Glycosylation of Conotoxins
Abstract
:1. Introduction

2. The ConoServer Database
3. General Structural Data of Mucin-Type Glycoconjugate O-Glycans
| α-D-GalpNAc-(1→O) | core 0 |
| β-D-Galp-(1→3)-α-D-GalpNAc-(1→O) | core 1 |
| β-D-GlcpNAc-(1→6)-[β-D-Galp-(1→3)-]α-D-GalpNAc-(1→O) | core 2 |
| β-D-GlcpNAc-(1→3)-α-D-GalpNAc-(1→O) | core 3 |
| β-D-GlcpNAc-(1→6)-[β-D-GlcpNAc-(1→3)-]α-D-GalpNAc-(1→O) | core 4 |
| α-D-GalpNAc-(1→3)-α-D-GalpNAc-(1→O) | core 5 |
| β-D-GlcpNAc-(1→6)-α-D-GalpNAc-(1→O) | core 6 |
| α-D-GalpNAc-(1→6)-α-D-GalpNAc-(1→O) | core 7 |
| α-D-Galp-(1→3)-α-D-GalpNAc-(1→O) | core 8 |
| α-D-GlcpNAc-(1→6)-[β-D-Galp-(1→3)-]α-D-GalpNAc-(1→O) | core 9 |
4. Glycosylated Conotoxins
4.1. Conus striatus
| Conus species | Diet | Conotoxin name(s) | Glycopeptide sequence | O-linkedresidue | O-glycan | References |
|---|---|---|---|---|---|---|
| C. striatus | P | κA-SIVA | ZKSLVPS*VITTCCGYDOGTMCOOCRCTNSC-NH2 | Ser-7 | Hex3HexNAc2 | [53,54,55,56,57] |
| (s4a) | ||||||
| C. striatus | P | κA-SIVB | ZKELVPS*VITTCCGYDOGTMCOOCRCTNSCOTKOKKO-NH2 | Ser-7 | Hex3HexNAc2 | [53,56,57] |
| (s4b) | ||||||
| C. stercusmuscarum | P | SmIVA | ZTWLVPS*T*ITTCCGYDOGTMCOTCMCDNTCKOKOKKS-NH2 | Ser-7 | No information | [57] |
| (κA-SmIVA) | Thr-8 | No information | ||||
| C. stercusmuscarum | P | SmIVB | AOWLVPS*T*ITTCCGYDOGSMCOOCMCNNTCKOKOKKS-NH2 | Ser-7 | No information | [57] |
| (κA-SmIVB) | Thr-8 | No information | ||||
| C. consors | P | CcTx | AOWLVPS*QITTCCGYNOGTMCOSCMCTNTC | Ser-7 | Hex2HexNAc2 | [10,11,15,52,58] |
| (κA-CcTx) | Gal3GlcNAcGalNAc | |||||
| C. magus | P | κA-MIVA | AOγLVVT*AT*TNCCGYNOMTICOOCMCTYSCOOKRKO-NH2 | Thr-7 | Hex4HexNAc2 as sum of both sites | [57] |
| Thr-9 | ||||||
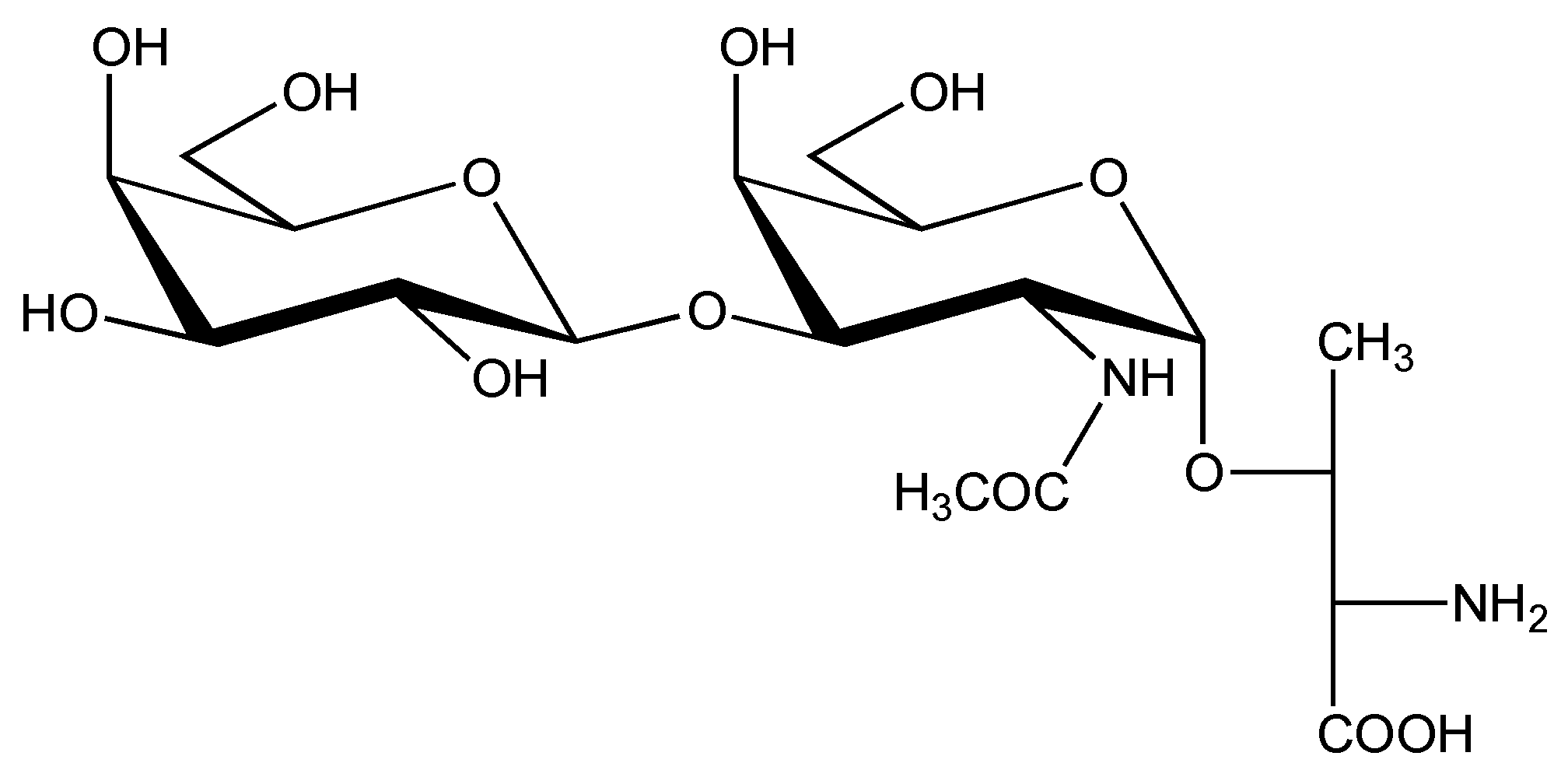
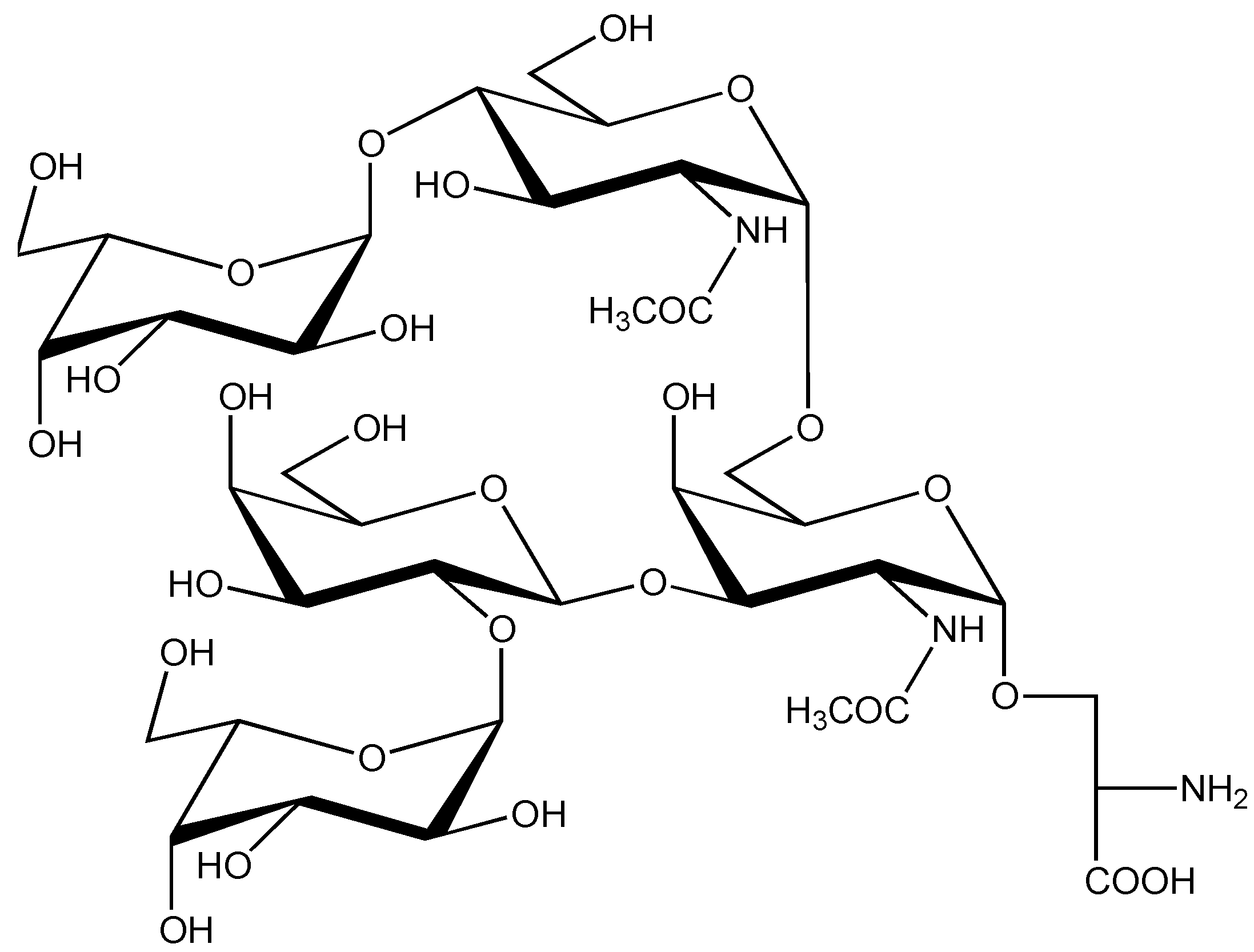
| C. geographus | P | contulakin-G | ZSEEGGSNAT*KKPYIL | Thr-10 | β-D-Galp-(1→3)-α-D-Galp NAc | [36,59,60,61,62,63] |
| SO4(HexHexNAc) Hex3 | ||||||
| (CGX-1160) | Hex2HexNAc2 | |||||
| C. textile | M | ε-TxIX | γCCγDGW*CCT*AAO | Thr-10 | α-D-Galp-(1→3)-α-D-GalpNAc | [41,64,65,66] |
| (tx5a, TxVa or Tx-012) |
4.2. Conus geographus
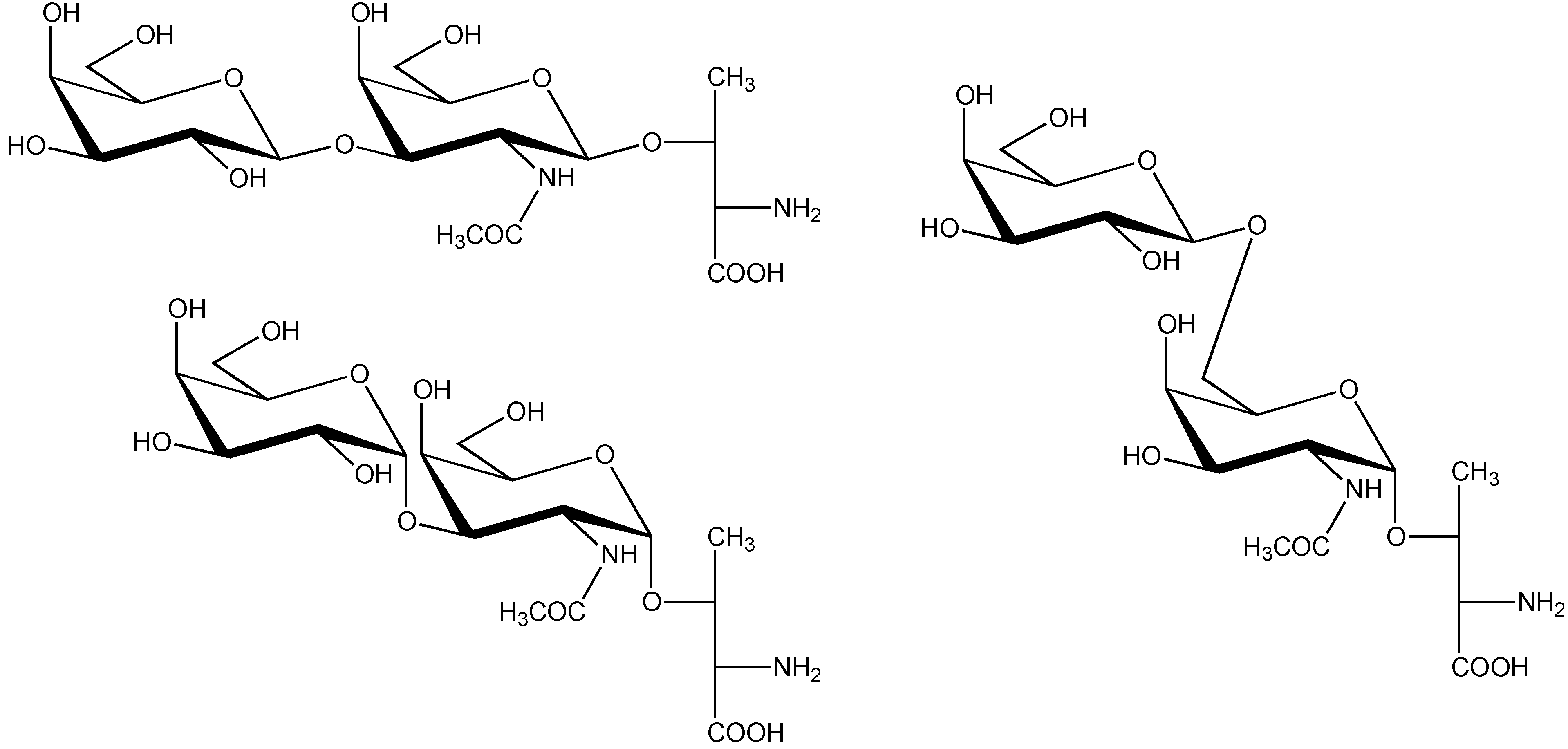
4.3. Conus textile
4.4. Conus magus
4.5. Conus stercusmuscarum
4.6. Conus consors
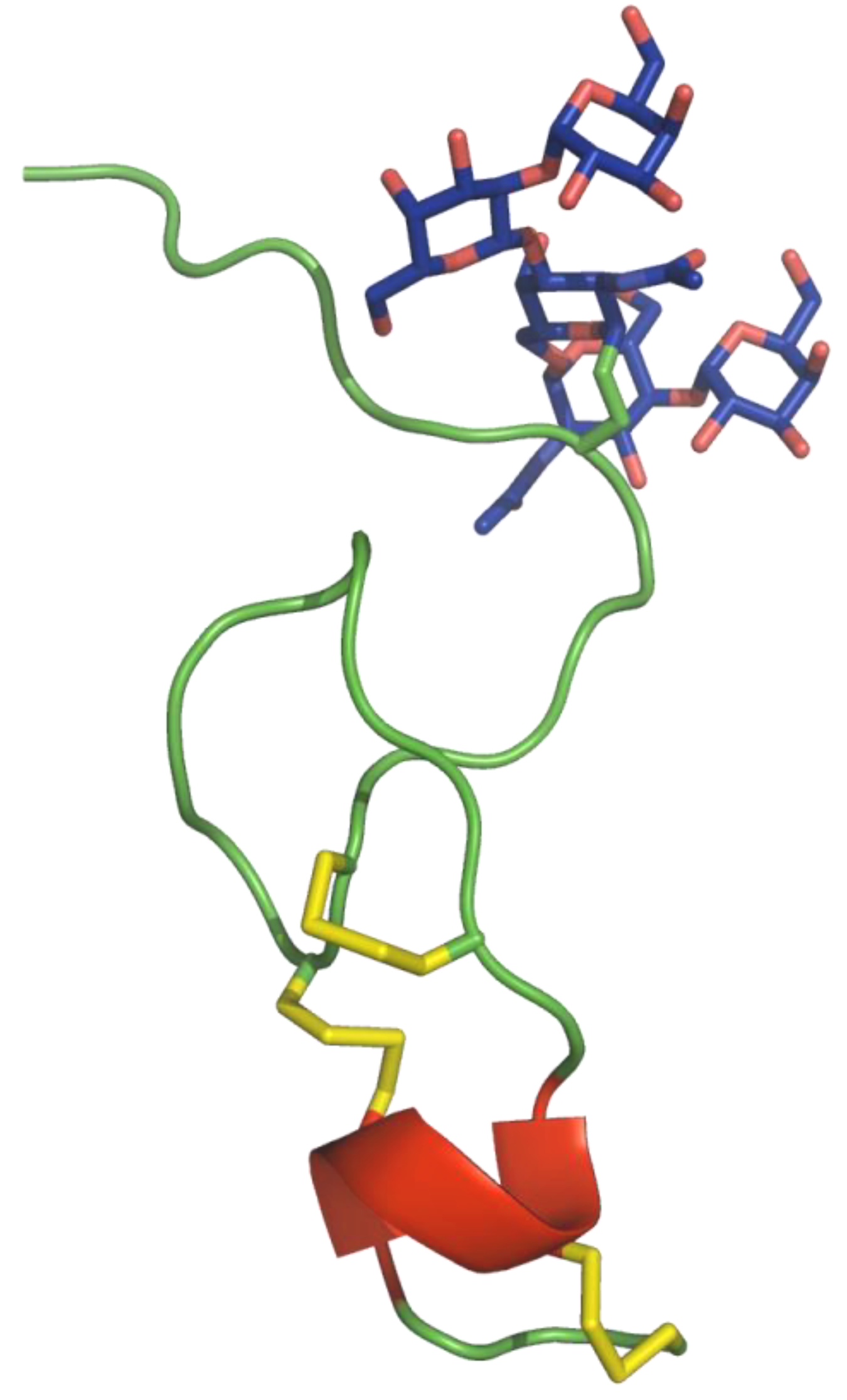
5. Peptide Sequence Comparison
6. Biosynthesis and Roles of O-Glycosylation in Conotoxins
7. Concluding Remarks
Acknowledgments
References
- Conchology Inc. Available online: http://www.conchology.be (accessed on 5 December 2012).
- CONCO, the Cone Snail Genome Project for Health. Available online: http://www.conco.eu/ (accessed on 5 December 2012).
- Terlau, H.; Shon, K.J.; Grilley, M.; Stocker, M.; Stühmer, W.; Olivera, B.M. Strategy for rapid immobilization of prey by a fish-hunting marine snail. Nature 1996, 381, 148–151. [Google Scholar]
- West, D.J.; Andrews, E.B.; Bowman, D.; McVean, A.R.; Thorndyke, M.C. Toxins from some poisonous and venomous marine snails. Comp. Biochem. Phys.Part C 1996, 113, 1–10. [Google Scholar]
- Olivera, B.M.; Rivier, J.; Clark, C.; Ramilo, C.A.; Corpuz, G.P.; Abogadie, F.C.; Mena, E.E.; Woodward, S.R.; Hillyard, D.R.; Cruz, L.J. Diversity of Conus neuropeptides. Science 1990, 249, 257–263. [Google Scholar]
- Rockel, D.; Korn, W.; Kohn, A.J. Manual of the Living Conidae; Verlag Christa Hemmen: Wiesbaden, Germany, 1995. [Google Scholar]
- Espiritu, D.J.; Watkins, M.; Dia-Monje, V.; Cartier, G.E.; Cruz, L.J.; Olivera, B.M. Venomous cone snails: Molecular phylogeny and the generation of toxin diversity. Toxicon 2001, 39, 1899–1916. [Google Scholar] [CrossRef]
- Kaas, Q.; Westermann, J.-C.; Craik, D.J. Conopeptide characterization and classifications: An analysis using ConoServer. Toxicon 2010, 55, 1491–1509. [Google Scholar] [CrossRef]
- Davis, J.; Jones, A.; Lewis, R.J. Remarkable inter- and intra-species complexity of conotoxins revealed by LC/MS. Peptides 2009, 30, 1222–1227. [Google Scholar] [CrossRef]
- Biass, D.; Dutertre, S.; Gerbault, A.; Menou, J.-L.; Offord, R.; Favreau, P.; Stöcklin, R. Comparative proteomic study of the venom of the piscivorous cone snail Conus consors. J. Proteomics 2009, 72, 210–218. [Google Scholar] [CrossRef]
- Dutertre, S.; Biass, D.; Stöcklin, R.; Favreau, P. Dramatic intraspecimen variations within the injected venom of Conus consors: An unsuspected contribution to venom diversity. Toxicon 2010, 55, 1453–1462. [Google Scholar] [CrossRef]
- Dutertre, S.; Jin, A.-H.; Kaas, Q.; Jones, A.; Alewood, P.F.; Lewis, R.J. Deep venomics reveals the mechanism for expanded peptide diversity in cone snail venom. Mol. Cell. Proteomics 2012, 12, 312–329. [Google Scholar]
- Violette, A.; Biass, D.; Dutertre, S.; Koua, D.; Piquemal, D.; Pierrat, F.; Stöcklin, R.; Favreau, P. Large-scale discovery of conopeptides and conoproteins in the injectable venom of a fish-hunting cone snail using a combined proteomic and transcriptomic approach. J. Proteomics 2012, 75, 5215–5225. [Google Scholar] [CrossRef]
- Leonardi, A.; Biass, D.; Kordiš, D.; Stöcklin, R.; Favreau, P.; Križaj, I. Conus consors snail venom proteomics proposes functions, pathways, and novel families involved in its venomic system. J. Proteome Res. 2012, 11, 5046–5058. [Google Scholar] [CrossRef]
- Le Gall, F.; Favreau, P.; Richard, G.; Letourneux, Y.; Molgó, J. The strategy used by some piscivorous cone snails to capture their prey: The effects of their venoms on vertebrates and on isolated neuromuscular preparations. Toxicon 1999, 37, 985–998. [Google Scholar] [CrossRef]
- Yoshiba, S. An estimation of the most dangerous species of cone shell, Conus (Gastridium) geographus Linne, 1758, venom’s lethal dose in humans. Nihon Eiseigaku Zasshi 1984, 39, 565–572. [Google Scholar] [CrossRef]
- Fegan, D.; Andresen, D. Conus geographus envenomation. Lancet 1997, 349, 1672. [Google Scholar] [CrossRef]
- McIntosh, J.M.; Jones, R.M. Cone venom—From accidental stings to deliberate injection. Toxicon 2001, 39, 1447–1451. [Google Scholar] [CrossRef]
- Jones, R.M.; Bulaj, G. Conotoxins—New vistas for peptide therapeutics. Curr. Pharm. Des. 2000, 6, 1249–1285. [Google Scholar] [CrossRef]
- Lewis, R.J.; Garcia, M.L. Therapeutic potential of venom peptides. Nat. Rev. Drug Discov. 2003, 2, 790–802. [Google Scholar] [CrossRef]
- Terlau, H.; Olivera, B.M. Conus venoms: A rich source of novel ion channel-targeted peptides. Physiol. Rev. 2004, 84, 41–68. [Google Scholar] [CrossRef]
- Han, T.S.; Teichert, R.W.; Olivera, B.M.; Bulaj, G. Conus venoms—A rich source of peptide-based therapeutics. Curr. Pharm. Des. 2008, 14, 2462–2479. [Google Scholar] [CrossRef]
- Favreau, P.; Stöcklin, R. Marine snail venoms: Use and trends in receptor and channel neuropharmacology. Curr. Opin. Pharmacol. 2009, 9, 594–601. [Google Scholar] [CrossRef]
- Molinski, T.F.; Dalisay, D.S.; Lievens, S.L.; Saludes, J.P. Drug development from marine natural products. Nat. Rev. Drug Discov. 2009, 8, 69–85. [Google Scholar]
- Lewis, R.J.; Dutertre, S.; Vetter, I.; Christie, M.J. Conus venom peptide pharmacology. Pharmacol. Rev. 2012, 64, 259–298. [Google Scholar] [CrossRef]
- Livett, B.G.; Gayler, K.R.; Khalil, Z. Drugs from the sea: Conopeptides as potential therapeutics. Curr. Med. Chem. 2004, 11, 1715–1723. [Google Scholar] [CrossRef]
- Fox, J.W.; Serrano, S.M.T. Approaching the golden age of natural product pharmaceuticals from venom libraries: An overview of toxins and toxin-derivatives currently involved in therapeutic or diagnostic applications. Curr. Pharm. Des. 2007, 13, 2927–2934. [Google Scholar] [CrossRef]
- Bingham, J.-P.; Mitsunaga, E.; Bergeron, Z.L. Drugs from slugs—Past, present and future perspectives of ω-conotoxin research. Chem. Biol. Interact. 2010, 183, 1–18. [Google Scholar] [CrossRef]
- Essack, M.; Bajic, V.B.; Archer, J.A.C. Conotoxins that confer therapeutic possibilities. Mar. Drugs 2012, 10, 1244–1265. [Google Scholar] [CrossRef]
- Bingham, J.-P.; Andrews, E.A.; Kiyabu, S.M.; Cabalteja, C.C. Drugs from slugs. Part II—Conopeptide bioengineering. Chem. Biol. Interact. 2012, 200, 92–113. [Google Scholar] [CrossRef]
- Miljanich, G.P. Ziconotide: Neuronal calcium channel blocker for treating severe chronic pain. Curr. Med. Chem. 2004, 11, 3029–3040. [Google Scholar] [CrossRef]
- Staats, P.S.; Yearwood, T.; Charapata, S.G.; Presley, R.W.; Wallace, M.S.; Byas-Smith, M.; Fisher, R.; Bryce, D.A.; Mangieri, E.A.; Luther, R.R.; et al. Intrathecal ziconotide in the treatment of refractory pain in patients with cancer or AIDS: A randomized controlled trial. JAMA 2004, 291, 63–70. [Google Scholar]
- Williams, J.A.; Day, M.; Heavner, J.E. Ziconotide: An update and review. Expert Opin. Pharmacother. 2008, 9, 1575–1583. [Google Scholar] [CrossRef]
- Shen, G.S.; Layer, R.T.; McCabe, R.T. Conopeptides: From deadly venoms to novel therapeutics. Drug Discov. Today 2000, 5, 98–106. [Google Scholar] [CrossRef]
- Daly, N.L.; Rosengren, K.J.; Henriques, S.T.; Craik, D.J. NMR and protein structure in drug design: Application to cyclotides and conotoxins. Eur. Biophys. J. 2011, 40, 359–370. [Google Scholar] [CrossRef]
- Craig, A.G.; Bandyopadhyay, P.; Olivera, B.M. Post-translationally modified neuropeptides from Conus venoms. Eur. J. Biochem. 1999, 264, 271–275. [Google Scholar] [CrossRef]
- Arias, H.R.; Blanton, M.P. α-Conotoxins. Int. J. Biochem. Cell Biol. 2000, 32, 1017–1028. [Google Scholar] [CrossRef]
- Craig, A.G. The characterisation of conotoxins. J. Toxicol. Toxin Rev. 2000, 19, 53–93. [Google Scholar] [CrossRef]
- Olivera, B.M.; Cruz, L.J. Conotoxins, in retrospect. Toxicon 2001, 39, 7–14. [Google Scholar] [CrossRef]
- Conticello, S.G.; Gilad, Y.; Avidan, N.; Ben-Asher, E.; Levy, Z.; Fainzilber, M. Mechanisms for evolving hypervariability: The case of conopeptides. Mol. Biol. Evol. 2001, 18, 120–131. [Google Scholar]
- Wang, C.-Z.; Chi, C.-W. Conus peptides—A rich pharmaceutical treasure. Acta Biochim. Biophys. Sin. (Shanghai) 2004, 36, 713–723. [Google Scholar] [CrossRef]
- Buczek, O.; Bulaj, G.; Olivera, B.M. Conotoxins and the posttranslational modification of secreted gene products. Cell. Mol. Life Sci. 2005, 62, 3067–3079. [Google Scholar] [CrossRef]
- Olivera, B.M. Conus peptides: Biodiversity-based discovery and exogenomics. J. Biol. Chem. 2006, 281, 31173–31177. [Google Scholar] [CrossRef]
- Norton, R.S.; Olivera, B.M. Conotoxins down under. Toxicon 2006, 48, 780–798. [Google Scholar] [CrossRef]
- Olivera, B.M.; Teichert, R.W. Diversity of the neurotoxic Conus peptides: A model for concerted pharmacological discovery. Mol. Interv. 2007, 7, 251–260. [Google Scholar] [CrossRef]
- Halai, R.; Craik, D.J. Conotoxins: Natural product drug leads. Nat. Prod. Rep. 2009, 26, 526–536. [Google Scholar] [CrossRef]
- Kaas, Q.; Westermann, J.-C.; Halai, R.; Wang, C.K.L.; Craik, D.J. ConoServer, a database for conopeptide sequences and structures. Bioinformatics 2007, 24, 445–446. [Google Scholar]
- Kaas, Q.; Yu, R.; Jin, A.-H.; Dutertre, S.; Craik, D.J. ConoServer: Updated content, knowledge, and discovery tools in the conopeptide database. Nucl. Acids Res. 2012, 40, D325–D330. [Google Scholar] [CrossRef]
- Brockhausen, I.; Schachter, H.; Stanley, P. O-GalNAc Glycans. In Essentials of Glycobiology, 2nd; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, New York, NY, USA, 2009; pp. 115–127. [Google Scholar]
- Brockhausen, I. Pathways of O-glycan biosynthesis in cancer cells. Biochim. Biophys. Acta 1999, 1473, 67–95. [Google Scholar] [CrossRef]
- Vliegenthart, J.F.G.; Kamerling, J.P. 1H NMR Structural-Reporter-Group Concepts in Carbohydrate Analysis. In Comprehensive Glycoscience—From Chemistry to Systems Biology, 1st; Kamerling, J.P., Boons, G.-J., Lee, Y.C., Suzuki, A., Taniguchi, N., Voragen, A.G.J., Eds.; Elsevier: Oxford, UK, 2007; Volume 2, pp. 133–191. [Google Scholar]
- Hocking, H.G.; Gerwig, G.J.; Dutertre, S.; Violette, A.; Favreau, P.; Stöcklin, R.; Kamerling, J.P.; Boelens, R. Structure of the O-glycosylated conopeptide CcTx from Conus consors venom. Chem. Eur. J. 2013, 19, 870–879. [Google Scholar] [CrossRef]
- Kelley, W.P.; Schulz, J.R.; Jakubowski, J.A.; Gilly, W.F.; Sweedler, J.V. Two Toxins from Conus striatus that individually induce tetanic paralysis. Biochemistry 2006, 45, 14212–14222. [Google Scholar]
- Craig, A.G.; Zafaralla, G.; Cruz, L.J.; Santos, A.D.; Hillyard, D.R.; Dykert, J.; Rivier, J.E.; Gray, W.R.; Imperial, J.; DelaCruz, R.G.; et al. An O-glycosylated neuroexcitatory Conus peptide. Biochemistry 1998, 37, 16019–16025. [Google Scholar]
- Jakubowski, J.A.; Kelley, W.P.; Sweedler, J.V. Screening for post-translational modifications in conotoxins using liquid chromatography/mass spectrometry: An important component of conotoxin discovery. Toxicon 2006, 47, 688–699. [Google Scholar] [CrossRef]
- Jakubowski, J.A.; Kelley, W.P.; Sweedler, J.V.; Gilly, W.F.; Schulz, J.R. Intraspecific variation of venom injected by fish-hunting Conus snails. J. Exp. Biol. 2005, 208, 2873–2883. [Google Scholar] [CrossRef]
- Santos, A.D.; McIntosh, J.M.; Hillyard, D.R.; Cruz, L.J.; Olivera, B.M. The A-superfamily of conotoxins: Structural and functional divergence. J. Biol. Chem. 2004, 279, 17596–17606. [Google Scholar]
- Le Gall, F.; Favreau, P.; Benoit, E.; Mattei, C.; Bouet, F.; Menou, J.-L.; Ménez, A.; Letourneux, Y.; Molgó, J. A new conotoxin isolated from Conus consors venom acting selectively on axons and motor nerve terminals through a Na+-dependent mechanism. Eur. J. Neurosci. 1999, 11, 3134–3142. [Google Scholar] [CrossRef]
- Craig, A.G.; Norberg, T.; Griffin, D.; Hoeger, C.; Akhtar, M.; Schmidt, K.; Low, W.; Dykert, J.; Richelson, E.; Navarro, V.; et al. Contulakin-G, an O-glycosylated invertebrate neurotensin. J. Biol. Chem. 1999, 274, 13752–13759. [Google Scholar]
- Kindahl, L.; Sandström, C.; Craig, A.G.; Norberg, T.; Kenne, L. 1H NMR studies on the solution conformation of contulakin-G and analogues. Can. J. Chem. 2002, 80, 1022–1031. [Google Scholar] [CrossRef]
- Kindahl, L.; Kenne, L.; Sandström, C. 1H NMR studies on the solution conformation of the [L-Ser10] and [D-Ser10] analogues of contulakin-G. Can. J. Chem. 2005, 83, 156–165. [Google Scholar] [CrossRef]
- Westerlind, U.; Norberg, T. Chemical synthesis of analogs of the glycopeptide contulakin-G, an analgetically active conopeptide from Conus geographus. Carbohydr. Res. 2006, 341, 9–18. [Google Scholar] [CrossRef]
- Craig, A.G.; Park, M.; Fischer, W.H.; Kang, J.; Compain, P.; Piller, F. Enzymatic glycosylation of contulakin-G, a glycopeptide isolated from Conus venom, with a mammalian ppGalNAc-transferase. Toxicon 2001, 39, 809–815. [Google Scholar] [CrossRef]
- Rigby, A.C.; Lucas-Meunier, E.; Kalume, D.E.; Czerwiec, E.; Hambe, B.; Dahlqvist, I.; Fossier, P.; Baux, G.; Roepstorff, P.; Baleja, J.D.; et al. A conotoxin from Conus textile with unusual posttranslational modifications reduces presynaptic Ca2+ influx. Proc. Natl. Acad. Sci. USA 1999, 96, 5758–5763. [Google Scholar]
- Kalume, D.E.; Stenflo, J.; Czerwiec, E.; Hambe, B.; Furie, B.C.; Furie, B.; Roepstorff, P. Structure determination of two conotoxins from Conus textile by a combination of matrix-assisted laser desorption/ionization time-of-flight and electrospray ionization mass spectrometry and biochemical methods. J. Mass Spectrom. 2000, 35, 145–156. [Google Scholar] [CrossRef]
- Kang, J.; Low, W.; Norberg, T.; Meisenhelder, J.; Hansson, K.; Stenflo, J.; Zhou, G.-P.; Imperial, J.; Olivera, B.M.; Rigby, A.C.; et al. Total chemical synthesis and NMR characterization of the glycopeptide tx5a, a heavily post-translationally modified conotoxin, reveals that the glycan structure is α-D-Gal-(1→3)-α-D-GalNAc. Eur. J. Biochem. 2004, 271, 4939–4949. [Google Scholar] [CrossRef]
- Kern, S.; Allen, J.; Wagstaff, J.; Shafer, S.L.; Yaksh, T. The pharmacokinetics of the conopeptide contulakin-G (CGX-1160) after intrathecal administration: An analysis of data from studies in beagles. Anesth. Analg. 2007, 104, 1514–1520. [Google Scholar] [CrossRef]
- Allen, J.W.; Hofer, K.; McCumber, D.; Wagstaff, J.D.; Layer, R.T.; McCabe, R.T.; Yaksh, T.L. An assessment of the antinociceptive efficacy of intrathecal and epidural contulakin-G in rats and dogs. Anesth. Analg. 2007, 104, 1505–1513. [Google Scholar] [CrossRef]
- Jones, R.M.; Bulaj, G. Conus peptides—Combinatorial chemistry at a cone snail’s pace. Curr. Opin. Drug Discov. Dev. 2000, 3, 141–154. [Google Scholar]
- NetOGlyc 3.1 Server. Available online: http://www.cbs.dtu.dk/services/NetOGlyc (accessed on 5 December 2012).
- Garrett, J.E.; Buczek, O.; Watkins, M.; Olivera, B.M.; Bulaj, G. Biochemical and gene expression analyses of conotoxins in Conus textile venom ducts. Biochem. Biophys. Res. Commun. 2005, 328, 362–367. [Google Scholar] [CrossRef]
- Bulaj, G.; Olivera, B.M. Folding of conotoxins: Formation of the native disulfide bridges during chemical synthesis and biosynthesis of Conus peptides. Antioxid. Redox Signal. 2008, 10, 141–155. [Google Scholar] [CrossRef]
- Safavi-Hemami, H.; Siero, W.A.; Gorasia, D.G.; Young, N.D.; Macmillan, D.; Williamson, N.A.; Purcell, A.W. Specialisation of the venom gland proteome in predatory cone snails reveals functional diversification of the conotoxin biosynthetic pathway. J. Proteome Res. 2011, 10, 3904–3919. [Google Scholar] [CrossRef]
- Safavi-Hemami, H.; Gorasia, D.G.; Steiner, A.M.; Williamson, N.A.; Karas, J.A.; Gajewiak, J.; Olivera, B.M.; Bulaj, G.; Purcell, A.W. Modulation of conotoxin structure and function is achieved through a multienzyme complex in the venom glands of cone snails. J. Biol. Chem. 2012, 287, 34288–34303. [Google Scholar]
- Tayo, L.L.; Lu, B.; Cruz, L.J.; Yates, J.R., III. Proteomic analysis provides insights on venom processing in Conus textile. J. Proteome Res. 2010, 9, 2292–2301. [Google Scholar] [CrossRef]
- Hu, H.; Bandyopadhyay, P.; Olivera, B.; Yandell, M. Elucidation of the molecular envenomation strategy of the cone snail Conus geographus through transcriptome sequencing of its venom duct. BMC Genomics 2012, 13, 284. [Google Scholar] [CrossRef]
- Dobson, R.; Collodoro, M.; Gilles, N.; Turtoi, A.; de Pauw, E.; Quinton, L. Secretion and maturation of conotoxins in the venom ducts of Conus textile. Toxicon 2012, 60, 1370–1379. [Google Scholar] [CrossRef]
- Van Halbeek, H.; Strang, A.M.; Lhermitte, M.; Rahmoune, H.; Lamblin, G.; Roussel, P. Structures of monosialyl oligosaccharides isolated from the respiratory mucins of a non-secretor (O, Lea+b−) patient suffering from chronic bronchitis. Characterization of a novel type of mucin carbohydrate core structure. Glycobiology 1994, 4, 203–219. [Google Scholar] [CrossRef]
- Dorland, L.; van Halbeek, H.; Vliegenthart, J.F.G. The identification of terminal α(1→3)-linked galactose in N-acetyllactosamine type of glycopeptides by means of 500-MHz 1H-NMR spectroscopy. Biochem. Biophys. Res. Commun. 1984, 122, 859–866. [Google Scholar] [CrossRef]
- Joziasse, D.H.; Shaper, J.H.; van den Eijnden, D.H.; van Tunen, A.J.; Shaper, N.L. Bovine α1→3-galactosyltransferase: Isolation and characterization of a cDNA clone. Identification of homologous sequences in human genomic DNA. J. Biol. Chem. 1989, 264, 14290–14297. [Google Scholar]
- Joziasse, D.H.; Shaper, N.L.; Kim, D.; van den Eijnden, D.H.; Shaper, J.H. Murine α1,3-galactosyltransferase. A single gene locus specifies four isoforms of the enzyme by alternative splicing. J. Biol. Chem. 1992, 267, 5534–5541. [Google Scholar]
- Varki, A.; Lowe, J.B. Biological roles of glycans. In Essentials of Glycobiology, 2nd; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, New York, NY, USA, 2009; pp. 75–88. [Google Scholar]
- Taylor, M.E.; Drickamer, K. Introduction to Glycobiology, 3rd ed; Oxford University Press: Oxford, UK, 2011; p. 4. [Google Scholar]
- Lopez-Vera, E.; Walewska, A.; Skalicky, J.J.; Olivera, B.M.; Bulaj, G. Role of hydroxyprolines in the in vitro oxidative folding and biological activity of conotoxins. Biochemistry 2008, 47, 1741–1751. [Google Scholar] [CrossRef]
- Marx, U.C.; Daly, N.L.; Craik, D.J. NMR of conotoxins: Structural features and an analysis of chemical shifts of post-translationally modified amino acids. Magn. Reson. Chem. 2006, 44, S41–S50. [Google Scholar] [CrossRef]
- Winchester, B. Lysosomal metabolism of glycoproteins. Glycobiology 2005, 15, 1R–15R. [Google Scholar] [CrossRef]
- Freeze, H.H. Genetic Disorders of Glycan Degradation. In Essentials of Glycobiology, 2nd; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, New York, NY, USA, 2009; pp. 567–600. [Google Scholar]
- Rebuffet, E.; Groisillier, A.; Thompson, A.; Jeudy, A.; Barbeyron, T.; Czjzek, M.; Michel, G. Discovery and structural characterization of a novel glycosidase family of marine origin. Environ. Microbiol. 2011, 13, 1253–1270. [Google Scholar] [CrossRef]
- Chi, W.-J.; Chang, Y.-K.; Hong, S.-K. Agar degradation by microorganisms and agar-degrading enzymes. Appl. Microbiol. Biotechnol. 2012, 94, 917–930. [Google Scholar] [CrossRef]
- Romeo, C.; Di Francesco, L.; Oliverio, M.; Palazzo, P.; Massilia, G.R.; Ascenzi, P.; Polticelli, F.; Schininà, M.E. Conus ventricosus venom peptides profiling by HPLC-MS: A new insight in the intraspecific variation. J. Sep. Sci. 2008, 31, 488–498. [Google Scholar]
- Rosengren, K.J.; Daly, N.L.; Craik, D.J. NMR of peptide toxins. Annu. Rep. NMR Spectrosc. 2009, 68, 89–147. [Google Scholar] [CrossRef]
- Blunt, J.W.; Copp, B.R.; Keyzers, R.A.; Munro, M.H.G.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2012, 29, 144–222. [Google Scholar] [CrossRef]
- Favreau, P.; Benoit, E.; Hocking, H.G.; Carlier, L.; D’ hoedt, D.; Leipold, E.; Markgraf, R.; Schlumberger, S.; Córdova, M.A.; Gaertner, H.; et al. A novel μ-conopeptide, CnIIIC, exerts potent and preferential inhibition of Nav1.2/1.4 channels and blocks neuronal nicotinic acetylcholine receptors. Br. J. Pharmacol. 2012, 166, 1654–1668. [Google Scholar] [CrossRef]
- Rivera-Ortiz, J.A.; Cano, H.; Marí, F. Intraspecies variability and conopeptide profiling of the injected venom of Conus ermineus. Peptides 2011, 32, 306–316. [Google Scholar] [CrossRef]
- Escoubas, P.; Quinton, L.; Nicholson, G.M. Venomics: Unravelling the complexity of animal venoms with mass spectrometry. J. Mass Spectrom. 2008, 43, 279–295. [Google Scholar] [CrossRef]
- Prashanth, J.R.; Lewis, R.J.; Dutertre, S. Towards an integrated venomics approach for accelerated conopeptide discovery. Toxicon 2012, 60, 470–477. [Google Scholar] [CrossRef]
- Samples Availability: Available from the authors.
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gerwig, G.J.; Hocking, H.G.; Stöcklin, R.; Kamerling, J.P.; Boelens, R. Glycosylation of Conotoxins. Mar. Drugs 2013, 11, 623-642. https://doi.org/10.3390/md11030623
Gerwig GJ, Hocking HG, Stöcklin R, Kamerling JP, Boelens R. Glycosylation of Conotoxins. Marine Drugs. 2013; 11(3):623-642. https://doi.org/10.3390/md11030623
Chicago/Turabian StyleGerwig, Gerrit J., Henry G. Hocking, Reto Stöcklin, Johannis P. Kamerling, and Rolf Boelens. 2013. "Glycosylation of Conotoxins" Marine Drugs 11, no. 3: 623-642. https://doi.org/10.3390/md11030623




