Two New Antibiotic Pyridones Produced by a Marine Fungus, Trichoderma sp. Strain MF106
Abstract
:1. Introduction
2. Results and Discussion
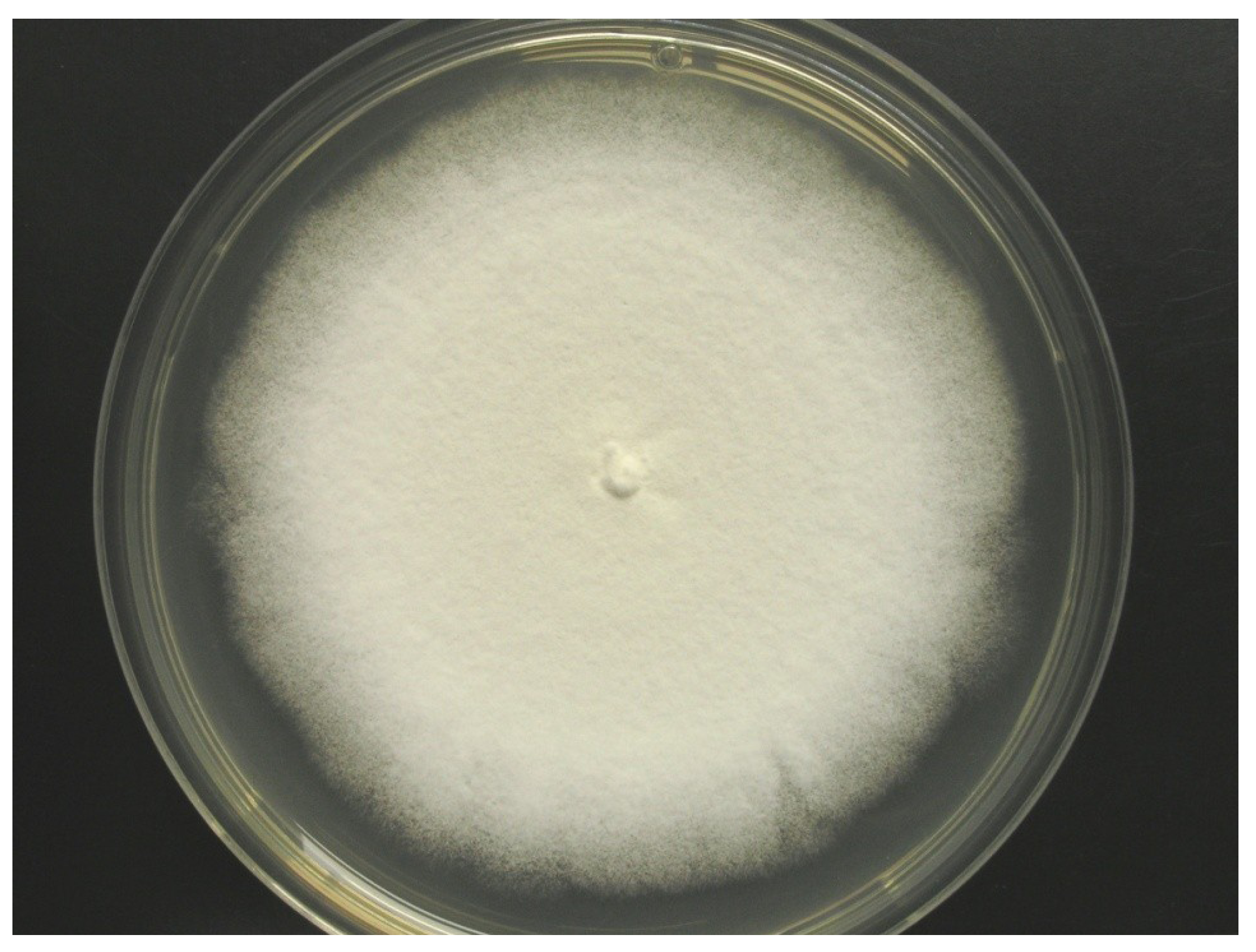
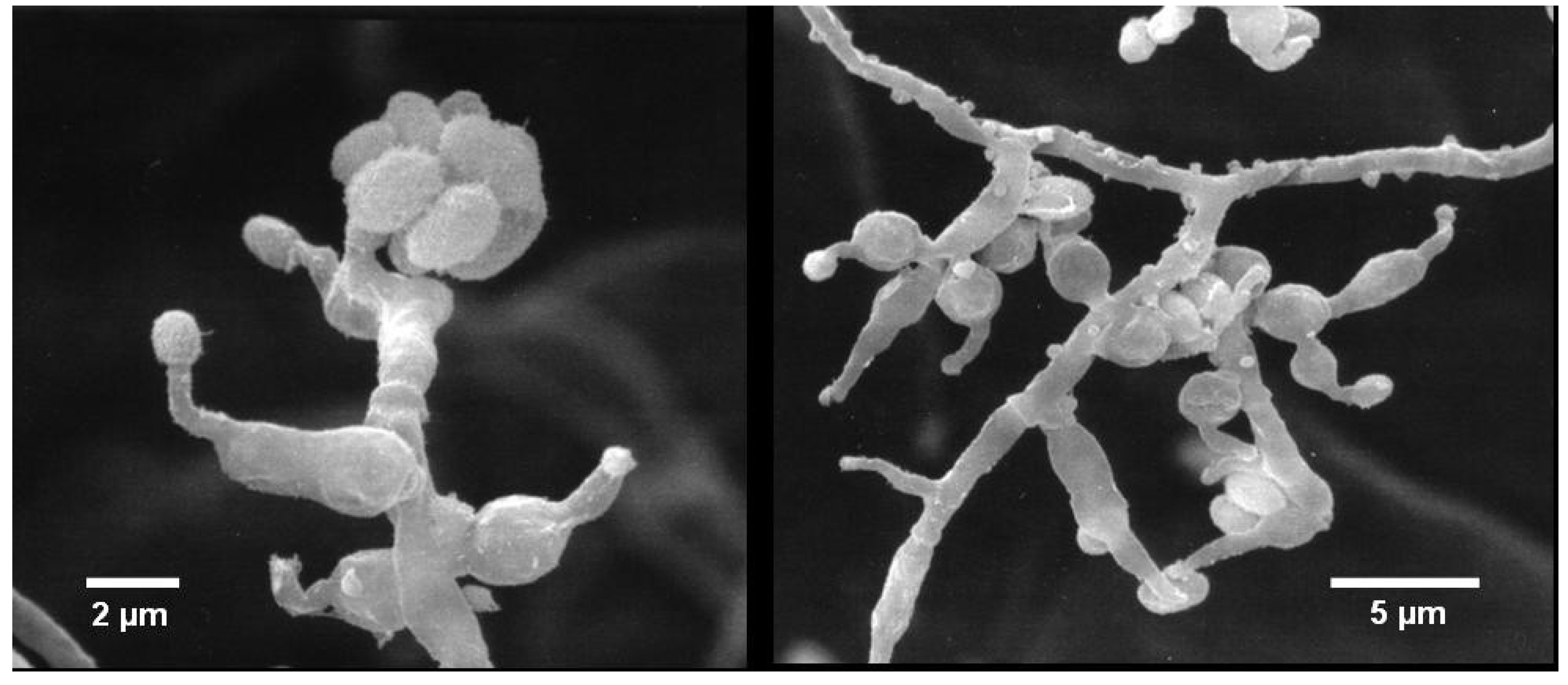
2.1. Identification of Strain MF106
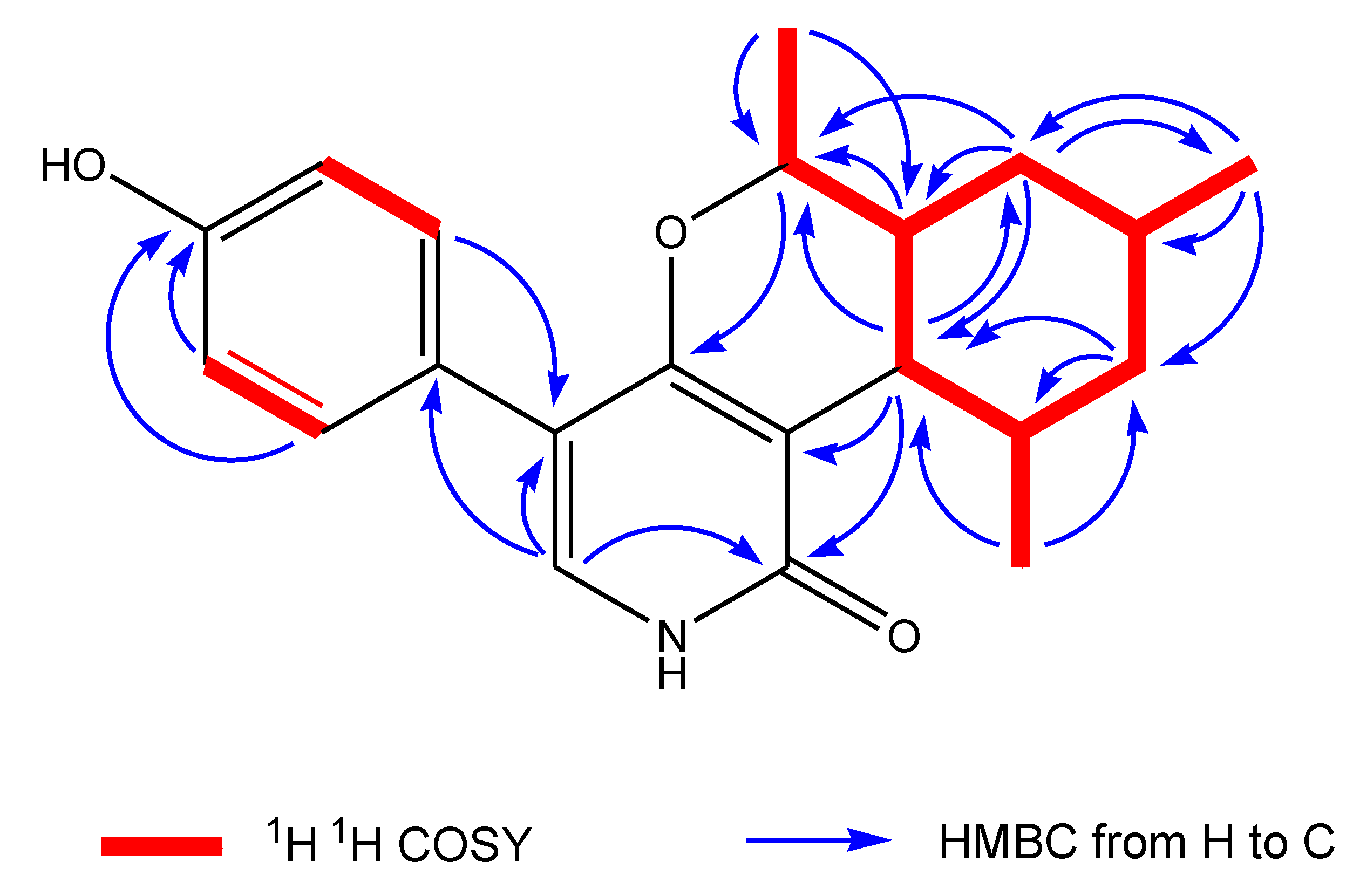
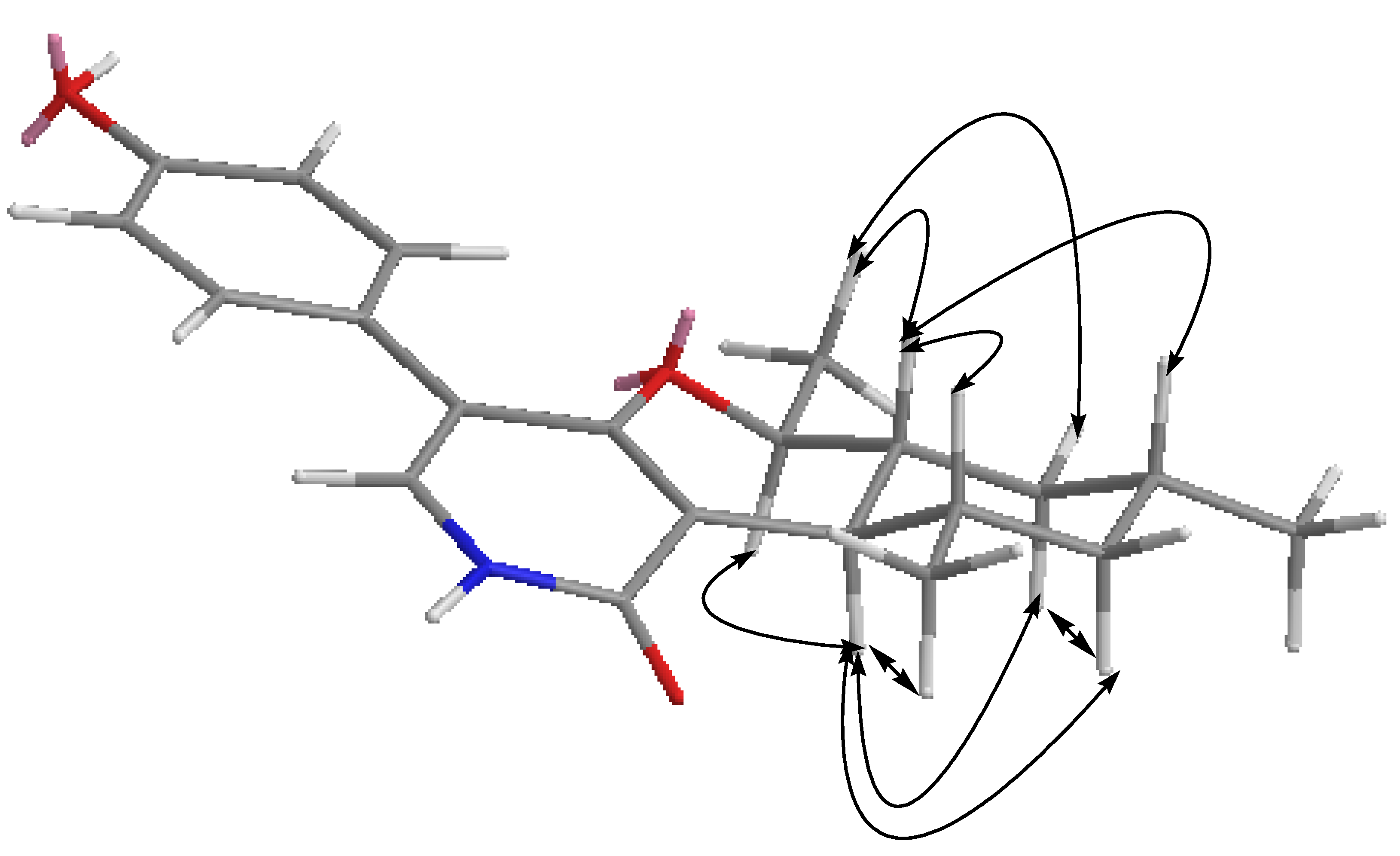
2.2. Structural Elucidation

| Position | 1 | 2 | 3 | |||
|---|---|---|---|---|---|---|
| δCa, b, type | δHc, multiplicities (J in Hz) | δCa, b, type | δHc, multiplicities (J in Hz) | δCa, b, type | δHc, multiplicities (J in Hz) | |
| 2 | 165.9, C | - | 165.8, C | - | 160.4, C | - |
| 3 | 113.0, C | - | 113.0, C | - | 114.7, C | - |
| 4 | 165.9, C | - | 166.0, C | - | 164.0, C | - |
| 5 | 117.6, C | - | 117.2, C | - | 99.0, CH | 5.94, d (8.2) |
| 6 | 131.3, CH | 7.13, d (0.5) | 131.6, CH | 7.15, s | 132.8, CH | 7.52, d (8.2) |
| 7 | 45.4, CH | 2.29, t (10.1) | 45.3, CH | 2.26, t (10.1) | 45.2, CH | 2.46, t (10.8) |
| 8 | 50.9, CH | 1.54, ddd (23.9, 10.2, 2.6) | 50.9, CH | 1.53, ddd (24.0, 10.2, 2.6) | 48.1, CH | 3.00, m |
| 9α | 38.5, CH2 | 0.89, dd (23.9, 12.1) | 38.5, CH2 | 0.89, dd (24.0, 12.0) | 43.6, CH2 | 1.73, m |
| 9β | 1.89, br d (14.9) | - | 1.87. br d (12.2) | - | 0.89, m | |
| 10 | 34.1 CH | 1.70, m | 34.1, CH | 1.68, m | 33.0, CH | 1.61, m |
| 11α | 47.1, CH2 | 1.07, dd (24.8, 11.8) | 47.1, CH2 | 1.06, dd (24.7, 11.8) | 45.9, CH2 | 1.71, m |
| 11β | - | 1.80, br d (13.2) | - | 1.78, br d (13.0) | - | 0.77, m |
| 12 | 41.9, CH | 1.75, m | 41.9, CH | 1.71, m | 33.0, CH | 2.37, m |
| 13 | 79.7, CH | 3.72, m | 79.7, CH | 3.68, m | 144.7, CH | 5.53, ddd (16.8, 10.0, 9.2) |
| 14 | 19.1, CH3 | 1.31, d (6.3) | 19.1, CH3 | 1.28, d (6.2) | 113.0, CH | 4.75, dd (16.5, 2.1), 4.61 dd (10.0, 2.1) |
| 15 | 22.8, CH3 | 1.02, d (6.6) | 22.8, CH3 | 0.99, d (6.4) | 23.2, CH3 | 0.92, d (6.6) |
| 16 | 23.2, CH3 | 1.15, d (6.8) | 23.2, CH3 | 1.12, d (6.7) | 21.0, CH3 | 0.71, dd (6.7) |
| 17 | 126.6, C | 129.3, C | - | - | - | |
| 18/22 | 131.3, CH | 7.25, d (8.8) | 131.2, CH | 7.33, d (8.7) | - | - |
| 19/21 | 115.9, CH | 6.80, d (8.8) | 117.9, CH | 7.13, d (8.7) | - | - |
| 20 | 157.9, C | - | 158.1, C | - | - | - |
| Ribose | - | - | - | - | - | - |
| 1 | - | - | 102.4, CH | 5.64, d (4.4) | - | - |
| 2 | - | - | 73.4, CH | 4.19, dd (6.4, 4.5) | - | - |
| 3 | - | - | 71.2, CH | 4.10, dd (6.5, 3.2) | - | - |
| 4 | - | - | 87.5, CH | 5.15, dd (7.0, 3.5) | - | - |
| 5a | - | - | 63.2, CH2 | 3.65, dd (12.1, 4.0) | - | - |
| 5b | - | - | - | 3.71, dd (12.1, 3.4) | - | - |
2.3. Biological Activities
3. Experimental Section
3.1. General Experimental Procedures
3.2. Isolation, Cultivation, Identification and Storage of the Producer Strain, MF106
3.3. Fermentation and Production of Extracts
3.4. Isolation of Compounds
3.5. Antimicrobial Activity Assays
4. Conclusions
Supplementary Files
Acknowledgments
Conflicts of Interest
References
- Debbab, A.; Aly, A.H.; Lin, W.H.; Proksch, P. Bioactive compounds from marine bacteria and fungi. Microb. Biotechnol. 2010, 3, 544–563. [Google Scholar] [CrossRef]
- Saleema, M.; Ali, M.S.; Hussain, S.; Jabbar, A.; Ashraf, M.; Lee, Y.S. Marine natural products of fungal origin. Nat. Prod. Rep. 2007, 24, 1142–1152. [Google Scholar] [CrossRef]
- Yamazaki, H.; Rotinsulu, H.; Kaneko, T.; Murakami, K.; Fujiwara, H.; Ukai, K.; Namikoshi, M. A new dibenz[b,e]oxepine derivative, 1-hydroxy-10-methoxy-dibenz[b,e]oxepin-6,11-dione, from a marine-derived fungus, Beauveria bassiana TPU942. Mar. Drugs 2012, 10, 2691–2697. [Google Scholar] [CrossRef]
- Jiang, W.; Ye, P.; Chen, C.-T.A.; Wang, K.; Liu, P.; He, S.; Wu, X.; Gan, L.; Ye, Y.; Wu, B. Two novel hepatocellular carcinoma cycle inhibitory cyclodepsipeptides from a hydrothermal vent crab-associated fungus Aspergillus clavatus C2WU. Mar. Drugs 2013, 11, 4761–4772. [Google Scholar] [CrossRef]
- Sun, L.; Li, D.; Tao, M.; Chen, Y.; Dan, F.; Zhang, W. Scopararanes C–G: New oxygenated pimarane diterpenes from the marine sediment-derived fungus Eutypella scoparia FS26. Mar. Drugs 2012, 10, 539–550. [Google Scholar] [CrossRef]
- Wu, B.; Wu, X.; Sun, M.; Li, M. Two novel tyrosinase inhibitory sesquiterpenes induced by CuCl2 from a marine-derived fungus Pestalotiopsis sp. Z233. Mar. Drugs 2013, 11, 2713–2721. [Google Scholar] [CrossRef] [Green Version]
- Bhadury, P.; Mohammad, B.T.; Wright, P.C. The current status of natural products from marine fungi and their potential as anti-infective agents. J. Ind. Microbiol. Biotechnol. 2006, 33, 325–337. [Google Scholar] [CrossRef]
- De Silva, E.D.; Geiermann, A.-S.; Mitova, M.I.; Kuegler, P.; Blunt, J.W.; Cole, A.L.J.; Munro, M.H.G. Isolation of 2-pyridone alkaloids from a new zealand marine-derived Penicillium species. J. Nat. Prod. 2009, 72, 477–479. [Google Scholar] [CrossRef]
- Teshima, Y.; Shin-ya, K.; Shimazu, A.; Furihata, K.; Chul, H.S.; Furihata, K.; Hayakawa, Y.; Nagai, K.; Seto, H. Isolation and structural elucidation of pyridoxatin, a free radical scavenger of microbial origin. J. Antibiot. 1991, 44, 685–687. [Google Scholar] [CrossRef]
- Snider, B.B.; Lu, Q. Total synthesis of (±)-pyridoxatin. J. Org. Chem. 1994, 59, 8065–8070. [Google Scholar] [CrossRef]
- Cai, P.; Smith, D.; Cunningham, B.; Brown-Shimer, S.; Katz, B.; Pearce, C.; Venables, D.; Houck, D. 8-Methyl-pyridoxatin: A novel N-hydroxy pyridone from fungus OS-F61800 that induces erythropoietin in human cells. J. Nat. Prod. 1999, 62, 397–399. [Google Scholar] [CrossRef]
- Druzhinina, I.S. Evolution, diversity and ecophysiology of Trichoderma. Habilitation Thesis, Vienna University of Technology, Vienna, Austria,.
- Song, F.; Dai, H.; Tong, Y.; Ren, B.; Chen, C.; Sun, N.; Liu, X.; Bian, J.; Liu, M.; Gao, H.; et al. Trichodermaketones A–D and 7-O-methylkoninginin D from the marine fungus Trichoderma koningii. J. Nat. Prod. 2010, 73, 806–810. [Google Scholar] [CrossRef]
- Harman, G.E. Overview of mechanisms and uses of Trichoderma spp. Phytopathology 2006, 96, 90–194. [Google Scholar] [CrossRef]
- Kredics, L.; Antal, Z.; Doczi, I.; Manczinger, L.; Kevei, F.; Nagy, E. Clinical importance of the genus Trichoderma. A review. Acta Microbiol. Immunol. Hung. 2003, 50, 105–117. [Google Scholar] [CrossRef]
- Thrane, U.; Poulsen, S.B.; Nirenberg, H.I.; Lieckfeldt, E. Identification of Trichoderma strains by image analysis of HPLC chromatograms. FEMS Microbiol. Lett. 2001, 203, 249–255. [Google Scholar] [CrossRef]
- Ruiz, N.; Roullier, C.; Petit, K.; Sallenave-Namont, C.; Grovel, O.; Pouchus, Y.F. Marine-Derived Trichoderma: A Source of New Bioactive Metabolites. In Trichoderma: Biology and Applications; Mukherjee, P.K., Horwitz, B.A., Singh, U.S., Mukherjee, M., Schmoll, M., Eds.; CABI: Wallingford, UK, 2013; pp. 247–279. [Google Scholar]
- Pruksakorn, P.; Arai, M.; Kotoku, N.; Vilchèze, C.; Baughn, A.D.; Moodley, P.; Jacobs, W.R., Jr.; Kobayashi, M. Trichoderins, novel aminolipopeptides from a marine sponge-derived Trichoderma sp., are active against dormant mycobacteria. Bioorg. Med. Chem. Lett. 2010, 20, 3658–3663. [Google Scholar] [CrossRef]
- Garo, E.; Starks, C.M.; Jensen, P.R.; Fenical, W.; Lobkovsky, E.; Clardy, J. Trichodermamides A and B, cytotoxic modified dipeptides from the marine-derived fungus Trichoderma virens. J. Nat. Prod. 2003, 66, 423–426. [Google Scholar] [CrossRef]
- Khamthong, N.; Rukachaisirikul, V.; Tadpetch, K.; Kaewpet, M.; Phongpaichit, S.; Preedanon, S.; Sakayaroj, J. Tetrahydroanthraquinone and xanthone derivatives from the marine-derived fungus Trichoderma aureoviride PSU-F95. Arch. Pharm. Res. 2012, 35, 461–468. [Google Scholar] [CrossRef]
- Crous, P.W.; Gams, W.; Stalpers, J.A.; Robert, V.; Stegehuis, G. MycoBank: An online initiative to launch mycology into the 21st century. Stud. Mycol. 2004, 50, 19–22. [Google Scholar]
- Yuan, W.; Zhang, L.-P.; Cheng, K.-D.; Zhu, P.; Wang, Q.; He, H.-X.; Zhu, H.X. Microbial O-demethylation, hydroxylation, sulfation, and ribosylation of a xanthone derivative from Halenia elliptic. J. Nat. Prod. 2006, 69, 811–814. [Google Scholar]
- Fütterer, D. Expeditionsprogramm Nr. 21. FS “Polarstern” ARKTIS VIII/1 und 2 1991. Alfred-Wegener-Institut für Polar- und Meeresforschung Bremerhaven. (in German). Available online: http://epic.awi.de/29797/1/PE_21.pdf (accessed on 19 February 2014).
- Wickerham, L.J. Taxonomy of yeasts. U.S. Dept. Tech. Bull. 1951, 29, 1–56. [Google Scholar]
- Ohlendorf, B.; Schulz, D.; Erhard, A.; Nagel, K.; Imhoff, J.F. Geranylphenazinediol, an acetylcholinesterase inhibitor produced by a Streptomyces species. J. Nat. Prod. 2012, 75, 1400–1404. [Google Scholar] [CrossRef]
- Silber, J.; Ohlendorf, B.; Labes, A.; Erhard, A.; Imhoff, J.F. Calcarides. Mar. Drugs 2013, 11, 3309–3323. [Google Scholar] [CrossRef]
- Jansen, N.; Ohlendorf, B.; Erhard, A.; Imhoff, J.F.; Helicusin, E. isochromophilone X and isochromophilone XI: New chloroazaphilones produced by the fungus Bartalinia robillardoides strain LF550. Mar. Drugs 2012, 11, 800–816. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wu, B.; Oesker, V.; Wiese, J.; Schmaljohann, R.; Imhoff, J.F. Two New Antibiotic Pyridones Produced by a Marine Fungus, Trichoderma sp. Strain MF106. Mar. Drugs 2014, 12, 1208-1219. https://doi.org/10.3390/md12031208
Wu B, Oesker V, Wiese J, Schmaljohann R, Imhoff JF. Two New Antibiotic Pyridones Produced by a Marine Fungus, Trichoderma sp. Strain MF106. Marine Drugs. 2014; 12(3):1208-1219. https://doi.org/10.3390/md12031208
Chicago/Turabian StyleWu, Bin, Vanessa Oesker, Jutta Wiese, Rolf Schmaljohann, and Johannes F. Imhoff. 2014. "Two New Antibiotic Pyridones Produced by a Marine Fungus, Trichoderma sp. Strain MF106" Marine Drugs 12, no. 3: 1208-1219. https://doi.org/10.3390/md12031208
APA StyleWu, B., Oesker, V., Wiese, J., Schmaljohann, R., & Imhoff, J. F. (2014). Two New Antibiotic Pyridones Produced by a Marine Fungus, Trichoderma sp. Strain MF106. Marine Drugs, 12(3), 1208-1219. https://doi.org/10.3390/md12031208








