCACCCCGGGTAAACATTCCTGCATATTCAGCATAGTAATC |
| cg0725-C | TGTTTACCCGGGGTGGATGGGTCCCTTAATAATGCACCATGGC |
| cg0725-D | CCAGTGAATTCGAGCTCGGTACCCCTTGTCACCACAGCACTACT |
| cg0725-E | GCGCGAAGATTTGATGGG |
| cg0725-F | ACTTGTCACCACAGCACTAC |
| crtY-Int1 | GCAGGTCGACTCTAGAGGATCCCCCAGTGAAGGATCGGTGCG |
| crtY-Int2 | CATTCGCAGGGTAACGGCCACCTATCTGCTGGCCGGTG |
| crtY-Int3 | CACCGGCCAGCAGATAGGTGGCCGTTACCCTGCGAATG |
| crtY-Int4 | CAGATCATAATGCGGTTGCATTGTATGTCCTCCTGGACTTC |
| crtY-Int5 | GAAGTCCAGGAGGACATACAATGCAACCGCATTATGATCTG |
| crtY-Int6 | TCTTACTACTTGCGCTAGGTACAGTTAACGATGAGTCGTCATAATGG |
| crtY-Int7 | CCATTATGACGACTCATCGTTAACTGTACCTAGCGCAAGTAGTAAGA |
| crtY-Int8 | CCAGTGAATTCGAGCTCGGTACCCCTGCTCATCCTTCAACAACGT |
| cgp1-E | GTGGTGCTCGAGAACATAAG |
| cgp1-F | CGGTCACCCGTAACAATCAG |
| crtEBI-Int1 | GCAGGTCGACTCTAGAGGATCCCCGTGCTTCGCATCGTCTATGTC |
| crtEBI-Int2 | CATTCGCAGGGTAACGGCCAATAGTTGGGGGAATTTATAAGGATTTG |
| crtEBI-Int3 | CAAATCCTTATAAATTCCCCCAACTATTGGCCGTTACCCTGCGAATG |
| crtEBI-Int4 | GATTGTCATGCCATTGTCCATTGTATGTCCTCCTGGACTTC |
| crtEBI-Int5 | GAAGTCCAGGAGGACATACAATGGACAATGGCATGACAATC |
| crtEBI-Int6 | CTAATGGACGGTGAAGTATCATTTATGTTAATGATCGTATGAGGTCTTTTGAG |
| crtEBI-Int7 | CTCAAAAGACCTCATACGATCATTAACATAAATGATACTTCACCGTCCATTAG |
| crtEBI-Int8 | CCAGTGAATTCGAGCTCGGTACCCCGCCGTATGTAACAAGATTTG |
| Cgp2-E | TCGCACCATCTACGACAACC |
| Cgp2-F | CTACGAAGCTGACGCCGAAG |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Henke, N.A.; Heider, S.A.E.; Peters-Wendisch, P.; Wendisch, V.F. Production of the Marine Carotenoid Astaxanthin by Metabolically Engineered Corynebacterium glutamicum. Mar. Drugs 2016, 14, 124. https://doi.org/10.3390/md14070124
Henke NA, Heider SAE, Peters-Wendisch P, Wendisch VF. Production of the Marine Carotenoid Astaxanthin by Metabolically Engineered Corynebacterium glutamicum. Marine Drugs. 2016; 14(7):124. https://doi.org/10.3390/md14070124
Chicago/Turabian StyleHenke, Nadja A., Sabine A. E. Heider, Petra Peters-Wendisch, and Volker F. Wendisch. 2016. "Production of the Marine Carotenoid Astaxanthin by Metabolically Engineered Corynebacterium glutamicum" Marine Drugs 14, no. 7: 124. https://doi.org/10.3390/md14070124








