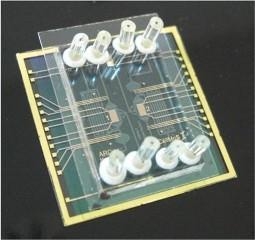
Microfluidic Systems for Pathogen Sensing: A Review
Abstract
:1. Introduction
2. Materials, Manufacturing and Detection Methods of LOC Devices
3. Nucleic Acid Based Microfluidic Pathogen Sensing
3.1. Sample preparation, isolation, amplification and detection of pathogenic DNA/RNA
4. Microfluidic Protein/Enzyme Based Pathogen Sensing
4.1. Microfluidic pathogen detection systems based antibody- and aptamer sensor
5. Microfluidic Cell Based Pathogen Sensing
6. Commercial LOC Based Pathogen Sensor Systems
7. Conclusions
References and Notes
- Yager, P.; Edwards, T.; Fu, E.; Helton, K.; Nelson, K.; Tam, M.R.; Weigl, B.H. Microfluidic Diagnostic Technologies for Global Public Health. Nat. Biotechnol. 2006, 442, 412–418. [Google Scholar]
- Lazcka, O.; Del Campo, F.J.; Muñoz, F.X. Pathogen Detection: A Perspective of Traditional Methods and Biosensors. Biosens. Bioelectron. 2007, 22, 1205–1207. [Google Scholar]
- Gomez, R.; Bashir, R.; Sarikaya, A. Microfluidic Biochip for Impedance Spectroscopy of Biological Species. Biomed. Microdevices 2001, 3, 201–209. [Google Scholar]
- Auroux, P.A.; Iossifidis, D.; Reyes, D.R.; Manz, A. Micro Total Analysis Systems. 2. Analytical Standard Operations and Applications. Anal. Chem. 2002, 74, 2637–2652. [Google Scholar]
- Reyes, D.R.; Iossifidis, D.; Auroux, P.A.; Manz, A. Micro Total Analysis Systems. 1. Introduction, Theory, and Technology. Anal. Chem. 2002, 74, 2623–2636. [Google Scholar]
- Dittrich, P.S.; Tachikawa, K.; Manz, A. Micro Total Analysis Systems. Latest Advancements and Trends. Anal. Chem. 2006, 78, 3887–3907. [Google Scholar]
- Vilkner, T.; Janasek, D.; Manz, A. Micro Total Analysis Systems. Recent Developments. Anal. Chem. 2004, 76, 3373–3385. [Google Scholar]
- Chin, C.D.; Linder, V.; Sia, S.K. Lab-on-a-Chip Devices for Global Health: Past Studies and Future Opportunities. Lab Chip 2007, 7, 41–57. [Google Scholar]
- Morens, D.M.; Folkers, G.K.; Fauci, A.S. The Challenge of Emerging and Re-emerging Infectious Diseases. Nature 2004, 430, 242–249. [Google Scholar]
- Harrison, D.J.; Manz, A.; Fan, Z.H.; Ludi, H.; Widmer, H.M. Capillary Electrophoresis and Sample Injection Systems Integrated on a Planar Glass Chip. Anal. Chem. 1992, 64, 1926–1932. [Google Scholar]
- Beebe, D.J.; Moore, J.S.; Yu, Q.; Liu, R.H.; Kraft, M.L.; Jo, B.H.; Devadoss, C. Microfluidic Tectonics: A Comprehensive Construction Platform for Microfluidic Systems. Proc. Nat. Acad. Sci. U.S.A. 2000, 97, 13488–13493. [Google Scholar]
- Lagally, E.T.; Simpson, P.C.; Mathies, R.A. Monolithic Integrated Microfluidic DNA Amplification and Capillary Electrophoresis Analysis System. Sens. Actuat. B 2000, 63, 138–146. [Google Scholar]
- Ertl, P.; Emrich, C.A.; Singhal, P.; Mathies, R.A. Capillary Electrophoresis Chips with a Sheath-flow Supported Electrochemical Detection System. Anal. Chem. 2004, 76, 3749–3755. [Google Scholar]
- Whitesides, G.M. The ‘Right’ Size in Nanobiotechnology. Nat. Biotechnol. 2003, 21, 1161–1165. [Google Scholar]
- Ionescu-Zanetti, C.; Shaw, R.M.; Seo, J.; Jan, Y.N.; Jan, L.Y.; Lee, L.P. Mammalian Electrophysiology on a Microfluidic Platform. Proc. Nat. Acad. Sci. U.S.A. 2005, 102, 9112–9117. [Google Scholar]
- Becker, H.; Gärtner, C. Polymer Microfabrication Technologies for Microfluidic Systems. Anal. Bioanal. Chem. 2008, 390, 89–111. [Google Scholar]
- Mukhopadhyay, R. When PDMS Isn't the Best. Anal. Chem. 2007, 9, 3248–3253. [Google Scholar]
- Duffy, D.C.; McDonald, J.C.; Schueller, O.J.A.; Whitesides, G.M. Rapid Prototyping of Microfluidc Systems in Poly(dimethylsiloxane). Anal. Chem. 1998, 70, 4974–4984. [Google Scholar]
- Abgrall, P.; Gue, A.M. Lab-on-a-Chip Technologies: Making a Microfluidic Network and Coupling It into a Complete Microsystem — A Review. J. Micromech. Microeng. 2007, 17, R15–R49. [Google Scholar]
- Zourob, M.; Mohr, S.; Brown, B.J.T.; Fielden, P.R.; McDonnell, M.B.; Goddard, N.J. Bacteria Detection Using Disposable Optical Leaky Waveguide Sensors. Biosens. Bioelectron. 2005, 21, 293–302. [Google Scholar]
- Mujika, M.; Arana, S.; Castano, E.; Tijero, M.; Vilares, R.; Ruano-Lopez, J.M. Magnetoresisitve Immunosensor for the Detection of Escherichia coli O157:H7. Biosens. Bioelectron. 2009, 24, 1253–1258. [Google Scholar]
- Li, Y.; Su, X. -L. Microfluidics-based Optical Biosensing Method for Rapid Detection of Escherichia coli O157:H7. J. Rapid Methods Autom. Microbiol. 2006, 14, 96–109. [Google Scholar]
- Godber, B.; Kevin, S.J.; Thompson, K.S.J.; Rehak, M.; Uludag, Y.; Kelling, S.; Sleptsov, M.; Frogley, M.; Wiehler, K.; Whalen, C.; Cooper, J.M. Direct Quantification of Analyte Concentration by Resonant Acoustic Profiling. Clin. Chem. 2005, 51, 1962–1972. [Google Scholar]
- Tamarin, O.; Comeau, S.; Dejous, C.; Moynet, D.; Riebre, D.; Beziam, J. Real Time Device for Biosensing: Design of a Bacteriophage Model Using Love Acoustic Waves. Biosens. Bioelectron. 2003, 18, 755–763. [Google Scholar]
- Mason, H.-Y.; Lloyd, C.; Dice, M.; Sinclair, R.; Ellis, W.; Powers, L. Taxonomic Identification of Microorganisms by Capture and Intrinsic Fluorescence Detection. Biosens. Bioelectron. 2003, 18, 521–527. [Google Scholar]
- Gomez-Sjoberg, R.; Morisette, D.T.; Bashir, R. Impedance Microbiology-on-a-Chip: Microfluidic Bioprocessor for Rapid Detection of Bacterial Metabolism. J. Microelectromech. Syst. 2005, 14, 829–838. [Google Scholar]
- Lee, S.J.; Park, J.S.; Im, H.T.; Jung, H.-I. A Microfluidic ATP-bioluminescence Sensor for the Detection of Airborne Microbes. Sens. Actuat. B 2008, 132, 443–448. [Google Scholar]
- Silley, P.; Forsythe, S. Impedance Microbiology — A Rapid Change for Microbiologists. J. Appl. Bacteriol. 1996, 80, 233–243. [Google Scholar]
- Boehm, D.A.; Gottlieb, P.A.; Hua, S.Z. On-chip Microfluidic Biosensors for Bacterial Detection and Identification. Sens. Actuat. B 2007, 126, 508–514. [Google Scholar]
- Rider, T.H.; Petrovick, M.S.; Nargi, R.E.; Harper, J.D.; Schwoebel, E.D.; Mathews, R.H.; Blanchard, D.J.; Bortolin, L.T.; Young, A.M.; Chen, J.; Hollis, M.A. A B Cell-based Sensor for Rapid Identification of Pathogens. Science 2003, 301, 213–215. [Google Scholar]
- Xiang, Q.; Hu, G.; Gao, Y.; Li, D. Miniaturized Immunoassay Microfluidic System with Electrokinetic Control. Biosens. Bioelectron. 2006, 21, 2006–2009. [Google Scholar]
- Yadavalli, V.K.; Pishko, M.V. Biosensing in Microfluidic Channels Using Fluorescence Polarization. Anal. Chim. Acta 2004, 507, 123–128. [Google Scholar]
- Ivnitski, D.; Abdel-Hamid, I.; Atanasov, P.; Wilkins, E.; Stricher, S. Application of Electrochemical Biosensors for Detection of Food Pathogenic Bacteria. Electroanalysis 2000, 12, 371–325. [Google Scholar]
- Yi, C.; Zhang, Q.; Li, C.W.; Yang, J.; Zhao, J.; Yang, M. Optical and Electrochemical Detection Techniques for Cell-based Microfluidic Systems. Anal. Bioanal.Chem. 2006, 384, 1259–1268. [Google Scholar]
- Gaits, F.; Hahn, K. Shedding Light on Cell Signaling: Interpretation of FRET Biosensors. Sci STKE 2003, 165, PE3. [Google Scholar]
- Chinowsky, T.M.; Quinn, J.G.; Bartholomew, D.U.; Kaiser, R.; Elkind, J.L. Performance of the Spreeta 2000 Integrated Surface Plasmon Resonance Affinity Sensor. Sens. Actuat. B 2003, 91, 266–274. [Google Scholar]
- Nakamura, H.; Murakami, Y.; Yokoyama, K.; Tamiya, E.; Karube, I. A Compactly Integrated Flow Cell with a Chemiluminescent FIA System for Determining Lactate Concentration in Serum. Anal. Chem. 2001, 73, 373–378. [Google Scholar]
- Yakovleva, J.; Davidsson, R.; Lobanova, R.; Bengtsson, M.; Eremin, S.; Laurell, T.; Emneus, J. Microfluidic Enzyme Immunoassay Using Silicon Microchip with Immobilized Antibodies and Chemiluminescence Detection. Anal. Chem. 2002, 74, 2994–3004. [Google Scholar]
- Wang, Y.; Vaidya, B.; Farquar, H.D.; Stryjewski, W.; Hammer, R.P.; McCarley, R.L.; Soper, S.A.; Cheng, Y.W.; Barany, R. Microarrays Assembled in Microfluidic Chips Fabricated from Poly(methyl methacrylate) for the Detection of Low-Abundant DNA Mutations. Anal. Chem. 2003, 75, 1130–1140. [Google Scholar]
- Spegel, C.; Heiskanen, A.; Skjolding, L.H.D.; Emneus, J. Chip Based Electroanalytical Systems for Cell Analysis. Electroanalysis 2008, 20, 680–702. [Google Scholar]
- Ertl, P.; Wagner, M.; Corton, E.; Mikkelsen, S.R. Rapid Identification of Viable Escherichia coli subspecies with an Electrochemical Screen-printed Biosensor Array. Biosens. Bioelectron. 2003, 18, 907–916. [Google Scholar]
- Sadik, O.A.; Aluoch, A.A.; Zhou, A. Status of Biomolecular Recognition Using Electrochemical Techniques. Biosens. Bioelectron. 2009, 24, 2749–2765. [Google Scholar]
- Sandifer, J.R.; V, J. Review of Biosensor and Industrial Applications of pH-ISFETs and an Evaluation of Honeywell's “DuraFET”. Microchim. Acta 1999, 131, 91–98. [Google Scholar]
- Gehring, A.G.; Patterson, D.L.; Tu, S.I. Use of a Light-addressable Potentiometric Sensor for the Detection of Escherichia coli O157:H7. Anal. Biochem. 1998, 258, 293–298. [Google Scholar]
- Taton, T.A.; Lu, G.; Mirkin, C.A. Two-color Labeling of Oligonucleotide Arrays via Size-Selective Scattering of Nanoparticle Probes. J. Am. Chem. Soc. 2001, 123, 5164–5165. [Google Scholar]
- Taton, T.A.; Mirkin, C.A.; Letsinger, R.L. Scanometric DNA Array Detection with Nanoparticle Probes. Science 2000, 289, 1757–1760. [Google Scholar]
- Shang, L.; Chen, H.; Deng, L.; Dong, S. Enhanced Resonance Light Scattering Based on Biocatalytic Growth of Gold Nanoparticles for Biosensors Design. Biosens. Bioelectron. 2008, 23, 1180–1184. [Google Scholar]
- Anderson, R.C.; Su, X.; Bodgan, G.J.; Fenton, J. A Miniature Integrated Device for Automated Multistep Genetic Assays. Nucleic Acids Res. 2000, 28, e60. [Google Scholar]
- Situma, C.; Hashimoto, M.; Soper, S.A. Mergin Microfluidics with Microarray-based Bioassays. Biomol. Eng. 2006, 23, 213–231. [Google Scholar]
- Liu, R.H.; Munro, S.B.; Nguyen, T.; Siuda, T.; Suciu, D.; Bizak, M.; Slota, M.; Fuji, H.S.; Danley, D.; McShea, A. Integrated Microfluidic Custom Array Device for Bacterial Genotyping and Identification. J. Assoc. Lab. Autom. 2006, 11, 360–367. [Google Scholar]
- Wang, L.; Li, P.C.H.; Yu, H.-Z.; Parameswaran, A.M. Fungal Pathogenic Nucleic Acid Detection Achieved with a Microfluidic Microarray Device. Anal. Chim. Acta 2008, 610, 97–104. [Google Scholar]
- Liu, R.H.; Lodes, M.J.; Nyuyen, T.; Siuda, T.; Slota, M.; Fuji, H.S.; McShea, A. Validation of a Fully Integrated Microfluidic Array Device for Influenza A subtype Identification and Sequencing. Anal. Chem. 2006, 78, 5184–5193. [Google Scholar]
- Myers, K.M.; Gaba, J.; Al-Khaldi, S.F. Molecular Identification of Yersinia enterocolitica Isolated from Pasteurized Whole Milk Using DNA Microarray Chip Hybridization. Mol. Cell. Probes 2006, 20, 71–80. [Google Scholar]
- Piliarik, M.; Paravoa, L.; Homola, J. High-throughput SPR Sensor for Food Safety. Biosens. Bioelectron. 2009, 24, 1399–1404. [Google Scholar]
- Mulvaney, S.P.; Cole, C.L.; Kniller, M.D.; Malito, M.; Tamanaha, C.R.; Rife, J.C.; Stanton, M.W. Rapid, Femtomolar Bioassays in Complex Matrices Combining Microfluidics and Magnetoelectronics. Biosens. Bioelectron. 2007, 23, 191–200. [Google Scholar]
- Gabig-Ciminiska, M.; Andresen, H.; Albers, J.; Hintsche, R.; Enfors, S.O. Identification of Pathogenic Microbial Cells and Spores by Electrochemcial Detection on a Biochip. Microbial Cell Factories 2004, 3, 2–11. [Google Scholar]
- Chang, H.-C. Nanobead Electrokinetics: the Enabling Microfluidic Platform for Rapid Multi-Target Pathogen Sensing. AICHE J. 2007, 53, 2486–2492. [Google Scholar]
- Edelstein, R.L.; Tamanaha, C.R.; Sheehan, P.D.; Miller, M.M.; Baselt, D.R.; Whitman, L.J.; Colton, R.J. The BARC Biosensor Applied to the Detection of Biological Warfare Agents. Biosens. Bioelectron. 2000, 14, 805–813. [Google Scholar]
- Zaytseva, N.V.; Goral, V.N.; Montagna, R.A.; Baeumner, A.J. Development of a Microfluidic Biosensor Module for Pathogen Detection. Lab Chip 2005, 5, 801–811. [Google Scholar]
- Wang, C.-H.; Chen, Y.Y.; Liao, C.S.; Hsieh, T.M.; Luo, C.H.; Wu, J.J.; Lee, H.H.; Lee, G.B. Circulating Polymerase Chain Reaction Chips utilizing Multiple-membrane Activation. J. Micromech. Microeng. 2007, 17, 367–375. [Google Scholar]
- Chang, Y.-H.; Lee, G.B.; Huang, F.C.; Chen, Y.Y.; Lin, L. Integrated Polymerase Chain Reaction Chips Utilizing Digital Microfluidics. Biomed. Microdevices 2006, 8, 215–225. [Google Scholar]
- Cheong, K.H.; Yi, D.K.; Lee, J.-G.; Park, J.M.; Kim, M.J.; Edel, J.B.; Ko, C. Gold Nanoparticles for One Step DNA Extraction and Real-time PCR of Pathogens in a Single Chamber. Lab Chip 2008, 8, 810–813. [Google Scholar]
- Christensen, T.B.; Bang, D.D.; Wolff, A. Multplex Polymerase Chain Rreaction (PCR) on a SU-8 Chip. Microelectron. Eng. 2008, 85, 1278–1281. [Google Scholar]
- Zhang, C.; Xu, J.; Ma, W.; Zheng, W. PCR Microfluidic Devices for DNA Amplification. Biotechnol. Adv. 2006, 24, 243–284. [Google Scholar]
- Muck, A.; Svatos, A. Chemical Modification of Polymeric Microchip Devices. Talanta 2007, 74, 333–341. [Google Scholar]
- Felbel, J.; Reichert, A.; Kielpinski, M.; Urban, M.; Henkel, T.; Häfner, N.; Dürst, M.; Weber, J. Reverse Transcription-Polyermerase Chain Reaction (RT-PCR) in Flow-through Micro-reactors: Thermal and Fluidic concepts. Chem. Eng. J. 2008, 135S, 298–302. [Google Scholar]
- Liu, Y.L.; Elsholz, B.; Enfors, S.-O.; Gabig-Ciminiska, M. Confirmative Electric DNA Array-based Test for Food Poisoning Bacillus cereus. J. Microbiol. Methods 2007, 70, 55–64. [Google Scholar]
- Easley, C.; Karlinseq, J.M.; Bienvenue, J.M.; Legendre, L.A.; Roper, M.G.; Feldman, S.H.; Hughes, M.A.; Hewlett, E.L.; Merkel, T.J.; Ferrance, J.P.; Landers, J.P. A Fully Integrated Microfuidic Genetic Analysis System with Sample-in-Answer-out Capability. Proc. Nat. Acad. Sci. U.S.A. 2006, 103, 19272–19277. [Google Scholar]
- Liao, J.C.; Mastali, M.; Gau, V.; Suchard, M.A.; Moller, A.K.; Bruckner, D.A.; Babbitt, J.T.; Gornbein, J.; Landaw, E.M.; McCabe, E.R.B.; Churchill, B.M.; Haake, D.A. Use of Electrochemical DNA Biosensors for Rapid Molecular Identification of Uropathogens in Clinical Urine Specimens. J. Clin. Microbiol. 2006, 44, 561–570. [Google Scholar]
- Kaigala, G.V.; Huskins, R.J.; Preiksaitis, J.; Pang, X.-L.; Pilarski, L.M.; Backhouse, C.J. Automated Screening Using Microfluidic Chip-based PCR and Product Detection to Assess Risk of BK Virus-associated Nephropathy in Renal Transplant Recipients. Electrophoresis. 2006, 27, 3753–3763. [Google Scholar]
- Gilbride, K.A.; Lee, D.-Y.; Beaudette, L.A. Molecular Techniques in Wastewater: Understanding Microbial Communities, Detecting Pathogens and Real-time Process Control. J. Microbiol. Methods 2006, 66, 1–20. [Google Scholar]
- Bhattacharya, S.; Salmat, S.; Morisette, D.; Banada, P.; Akin, D.; Liu, Y.-S.; Bhunia, A.K.; Ladisch, M.; Bashir, R. PCR-based Detection in a Micro-fabricated Platform. Lab Chip 2008, 8, 1130–1136. [Google Scholar]
- Lee, J.-G.; Cheong, K.H.; Huh, N.; Kim, S.; Choi, J.W.; Ko, C. Microchip-Based One Step DNA Extraction and Real-time PCR in one Chamber for Rapid Pathogen Identification. Lab Chip 2006, 6, 886–895. [Google Scholar]
- Anzai, Y.; Saito, S.; Fujimoto, K.; Kinoshita, K.; Kato, F. Detection and Identification of Species with Bacterial Cells Using a Plastic DNA Array. J. Health Sci. 2008, 54, 229–234. [Google Scholar]
- Lien, K.-Y.; Lee, W.C.; Lei, H.Y.; Lee, G.B. Integrated Reverse Transcription Polymerase Chain Reaction Systems for Virus Detection. Biosens. Bioelectron. 2007, 22, 1739–1748. [Google Scholar]
- Gascoyne, P.; Satayavivad, J.; Ruchirawat, M. Microfluidic Approaches to Malaria Detection. Acta Tropica 2004, 89, 357–369. [Google Scholar]
- Liao, C.-S.; Lee, G.B.; Liu, H.S.; Hsieh, T.M.; Luo, C.H. Minaturized RT-PCR System for Diagnosis of RNA-based Viruses. Nucleic Acids Res. 2005, 33, e156. [Google Scholar]
- Roper, M.G.; Easley, C.J.; Landers, J.P. Advances on Polymerase Chain Reaction on Microfluidic Chips. Anal. Chem. 2005, 77, 3887–3894. [Google Scholar]
- Beer, N.R.; Wheeler, E.K.; Lee-Houghton, L.; Watkins, N.; Nasarabidi, S.; Hebert, N.; Leung, P.; Arnold, D.W.; Bailey, C.G.; Colston, B.W. On-chip Single-copy Real-time Reverse-Transcription PCR in Isolated Picoliter Droplets. Anal. Chem. 2008, 80, 1854–1858. [Google Scholar]
- Kiss, M.M.; Ortoleva-Donnelly, L.; Beer, N.R.; Warner, J.; Bailey, C.G.; Colston, B.W.; Rothberg, J.M.; Link, D.R.; Leamon, J.H. High-throughput Quantitative Polymerase Chain Reaction in Picoliter Droplets. Anal. Chem. 2008, 80, 8975–8981. [Google Scholar]
- Csordas, A.; Delwiche, M.J.; Barak, J.D. Nucleic Acid Sensor and Fluid Handling for Detection of Bacterial Pathogens. Sens. Actuat. B 2008, 134, 1–8. [Google Scholar]
- Mitterer, G.; Huber, M.; Leidinger, E.; Kirisits, C.; Lubitz, W.; Mueller, M.W.; Schmidt, W.M. Microarray-based Identification of Bacteria in Clinical Samples by Solid-Phase PCR Amplification of 23S ribosomal DNA Sequences. J. Clin. Microbiol. 2004, 42, 1048–1057. [Google Scholar]
- Hou, C.-S.J.; Godin, M.; Payer, K.; Chakarabarti, R.; Manalis, S.R. Integrated Microelectronic Device for Label-free Nucleic Acid Amplification and Detection. Lab Chip 2007, 7, 347–354. [Google Scholar]
- Huang, F.-C.; Liao, C.S.; Lee, G.B. An Integrated Microfluidic Chip for DNA/RNA Amplification, Electrophoresis, Separation and On-line Optical Detection. Electrophoresis. 2006, 27, 3297–3305. [Google Scholar]
- Möller, R.; Schüler, T.; Günther, S.; Carlsohn, R.; Munder, T.; Fritzsche, W. Electrical DNA-Chip-based Identification of Different Species of the Genus Kitasatospora. Appl. Microbiol. Biotechnol. 2008, 77, 1181–1188. [Google Scholar]
- Yeung, S.-W.; Lee, T.M.-H.; Cai, H.; Hsing, I.M. A DNA Biochip for On-the-spot Multiplexed Pathogen Identification. Nucleic Acids Res. 2006, 34, e118. [Google Scholar]
- Holliger, P.; Hudson, P.J. Engineered Antibody Fragments and the Rise of Single Domains. Nat. Biotechnol. 2005, 23, 1126–1136. [Google Scholar]
- Liu, J.L.; Anderson, G.P.; Goldman, E.R. Isolation of Anti-toxin Single Domain Antibodies from a Semi-synthetic Spiny Dogfish Shark Display Library. BMC Biotechnol. 2007, 7, 78. [Google Scholar]
- Saerens, D.; Huang, L.; Bonroy, K.; Muyldermans, S. Antibody Fragments as Probe in Biosensor Development. Sensors. 2008, 8, 4669–4686. [Google Scholar]
- Makamba, H.; Kim, J.H.; Lim, K.; Park, N.; Hahn, J.H. Surface Modification of Poly(dimethylsiloxane) Microchannels. Electrophoresis. 2003, 24, 3607–3619. [Google Scholar]
- Talapatra, A.; Rouse, R.; Hardiman, G. Protein Microarrays: Challenges and Promises. Pharmacogenomics. 2002, 3, 527–536. [Google Scholar]
- Seurynck-Servoss, S.L.; Baird, C.L.; Rodland, K.D.; Zangar, R.C. Surface Chemistries for Antibody Microarrays. Front. Biosci. 2007, 12, 3956–3964. [Google Scholar]
- Dong, Y.; Phillips, K.S.; Cheng, Q. Immunosensing of Staphylococcus Enterotoxin B (SEB) in Milk with PDMS Microfluidic Systems Using Reinforced Supported Bi-layer Membranes (r-SBMs). Lab Chip 2006, 6, 675–681. [Google Scholar]
- Ramachandran, N.; Hainsworth, E.; Bhullar, B.; Eisenstein, S.; Rosen, B.; Lau, A.Y.; Walter, J.C.; LaBaer, J. Self-Assembling Protein Microarrays. Science 2004, 305, 86–90. [Google Scholar]
- Milovanova, T.; Chatterjee, S.; Hawkins, B.J.; Hong, N.; Sorokina, E.M.; DeBolt, K.; Moore, J.S.; Madesh, M.; Fisher, A.B. Caveolae Are an Essential Component of the Pathway for Endothelial Cell Signaling Associated with Abrupt Reduction of Shear Stress. Biochim. Biophys. Acta 2008, 1783, 1866–1875. [Google Scholar]
- Sivagnanam, V.; Bouhmad, A.; Lacharme, F.; Vandevyver, C.; Gijs, M.A.M. Sandwich Immunoassay on a Microfluidic Chip Using Patterns of Electrostatically Self-assembled Streptavidin-coated Beads. Microelectron. Eng. 2008. In press. [Google Scholar]
- He, M.; Stoevesandt, O.; Taussig, M.J. In Situ Synthesis of Protein Arrays. Curr. Opin. Biotechnol. 2008, 19, 4–9. [Google Scholar]
- Cha, T.; Guo, A.; Zhu, X.-Y. Enzymatic Activity on a Chip: The Critical Role of Protein Orientation. Proteomics. 2005, 5, 416–419. [Google Scholar]
- Talasaz, A.H.; Nemat-Gorgani, M.; Liu, Y.; Ståhl, P.; Dutton, R.W.; Ronaghi, M.; Davis, R.W. Prediction of Protein Orientation upon Immobilization on Biological and non-biological Surfaces. Proc. Nat. Acad. Sci. U.S.A. 2006, 103, 14773–14778. [Google Scholar]
- Hall, R.H. Biosensor Technologies for Detecting Microbiological Food Borne Hazards. Microb. Infect. 2002, 4, 425–432. [Google Scholar]
- Varnum, S.M.; Warner, M.G.; Dockendorff, B.; Anheier, N.C., Jr.; Lou, J.; Marks, J.D.; Smith, L.A.; Feldhaus, M.J.; Grate, J.W.; Bruckner-Lea, C.J. Enzyme-Amplified Protein Microarray and a Fluidic Renewable Surface Fluorescence Immunoassay for Botulinum Neurotoxin Detection Using High-Affinity Recombinant Antibodies. Anal. Chim. Acta 2006, 570, 137–143. [Google Scholar]
- Johnson, T.J.; Locascio, L.E. Characterization and Optimization of Slanted Well Designs for Microfluidic Mixing under Electroosmotic Flow. Lab Chip 2002, 2, 135–140. [Google Scholar]
- Chen, H.; Zheng, Y.; Jiang, J.-H.; Wu, H.L.; Shen, G.L.; Yu, R.Q. An Ultrasensitive Chemiluminescence Biosensor for Cholera Toxin Based on Ganglioside-functionalized Supported Lipid Membrane and Liposome. Biosens. Bioelectron. 2008, 24, 684–689. [Google Scholar]
- Viswanathan, S.; Wu, L.-C.; Huang, M.R.; Ho, J.A. Electrochemical Immunosensor for Cholera Toxin Using Liposomes and Poly(3,4-ethylenedioxythiophene)-coated Carbon Nanotubes. Anal. Chem. 2006, 78, 1115–1121. [Google Scholar]
- Quinn, J.; Patel, P.; Fitzpatrick, B.; Manning, B.; Dillon, P.; Daly, S.; O'Kennedy, R.; Alcocer, M.; Lee, H.; Morgan, M.; Lang, K. The Use of Regenerable, Affinity Ligand-based Surfaces for Immunosensor Applications. Biosens. Bioelectron. 1999, 14, 587–595. [Google Scholar]
- Mayes, A.G.; Whitcombe, M.J. Synthetic Strategies for the Generation of Molecularly Imprinted Organic Polymers. Adv. Drug Deliv. Rev. 2005, 57, 1742–1778. [Google Scholar]
- Rydholm, S.; Frisk, T.; Kowalewski, J.M.; Andersson Svahn, H.; Stemme, G.; Brismar, H. Microfluidic Devices for Studies of Primary Cilium Mediated Cellular Response to Dynamic Flow Conditions. Biomed. Microdevices 2008, 10, 555–560. [Google Scholar]
- Tan, H.Y.; Loke, W.K.; Tan, Y.T.; Nguyen, N.T. A Lab-on-a-Chip for Detection of Nerve Agent Sarin in Blood. Lab Chip 2008, 8, 885–891. [Google Scholar]
- Le Nel, A.; Minc, N.; Smadja, C.; Slovakova, M.; Bilkova, Z.; Peyrin, J.M.; Viovy, J.L.; Taverna, M. Controlled Proteolysis of Normal and Pathological Prion Protein in a Microfluidic Chip. Lab Chip 2008, 8, 294–301. [Google Scholar]
- Ellington, A.D.; Szostak, J.W. In Vitro Selection of RNA Molecules that Bind Specific Ligands. Nature 1990, 346, 818–822. [Google Scholar]
- Tuerk, C.; Gold, L. Systematic Evolution of Ligands by Exponential Enrichment: RNA Ligands to Bacteriophage T4 DNA Polymerase. Science 1990, 249, 505–510. [Google Scholar]
- Torres-Chavolla, E.; Alocilja, E.C. Aptasensors for Detection of Microbial and Viral Pathogens. Biosens. Bioelectron. 2008. (online e-pub ahead of print). [Google Scholar]
- So, H.-M.; Won, K.; Kim, Y.H.; Kim, B.K.; Ryu, B.H.; Na, P.S.; Kim, H.; Lee, J.O. Single-walled Carbon Nanotube Biosensors Using Aptamers as Molecular Recognition Elements. J. Am. Chem. Soc. 2005, 127, 11906–11907. [Google Scholar]
- Klostranec, J.M.; Xiang, Q.; Farcas, G.A.; Lee, J.A.; Rhee, A.; Lafferty, E.I.; Perrault, S.D.; Kain, K.C.; Chan, W.C. Convergence of Quantum Dot Barcodes with Microfluidics and Signal Processing for Multiplexed High-Throughput Infectious Disease Diagnostics. Nano Lett. 2007, 7, 2812–2818. [Google Scholar]
- Blicharz, T.M.; Siqueira, W.L.; Helmerhorst, E.J.; Oppenheim, F.G.; Wexler, P.J.; Little, F.F.; Walt, D.R. Fiber-Optic Microsphere-Based Antibody Array for the Analysis of Inflammatory Cytokines in Saliva. Anal. Chem. 2009, 81, 2106–2114. [Google Scholar]
- Lucas, L.J.; Chesler, J.N.; Yoon, J.-Y. Lab-on-a-Chip Immunoassay for Multiple Antibodies Using Microsphere Light Scattering and Quantum Dot Emission. Lab Chip 2007, 23, 675–681. [Google Scholar]
- Bunyakul, N.; Edwards, K.A.; Promptmas, C.; Baeumner, A.J. Cholera Toxin Subunit B Detection in Microfluidic Devices. Anal. Bioanal.Chem. 2009, 393, 177–186. [Google Scholar]
- Delehanty, J.B.; Ligler, F.S. A Microarray Immunoassay for Simultaneous Detection of Proteins and Bacteria. Anal. Chem. 2002, 74, 5681–5687. [Google Scholar]
- Pal, S.; Alocilja, E.C.; Downes, F.P. Nanowire Labeled Direct-charge Transfer Biosensor for Detecting Bacillus Species. Biosens. Bioelectron. 2007, 22, 2329–2336. [Google Scholar]
- Diercks, A.H.; Ozinsky, A.; Hansen, C.L.; Spotts, J.M.; Rodriguez, D.J.; Aderem, A. A Microfluidic Device for Multiplexed Protein Detection in Nano-liter Volumes. Anal. Biochem. 2009, 386, 30–35. [Google Scholar]
- Huh, Y.S.; Park, T.J.; Lee, E.Z.; Hong, W.H.; Lee, S.Y. Development of a Fully Integrated Microfluidic System for Sensing Infectious Viral Disease. Electrophoresis. 2008, 29, 2960–2969. [Google Scholar]
- Patolsky, F.; Zheng, G.; Hayden, O.; Lakadamyali, M.; Zhuang, X.; Lieber, C.M. Electrical Detection of Single Viruses. Proc. Nat. Acad. Sci. U.S.A. 2004, 101, 14017–14022. [Google Scholar]
- Ymeti, A.; Subramaniam, V.; Beumer, T.A.; Kanger, J.S. An Ultrasensitive Young Interferometer Handheld Sensor for Rapid Virus Detection. Expert Rev. Med. Devices. 2007, 4, 447–454. [Google Scholar]
- Kirschbaum, M.; Jaeger, M.S.; Schenkel, T.; Breinig, T.; Meyerhans, A.; Duschl, C. T Cell Activation on a Single-cell Level in Dielectrophoresis-based Microfluidic Devices. J. Chromatogr. A. 2008, 1202, 83–89. [Google Scholar]
- Lee, J.; Jang, J.; Akin, D.; Savran, C.A.; Bashir, R. Real-time Detection of Airborne Viruses on a Mass-sensitive Device. Appl. Phys. Lett. 2008, 93, 013901–3. [Google Scholar]
- McClain, M.A.; Culbertson, C.T.; Jacobson, S.C.; Ramsey, J.M. Flow Cytometry of Escherichia coli on Mirofluidic Devices. Anal. Chem. 2001, 73, 5334–5338. [Google Scholar]
- Ateya, D.A.; Sachs, R.; Besch, S.; Gottlieb, P.; Hua, S.Z. Volume Cytometry: Microfluidic Sensor for High-throughput Screening in Real-time. Anal. Chem. 2005, 77, 1290–1294. [Google Scholar]
- Shelby, J.P.; White, J.; Ganesan, K.; Rathod, P.K.; Chiu, D.T. A Microfluidic Model for Single-cell Capillary Obstruction by Plasmodium Falciparum-infected Erythrocytes. Proc. Nat. Acad. Sci. U.S.A. 2003, 309, 137–140. [Google Scholar]
- Yacoub-George, E.; Hell, W.; Meixner, L.; Wenninger, R.; Bock, K.; Lindner, P.; Wolf, H.; Kloth, T.; Feller, K.A. Automated 10-channel Capillary Chip Immunodetector for Biological Agents Detection. Biosens. Bioelectron. 2007, 22, 1368–1375. [Google Scholar]
- Huh, D.; Gu, W.; Kamotani, Y.; Grotberg, J.B.; Takayama, S. Microfluidics for Flow Cytometric Analysis of Cells and Particles. Physiol. Meas. 2005, 26, R73–98. [Google Scholar]
- Doh, I.; Cho, Y.-H. A Continuous Cell Separation Chip Using Hydrodynamic Dielectrophoresis (DEP) Process. Sens. Actuat. B 2005, 121, 59–65. [Google Scholar]
- Suehiro, J.; Hamada, R.; Noutomi, D.; Shutou, M.; Hara, M. Selective Detection of Viable Bacteria Using Dielectrophoretic Impedance Measurement Method. J. Electrostat. 2003, 57, 157–168. [Google Scholar]
- Suehiro, J.; Ohtsubo, A.; Hatano, T.; Hara, M. Selective Detection of Bacteria by a Dielectrophoretic Impedance Measurement Method Using an Anitbody-immobilized Electrode Chip. Sens. Actuat. B 2006, 119, 319–326. [Google Scholar]
- Balagadde, F.K.; You, L.C.; Hansen, C.L.; Arnold, F.H.; Quake, S.R. Long-term Monitoring of Bacteria Undergoing Programmed Population Control in a Microchemostat. Science 2005, 309, 137–140. [Google Scholar]
- Futai, N.; Gu, W.; Song, J.W.; Takayama, S. Handheld Recirculation System and Customized Media for Microfluidic Cell Culture. Lab Chip 2006, 6, 149–154. [Google Scholar]
- Ertl, P.; Robello, E.; Battaglini, F.; Mikkelsen, S.R. Rapid Antibiotic Susceptibility Testing via Electrochemical Measurement of Ferricyanide Reduction by Escherichia coli and Clostridium sporogenes. Anal. Chem. 2000, 72, 4957–4964. [Google Scholar]
- Ertl, P.; Mikkelsen, S.R. Electrochemical Biosensor Array for the Identification of Microorganisms Based on Lectin-lipopolysaccharide Recognition. Anal. Chem. 2001, 73, 4241–4248. [Google Scholar]
- Peterfi, Z.; Kustos, I.; Kilar, R.; Kocsis, B. Microfluidic Chip Analysis of Outer Membrane Proteins Responsible for Serological Cross-reaction Between Three Gram-negative Bacteria: Proteus morganii O34, Escherichia coli O111 and Salmonella Adelaide O35. J. Chromatogr. A. 2007, 1155, 214–217. [Google Scholar]
- Ekins, R.P. Immunoassay, DNA Analysis and Other Ligand Binding Assay Techniques: From Electrophrograms to Multiplexed, Ultrasensitive Microarrays on a Chip. J. Chem. Educ. 1999, 76, 769–780. [Google Scholar]
- Minerick, A.R. The Rapidly Growing Field of Micro and Nanotechnology to Measure Living Cells. AICHE J. 2008, 54, 2230–2237. [Google Scholar]
- Leonard, P.; Hearty, S.; Quinn, J.; O'Kennedy, R. A Generic Approach for the Detection of Whole Listeria Monocytogenes Cells in Contaminated Samples Using Surface Plasmon Resonance. Biosens. Bioelectron. 2004, 19, 1331–1335. [Google Scholar]
- Gomez, R.; Bashir, R.; Bhunia, A.K. Microscale Electronic Detection of Bacterial Metabolism. Sens. Actuat. B 2002, 86, 198–208. [Google Scholar]
- Muhammead-Tahir, Z.; Alocilja, E.C. Fabrication of a Disposable Biosensor for Escherichia coli O157:H7 Detection. IEEE Sens. J. 2003, 4, 345–351. [Google Scholar]
- Hawkes, J.J.; Long, J.M.; Coakley, W.T.; McDonnell, M.B. Ultrasonic Deposition of Cells on a Surface. Biosens. Bioelectron. 2004, 19, 1021–1028. [Google Scholar]
- Verporte, E. Beads and Chips: New Recipes of Analyses. Lab Chip 2003, 3, 60N–68N. [Google Scholar]
- Masafumi, I.; Nobuyasu, Y.; Katsuji, T.; Masao, N. Rapid and Simple Detection of Food Poisoning Bacteria by Bead Assay with a Microfluidic Chip-based System. J. Microbiol. Methods 2006, 67, 241–247. [Google Scholar]
- Floriano, P.N.; Christoduolides, N.; Romanovicz, D.; Bernard, B.; Simmons, G.W.; Cavell, M.; McDevitt, T.T. Membrane-based On-line Optical Analysis System for Rapid Detection of Bacteria and Spores. Biosens. Bioelectron. 2005, 20, 2079–2088. [Google Scholar]
| Company | Target | Website |
|---|---|---|
| Advanced Liquide Logics | Immunoassay | www.liquid-logic.com |
| Cepheid | DNA | www.cepheid.com |
| CombiMatrix CustomArray | DNA, biological threads | www.combimatrix.com |
| Invitrogen | DNA | www.invitrogen.com |
| Affymetrix | DNA | www.affymetrix.com |
| Caliper | DNA | www.caliperls.com |
| Febit | DNA/RNA/Proteins | www.febit.com |
| Claros Diagnostics | Proteins | www.clarosdx.com |
| HandyLab | DNA, proteins | www.handylab.com |
| Abbott Diagnostics (iStat) | Markers | www.istat.com |
| LabNow | HIV/AIDS | www.labnow.com |
| Micronics | Enteric pathogens | www.micronics.net |
| Nanogen | DNA/RNA | www.nanogen.com |
| Nanosphere | DNA, proteins | www.nanosphere-inc.com |
| Sensata (Spreeta) | Viruses, bacteria | www.sensata.com |
| Sequella | Proteins | www.sequella.com |
| BIAcore | Bacteria, viruses | www.biacore.com |
| Canary | B-lymphocytes | www.canarysystem.com |
| Rapid Plex (ICx Biosystems) | Bacteria, protein | www.inivtrogen.com |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Mairhofer, J.; Roppert, K.; Ertl, P. Microfluidic Systems for Pathogen Sensing: A Review. Sensors 2009, 9, 4804-4823. https://doi.org/10.3390/s90604804
Mairhofer J, Roppert K, Ertl P. Microfluidic Systems for Pathogen Sensing: A Review. Sensors. 2009; 9(6):4804-4823. https://doi.org/10.3390/s90604804
Chicago/Turabian StyleMairhofer, Jürgen, Kriemhilt Roppert, and Peter Ertl. 2009. "Microfluidic Systems for Pathogen Sensing: A Review" Sensors 9, no. 6: 4804-4823. https://doi.org/10.3390/s90604804





