Two Phase Non-Rigid Multi-Modal Image Registration Using Weber Local Descriptor-Based Similarity Metrics and Normalized Mutual Information
Abstract
: Non-rigid multi-modal image registration plays an important role in medical image processing and analysis. Existing image registration methods based on similarity metrics such as mutual information (MI) and sum of squared differences (SSD) cannot achieve either high registration accuracy or high registration efficiency. To address this problem, we propose a novel two phase non-rigid multi-modal image registration method by combining Weber local descriptor (WLD) based similarity metrics with the normalized mutual information (NMI) using the diffeomorphic free-form deformation (FFD) model. The first phase aims at recovering the large deformation component using the WLD based non-local SSD (wldNSSD) or weighted structural similarity (wldWSSIM). Based on the output of the former phase, the second phase is focused on getting accurate transformation parameters related to the small deformation using the NMI. Extensive experiments on T1, T2 and PD weighted MR images demonstrate that the proposed wldNSSD-NMI or wldWSSIM-NMI method outperforms the registration methods based on the NMI, the conditional mutual information (CMI), the SSD on entropy images (ESSD) and the ESSD-NMI in terms of registration accuracy and computation efficiency.1. Introduction
Non-rigid image registration is one of the most challenging problems in medical image processing. Given two medical images, the objective of the registration process is to find a reasonable non-rigid transformation, such that a transformed version of the float image is similar to the reference one. Despite phenomenal progress in medical image resolution, one modality is often not sufficient to produce a precise diagnosis since different imaging modalities differ in interpreting the anatomy, tissue and organ that they may capture, so multi-modal medical image registration is useful for relating clinically significant information from different images. For example, it can be used to improve the diagnostic tasks and image-guided interventions. However, accurate non-rigid multi-modal registration is highly challenging because of intensity variations and non-rigid transformations between images.
In general, image registration involves three main components: deformation model, similarity metric and optimization strategy. In the non-rigid image registration, the deformation model can be divided into two main categories [1]. The first category is originated from physical models of materials including elastic body models [2], fluid flow models [3] and diffusion models [4]. Another category is relevant to interpolation and approximation theory including radial basis functions [5], elastic body splines [6], free-form deformations (FFD) [7,8].
As regards optimization strategy, numerous optimization methods have been proposed to optimize the parameters of the deformation model in the non-rigid image registration. Examples include gradient descent, Newton's method, Powell's method and discrete optimization [9,10]. Especially, the Broyden–Fletcher–Goldfarb–Shanno (BFGS) algorithm [11] has been used to handle the large number of variables and constraints of registration and is included in the Insight Segmentation and Registration Toolkit (ITK) [12–14].
Apart from deformation model and optimization strategy, similarity metrics have received much attention in the field of image registration. Many similarity metrics have been proposed for different applications [15]. The well-known metrics such as sum of squared differences (SSD) and sum of absolute difference (SAD) have been successfully used for mono-modal image registration, but they are not appropriate for direct application to multi-modal image registration because it is required that the image intensities at corresponding points of two images should be similar [16]. To address this problem, the generalized divergence measure based on Renyi Entropy [17], Kullback-Leibler Divergence [18], mutual information (MI) and cross-cumulative residual entropy (CCRE) [19] have been proposed. Among these metrics, MI has been investigated in-depth and widely applied to multi-modal image registration. MI was firstly introduced to realize the rigid registration of multi-modal scans by Maes et al. [20] and Viola et al [21]. Rueckert et al. [7] extended this similarity metric to non-rigid image registration. Studholme et al. proposed the normalized mutual information (NMI) as an overlap invariant generalization of mutual information [22]. Conditional mutual information (CMI) was introduced for non-rigid multi-modal image registration by Loeckx et al. to reduce the negative influence of bias fields [23]. However, the traditional global MI approach involves such disadvantages as high computational complexity, tendency to get trapped in local minima and suffering from erroneous transformations of anatomical images even though the MI between the fixed image and the float image achieves the maximum value with this transform [24,25].
In addition to information theory measures-based registration methods, structural representation methods have been investigated for multi-modal medical image registration. Pluim et al. [26] utilized the local gradient orientation and Nigris et al. [27] used gradient orientations of minimal uncertainty for image registration. In [28], Liu et al. proposed a registration method based on the local frequency estimated by calculating the local phase gradient of the most significant Gabor filter response. This method has the advantage that the local frequency is the same for corresponding structures in the two images, even when edge strength and contrast have significant differences. However, the above three methods were used for rigid registration, and not discussed for the non-rigid registration in [26–28]. Heinrich et al. [29] utilized modality independent neighbourhood descriptor (MIND) which is not a scalar representation but a vector-valued image descriptor for multi-modal deformable registration. Wachinger et al. proposed two structural representation methods. One method utilized the entropy of an image patch to assign a new intensity value and used the SSD on entropy images (ESSD) as similarity metric. Another method firstly used Laplacian Eigenmaps to embed image patches in a lower-dimensional manifold that preserves local distances and then computed L2 distance of Laplacian images. However, the entropy image based method can only meet certain requirements of a relaxed version of the theoretical properties, and the Laplacian image based method involves high computational complexity [30].
To achieve both accuracy and efficiency of the non-rigid multi-modal registration method, a similarity metric based on the Weber local descriptor (WLD) proposed in [31] is combined with the NMI to realize a two phase image registration in this paper. The introduction of the WLD in this paper results from the fact that it can extract image features more effectively than the well-known scale-invariant feature transform and local binary patterns. In the proposed method, the first phase aims at obtaining the parameters (i.e., the control points) relevant to the large deformation, ensuring that the anatomical features of the reference and float images are not destroyed, and reducing the opportunities of getting trapped in local minima by integrating the WLD with the non-local SSD or the structural similarity (SSIM) proposed in [32]. The second phase is focused on getting the parameters related to the small deformation using the NMI to obtain high registration accuracy.
This paper is structured as follows: Section 2 presents the novel two phase non-rigid multi-modal image registration method. Section 3 provides discussions of key parameters in the proposed method and comparisons of registration accuracy and efficiency among our method and the NMI, CMI, ESSD and ESSD-NMI methods. Finally, the conclusion is given in Section 4.
2. Methods
2.1. Registration Framework
In general, image registration is stated as the following minimization problem:
In Equation (1), it is highly challenging to tackle large deformations. On the one hand, coarse-to-fine deformation schemes have been commonly applied. Wu et al. [33] used a wavelet-based deformation model to treat global and local information in a coarse-to-fine approach. The combination of coarse-scale landmark and B-splines deformable registration techniques was proposed in [34]. A hybrid method that combined surface and volume information to register cortical structures was proposed in [35]. Postelnicu et al. [36] started with a geometric registration which was used as the initialization and refined it with a non-linear optical flow registration method. On the other hand, to prevent from folding artifacts and preserve local orientation in case of large deformations, the deformation field should be diffeomorphisms. Rueckert et al. [8] enforced the transformation of FFD model to be a diffeomorphism by limiting the control points displacement.
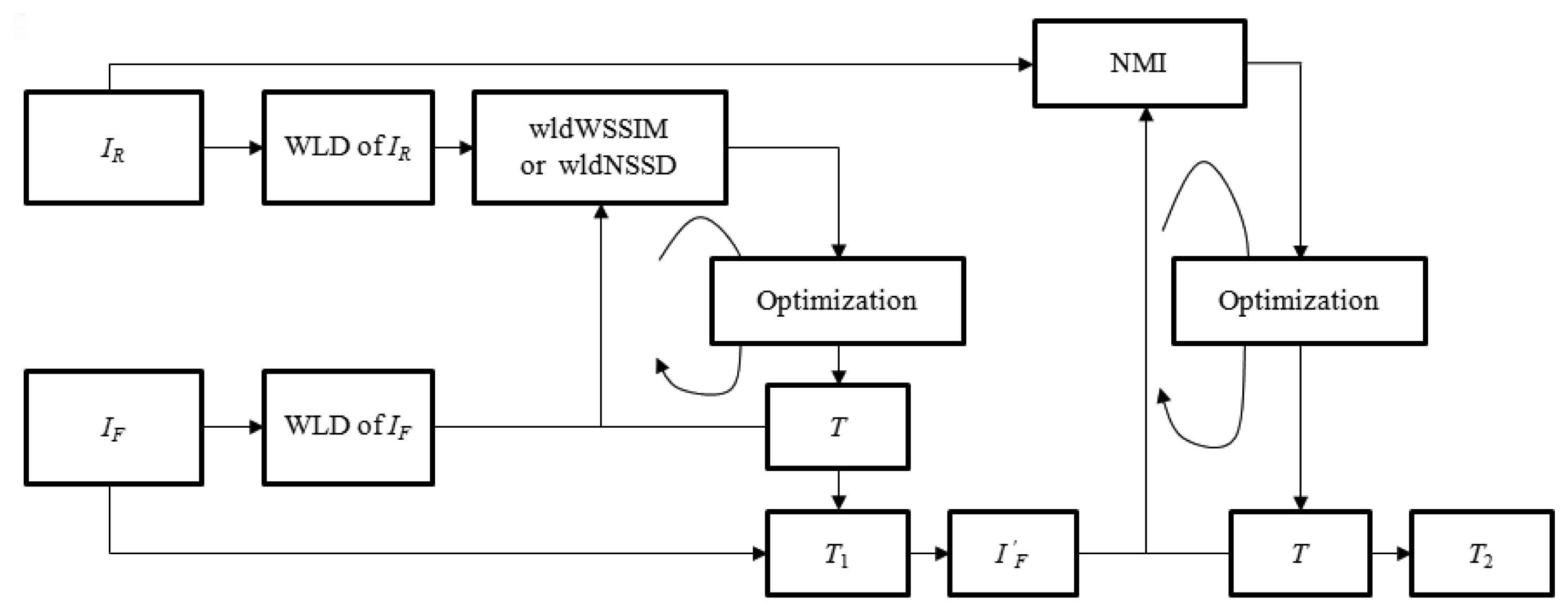
Different from above-mentioned coarse-to-fine deformation registration methods, the proposed two phase image registration method aims at resolving the above minimization problem by using different similarity metrics in two registration phases as shown in Figure 1. In the large deformation phase, the structural representations of IR and IF are firstly obtained using the WLD to compute similarity metric wldNSSD or wldWSSIM. Then, the iterative optimization of the objective function f1 defined by the wldNSSD or the wldWSSIM is realized to obtain a relatively good initial transformation T1 while reducing the opportunities of getting trapped in the local minima. However, only using the wldNSSD or the wldWSSIM cannot ensure accurate registration results especially for medical images with complicated deformations because some useful image information may be lost when using the WLD to extract image features. Therefore, a refined registration is implemented by using the original image intensities. In the small deformation phase, the float image IF is firstly deformed to generate the image using T1 which can provide a good initial value, and then the small deformation for is processed for minimizing the objective function f2 defined by the NMI to obtain the final transformation T2.
In our method, we used the diffeomorphic FFD model as the deformation model which uses a multi-resolution way by concatenating the FFDs with different grid sizes and limiting the control points displacement less than 0.4 × the grid sizes [8]. Meanwhile, we selected the BFGS algorithm as the optimization method because it is particularly suited to the registration problem with very large numbers of variables without requiring explicit computations.
2.2. Large Deformation Phase
2.2.1. Weber Local Descriptor
Ernst Weber observed that the ratio of the increment threshold to the background intensity is a constant in human perception [37]. This relationship between the physical magnitudes of stimuli and the perceived intensity of the stimuli was known as Weber's Law which can be expressed as:
Although Weber's Law describes fundamental relationships of the human perception, i.e., in a biological setting rather than for digital images, several researchers have extended its application to signal and image processing. Dabeer and Onkar introduced a regularized Weber sampler for smooth deterministic signals [38]. Bruni et al. used Weber's Law for scratch detection on digital film materials [39]. By using Weber's Law, Chen proposed a Weber local descriptor (WLD) which was used to extract local features [31].
The WLD has two components: differential excitation (ξ) and orientation (θ). In our paper, only differential excitation is used for registration because using both differential excitation and orientation result in increased computational complexity, and differential excitation has such advantages as detecting edges elegantly, robustness to noise and illumination change, and its powerful representation ability for textures [31]. Specifically, a differential excitation ξ(xc)of a current pixel xc is computed as illustrated in Figure 2. The symbols and are the outputs of the filters f00 and f01. It is easy to understand that . The difference between the center point xc and its neighbors is given by:
By applying the arctangent function which can limit the output to prevent from increasing or decreasing too quickly when the input becomes larger or smaller [31], the differential excitation ξ(xc) of the pixel xc is computed as:
To tailor the WLD to our non-rigid multi-modal registration method, it is expressed as:
Actually, WLD features can be extracted from a square symmetric neighborhood of size (2R + 1) × (2R + 1). Figure 3 shows examples of the neighborhood with R = 1 and R = 2. It should be noted that for extracting WLD features, only the border pixels (highlighted in blue in Figure 3) are used instead of all the pixels in the square neighborhood to reduce computation time.
To demonstrate the performance of the WLD with different R and its advantage over the entropy image for structural representation, the WLD and the entropy image for deformed T1, PD and original T2 weighted MR images are shown in Figure 4. The comparison between Figure 4(b) and Figure 4(c–e) shows that the detail information in the entropy image of T1 weighted MR images is blurrier than that in the WLD, which indicates that the WLD can provide better structural representation than the entropy image for registration. It should be noted that there exist the black squares and dots in Figure 4(b) because the considered pixel is set to be zero when the minimum intensity is equal to the maximum one in the image patch centered at this pixel. Meanwhile, it is easy to understand that the performance of the WLD changes with different R.
From Figure 4(c–e), (h–j) and (m–o), we can see that the WLD with a small R (e.g., R = 1) generates relatively weak but thin edges and thus facilitates the accurate localization of relatively strong edges. By comparison, the WLD with a larger R (e.g., R = 2) produces thicker but stronger edges and thus assists with detecting the weak edges while a much larger R (e.g., R = 3) results in poor edge localization. The zooming of features marked with the red square in Figure 4(c–d) and those marked with the blue square in Figure 4(d–e) can further illustrate the above characteristics. Obviously, the combination of multiple selected R for the WLD produces “multi-granuality” features especially in regions with rich textures and discontinuities. Therefore, using the various radii in the WLD is preferable to using a single radius for the effective structural representation of medical images with complicated features. Based on the above analysis, we use the WLD with R = 1 and R = 2 for structural representation of images.
2.2.2. Weber Local Descriptor Based Non-Local Sum of Squared Differences
To describe the difference between the extracted WLD feature of IR and that of IF, the traditional SSD is improved to resist the disadvantageous influence of noise in the medical images. Inspired by the idea of non-local means proposed by Buades et al. [41], which aims at image denoising, that the pixel similarity can be represented more effectively using image patches than using individual pixels, the non-local SSD (NSSD) is introduced to represent the difference between the WLD features. The NSSD between two images IA and IB is presented as:
As discussed in Section 2.2.1, by combining the WLD using the neighborhoods of different radii R1 = 1 and R2 = 2 with the NSSD, we can obtain the following similarity metric wldNSSD:
Based on the wldNSSD, we define the objective function f1 as:
2.2.3. Weber Local Descriptor Based Weighted Structural Similarity
Given two images IA and IB to be compared, the SSIM involves such components as a luminance (mean) distortion term l(IA, IB), a contrast (variance) distortion term c(IA, IB) and a correlation term s(IA, IB). By combining the three comparisons, the resultant SSIM index between IA and IB is presented in a simplified form as [32]:
The SSIM index is not a metric. However, the distance is a single scalar-valued distance measure [42]. In [32], a mean SSIM has been used for image quality assessment. However, it is helpful to improve the performance of image quality assessment algorithms by giving different weights to different image patches [43]. Therefore, the WLD difference based weighted SSIM (wldWSSIM) is adopted as a metric:
Similar to the wldNSSD, the similarity metric wldWSSIM is defined as:
Based on the wldWSSIM, the objective function f1 is defined as:
2.2.4. Analysis of the wldNSSD and the wldWSSIM
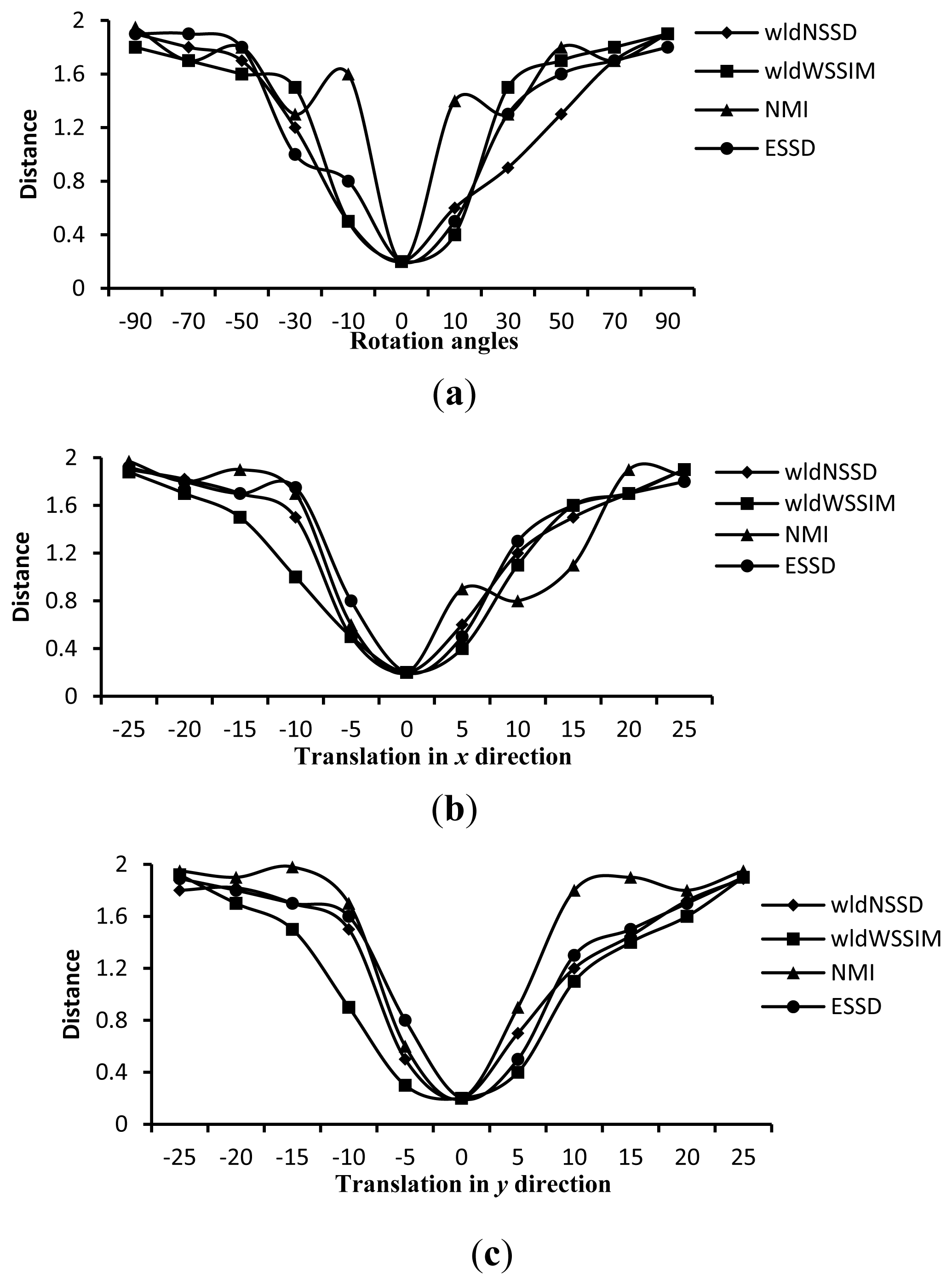
To demonstrate the advantage of the wldNSSD and the wldWSSIM over such metrics as the NMI and the ESSD, we compute the four distance measures on ten pairs of T1 and T2 weighted MR images rotated around the domain center with different angles and translated in x and y directions. Figure 5 shows an excerpt of the corresponding computation results. We can see from Figure 5 that the wldNSSD and wldWSSIM have no local extremes while the NMI has local extremes in Figure 5(a–c), and the ESSD involves local extremes in Figure 5(b). Similar findings also exist for other computation results, which indicate that the wldNSSD and the wldWSSIM are more effective similarity metrics for registration than the NMI and the ESSD.
2.3. Small Deformation Phase
To obtain more accurate registration results, the small deformation phase is needed for the refined registration. In this phase, the float image IF is firstly deformed using T1 which is the output of the large deformation phase. So, we have . The small deformation is processed by minimizing the objective function f2 defined by the NMI between and IR:
3. Experiments
In this section, to determine the key parameters in the proposed method and make comparisons of registration performance among our method, the NMI method, the CMI method, the ESSD method and the ESSD-NMI method, extensive experiments have been performed on thirty T1, T2 and PD-weighted MR images of size 256 × 212 pixels from the BrainWeb database [45]. Registration efficiency of all these evaluated methods is appreciated by their computation time (in seconds) when implemented in a multi-resolution way by rescaling the FFD grid spacing 2k × 2k (each image is rescaled to a square of size 2 to the kth power) [8] using MATLAB 2010 on a personal computer with 2.40 GHz CPU and 4 GB RAM. Registration accuracy is appreciated by target registration error (TRE) [46] with simulated deformation and expert landmark annotations as ground truth, respectively.
As regards the simulated deformation Ts, it is used as the ground truth to deform one of two different weighted MR images (e.g., T1 and T2). By implementing the various registration methods on the float image and the generated reference image, the estimated deformation Tc will be obtained. Based on the whole image domain, TREd is computed as:
When we use expert landmark annotations as ground truth, for a estimated deformation Tc and a set of anatomical landmark pairs (i=1, 2,···, m, where m is the number of anatomical landmarks), TREl is defined as:
3.1. Choice of Parameters
3.1.1. Choice of the patch size
The patch sizes SP1 and SP2 are very important for the similarity metrics wldNSSD and wldWSSIM. To quantitatively determine these parameters, we discuss their influence on registration accuracy and efficiency by using fifteen images.
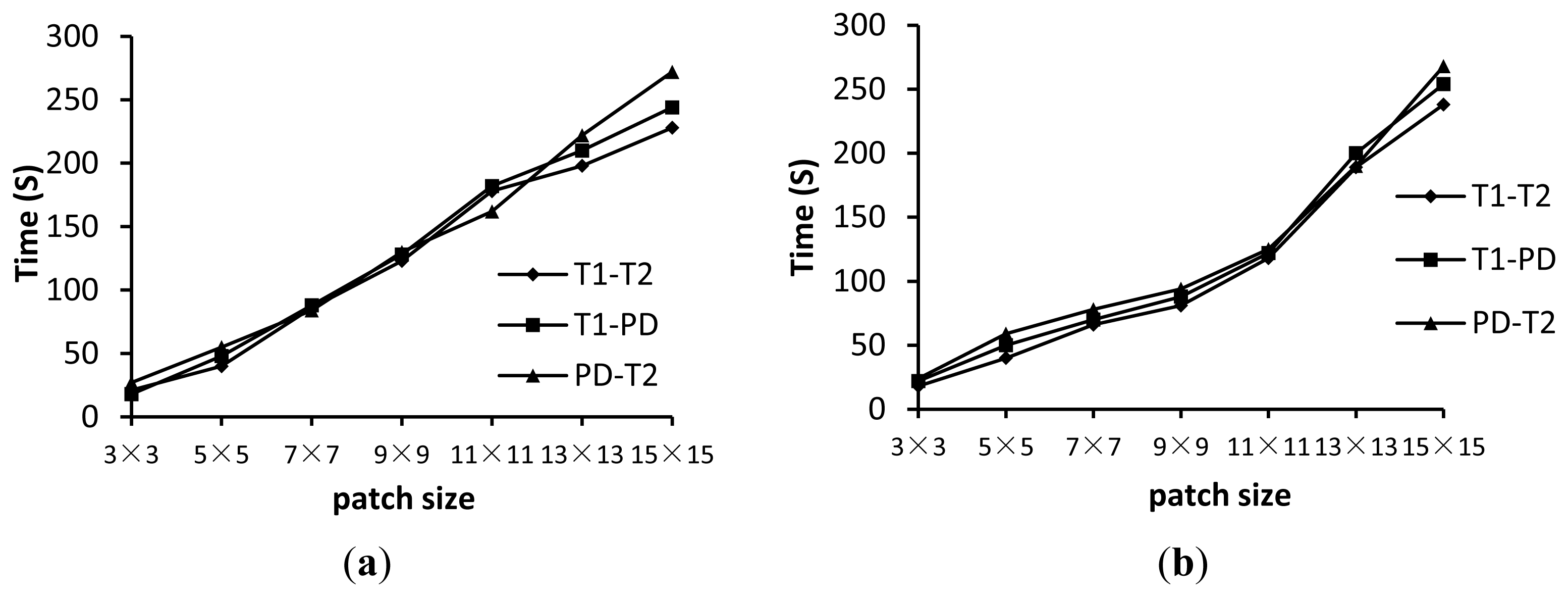
Figure 7 shows the TREl for the wldNSSD and wldWSSIM methods using various patch sizes. Figure 8 shows the computation time for the two metrics using various patch sizes. We can see from Figure 7 that a too small or too large patch size has a disadvantageous influence on registration accuracy. For the wldNSSD method, the TREl can achieve the minimum value of 7.5 mm when the patch size is 7 × 7 or 9 × 9 while the optimal patch size is 11 × 11 for the wldWSSIM method. Meanwhile, it is shown in Figure 8 that the computation time increases with the increasing patch size because a larger patch size means that more pixels need to be processed. Therefore, to achieve the tradeoff between registration precision and efficiency, the patch size is chosen to be 7 × 7 for the wldNSSD method and 11 × 11 for the wldWSSIM method in our experiments. It should be noted that the wldWSSIM method needs bigger patches for higher registration performance than the wldNSSD. The reason can be explained in this way. The wldNSSD depends on the difference of intensities of two WLD feature images while the wldSSIM takes into account not only the local mean and the local variance, but also the local covariance of two WLD feature images for representing their structural similarity. Therefore, for the wldWSSIM method, a larger patch size is needed to ensure the statistical significance.
3.1.2. Choice of the Weighting Term γ
The weighting term γ is specific to the processed images. To quantitatively determine γ, we make some tests on γ with fifteen T1, T2 and PD-weighted MR images.
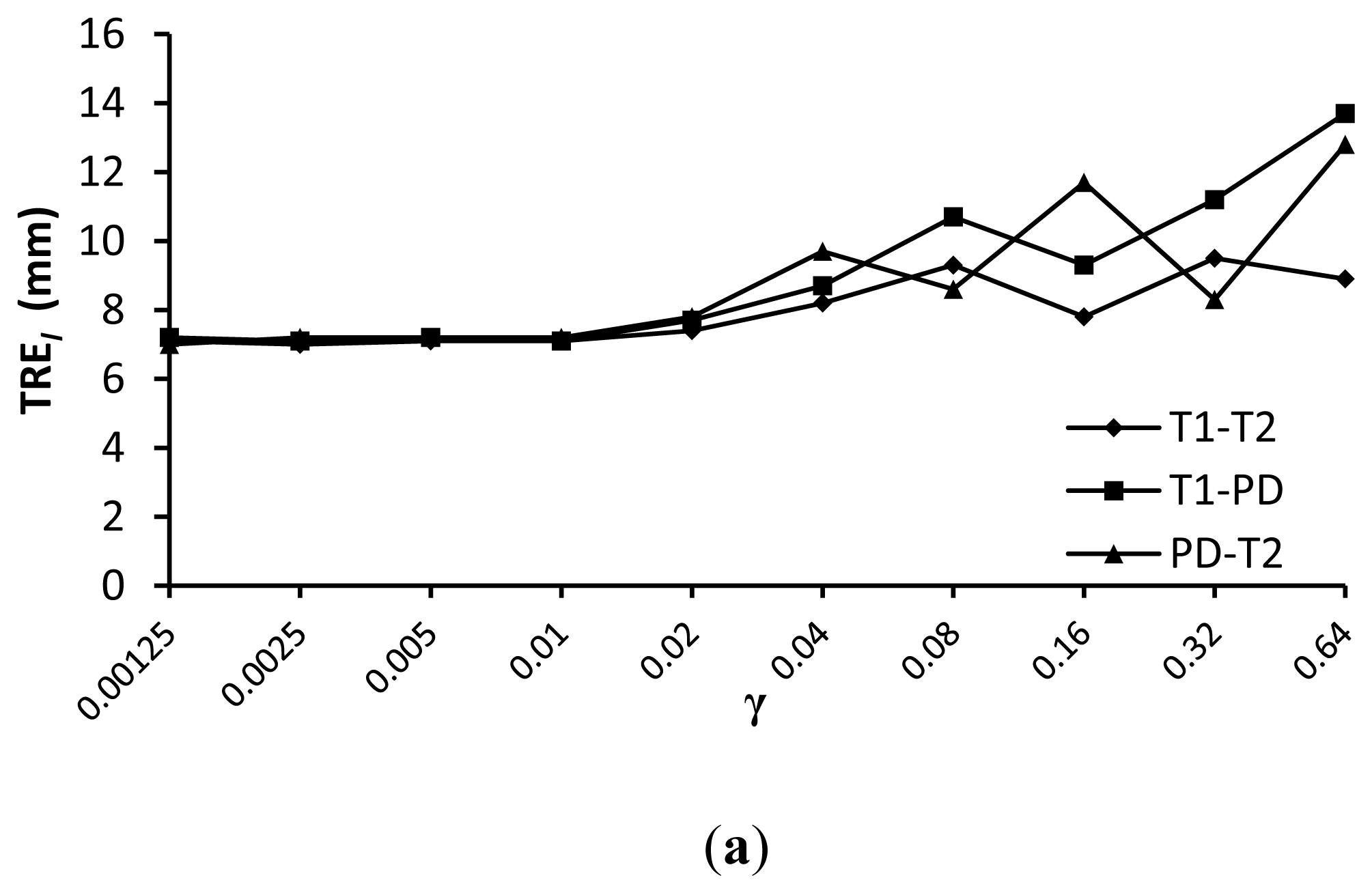
The comparison results of registration precision with different γ for similarity measures wldWSSIM and wldNSSD are shown in Figure 9. We can see from this figure that a value of γ =0.01 provides good registration results for the various weighted MR images. The reason lies in two aspects. On the one hand, the target registration error TREl becomes unstable with a larger γ because it will reduce the impact of the similarity measure which is important for registration precision. On the other hand, the regularization term which is used to constrain the FFD transformation to be smooth is less important because we have limited the control point displacement less than 0.4 × the grid sizes in the FFD model.
3.2. Comparison of Registration Performance
To demonstrate the advantage of the wldNSSD and wldWSSIM, wldNSSD-NMI and wldWSSIM-NMI methods, they are compared with other evaluated methods in terms of registration accuracy and efficiency.
The mean and standard deviation (std) of TREd and TREl for all the evaluated methods are shown in Table 1 and Table 2. Here, “/” in Tables 1 and 2 means that no registration is implemented. Obviously, the wldNSSD and wldWSSIM methods can achieve a smaller TREd and TREl than the NMI, CMI and ESSD methods. Meanwhile, we can see that the wldNSSD-NMI and wldWSSIM-NMI methods for two deformation phases can achieve higher registration accuracy than the corresponding wldWSSIM and wldNSSD methods, which demonstrates the advantage of combining the WLD similarity metrics with the NMI. It should be noted that the NMI method can be seen as a two phase method since it was implemented in a multi-resolution way in our test, but this method achieves a bigger TREd and TREl than the wldWSSIM-NMI and wldNSSD-NMI methods because the NMI method is easy to get trapped into local minima only using the image intensity information. Besides, the intra-observer errors and inter-observer errors are listed in Table 2 for reference. It is shown that the smallest registration error for the wldWSSIM-NMI method is 6.1 mm and it is still higher than the inter-observer errors. The main reason is that experts were requested to repeat the marking procedure if necessary to ensure that the inter-observer errors are less than 5.0 mm in the case of the large simulated deformation among different weighted MR images.
Table 3 lists the computation time of all these methods. The observation from Table 3 shows that our methods have less computation time than the NMI, CMI, ESSD and ESSD-NMI methods and thus they outperform these compared methods in terms of registration efficiency. Moreover, the wldWSSIM-NMI method is of lower registration efficiency than the wldNSSD-NMI while the former can provide higher registration accuracy. Therefore, our two phase methods can be applied according to the clinical requirements. If one application attaches more significance to registration accuracy, the wldWSSIM-NMI method is better choice. If registration efficiency is more important, the wldNSSD-NMI method is preferable.
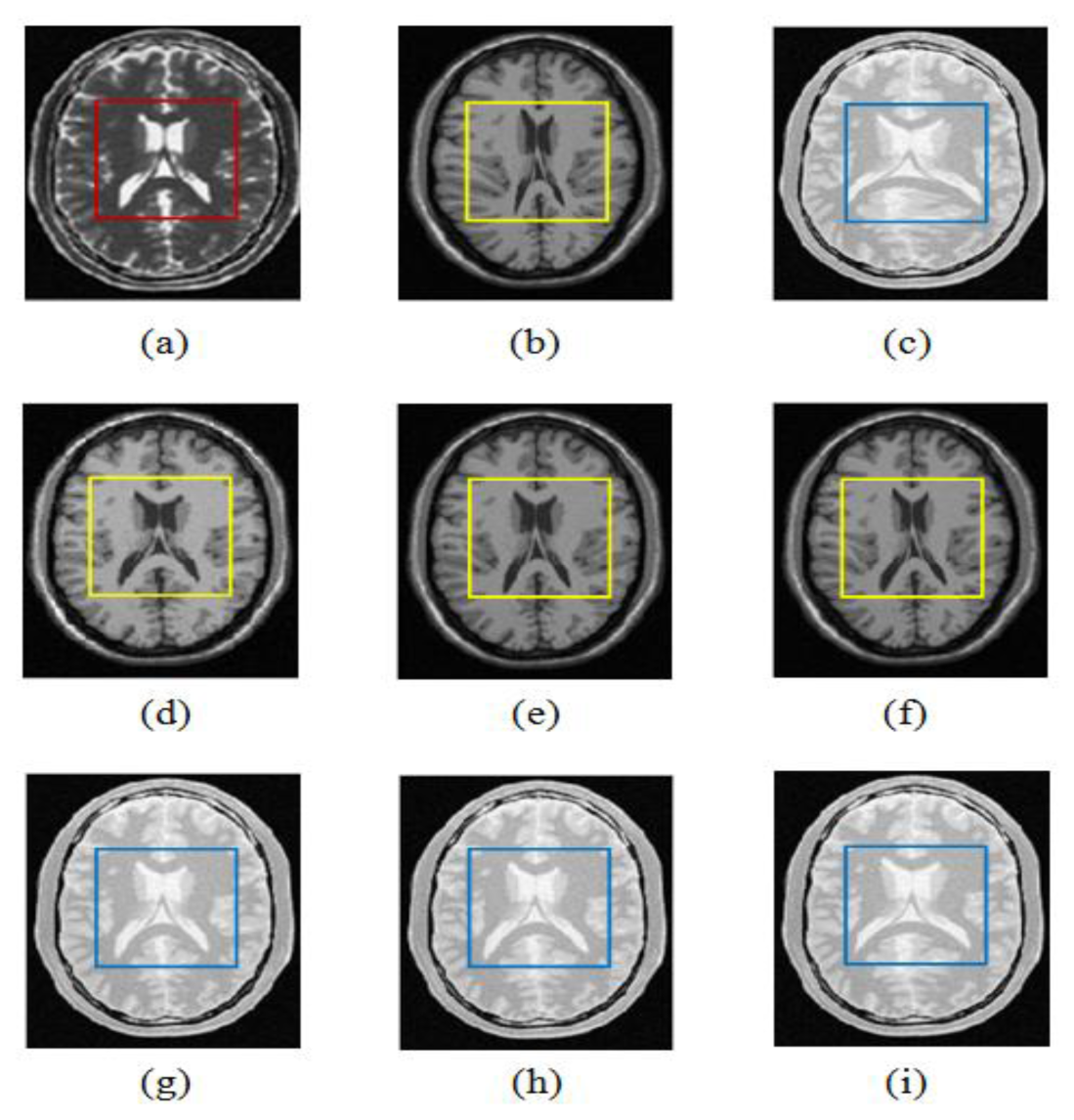
Figure 10 shows an example of T1-T2 and PD-T2 weighted MR images and registration results for the wldWSSIM-NMI, wldNSSD-NMI and ESSD-NMI methods. From Figure 10(d–f), we can see that the results using the wldWSSIM-NMI and wldNSSD-NMI methods are anatomically more similar to the reference T2 weighted MR image than those using the ESSD-NMI, especially in the lateral ventricle which is marked with a yellow square. Similar results can be seen in Figure 10(g–i).
4. Conclusions
In this paper, we have proposed a two phase non-rigid multi-modal medical image registration method using the Weber local descriptor based similarity metrics and the normalized mutual information. In the first phase, the parameters relevant to the large deformation are obtained by minimizing the objective function defined by the novel similarity metric wldNSSD or wldWSSIM which is focused not on intensities of individual pixels but on structural information of images. With the good initial deformation value provided by the output of the large deformation phase, the parameters related to the small deformation can be accurately obtained using the NMI in the second phase. The non-rigid image registration experiments on the T1, T2 and PD weighted MR images demonstrate that compared with the NMI, CMI, ESSD and ESSD-NMI methods, our method can obtain smaller registration errors and higher computational efficiency. Future work will be focused on extending our method to multi-modal 3D medical image registration.
Acknowledgments
This work was partly supported by the National 973 project (Grant No.: 2011CB933103), the Project of the National 12th-Five Year Research Program of China (Grant No.: 2012BAI13B02), the National Natural Science Foundation of China (NSFC) (Grant No.: 30911120497) and the Biology and Bioinformatics Ph.D. Programs Construction Project of Guangxi University, China.
Conflict of Interest
The authors declare no conflict of interest.
References
- Holden, M. A review of geometric transformations for nonrigid body registration. IEEE Trans. Med. Imaging 2008, 27, 111–128. [Google Scholar]
- Kohlrausch, J.; Rohr, K.; Stiehl, H.S. A new class of elastic body splines for nonrigid registration of medical images. J. Math. Imag Vis. 2005, 23, 253–280. [Google Scholar]
- Agostino, E.D.; Maes, F.; Vandermeulen, D.; Suetens, P. A viscous fluid model for multimodal non-rigid image registration using mutual information. Med. Image Anal. 2003, 7, 565–575. [Google Scholar]
- Vercauteren, T.; Pennec, X.; Malis, E.; Perchant, A.; Ayache, N. Insight into efficient image registration techniques and the demons algorithm. Inf. Process. Med. Imaging. 2007, 20, 495–506. [Google Scholar]
- Yang, X.; Xue, Z.; Liu, X.; Xiong, D. Topology preservation evaluation of compact-support radial basis functions for image registration. Pattern Recogn. Lett. 2011, 32, 1162–1177. [Google Scholar]
- Rohr, K.; Wörz, S. An Extension of Thin-plate Splines for Image Registration with Radial Basis Functions. Proceedings of 9th IEEE International Symposium on Biomedical Imaging (ISBI), Barcelona, Spain, 2–5 May 2012; pp. 442–445.
- Rueckert, D.; Sonoda, L.; Hayes, C.; Hill, D.; Leach, M.; Hawkes, D. Nonrigid registration using free-form deformations: Application to breast MR images. IEEE Trans. Med. Imaging. 1999, 18, 712–721. [Google Scholar]
- Rueckert, D.; Aljabar, P.; Heckemann, R.A.; Hajnal, J.V.; Hammers, A. Diffeomorphic Registration using B-splines. Proceedings of MICCAI 2006, Copenhagen, Denmark, 1–6 October 2006; pp. 702–709.
- Komodakis, N.; Tziritas, G.; Paragios, N. Performance vs computational efficiency for optimizing single and dynamic mrfs: Setting the state of the art with primal-dual strategies. Comput. Vis. Image Underst. 2008, 112, 14–29. [Google Scholar]
- Glocker, B.; Komodakis, N.; Tziritas, G.; Navab, N.; Paragios, N. Dense image registration through MRFs and efficient linear programming. Med. Image Anal. 2008, 12, 731–741. [Google Scholar]
- Morales, J.L.; Nocedal, J. Remark on “algorithm 778: L-BFGS-B: Fortran subroutines for large-scale bound constrained optimization”. ACM Trans. Math. Softw. 2011, 38, 71–74. [Google Scholar]
- Sdika, M. A fast nonrigid image registration with constraints on the Jacobian using large scale constrained optimization. IEEE Trans. Med. Imaging 2008, 27, 271–281. [Google Scholar]
- So, R.W.K.; Chung, A.C.S. Multi-modal non-rigid Image Registration based on Similarity and Dissimilarity with the Prior Joint Intensity Distributions. Proceedings of 7th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Rotterdam, The Netherlands, 14–17 April 2010; pp. 368–371.
- ITK. Available online: http://www.itk.org/Doxygen/html/classitk_1_1LBFGSBOptimizer.html (accessed on 6 June 2013).
- Klein, A.; Andersson, J.; Ardekani, B.A.; Ashburner, J.; Avants, B.; Chiang, M.C.; Christensen, G.E.; Collins, D.L.; Gee, J.; Hellier, P.; et al. Evaluation of 14 nonlinear deformation algorithms applied to human brain MRI registration. NeuroImage 2009, 46, 786–802. [Google Scholar]
- Clarysse, P.; Sipila, O.; Pauna, N.; Pham, Quoc Cuong; Katila, T.; Magnin, I.E. A review of cardiac image registration methods. IEEE Trans. Med. Imaging 2002, 21, 1011–1021. [Google Scholar]
- Hamza, A.B.; Krim, H. A generalized divergence measure for robust image registration. IEEE Trans. Signal Process. 2003, 51, 1211–1220. [Google Scholar]
- Guetter, C.; Xu, C.; Sauer, F.; Hornegger, J. Learning Based Non-rigid Multi-Modal Image Registration Using Kullback-Leibler Divergence. Proceedings of MICCAI 2005, Palm Springs, CA, USA, 26–30 October 2005; pp. 255–262.
- Fei, W.; Baba, C.V. Non-rigid multi-modal image registration using cross-cumulative residual entropy. Int. J. Comput. Vis. 2007, 74, 201–215. [Google Scholar]
- Maes, F.; Collignon, A.; Vandermeulen, D.; Marchal, G.; Suetens, P. Multimodality image registration by maximization of mutual information. IEEE Trans. Med. Imaging 1997, 16, 187–198. [Google Scholar]
- Viola, P.; Wells, W., III. Alignment by maximization of mutual information. Int. J. Comput. Vis. 1997, 24, 137–154. [Google Scholar]
- Studholme, C.; Hill, D.; Hawkes, D. An overlap invariant entropy measure of 3D medical image alignment. Pattern Recogn. 1999, 32, 71–86. [Google Scholar]
- Loeckx, D.; Slagmolen, P.; Maes, F.; Vandermeulen, D.; Suetens, P. Nonrigid image registration using conditional mutual information. IEEE Trans. Med. Imaging 2010, 29, 19–29. [Google Scholar]
- Haber, E.; Modersitzki, J. Intensity Gradient Based Registration and Fusion of Multi-modal Images. Proceedings of MICCAI 2006, Copenhagen, Denmark, 1–6 October 2006; pp. 26–733.
- Rohlfing, T. Image similarity and tissue overlaps as surrogates for image registration accuracy: Widely used but unreliable. IEEE Trans. Med. Imaging 2012, 31, 153–163. [Google Scholar]
- Pluim, J.; Maintz, J.B.; Viergever, M. Image Registration by Maximization of Combined Mutual Information and Gradient Information. Proceedings of MICCAI 2000, Pittsburgh, PA, USA, 11–14 October 2000; pp. 103–129.
- De Nigris, D.; Collins, D.L.; Arbel, T. Multi-modal image registration based on gradient orientations of minimal uncertainty. IEEE Trans. Med. Imaging 2012, 31, 2343–2354. [Google Scholar]
- Liu, J.; Vemuri, B.C.; Marroquin, J.L. Local frequency representations for robust multimodal image registration. IEEE Trans. Med. Imag. 2002, 21, 462–469. [Google Scholar]
- Heinrich, M.P.; Jenkinson, M.; Bhushan, M.; Matin, T.; Gleeson, F.V.; Brady, S.M.; Schnabel, J.A. MIND: Modality independent neighbourhood descriptor for multi-modal deformable registration. Med. Image Anal. 2012, 16, 1423–1435. [Google Scholar]
- Wachinger, C.; Navab, N. Entropy and laplacian images: Structural representations for multi-modal registration. Med. Image Anal. 2012, 16, 1–17. [Google Scholar]
- Chen, J.; Shan, S.; He, C.; Zhao, G.; Pietikäinen, M.; Chen, X.; Gao, W. WLD: A robust local image descriptor. IEEE Trans. Pattern Anal. Mach. Intell. 2010, 32, 1705–1720. [Google Scholar]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar]
- Wu, Y.; Kanade, T.; Li, C.; Cohn, J. Image registration using wavelet-based motion model. Int. J. Comput. Vis. 2000, 38, 129–152. [Google Scholar]
- Paquin, D.; Levy, D.; Xing, L. Hybrid multiscale landmark and deformable image registration. Math. Biosci. Eng. 2007, 4, 711–737. [Google Scholar]
- Liu, T.; Shen, D.; Davatzikos, C. Deformable registration of cortical structures via hybrid volumetric and surface warping. NeuroImage 2004, 22, 1790–1801. [Google Scholar]
- Postelnicu, G.; Zollei, L.; Fischl, B. Combined volumetric and surface registration. IEEE Trans. Med. Imaging 2009, 28, 508–522. [Google Scholar]
- Murray, D.J. A perspective for viewing the history of psychophysics. Behav. Brain Sci. 1993, 16, 115–186. [Google Scholar]
- Dabeer, O.; Chaudhuri, S. Analysis of an adaptive sampler based on Weber's Law. IEEE Trans. Signal Process. 2011, 59, 1868–1878. [Google Scholar]
- Bruni, V.; Vitulano, D. A generalized model for scratch detection. IEEE Trans. Image Process. 2004, 13, 44–50. [Google Scholar]
- Papenberg, N.; Bruhn, A.; Brox, T. Highly accurate optic flow computation with theoretically justified warping. Int. J. Comput. Vis. 2006, 67, 141–158. [Google Scholar]
- Buades, A.; Coll, B.; Morel, J.M. A Non-Local Algorithm for Image Denoising. Proceedings of CVPR 2005, San Diego, CA, USA, 20–26 June 2005; pp. 60–65.
- Brunet, D.; Vrscay, E.R.; Wang, Z. On the mathematical properties of the structural similarity index. IEEE Trans. Image Process. 2012, 21, 1488–1499. [Google Scholar]
- Wang, Z.; Li, Q. Information content weighting for perceptual image quality assessment. IEEE Trans. Image Process. 2011, 20, 1185–1198. [Google Scholar]
- Pluim, J.P.W.; Maintz, J.B.A.; Viergever, M.A. Mutual-information-based registration of medical images: A survey. IEEE Trans. Med. Imaging 2003, 22, 986–1004. [Google Scholar]
- Brainweb. Available online: http://www.bic.mni.mcgill.ca/brainweb/ (accessed on 6 June 2013).
- Maurer, C.R.J.; Fitzpatrick, J.; Wang, M.; Galloway, R.L.J.; Maciunas, R.; Allen, G. Registration of head volume images using implantable fiducial markers. IEEE Trans. Med. Imaging. 1997, 16, 447–462. [Google Scholar]






| Similarity Metric | TREd (mm) | |||||
|---|---|---|---|---|---|---|
| T1-T2 | T1-PD | PD-T2 | ||||
| mean | std | mean | std | mean | std | |
| / | 2.8 | 2.2 | 3.0 | 2.4 | 2.9 | 2.4 |
| NMI | 1.5 | 1.1 | 1.7 | 1.2 | 1.7 | 1.3 |
| CMI | 1.4 | 1.0 | 1.5 | 1.2 | 1.6 | 1.2 |
| ESSD | 1.4 | 0.9 | 1.6 | 1.1 | 1.5 | 1.1 |
| ESSD-NMI | 1.4 | 0.9 | 1.5 | 1.1 | 1.5 | 1.1 |
| wldNSSD | 1.2 | 0.7 | 1.3 | 0.8 | 1.3 | 0.9 |
| wldNSSD-NMI | 0.9 | 0.5 | 1.0 | 0.7 | 1.0 | 0.6 |
| wldWSSIM | 1.0 | 0.6 | 1.1 | 0.7 | 1.1 | 0.7 |
| wldWSSIM-NMI | 0.8 | 0.4 | 0.9 | 0.5 | 0.9 | 0.5 |
| Similarity Metric | TREl (mm) | |||||
|---|---|---|---|---|---|---|
| T1-T2 | T1-PD | PD-T2 | ||||
| mean | std | mean | std | mean | std | |
| Intra-observer errors | 0.9 | 0.7 | 1.0 | 0.8 | 1.0 | 0.8 |
| Inter-observer errors | 4.9 | 4.2 | 5.0 | 4.4 | 4.9 | 4.2 |
| / | 15.4 | 9.2 | 15.7 | 9.3 | 15.6 | 9.5 |
| NMI | 9.5 | 6.2 | 9.8 | 6.3 | 9.6 | 6.8 |
| CMI | 8.9 | 5.8 | 8.6 | 5.6 | 9.1 | 6.2 |
| ESSD | 8.6 | 5.7 | 8.6 | 5.6 | 8.9 | 6.2 |
| ESSD-NMI | 8.4 | 5.6 | 8.4 | 5.6 | 8.7 | 5.8 |
| wldNSSD | 7.4 | 4.9 | 7.7 | 5.0 | 7.6 | 5.0 |
| wldNSSD-NMI | 6.7 | 3.8 | 6.8 | 3.9 | 6.8 | 3.9 |
| wldWSSIM | 7.1 | 4.0 | 7.1 | 4.1 | 7.2 | 4.2 |
| wldWSSIM-NMI | 6.1 | 3.3 | 6.2 | 3.4 | 6.2 | 3.5 |
| Similarity Metric | Time (s) | ||
|---|---|---|---|
| T1-T2 | T1-PD | PD-T2 | |
| NMI | 248 ± 20 | 268 ± 22 | 280 ± 31 |
| CMI | 350 ± 28 | 346 ± 26 | 366 ± 26 |
| ESSD | 224 ± 16 | 230 ± 15 | 246 ± 16 |
| ESSD-NMI | 284 ± 24 | 292 ± 21 | 294 ± 28 |
| wldNSSD | 86 ± 7 | 88 ± 9 | 84 ± 9 |
| wldNSSD-NMI | 119 ± 12 | 118 ± 12 | 123 ± 14 |
| wldWSSIM | 118 ± 11 | 122 ± 10 | 125 ± 11 |
| wldWSSIM-NMI | 148 ± 14 | 152 ± 15 | 157 ± 15 |
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Yang, F.; Ding, M.; Zhang, X.; Wu, Y.; Hu, J. Two Phase Non-Rigid Multi-Modal Image Registration Using Weber Local Descriptor-Based Similarity Metrics and Normalized Mutual Information. Sensors 2013, 13, 7599-7617. https://doi.org/10.3390/s130607599
Yang F, Ding M, Zhang X, Wu Y, Hu J. Two Phase Non-Rigid Multi-Modal Image Registration Using Weber Local Descriptor-Based Similarity Metrics and Normalized Mutual Information. Sensors. 2013; 13(6):7599-7617. https://doi.org/10.3390/s130607599
Chicago/Turabian StyleYang, Feng, Mingyue Ding, Xuming Zhang, Yi Wu, and Jiani Hu. 2013. "Two Phase Non-Rigid Multi-Modal Image Registration Using Weber Local Descriptor-Based Similarity Metrics and Normalized Mutual Information" Sensors 13, no. 6: 7599-7617. https://doi.org/10.3390/s130607599




