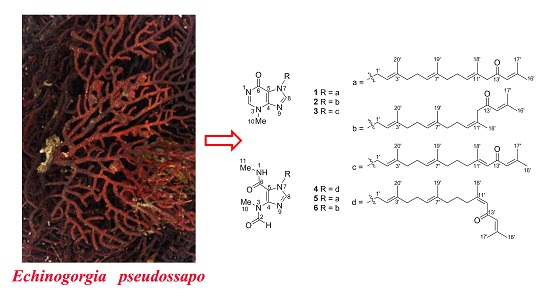
Six New Tetraprenylated Alkaloids from the South China Sea Gorgonian Echinogorgia pseudossapo
Abstract
:1. Introduction
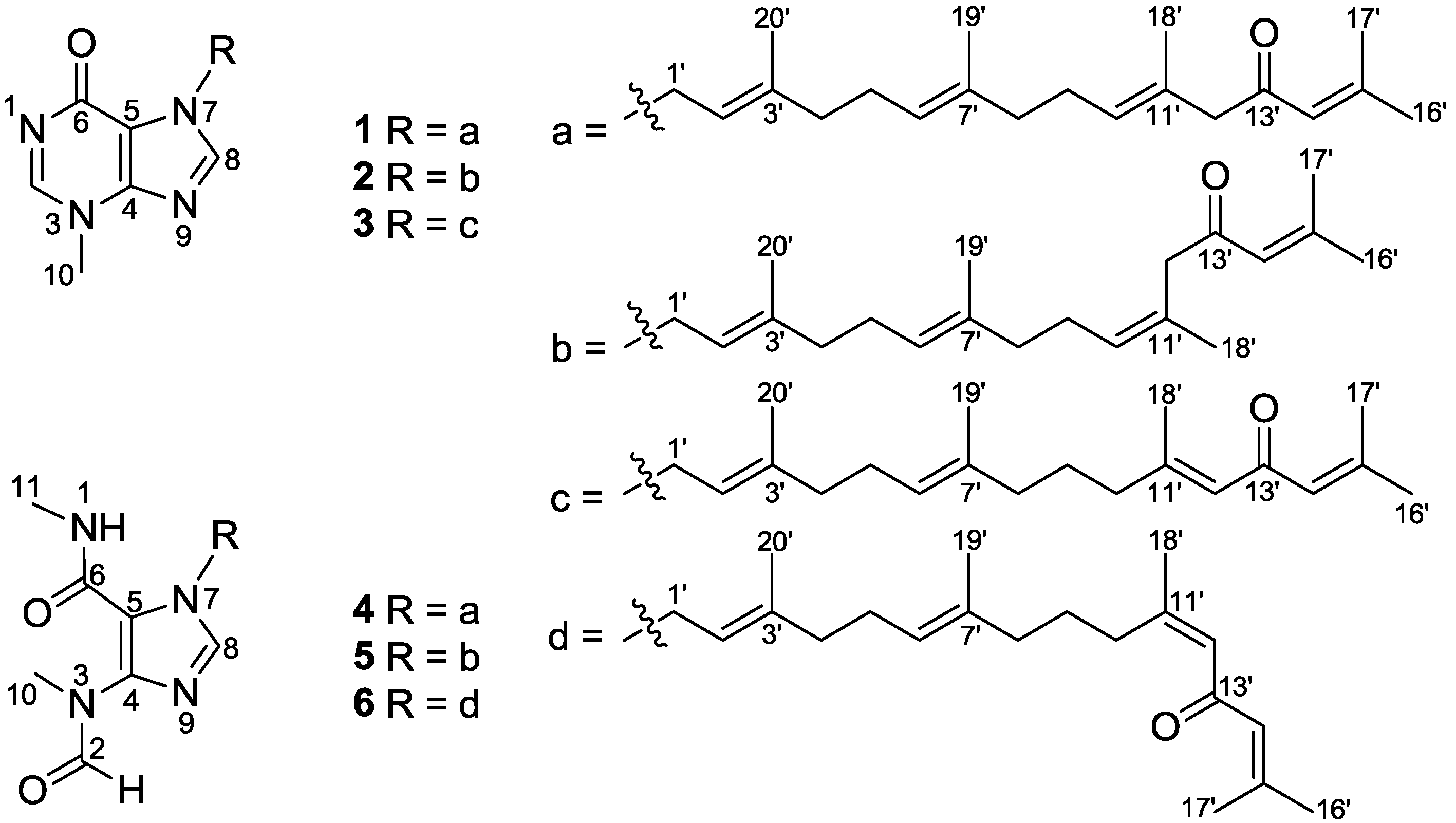
2. Results and Discussion
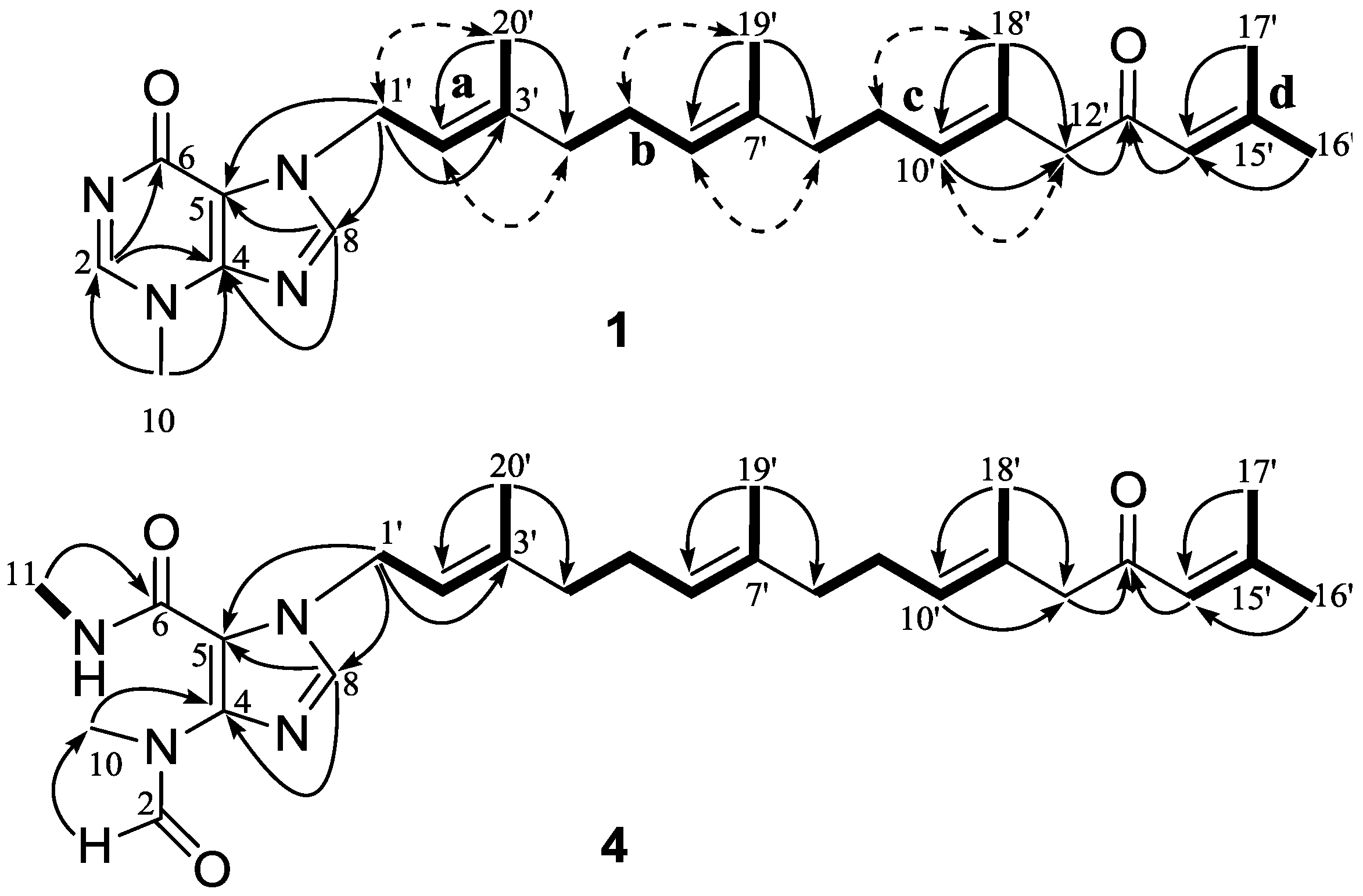
2.1. Structural Elucidation of New Compounds
| No. | 1 a | 2 a | 3 a | 4 b | 5 b | 6 b |
|---|---|---|---|---|---|---|
| 1 | 7.23, brs | 7.22, brs | ||||
| 2 | 8.26, br s | 8.57, br s | 8.62, br s | 8.23, s | 8.23, s | 8.23, s |
| 8 | 7.69, s | 7.73, s | 7.73, s | 7.59, s | 7.59, s | 7.57, s |
| 10 | 3.86, s | 3.94, s | 3.95, s | 3.13, s | 3.13, s | 3.13, s |
| 11 | 2.84, s | 2.84, s | 2.84, d (4.7) | |||
| 1′ | 5.07, d (7.3) | 5.09, d (7.2) | 5.08, d (7.4) | 4.90, d (7.0) | 4.90, d (7.0) | 4.89, d (7.1) |
| 2′ | 5.45, t (7.0) | 5.47, t (6.9) | 5.47, t (6.9) | 5.40, t (7.1) | 5.41, t (7.0) | 5.39, t (7.0) |
| 4′ | 2.08, m | 2.11, m | 2.11, m | 2.07, m | 2.08, m | 2.09, m |
| 5′ | 2.09, m | 2.11, m | 2.12, m | 2.12, m | 2.13, m | 2.13, m |
| 6′ | 5.05, m | 5.08, m | 5.07, m | 5.14, t (6.2) | 5.14, t (6.1) | 5.14, t (6.3) |
| 8′ | 1.98, m | 1.99, m | 1.97, m | 2.01, m | 2.01, m | 2.02, m |
| 9′ | 2.04, m | 2.07, m | 1.55, m | 2.12, m | 2.12, m | 1.56, m |
| 10′ | 5.19, t (6.4) | 5.32, t (6.6) | 2.06, m | 5.26, t (6.8) | 5.30, t (6.3) | 2.55, br t, (7.9) |
| 12′ | 2.99, s | 3.09, s | 6.00, br s | 3.01, s | 3.12, s | 6.08, br s |
| 14′ | 6.07, br s | 6.06, br s | 6.04, br s | 6.17, br s | 6.15, br s | 6.10, br s |
| 16′ | 1.84, s | 1.87, s | 1.87, d (0.9) | 1.86, d (1.1) | 1.87, d (1.1) | 1.86, d (1.0) |
| 17′ | 2.10, s | 2.13, s | 2.13, d (1.0) | 2.09, d (1.1) | 2.09, d (1.0) | 2.12, d (1.0) |
| 18′ | 1.58, s | 1.69, s | 2.15, d (1.2) | 1.61, s | 1.66, d (1.2) | 1.87, d (1.3) |
| 19′ | 1.56, s | 1.58, s | 1.58, s | 1.59, s | 1.60, s | 1.62, s |
| 20′ | 1.77, s | 1.79, s | 1.80, s | 1.79, s | 1.79, s | 1.79, s |
| No. | 1 a | 2 a | 3 a | 4 b | 5 b | 6 b |
|---|---|---|---|---|---|---|
| 2 | 147.7 | 148.1 | 148.1 | 163.3 | 163.3 | 163.3 |
| 4 | 147.3 | 147.2 | 147.2 | 141.8 | 141.8 | 141.8 |
| 5 | 115.0 | 115.2 | 115.2 | 118.2 | 118.2 | 118.2 |
| 6 | 162.0 | 160.4 | 160.3 | 160.9 | 160.9 | 161.0 |
| 8 | 140.3 | 140.7 | 140.8 | 137.6 | 137.6 | 137.6 |
| 10 | 35.0 | 35.5 | 35.5 | 31.9 | 31.9 | 31.9 |
| 11 | 26.1 | 26.1 | 26.3 | |||
| 1′ | 44.5 | 44.6 | 44.7 | 45.2 | 45.2 | 45.2 |
| 2′ | 117.6 | 117.3 | 117.4 | 120.5 | 120.5 | 120.4 |
| 3′ | 143.4 | 143.7 | 143.7 | 142.0 | 142.0 | 142.0 |
| 4′ | 39.4 | 39.5 | 39.4 | 40.1 | 40.2 | 40.2 |
| 5′ | 26.1 | 26.1 | 26.1 | 27.0 | 26.9 | 26.9 |
| 6′ | 123.5 | 123.6 | 123.9 | 124.8 | 124.8 | 124.8 |
| 7′ | 135.5 | 135.5 | 135.4 | 135.8 | 135.8 | 136.0 |
| 8′ | 39.3 | 39.5 | 39.1 | 40.1 | 40.3 | 40.6 |
| 9′ | 26.7 | 26.9 | 25.8 | 27.4 | 27.7 | 27.2 |
| 10′ | 129.0 | 128.2 | 40.8 | 129.4 | 128.7 | 33.8 |
| 11′ | 129.6 | 129.2 | 157.8 | 130.7 | 130.2 | 158.6 |
| 12′ | 55.3 | 47.9 | 125.7 | 55.8 | 48.3 | 126.9 |
| 13′ | 199.3 | 198.6 | 191.7 | 198.8 | 198.1 | 191.0 |
| 14′ | 122.9 | 123.0 | 126.3 | 123.7 | 124.0 | 127.0 |
| 15′ | 155.5 | 155.8 | 154.2 | 155.0 | 155.3 | 154.2 |
| 16′ | 27.7 | 27.7 | 27.7 | 27.5 | 27.5 | 27.6 |
| 17′ | 20.6 | 20.7 | 20.5 | 20.5 | 20.5 | 20.4 |
| 18′ | 16.4 | 24.1 | 19.1 | 16.5 | 24.4 | 25.5 |
| 19′ | 16.0 | 16.0 | 15.9 | 16.1 | 16.1 | 16.0 |
| 20′ | 16.5 | 16.6 | 16.6 | 16.5 | 16.5 | 16.5 |
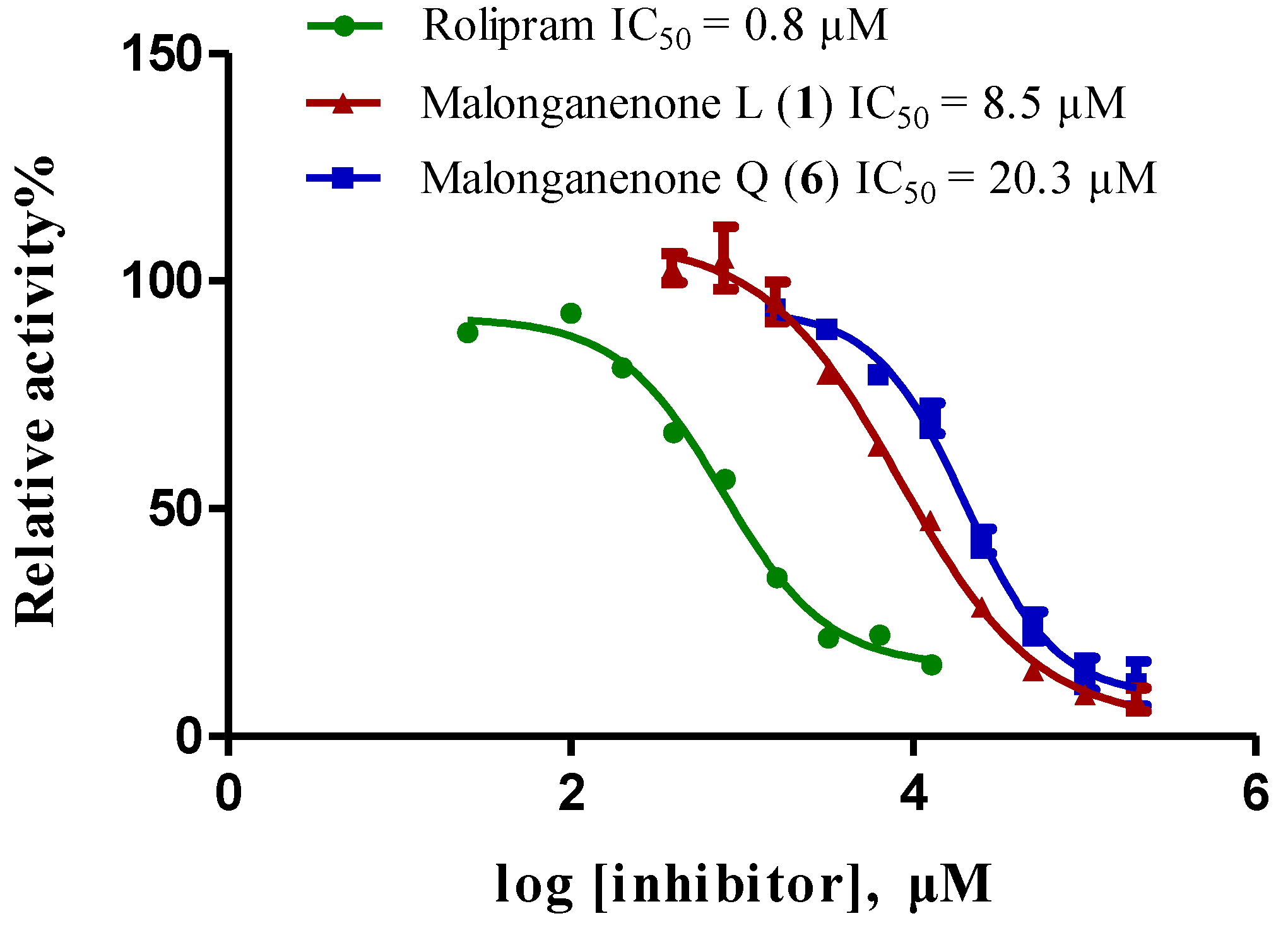
2.2. In Vitro Inhibitory Activity Screening against PDEs
| Compound | Inhibition (%) of Compounds at 50 µM | Inhibition (%) of Compounds at 5 µM | |||||
|---|---|---|---|---|---|---|---|
| PDE4D | PDE5A | PDE9A | PDE4D | PDE5A | PDE9A | ||
| 1 | 85 | 53 | 18 | 17 | 11 | <10 | |
| 2 | 72 | 32 | 15 | <10 | <10 | <10 | |
| 3 | 81 | 35 | 27 | 17 | <10 | 10 | |
| 4 | 79 | 38 | <10 | 18 | <10 | <10 | |
| 5 | 75 | 36 | 11 | 14 | <10 | <10 | |
| 6 | 85 | 38 | 15 | 18 | <10 | <10 | |
3. Experimental Section
3.1. General Experimental Procedures
3.2. Animal Material
3.3. Extraction and Isolation
4. Conclusions
Supplementary Files
Acknowledgments
Conflicts of Interest
References
- Berrue, F.; Kerr, R.G. Diterpenes from gorgonian corals. Nat. Prod. Rep. 2009, 26, 681–710. [Google Scholar] [CrossRef]
- Wang, L.H.; Sheu, J.H.; Kao, S.Y.; Su, J.H.; Chen, Y.H.; Chen, Y.H.; Su, Y.D.; Chang, Y.C.; Fang, L.S.; Wang, W.H.; et al. Natural product chemistry of gorgonian corals of the family Plexauridae distributed in the Indo-Pacific Ocean. Mar. Drugs 2012, 10, 2415–2434. [Google Scholar] [CrossRef]
- Keyzers, R.A.; Gray, C.A.; Schleyer, M.H.; Whibley, C.E.; Hendricks, D.T.; Davies-Coleman, M.T. Malonganenones A–C, novel tetraprenylated alkaloids from the Mozambique gorgonian Leptogorgia gilchristi. Tetrahedron 2006, 62, 2200–2206. [Google Scholar] [CrossRef]
- Sorek, H.; Rudi, A.; Benayahu, Y.; Ben-Califa, N.; Neumann, D.; Kashman, Y. Nuttingins A–F and malonganenones D–H, tetraprenylated alkaloids from the Tanzanian gorgonian Euplexaura nuttingi. J. Nat. Prod. 2007, 70, 1104–1109. [Google Scholar] [CrossRef]
- Zhang, J.R.; Li, P.L.; Tang, X.L.; Qi, X.; Li, G.Q. Cytotoxic tetraprenylated alkaloids from the South China Sea gorgonian Euplexaura robusta. Chem. Biodivers. 2012, 9, 2218–2224. [Google Scholar] [CrossRef]
- Gao, C.; Qi, S.; Luo, X.; Wang, P.; Zhang, S. Sterols from the South China Sea gorgonian Echinogorgia pseudossapo. Tianran Chanwu Yanjiu Yu Kaifa 2011, 23, 1006–1010. (in Chinese). [Google Scholar]
- Xu, W.; Liao, X.; Xu, S.; Liao, L. Isolation and identification of two polyhydroxylated sterols. Guangdong Huagong 2012, 39, 305–306. (in Chinese). [Google Scholar]
- Qou, Y.; Qi, S.; Zhang, S.; Yang, J.; Xiao, Z. New polyoxygenated steroids from the South China Sea gorgonian Echinogorgia aurantiaca. Pharmazie 2006, 61, 645–647. [Google Scholar]
- Zeng, L.; Wu, J. Structure determination of echifloristerol. Zhongshan Daxue Xuebao Ziran Kexueban 1989, 28, 28–31. (in Chinese). [Google Scholar]
- Li, R.; Long, K.; Zhou, Y.; Chen, F. Studies on the Chinese gorgonian. Zhongshan Daxue Xuebao Ziran Kexueban 1982, 4, 65–69. (in Chinese). [Google Scholar]
- Guo, Q.; Wei, Y.; Wang, C.; Shao, C.; Li, L.; Liu, X.; Guo, Q. Study on chemical constituents of Echinogorgia sassapo reticulata (esper). Zhongguo Haiyang Yaowu 2010, 29, 32–35. [Google Scholar]
- Su, J.; Long, K.; Jian, Z. Chemical constituents of Chinese gorgonia. (V). A new marine C29-sterol from Echinogorgia pseudossapo (Koelliker). Zhongshan Daxue Xuebao Ziran Kexueban 1984, 1, 97–101. (in Chinese). [Google Scholar]
- Liao, L.; Liao, X.J.; Xu, S.H. Compounds with nitrogen in coral Echinogorgia sp. Tianran Chanwu Yanjiu Yu Kaifa 2010, 22, 392–394. [Google Scholar]
- Gao, C.H.; Wang, Y.F.; Li, S.; Qian, P.Y.; Qi, S.H. Alkaloids and sesquiterpenes from the South China Sea gorgonian Echinogorgia pseudossapo. Mar. Drugs 2011, 9, 2479–2487. [Google Scholar]
- Tanaka, J.; Miki, H.; Higa, T. Echinofuran, a new furanosesquiterpene from the gorgonian Echinogorgia praelonga. J. Nat. Prod. 1992, 55, 1522–1524. [Google Scholar] [CrossRef]
- Manzo, E.; Ciavatta, M.L.; Lopez, G.M.P.; Gavagnin, M.; Villani, G.; Naik, C.G.; Cimino, G. New bioactive hydrogenated linderazulene-derivatives from the gorgonian Echinogorgia complexa. Tetrahedron Lett. 2007, 48, 2569–2571. [Google Scholar]
- Liao, L.; Wang, N.; Liang, Q.; Liao, X.J.; Xu, S.H. Three ceramides from gorgonian Echinogorgia sp. Zhongcaoyao 2010, 41, 851–854. (in Chinese). [Google Scholar]
- Tillekeratne, L.M.V.; de Silva, E.D.; Mahindaratne, M.P.D.; Schmitz, F.J.; Gunasekera, S.P.; Alderslade, P. Xanthyletin and xanthoxyletin from a gorgonian, Echinogorgia sp. J. Nat. Prod. 1989, 52, 1303–1304. [Google Scholar] [CrossRef]
- Han, Q.H.; Fan, C.Q.; Lu, Y.N.; Wu, J.P.; Liu, X.; Yin, S. Chemical constituents from the ascidian Aplidium constellatum. Biochem. Syst. Ecol. 2013, 48, 6–8. [Google Scholar] [CrossRef]
- Chen, S.K.; Zhao, P.; Shao, Y.X.; Li, Z.; Zhang, C.; Liu, P.; He, X.; Luo, H.B.; Hu, X. Moracin M from Morus alba L. is a natural phosphodiesterase-4 inhibitor. Bioorg. Med. Chem. Lett. 2012, 22, 3261–3264. [Google Scholar] [CrossRef]
- Li, Z.; Cai, Y.H.; Cheng, Y.K.; Lu, X.; Shao, Y.X.; Li, X.; Liu, M.; Liu, P.; Luo, H.B. Identification of novel phosphodiesterase-4D inhibitors prescreened by molecular dynamics-augmented modeling and validated by bioassay. J. Chem. Inf. Model. 2013, 53, 972–981. [Google Scholar] [CrossRef]
- Zhao, P.; Chen, S.K.; Cai, Y.H.; Lu, X.; Li, Z.; Cheng, Y.K.; Zhang, C.; Hu, X.; He, X.; Luo, H.B. The molecular basis for the inhibition of phosphodiesterase-4D by three natural resveratrol analogs. Isolation, molecular docking, molecular dynamics simulations, binding free energy, and bioassay. Biochim. Biophys. Acta 2013, 1834, 2089–2096. [Google Scholar] [CrossRef]
- Zhong, J.; Huang, Y.; Ding, W.; Wu, X.; Wan, J.; Luo, H. Chemical constituents of Aloe barbadensis Miller and their inhibitory effects on phosphodiesterase-4D. Fitoterapia 2013, 91, 159–165. [Google Scholar] [CrossRef]
- Burgin, A.B.; Magnusson, O.T.; Singh, J.; Witte, P.; Staker, B.L.; Bjornsson, J.M.; Thorsteinsdottir, M.; Hrafnsdottir, S.; Hagen, T.; Kiselyov, A.S.; et al. Design of phosphodiesterase 4D (PDE4D) allosteric modulators for enhancing cognition with improved safety. Nat. Biotechnol. 2010, 28, 63–72. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sun, Z.-H.; Cai, Y.-H.; Fan, C.-Q.; Tang, G.-H.; Luo, H.-B.; Yin, S. Six New Tetraprenylated Alkaloids from the South China Sea Gorgonian Echinogorgia pseudossapo. Mar. Drugs 2014, 12, 672-681. https://doi.org/10.3390/md12020672
Sun Z-H, Cai Y-H, Fan C-Q, Tang G-H, Luo H-B, Yin S. Six New Tetraprenylated Alkaloids from the South China Sea Gorgonian Echinogorgia pseudossapo. Marine Drugs. 2014; 12(2):672-681. https://doi.org/10.3390/md12020672
Chicago/Turabian StyleSun, Zhang-Hua, Ying-Hong Cai, Cheng-Qi Fan, Gui-Hua Tang, Hai-Bin Luo, and Sheng Yin. 2014. "Six New Tetraprenylated Alkaloids from the South China Sea Gorgonian Echinogorgia pseudossapo" Marine Drugs 12, no. 2: 672-681. https://doi.org/10.3390/md12020672