Influence of Lipid A Acylation Pattern on Membrane Permeability and Innate Immune Stimulation
Abstract
:1. Introduction
2. Results and Discussion
2.1. Structure Analysis of Lipid A Extracted from Six E. coli Strains
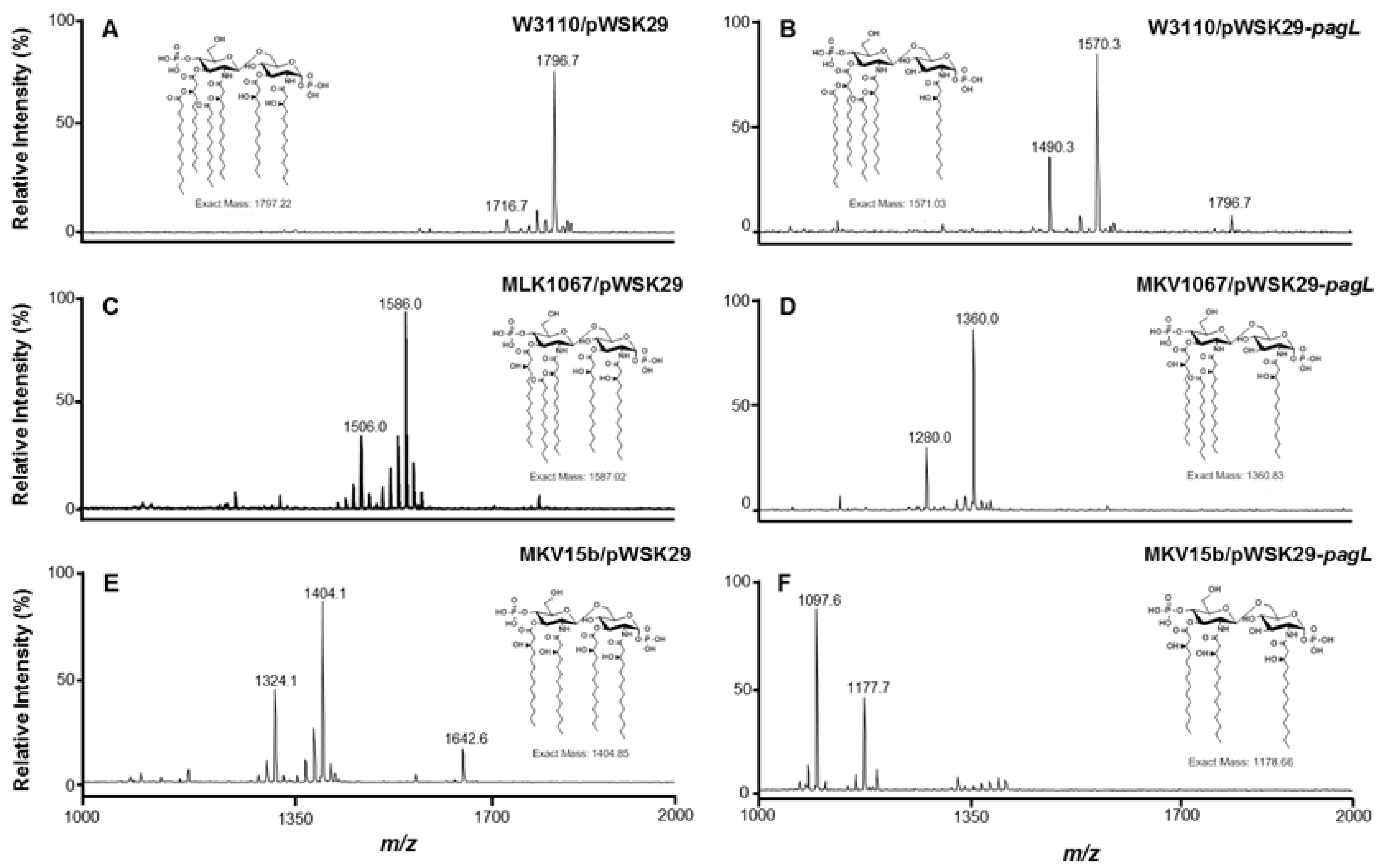
2.2. Effects of Lipid A Acylation Pattern on the Membrane Permeability of Different E. coli Strains
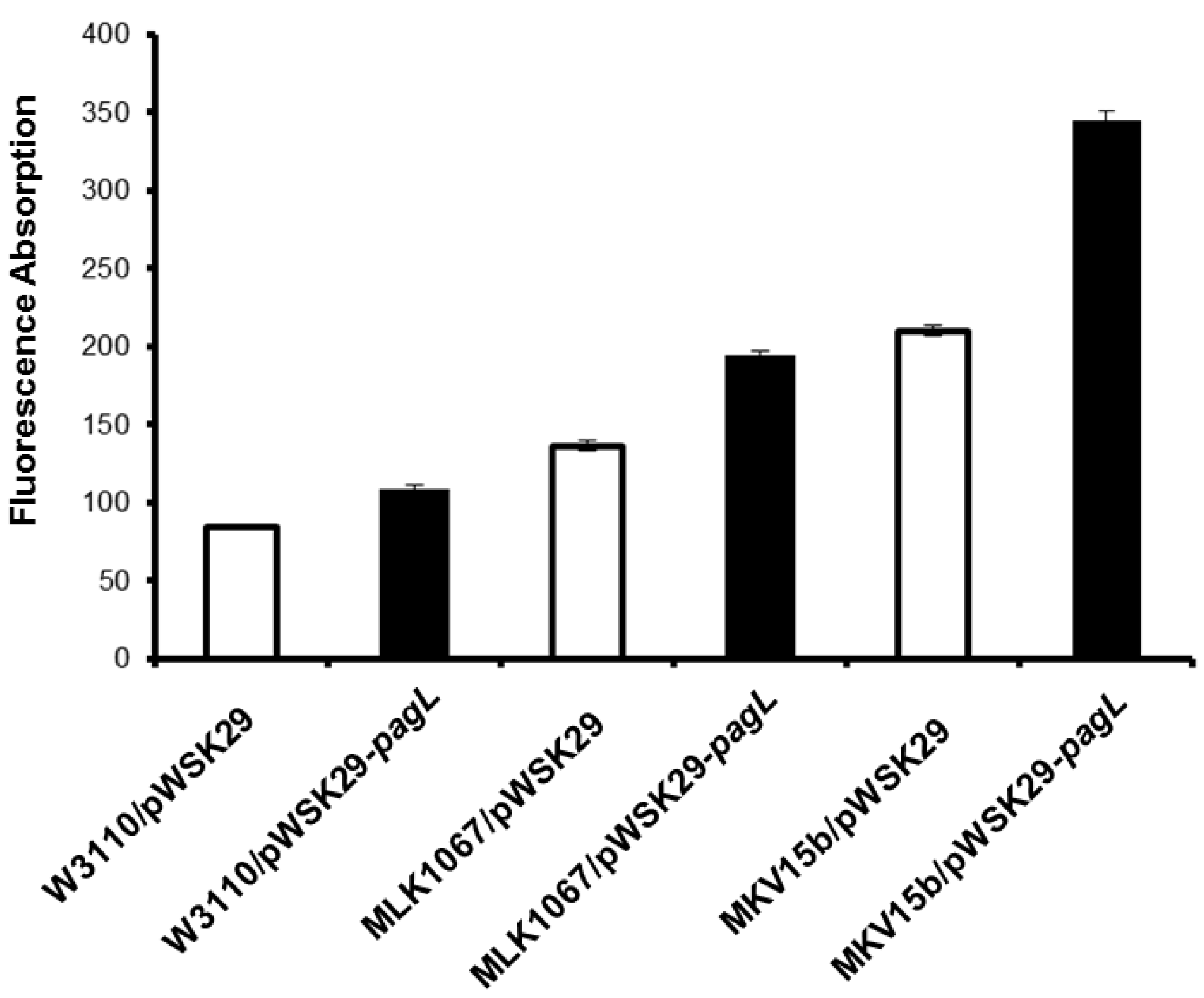
2.3. Effects of Lipid A Acylation Pattern on Innate Immune System Recognition
| Antibiotics | W3110/pWSK29 | W3110/pWSK29-pagL | MLK1067/pWSK29 | MLK1067/pWSK29-pagL | MKV15b/pWSK29 | MKV15b/pWSK29-pagL |
|---|---|---|---|---|---|---|
| Trimethoprim | 25 | 24 | 26 | 25 | 28 | 30 |
| Gentamicin | 23 | 23 | 24 | 26 | 25 | 28 |
| Cefotaxime | 34 | 34 | 35 | 36 | 36 | 38 |
| Imipenem | 28 | 29 | 31 | 32 | 32 | 35 |
| Erythromycin | 8 | 10 | 16 | 16 | 17 | 18 |
| Aztreonam | 34 | 34 | 34 | 34 | 36 | 37 |
| Minocycline | 16 | 16 | 17 | 18 | 18 | 18 |
| Polymyxin B | 17 | 17 | 18 | 18 | 18 | 20 |
| Vancomycin | 6 | 6 | 6 | 6 | 7 | 8 |
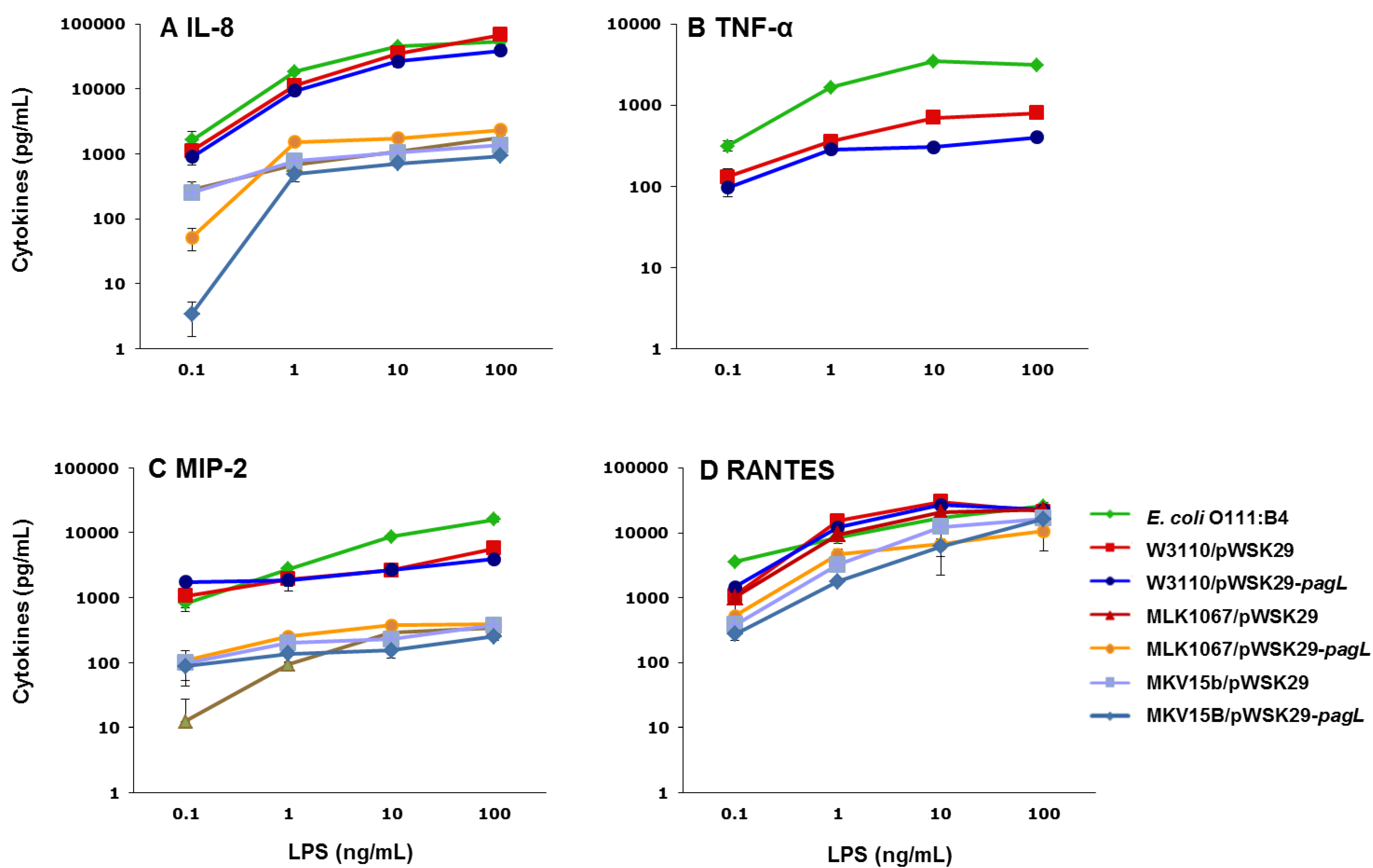
3. Experimental Section
3.1. Bacterial Strains, Plasmid and Growth Conditions
| Strains or plasmids | Relevant characteristics | Reference or source |
|---|---|---|
| Strains | ||
| DH5α | E. coli component cells | Invitrogen |
| E. coli W3110 | E. coli F−rph-1 IN(rrnD–rrnE)1 λ− | [19] |
| E. coli MLK1067 | E. coli W3110 msbB::Ωcam | [19] |
| E. coli MKV15b | E. coli W3110 lpxM::Ωcam lpxP::Ωkan lpxL::Tn10 | [20,21] |
| W3110/pWSK29 | W3110 harboring pWSK29 | This work |
| MLK1067/pWSK29 | MLK1067 harboring pWSK29 | This work |
| MKV15b/pWSK29 | MKV15b harboring pWSK29 | This work |
| W3110/pWSK29-pagL | W3110 harboring pWSK29-pagL | This work |
| MLK1067/pWSK29-pagL | MLK1067 harboring pWSK29-pagL | This work |
| MKV15b/pWSK29-pagL | MKV15b harboring pWSK29-pagL | This work |
| Plasmids | ||
| pWSK29 | Low copy vector, Ampr | [25] |
| pWSK29-pagL | pWSK29 containing pagL | This work |
3.2. LPS and Lipid A Isolation
3.3. Mass Spectrometry Procedures
3.4. Outer Membrane NPN Permeability Assay
3.5. Disk Diffusion Assay
3.6. THP-1 Cell Stimulations and Measurement of Cytokines
3.7. Stimulation of Mouse MH-S Alveolar Macrophages
3.8. Measurement of Cytokines
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Nikaido, H.; Vaara, M.C. Molecular basis of bacterial outer membrane permeability. Microbiol. Rev. 1985, 49, 1–32. [Google Scholar]
- Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar]
- Raetz, C.R.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Raetz, C.R.; Reynolds, C.M.; Trent, M.S.; Bishop, R.E. Lipid A modification systems in gram-negative bacteria. Annu. Rev. Biochem. 2007, 76, 295–329. [Google Scholar] [CrossRef]
- Raetz, C.R.; Garrett, T.A.; Reynolds, C.M.; Shaw, W.A.; Moore, J.D.; Smith, D.C., Jr.; Ribeiro, A.A.; Murphy, R.C.; Ulevitch, R.J.; Fearns, C.; et al. Kdo2-Lipid A of Escherichia coli, a defined endotoxin that activates macrophages via TLR-4. J. Lipid Res. 2006, 47, 1097–1111. [Google Scholar] [CrossRef]
- Wang, X.; Ribeiro, A.A.; Guan, Z.; Raetz, C.R. Identification of undecaprenyl phosphate-beta-d-galactosamine in Francisella novicida and its function in lipid A modification. Biochemistry 2009, 48, 1162–1172. [Google Scholar] [CrossRef]
- Li, Y.; Powell, D.A.; Shaffer, S.A.; Rasko, D.A.; Pelletier, M.R.; Leszyk, J.D.; Scott, A.J.; Masoudi, A.; Goodlett, D.R.; Wang, X.; et al. LPS remodeling is an evolved survival strategy for bacteria. Proc. Natl. Acad. Sci. USA 2012, 109, 8716–8721. [Google Scholar] [CrossRef]
- Rebeil, R.; Ernst, R.K.; Jarrett, C.O.; Adams, K.N.; Miller, S.I.; Hinnebusch, B.J. Characterization of late acyltransferase genes of Yersinia pestis and their role in temperature-dependent lipid A variation. J. Bacteriol. 2006, 188, 1381–1388. [Google Scholar] [CrossRef]
- Hajjar, A.M.; Ernst, R.K.; Fortuno, E.S., III; Brasfield, A.S.; Yam, C.S.; Newlon, L.A.; Kollmann, T.R.; Miller, S.I.; Wilson, C.B. Humanized TLR4/MD-2 mice reveal LPS recognition differentially impacts susceptibility to Yersinia pestis and Salmonella enterica. PLoS Pathog. 2012, 8. [Google Scholar] [CrossRef]
- Guo, L.; Lim, K.B.; Gunn, J.S.; Bainbridge, B.; Darveau, R.P.; Hackett, M.; Miller, S.I. Regulation of lipid A modifications by Salmonella typhimurium virulence genes phoP-phoQ. Science 1997, 276, 250–253. [Google Scholar] [CrossRef]
- Groisman, E.A. The pleiotropic two-component regulatory system PhoP-PhoQ. J. Bacteriol. 2001, 183, 1835–1842. [Google Scholar] [CrossRef]
- Gunn, J.S.; Ryan, S.S.; van Velkinburgh, J.C.; Ernst, R.K.; Miller, S.I. Genetic and functional analysis of a PmrA-PmrB-regulated locus necessary for lipopolysaccharide modification, antimicrobial peptide resistance, and oral virulence of Salmonella enterica serovar typhimurium. Infect. Immun. 2000, 68, 6139–6146. [Google Scholar] [CrossRef]
- Trent, M.S.; Pabich, W.; Raetz, C.R.; Miller, S.I. A PhoP/PhoQ-induced lipase (PagL) that catalyzes 3-O-deacylation of lipid A precursors in membranes of Salmonella typhimurium. J. Biol. Chem. 2001, 276, 9083–9092. [Google Scholar] [CrossRef]
- Kawasaki, K.; Ernst, R.K.; Miller, S.I. 3-O-Deacylation of lipid A by PagL, a PhoP/PhoQ-regulated deacylase of Salmonella typhimurium, modulates signaling through Toll-like receptor 4. J. Biol. Chem. 2004, 279, 20044–20048. [Google Scholar] [CrossRef]
- Casella, C.R.; Mitchell, T.C. Putting endotoxin to work for us: Monophosphoryl lipid A as a safe and effective vaccine adjuvant. Cell Mol. Life Sci. 2008, 65, 3231–3240. [Google Scholar] [CrossRef]
- Needham, B.D.; Carroll, S.M.; Giles, D.K.; Georgiou, G.; Whiteley, M.; Trent, M.S. Modulating the innate immune response by combinatorial engineering of endotoxin. Proc. Natl. Acad. Sci. USA 2013, 110, 1464–1469. [Google Scholar] [CrossRef]
- Mata-Haro, V.; Cekic, C.; Martin, M.; Chilton, P.M.; Casella, C.R.; Mitchell, T.C. The vaccine adjuvant monophosphoryl lipid A as a TRIF-biased agonist of TLR4. Science 2007, 316, 1628–1632. [Google Scholar] [CrossRef]
- Miller, S.I.; Ernst, R.K.; Bader, M.W. LPS, TLR4 and infectious disease diversity. Nat. Rev. Microbiol. 2005, 3, 36–46. [Google Scholar] [CrossRef]
- Karow, M.; Georgopoulos, C. Isolation and characterization of the Escherichia coli msbB gene, a multicopy suppressor of null mutations in the high-temperature requirement gene htrB. J. Bacteriol. 1992, 174, 702–710. [Google Scholar]
- Vorachek-Warren, M.K.; Ramirez, S.; Cotter, R.J.; Raetz, C.R. A triple mutant of Escherichia coli lacking secondary acyl chains on lipid A. J. Biol. Chem. 2002, 277, 14194–14205. [Google Scholar] [CrossRef]
- Vuorio, R.; Vaara, M. Comparison of the phenotypes of the lpxA and lpxD mutants of Escherichia coli. FEMS Microbiol. Lett. 1995, 134, 227–232. [Google Scholar] [CrossRef]
- Vaara, M.; Nurminen, M. Outer membrane permeability barrier in Escherichia coli mutants that are defective in the late acyltransferases of lipid A biosynthesis. Antimicrob. Agents Chemother. 1999, 43, 1459–1462. [Google Scholar]
- Park, B.S.; Song, D.H.; Kim, H.M.; Choi, B.S.; Lee, H.; Lee, J.O. The structural basis of lipopolysaccharide recognition by the TLR4-MD-2 complex. Nature 2009, 458, 1191–1195. [Google Scholar] [CrossRef]
- Folch, J.; Lees, M.; Sloane-Stanley, G.H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 1957, 226, 497–509. [Google Scholar]
- Hirschfeld, M.; Ma, Y.; Weis, J.H.; Vogel, S.N.; Weis, J.J. Cutting edge: Repurification of lipopolysaccharide eliminates signaling through both human and murine toll-like receptor 2. J. Immunol. 2000, 165, 618–622. [Google Scholar]
- Caroff, M.; Tacken, A.; Szabo, L. Detergent-accelerated hydrolysis of bacterial endotoxins and determination of the anomeric configuration of the glycosyl phosphate present in the “isolated lipid A” fragment of the Bordetella pertussis endotoxin. Carbohydr. Res. 1988, 175, 273–282. [Google Scholar] [CrossRef]
- Li, Y.; Wang, X.; Ernst, R.K. A rapid one-step method for the characterization of membrane lipid remodeling in Francisella using matrix-assisted laser desorption ionization time-of-flight tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2011, 25, 2641–2648. [Google Scholar]
- Helander, I.M.; Mattila-Sandholm, T. Fluorometric assessment of gram-negative bacterial permeabilization. J. Appl. Microbiol. 2000, 88, 213–219. [Google Scholar] [CrossRef]
- Ernst, R.K.; Guina, T.; Miller, S.I. How intracellular bacteria survive: Surface modifications that promote resistance to host innate immune responses. J. Infect. Dis. 1999, 179, S326–S330. [Google Scholar]
- Wang, R.F.; Kushner, S.R. Construction of versatile low-copy-number vectors for cloning, sequencing and gene expression in Escherichia coli. Gene 1991, 100, 195–199. [Google Scholar] [CrossRef]
- Westphal, O.; Jann, K. Bacterial lipopolysaccharides: Extraction with phenol-water and further applications of the procedure. Methods Carbohydr. Chem. 1965, 5, 83–91. [Google Scholar]
- Six, D.; Carty, S.M.; Guan, Z.; Raetz, C.R. Purification and mutagenesis of LpxL, the lauroyltransferase of Escherichia coli lipid A biosynthesis. Biochemistry 2008, 47, 8623–8637. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, Y.; Wang, Z.; Chen, J.; Ernst, R.K.; Wang, X. Influence of Lipid A Acylation Pattern on Membrane Permeability and Innate Immune Stimulation. Mar. Drugs 2013, 11, 3197-3208. https://doi.org/10.3390/md11093197
Li Y, Wang Z, Chen J, Ernst RK, Wang X. Influence of Lipid A Acylation Pattern on Membrane Permeability and Innate Immune Stimulation. Marine Drugs. 2013; 11(9):3197-3208. https://doi.org/10.3390/md11093197
Chicago/Turabian StyleLi, Yanyan, Zhou Wang, Jiuzhou Chen, Robert K. Ernst, and Xiaoyuan Wang. 2013. "Influence of Lipid A Acylation Pattern on Membrane Permeability and Innate Immune Stimulation" Marine Drugs 11, no. 9: 3197-3208. https://doi.org/10.3390/md11093197




