Nigribactin, a Novel Siderophore from Vibrio nigripulchritudo, Modulates Staphylococcus aureus Virulence Gene Expression
Abstract
:1. Introduction
2. Results and Discussion
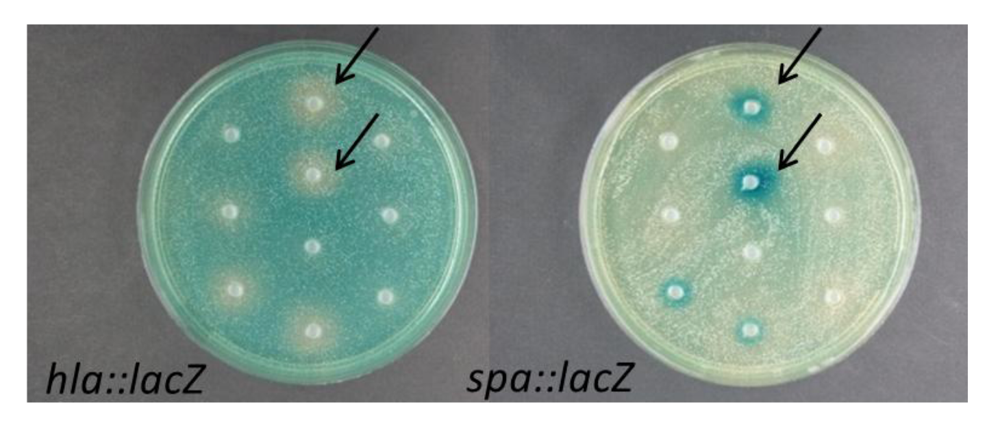
2.1. Modulation of S. aureus Virulence Gene Expression by Marine Bacteria
| Genus/family | Number of strains tested | hla interference | hla/spa interference | ||||
|---|---|---|---|---|---|---|---|
| Colony material | Extract | Supernatant | Colony material | Extract | Supernatant | ||
| Pseudoalteromonas | 41 | 37 | 15 | 0 | 19 | 0 | 0 |
| Vibrionaceae | 37 | 30 | 15 | 1 | 8 | 9 | 0 |
| Ruegeria | 5 | 0 | 0 | 0 | 0 | 0 | 0 |
2.2. Interference of Virulence Gene Expression by Vibrio nigripulchritudo
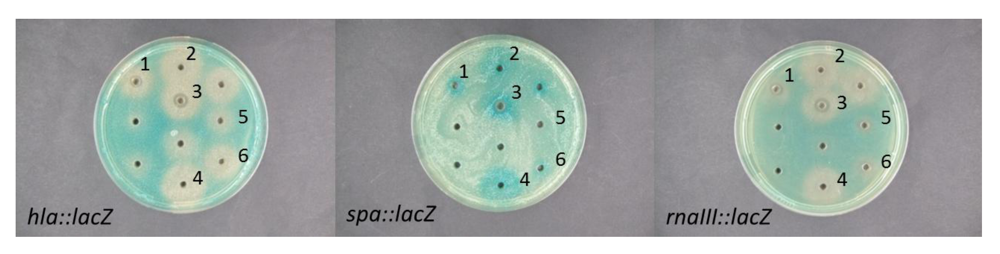
2.3. Nigribactin, a Novel Siderophore from Vibrio nigripulchritudo S2604, Enhances Spa Transcription
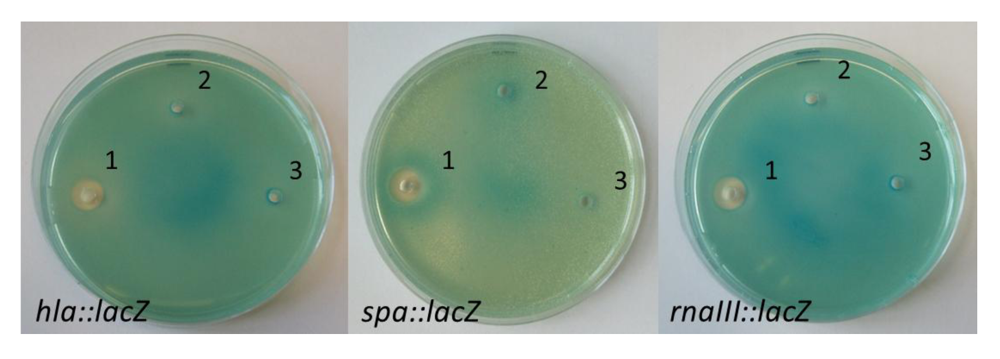
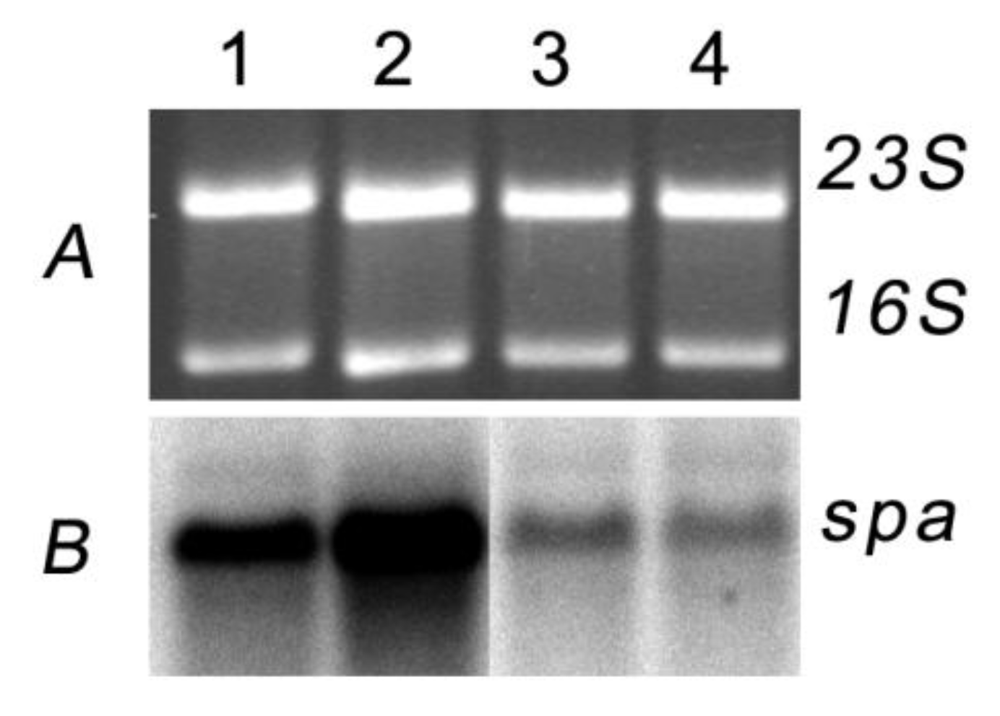
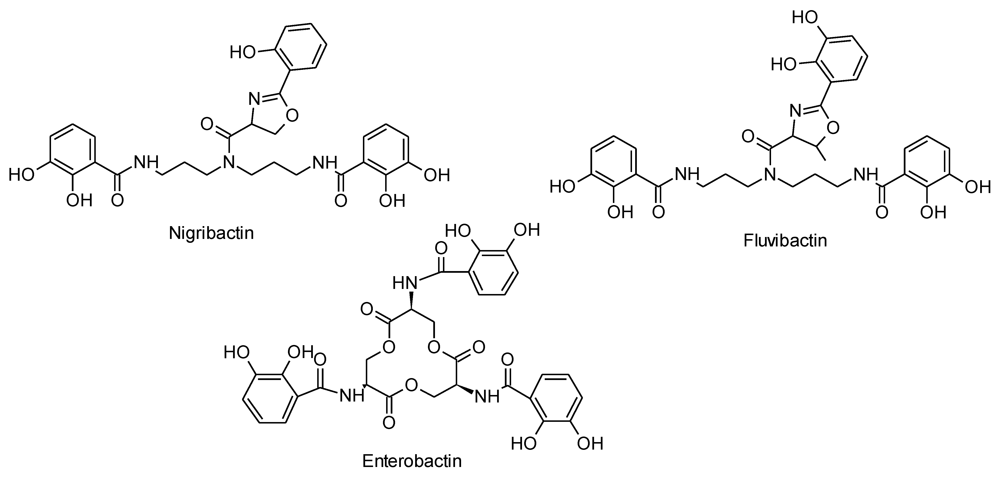
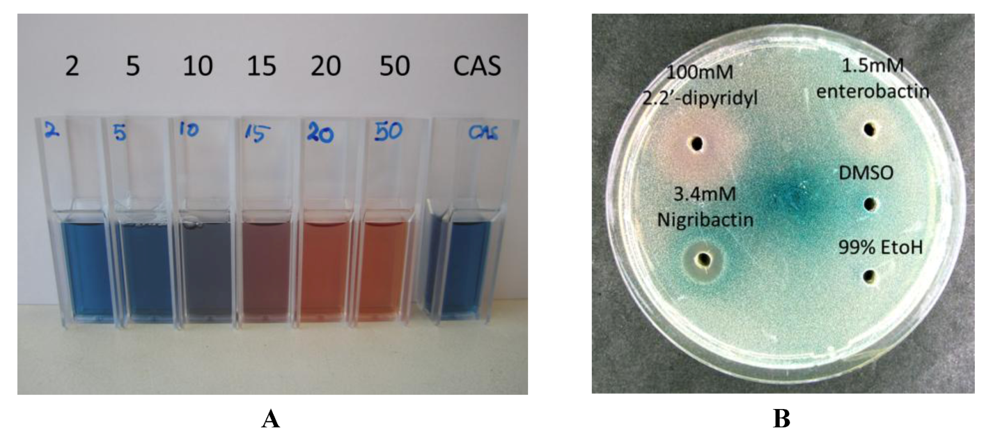
3. Experimental Section
3.1. Bacterial Strains
3.2. Bacterial Growth Conditions, Culture Extraction and Virulence Gene Expression Assay
3.3. Northern Blot Analysis
3.4. Purification and Structural Elucidation of Nigribactin
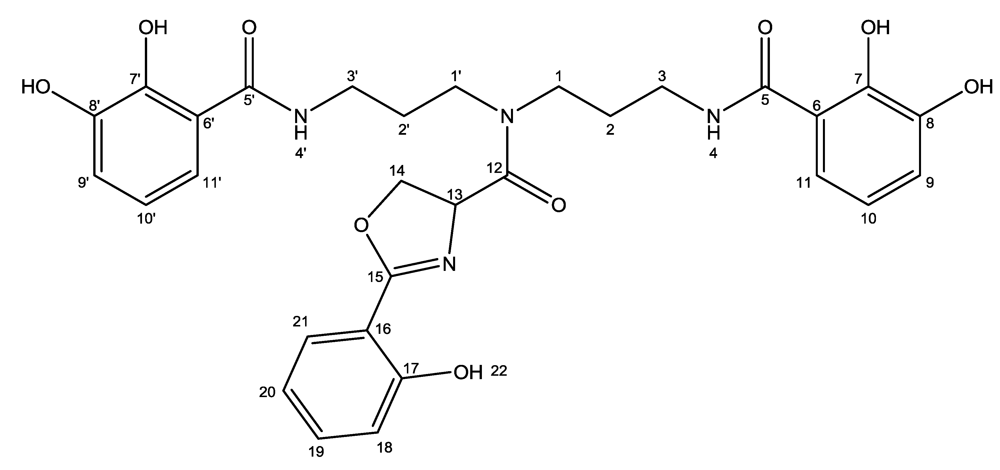
| Atom | δC (ppm) | δH (ppm) (multiplicity, J (Hz)) | HMBC |
|---|---|---|---|
| 1a | 43.2 | 3.38 (1H, m) | 1′, 2, 3, 12 |
| 1b | 43.2 | 3.36 (1H, m) | 1′, 2, 3, 12 |
| 2 | 26.8 | 1.77 (2H, m) | - |
| 3 | 36.3 | 3.26 (2H, m) | 1, 2, 5 |
| 4 | - | 8.74 | 5 |
| 5 | 169.4 | - | - |
| 6 | 114.8 | - | - |
| 7 | 149.4 | - | - |
| 8 | 145.9 | - | - |
| 9 | 118.6 | 6.88 (1H, d, 7.6) | 7, 8, 11 |
| 10 | 117.7 | 6.65 (1H, t, 7.6) | 6, 8 |
| 11 | 116.9 | 7.22 (1H, d, 8.0) | 5, 7, 9 |
| 1′a | 44.6 | 3.68 (1H, m) | 2′, 12 |
| 1′b | 44.6 | 3.56 (1H, m) | 2′, 12 |
| 2′ | 28.2 | 1.95 (2H, p, 7.2) | 1′, 3′ |
| 3′ | 36.3 | 3.38 (2H, m) | 1′, 2′, 5′ |
| 4′ | - | 8.83 | 5′ |
| 5′ | 169.6 | - | - |
| 6′ | 114.8 | - | - |
| 7′ | 149.4 | - | - |
| 8′ | 145.9 | - | - |
| 9′ | 118.6 | 6.88 (1H, d, 7.6) | 7′, 8′, 11′ |
| 10′ | 117.7 | 6.65 (1H, t, 7.6) | 6′, 8′ |
| 11′ | 116.9 | 7.26 (1H, d, 8.0) | 5′, 7′, 9′ |
| 12 | 168.3 | - | - |
| 13 | 64.2 | 5.36 (dd, 9.5, 6.7) | 12, 15 |
| 14a | 69.2 | 4.77 (1H, t, 7.5) | 12, 13, 15 |
| 14b | 69.2 | 4.54 (1H, t, 8.9) | 12, 15 |
| 15 | 165.3 | - | - |
| 16 | 109.6 | - | - |
| 17 | 158.6 | - | - |
| 18 | 116.4 | 6.97 (1H, d, 8.3) | 16, 17, 20 |
| 19 | 133.9 | 7.44 (1H, t, 7.9) | 17, 21 |
| 20 | 118.9 | 6.93 (1H, t, 7.6) | 16, 18 |
| 21 | 127.9 | 7.62 (1H, dd, 7.8, 1.1) | 17, 19 |
| 22 (–OH) | - | 11.7 | 16, 17, 18 |
| –OH | - | 12.7 | - |
| –OH | - | 12.6 | - |
| –OH | - | 9.12 | - |
| –OH | - | 9.09 | - |
4. Conclusions
Supplementary Files
Acknowledgments
References
- Rahman, H.; Austin, B.; Mitchell, W.J.; Morris, P.C.; Jamieson, D.J.; Adams, D.R.; Spragg, A.M.; Schweizer, M. Novel anti-infective compounds from marine bacteria. Mar. Drugs 2010, 8, 498–518. [Google Scholar] [CrossRef]
- Nagai, K.; Kamigiri, K.; Arao, N.; Suzumura, K.; Kawano, Y.; Yamaoka, M.; Zhang, H.; Watanabe, M.; Suzuki, K. YM-266183 and YM-266184, novel thiopeptide antibiotics produced by Bacillus cereus isolated from a marine sponge. I. taxonomy, fermentation, isolation, physico-chemical properties and biological properties. J. Antibiot. 2003, 56, 123–128. [Google Scholar] [CrossRef]
- Wietz, M.; Mansson, M.; Gotfredsen, C.H.; Larsen, T.O.; Gram, L. Antibacterial compounds from marine Vibrionaceae isolated on a global expedition. Mar. Drugs 2010, 8, 2946–2960. [Google Scholar] [CrossRef]
- Hidron, A.I.; Edwards, J.R.; Patel, J.; Horan, T.C.; Sievert, D.M.; Pollock, D.A.; Fridkin, S.K. NHSN annual update: Antimicrobial-Resistant pathogens associated with healthcare-associated infections: Annual summary of data reported to the national healthcare safety network at the centers for disease control and prevention, 2006–2007. Infect. Control Hosp. Epidemiol. 2008, 29, 996–1011. [Google Scholar] [CrossRef]
- Moran, G.J.; Krishnadasan, A.; Gorwitz, R.J.; Fosheim, G.E.; McDougal, L.K.; Carey, R.B.; Talan, D.A. EMERGEncy ID Net Study Group. Methicillin-Resistant S. aureus infections among patients in the emergency department. N. Engl. J. Med. 2006, 355, 666–674. [Google Scholar]
- Shoham, M. Antivirulence agents against MRSA. Future Med. Chem. 2011, 3, 775–777. [Google Scholar]
- Gotoh, Y.; Eguchi, Y.; Watanabe, T.; Okamoto, S.; Doi, A.; Utsumi, R. Two-Component signal transduction as potential drug targets in pathogenic bacteria. Curr. Opin. Microbiol. 2010, 13, 232–239. [Google Scholar] [CrossRef]
- Rasmussen, T.B.; Givskov, M. Quorum-Sensing inhibitors as anti-pathogenic drugs. Int. J. Med. Microbiol. 2006, 296, 149–161. [Google Scholar]
- Skindersoe, M.E.; Ettinger-Epstein, P.; Rasmussen, T.B.; Bjarnsholt, T.; De Nys, R.; Givskov, M. Quorum sensing antagonism from marine organisms. Mar. Biotechnol. 2008, 10, 56–63. [Google Scholar] [CrossRef]
- Jakobsen, T.H.; Van Gennip, M.; Phipps, R.K.; Shanmugham, M.S.; Christensen, L.D.; Alhede, M.; Skindersoe, M.E.; Rasmussen, T.B.; Friedrich, K.; Uthe, F.; et al. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrob. Agents Chemother. 2012, 56, 2314–2325. [Google Scholar]
- Novick, R.P.; Projan, S.J.; Kornblum, J.; Ross, H.F.; Ji, G.; Kreiswirth, B.; Vandenesch, F.; Moghazeh, S. The agr P2 operon: An autocatalytic sensory transduction system in Staphylococcus aureus. Mol. Gen. Genet. 1995, 248, 446–458. [Google Scholar] [CrossRef]
- Stefani, S.; Goglio, A. Methicillin-Resistant Staphylococcus aureus: Related infections and antibiotic resistance. Int. J. Infect. Dis. 2010, 14, 19–22. [Google Scholar] [CrossRef]
- Novick, R.P.; Geisinger, E. Quorum sensing in staphylococci. Annu. Rev. Genet. 2008, 42, 541–564. [Google Scholar] [CrossRef]
- Novick, R.P. Autoinduction and signal transduction in the regulation of staphylococcal virulence. Mol. Microbiol. 2003, 48, 1429–1449. [Google Scholar] [CrossRef]
- Mansson, M.; Nielsen, A.; Kjærulff, L.; Gotfredsen, C.H.; Wietz, M.; Ingmer, H.; Gram, L.; Larsen, T.O. Inhibition of virulence gene expression in Staphylococcus aureus by novel depsipeptides from a marine Photobacterium. Mar. Drugs 2011, 9, 2537–2552. [Google Scholar] [CrossRef]
- Gram, L.; Melchiorsen, J.; Bruhn, J.B. Antibacterial activity of marine culturable bacteria collected from a global sampling of ocean surface waters and surface swabs of marine organisms. Mar. Biotechnol. 2010, 12, 439–451. [Google Scholar] [CrossRef]
- Nielsen, A.; Nielsen, K.F.; Frees, D.; Larsen, T.O.; Ingmer, H. Method for screening compounds that influence virulence gene expression in Staphylococcus aureus. Antimicrob. Agents Chemother. 2010, 54, 509–512. [Google Scholar] [CrossRef]
- Musthafa, K.S.; Saroja, V.; Pandian, S.K.; Ravi, A.V. Antipathogenic potential of marine Bacillus sp. SS4 on N-acyl-homoserine-lactone-mediated virulence factors production in Pseudomonas aeruginosa (PAO1). J. Biosci. 2011, 36, 55–67. [Google Scholar] [CrossRef]
- George, E.A.; Muir, T.W. Molecular mechanisms of agr quorum sensing in virulent Staphylococci. Chembiochem 2007, 8, 847–855. [Google Scholar] [CrossRef]
- Mansson, M.; Phipps, R.K.; Gram, L.; Munro, M.H.; Larsen, T.O.; Nielsen, K.F. Explorative solid-phase extraction (E-SPE) for accelerated microbial natural product discovery, dereplication, and purification. J. Nat. Prod. 2010, 73, 1126–1132. [Google Scholar] [CrossRef]
- Yamamoto, S.; Okujo, N.; Fujita, Y.; Saito, M.; Yoshida, T.; Shinoda, S. Structures of two polyamine-containing catecholate siderophores from Vibrio fluvialis. J. Biochem. 1993, 113, 538–544. [Google Scholar]
- Payne, S.M. Iron acquisition in microbial pathogenesis. Trends Microbiol. 1993, 1, 66–69. [Google Scholar]
- Abergel, R.J.; Warner, J.A.; Shuh, D.K.; Raymond, K.N. Enterobactin protonation and iron release: Structural characterization of the salicylate coordination shift in ferric enterobactin. J. Am. Chem. Soc. 2006, 128, 8920–8931. [Google Scholar] [CrossRef]
- Schwyn, B.; Neilands, J.B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar]
- Raymond, K.N.; Dertz, E.A.; Kim, S.S. Enterobactin: An archetype for microbial iron transport. Proc. Natl. Acad. Sci. USA 2003, 100, 3584–3588. [Google Scholar] [CrossRef]
- Chan, P.F.; Foster, S.J. The role of environmental factors in the regulation of virulence-determinant expression in Staphylococcus aureus 8325-4. Microbiology 1998, 144, 2469–2479. [Google Scholar] [CrossRef]
- Horsburgh, M.J.; Aish, J.L.; White, I.J.; Shaw, L.; Lithgow, J.K.; Foster, S.J. SigmaB modulates virulence determinant expression and stress resistance: Characterization of a functional rsbU strain derived from Staphylococcus aureus 8325-4. J. Bacteriol. 2002, 184, 5457–5467. [Google Scholar]
- Fairhead, H. University of Sheffield, Sheffield, UK. Personal communication, 1998.
- Novick, R. Properties of a cryptic high-frequency transducing phage in Staphylococcus aureus. Virology 1967, 33, 155–166. [Google Scholar] [CrossRef]
- Jelsbak, L.; Ingmer, H.; Valihrach, L.; Cohn, M.T.; Christiansen, M.H.; Kallipolitis, B.H.; Frees, D. The chaperone ClpX stimulates expression of Staphylococcus aureus protein A by Rot dependent and independent pathways. PLoS One 2010, 5, e12752. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Nielsen, A.; Mansson, M.; Wietz, M.; Varming, A.N.; Phipps, R.K.; Larsen, T.O.; Gram, L.; Ingmer, H. Nigribactin, a Novel Siderophore from Vibrio nigripulchritudo, Modulates Staphylococcus aureus Virulence Gene Expression. Mar. Drugs 2012, 10, 2584-2595. https://doi.org/10.3390/md10112584
Nielsen A, Mansson M, Wietz M, Varming AN, Phipps RK, Larsen TO, Gram L, Ingmer H. Nigribactin, a Novel Siderophore from Vibrio nigripulchritudo, Modulates Staphylococcus aureus Virulence Gene Expression. Marine Drugs. 2012; 10(11):2584-2595. https://doi.org/10.3390/md10112584
Chicago/Turabian StyleNielsen, Anita, Maria Mansson, Matthias Wietz, Anders N. Varming, Richard K. Phipps, Thomas O. Larsen, Lone Gram, and Hanne Ingmer. 2012. "Nigribactin, a Novel Siderophore from Vibrio nigripulchritudo, Modulates Staphylococcus aureus Virulence Gene Expression" Marine Drugs 10, no. 11: 2584-2595. https://doi.org/10.3390/md10112584




