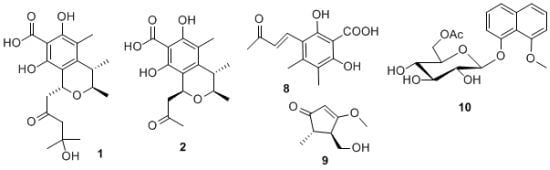
Polyketides from a Marine-Derived Fungus Xylariaceae sp.
Abstract
:1. Introduction
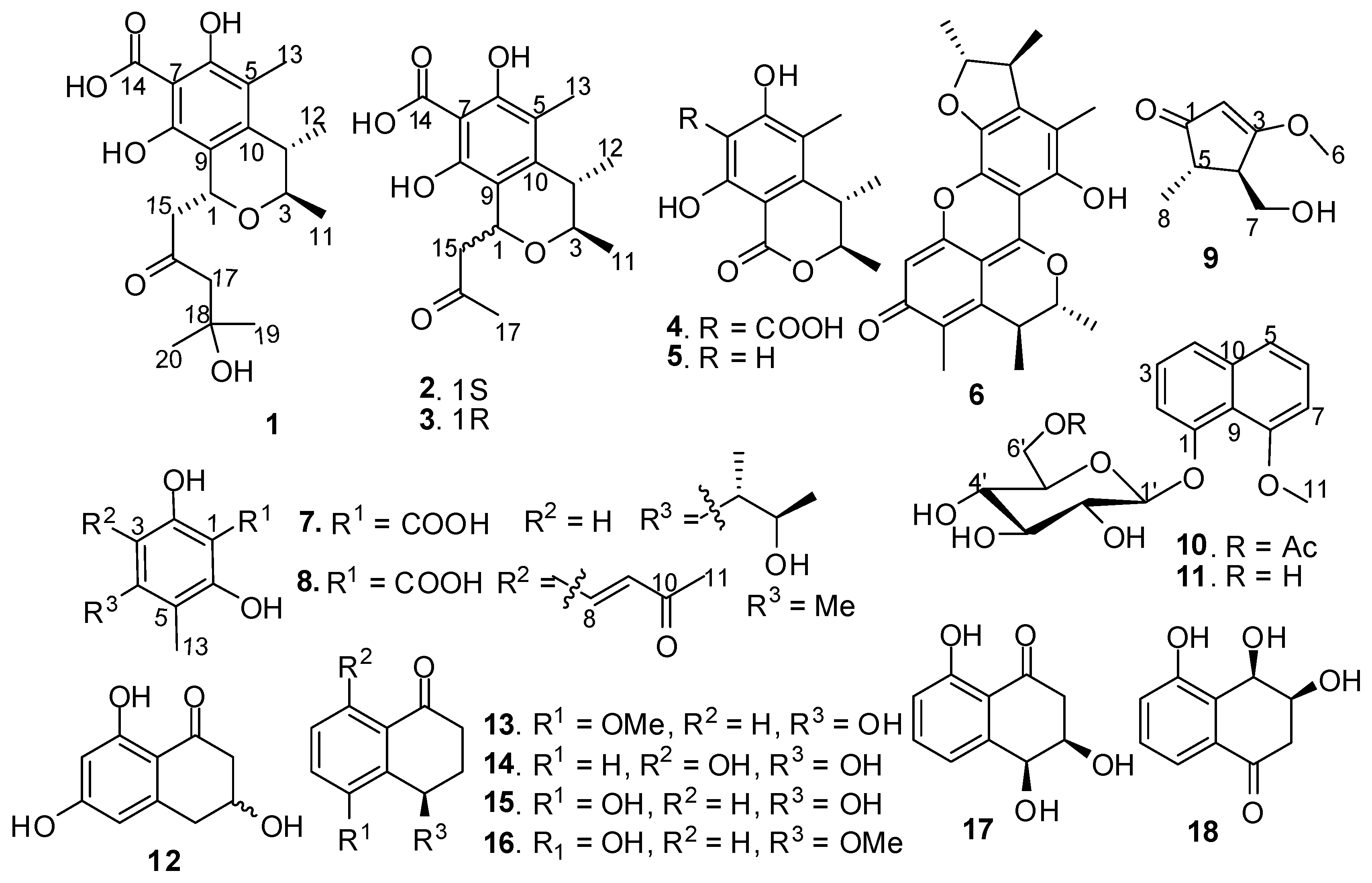
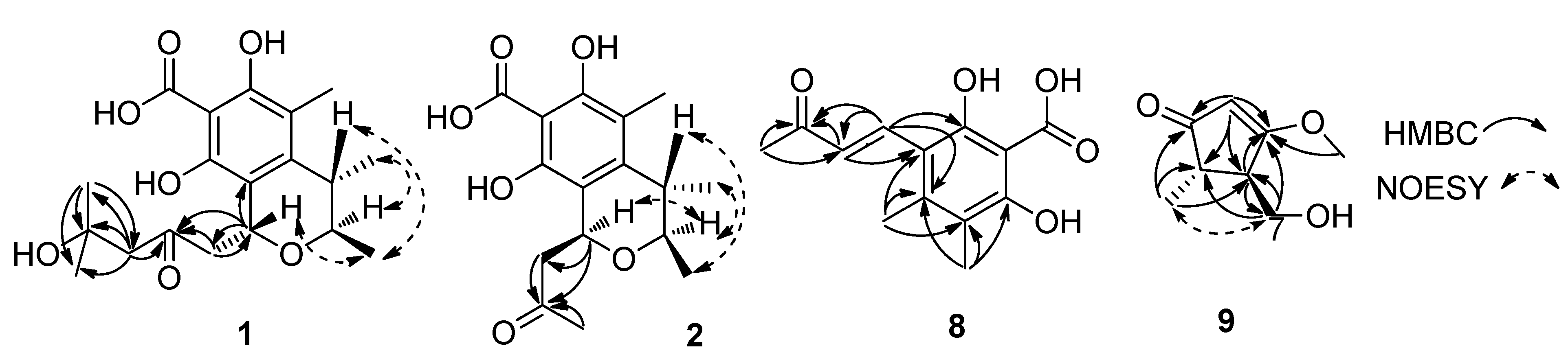
2. Results and Discussion
3. Experimental Section
3.1. General Experimental Procedures
3.2. Fungal Material
3.3. Fermentation and Extraction
3.4. Isolation and Purification
3.5. Enzyme-Inhibitory Activity Assays
3.6. Larval Settlement Assays
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Du, L.; Zhu, T.; Fang, Y.; Liu, H.; Gu, Q.; Zhu, W. Aspergiolide A, a novel anthraquinone derivative with naphtho 1,2,3-de chromene-2,7-dione skeleton isolated from a marine-derived fungus Aspergillus glaucus. Tetrahedron 2007, 63, 1085–1088. [Google Scholar] [CrossRef]
- Ishino, M.; Kiyomichi, N.; Takatori, K.; Sugita, T.; Shiro, M.; Kinoshita, K.; Takahashi, K.; Koyama, K. Phomactin I, 13-epi-Phomactin I, and Phomactin J, three novel diterpenes from a marine-derived fungus. Tetrahedron 2010, 66, 2594–2597. [Google Scholar] [CrossRef]
- Shao, C.L.; Wang, C.Y.; Gu, Y.C.; Wei, M.Y.; Pan, J.H.; Deng, D.S.; She, Z.G.; Lin, Y.C. Penicinoline, a new pyrrolyl 4-quinolinone alkaloid with an unprecedented ring system from an endophytic fungus Penicillium sp. Bioorg. Med. Chem. Lett. 2010, 20, 3284–3286. [Google Scholar] [CrossRef]
- Shao, C.L.; Wu, H.X.; Wang, C.Y.; Liu, Q.A.; Xu, Y.; Wei, M.Y.; Qian, P.Y.; Gu, Y.C.; Zheng, C.J.; She, Z.G.; et al. Potent antifouling resorcylic acid lactones from the gorgonian-derived fungus Cochliobolus lunatus. J. Nat. Prod. 2011, 74, 629–633. [Google Scholar] [CrossRef]
- Wei, M.Y.; Wang, C.Y.; Liu, Q.A.; Shao, C.L.; She, Z.G.; Lin, Y.C. Five sesquiterpenoids from a marine-derived fungus Aspergillus sp. isolated from a gorgonian Dichotella gemmacea. Mar. Drugs 2010, 8, 941–949. [Google Scholar]
- Xu, L.; Xue, J.; Xu, H.; Liu, X.; Ma, W.; Wei, X. Three new isochromans from the mycelial culture of a Cylindrocarpon fungus. Heterocycles 2006, 68, 1955–1959. [Google Scholar] [CrossRef]
- Xin, Z.H.; Li, T.; Zhu, T.J.; Wang, W.L.; Du, L.; Fang, Y.C.; Gu, Q.Q.; Zhu, W.M. Isocoumarin derivatives from the sea squirt-derived fungus Penicillium stoloniferum QY2-10 and the halotolerant fungus Penicillium notatum B-52. Arch. Pharm. Res. 2007, 30, 816–819. [Google Scholar] [CrossRef]
- Wakana, D.; Hosoe, T.; Itabashi, T.; Okada, K.; de Campos Takaki, G.M.; Yaguchi, T.; Fukushima, K.; Kawai, K.-I. New citrinin derivatives isolated from Penicillium citrinum. J. Nat. Med. 2006, 60, 279–284. [Google Scholar]
- Clark, B.R.; Capon, R.J.; Lacey, E.; Tennant, S.; Gill, J.H. Citrinin revisited: From monomers to dimers and beyond. Org. Biomol. Chem. 2006, 4, 1520–1528. [Google Scholar] [CrossRef]
- Cameron, D.W.; Craik, J.C.A. Colouring matters of the aphididae. Part XXXVI. The configuration of the glucoside linkage in protoaphins. J. Chem. Soc. C 1968. [Google Scholar] [CrossRef]
- Sankawa, U.; Shimada, H.; Sato, T.; Kinoshita, T.; Yamasaki, K. Biosynthesis of scytalone. Chem. Pharm. Bull. 1981, 29, 3536–3542. [Google Scholar] [CrossRef]
- Yamamoto, K.; Hatano, H.; Arai, M.; Shiomi, K.; Tomoda, H.; Omura, S. Structure elucidation of new monordens produced by Humicola sp FO-2942. J. Antibiot. 2003, 56, 533–538. [Google Scholar] [CrossRef]
- Inacio, M.L.; Silva, G.H.; Teles, H.L.; Trevisan, H.C.; Cavalheiro, A.J.; Bolzani, V.D.S.; Young, M.C.M.; Pfenning, L.H.; Araujo, A.R. Antifungal metabolites from Colletotrichum gloeosporioides, an endophytic fungus in Cryptocarya mandioccana Nees (Lauraceae). Biochem. Syst. Ecol. 2006, 34, 822–824. [Google Scholar] [CrossRef]
- Kokubun, T.; Veitch, N.C.; Bridge, P.D.; Simmonds, M.S.J. Dihydroisocoumarins and a tetralone from Cytospora eucalypticola. Phytochemistry 2003, 62, 779–782. [Google Scholar] [CrossRef]
- Cimmino, A.; Villegas-Fernandez, A.M.; Andolfi, A.; Melck, D.; Rubiales, D.; Evidente, A. Botrytone, a new naphthalenone pentaketide produced by Botrytis fabae, the causal agent of chocolate spot disease on Vicia faba. J. Agric. Food Chem. 2011, 59, 9201–9206. [Google Scholar] [CrossRef]
- Machida, K.; Matsuoka, E.; Kasahara, T.; Kikuchi, M. Studies on the constituents of Juglans species. I. Structural determination of (4S)- and (4R)-4-hydroxy-alpha-tetralone derivatives from the fruit of Juglans mandshurica MAXIM. var. sieboldiana MAKINO. Chem. Pharm. Bull. 2005, 53, 934–937. [Google Scholar]
- Krohn, K.; Biele, C.; Drogies, K.H.; Steingrover, K.; Aust, H.J.; Draeger, S.; Schulz, B. Fusidilactones, a new group of polycyclic lactones from an endophyte, Fusidium sp. Eur. J. Org. Chem. 2002, 14, 2331–2336. [Google Scholar]
- Yan, S.; Li, S.; Wu, W.; Zhao, F.; Bao, L.; Ding, R.; Gao, H.; Wen, H.A.; Song, F.; Liu, H.W. Terpenoid and phenolic metabolites from the fungus Xylaria sp associated with Termite Nests. Chem. Biodivers. 2011, 8, 1689–1700. [Google Scholar] [CrossRef]
- Lu, Z.Y.; Lin, Z.J.; Wang, W.L.; Du, L.; Zhu, T.J.; Fang, Y.C.; Gu, Q.Q.; Zhu, W.M. Citrinin dimers from the halotolerant fungus Penicillium citrinum B-57. J. Nat. Prod. 2008, 71, 543–546. [Google Scholar] [CrossRef]
- Gerwick, W.H.; Nagle, D.G.; Proteau, P.J. Oxylipins from marine-invertebrates. Top. Curr. Chem. 1993, 167, 117–180. [Google Scholar] [CrossRef]
- Orsini, F.; Pelizzoni, F.; Verotta, L.; Aburjai, T.; Rogers, C.B. Isolation, synthesis, and antiplatelet aggregation activity of resveratrol 3-O-beta-d-glucopyranoside and related compounds. J. Nat. Prod. 1997, 60, 1082–1087. [Google Scholar] [CrossRef]
- Pittayakhajonwut, P.; Suvannakad, R.; Thienhirun, S.; Prabpai, S.; Kongsaeree, P.; Tanticharoen, M. An anti-herpes simplex virus-type 1 agent from Xylaria mellisii (BCC 1005). Tetrahedron Lett. 2005, 46, 1341–1344. [Google Scholar] [CrossRef]
- Jimenez-Teja, D.; Daoubi, M.; Collado, I.G.; Hernandez-Galan, R. Lipase-catalyzed resolution of 5-acetoxy-1,2-dihydroxy-1,2,3,4-tetrahydronaphthalene. Application to the synthesis of (+)-(3R,4S)-cis-4-hydroxy-6-deoxyscytalone, a metabolite isolated from Colletotrichum acutatum. Tetrahedron 2009, 65, 3392–3396. [Google Scholar] [CrossRef]
- Qian, P.Y.; Xu, Y.; Fusetani, N. Natural products as antifouling compounds: Recent progress and future perspectives. Biofouling 2010, 26, 223–234. [Google Scholar] [CrossRef]
- Halangk, W.; Lerch, M.M.; Brandt-Nedelev, B.; Roth, W.; Ruthenbuerger, M.; Reinheckel, T.; Domschke, W.; Lippert, H.; Peters, C.; Deussing, J. Role of cathepsin B in intracellular trypsinogen activation and the onset of acute pancreatitis. J. Clin. Invest. 2000, 106, 773–781. [Google Scholar] [CrossRef]
- Nimmesgern, E.; Black, J.; Futer, O.; Fulghum, J.R.; Chambers, S.P.; Brummel, C.L.; Raybuck, S.A.; Sintchak, M.D. Biochemical analysis of the modular enzyme inosine 5′-monophosphate dehydrogenase. Protein Expr. Purif. 1999, 17, 282–289. [Google Scholar] [CrossRef]
- Montalibet, J.; Skorey, K.I.; Kennedy, B.P. Protein tyrosine phosphatase: Enzymatic assays. Methods 2005, 35, 2–8. [Google Scholar] [CrossRef]
- Nakajima, K.; Powers, J.C.; Ashe, B.M.; Zimmerman, M. Mapping the extendedsubstrate binding-site of cathepsing and human-leukocyte elastase—Studies with peptide-substrates related to the alpha-1-protease inhibitor reactive site. J. Biol. Chem. 1979, 254, 4027–4032. [Google Scholar]
- Qi, S.H.; Xu, Y.; Xiong, H.R.; Qian, P.Y.; Zhang, S. Antifouling and antibacterial compounds from a marine fungus Cladosporium sp F14. World J. Microbiol. Biotechnol. 2009, 25, 399–406. [Google Scholar] [CrossRef]
Supplementary Files
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Nong, X.-H.; Zheng, Z.-H.; Zhang, X.-Y.; Lu, X.-H.; Qi, S.-H. Polyketides from a Marine-Derived Fungus Xylariaceae sp. Mar. Drugs 2013, 11, 1718-1727. https://doi.org/10.3390/md11051718
Nong X-H, Zheng Z-H, Zhang X-Y, Lu X-H, Qi S-H. Polyketides from a Marine-Derived Fungus Xylariaceae sp. Marine Drugs. 2013; 11(5):1718-1727. https://doi.org/10.3390/md11051718
Chicago/Turabian StyleNong, Xu-Hua, Zhi-Hui Zheng, Xiao-Yong Zhang, Xin-Hua Lu, and Shu-Hua Qi. 2013. "Polyketides from a Marine-Derived Fungus Xylariaceae sp." Marine Drugs 11, no. 5: 1718-1727. https://doi.org/10.3390/md11051718
APA StyleNong, X.-H., Zheng, Z.-H., Zhang, X.-Y., Lu, X.-H., & Qi, S.-H. (2013). Polyketides from a Marine-Derived Fungus Xylariaceae sp. Marine Drugs, 11(5), 1718-1727. https://doi.org/10.3390/md11051718