A Simple Sequence Repeat (SSR) Marker Comparison of a Large In- and Ex-situ Potato Landrace Cultivar Collection from Peru Reaffirms the Complementary Nature of both Conservation Strategies
Abstract
:1. Introduction
2. Experimental Section
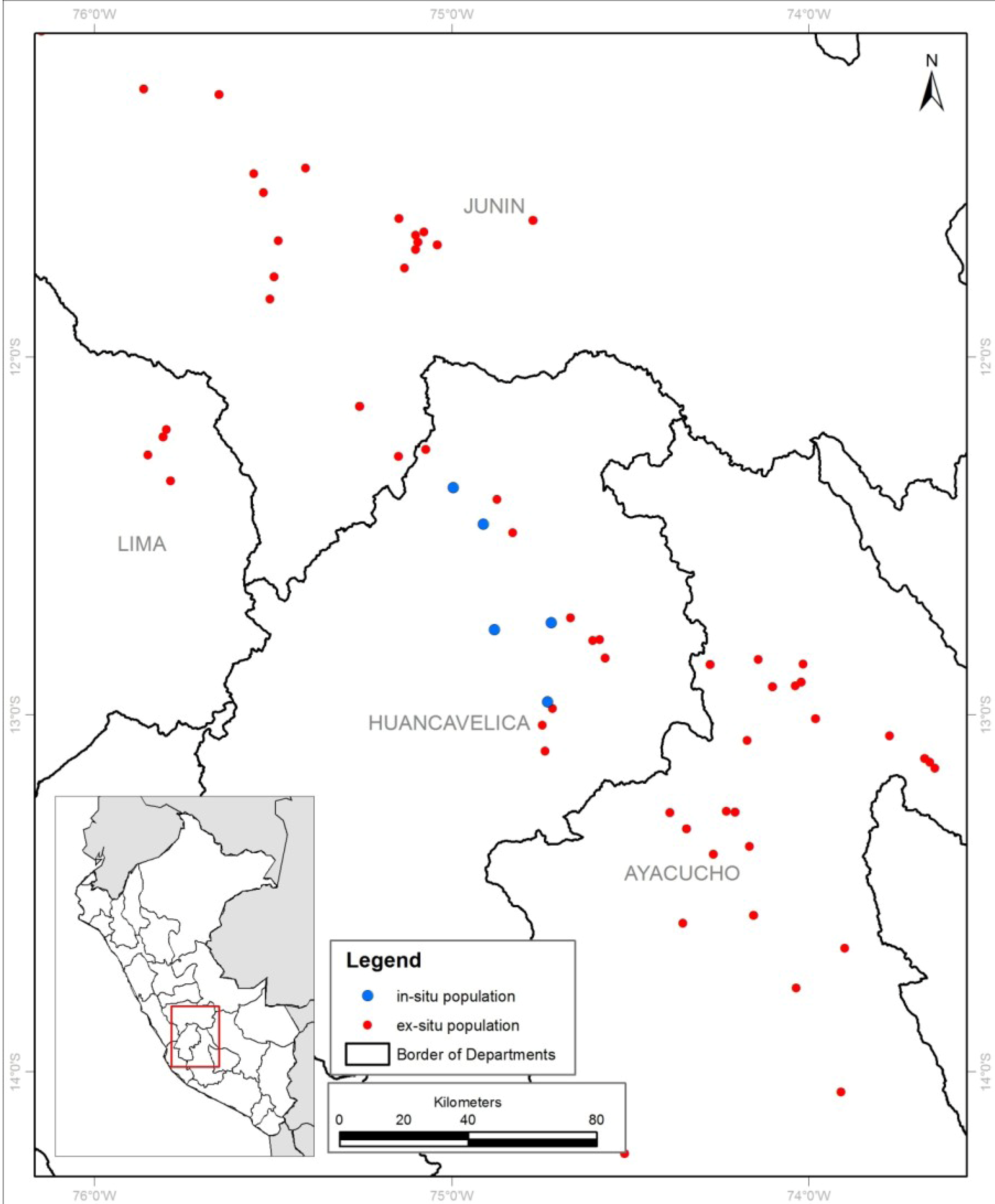
2.1. Site Description
2.2. Materials
2.3. Methods
| SSR Name | Source of markers (*) | Repeat motif | Map location (chromosome ) |
|---|---|---|---|
| STM5127 | SCRI | (TCT)n | I |
| STG0006 | TIGR | (AC)n | II |
| STI0036 | IDAHO | (AC)n (TC)imp | II |
| STG0010 | TIGR | (TG)n | III |
| STI0032 | IDAHO | (GGA)n | V |
| STM0031 | SCRI | (AC)n (AC)n GCAC (AC)n (GCAC)n | VII |
| STI0003 | IDAHO | (ACC)n | VIII |
| STI0022 | IDAHO | (ACCCG)n | VIII |
| STGBSS | SCRI | (TCT)n | VIII |
| STM1052 | SCRI | (AT)n GT (AT)n (GT)n | IX |
| STI0014 | IDAHO | (TGG)n (AGG)n | IX |
| STI0023 | IDAHO | (CAG)n | X |
| STM1106 | SCRI | (ATT)n | X |
| STM0037 | SCRI | (TC)n (AC)n AA (AC)n (AT)n | XI |
| STI0030 | IDAHO | (ATT)n | XII |
2.4. Statistical Analysis UPGMA
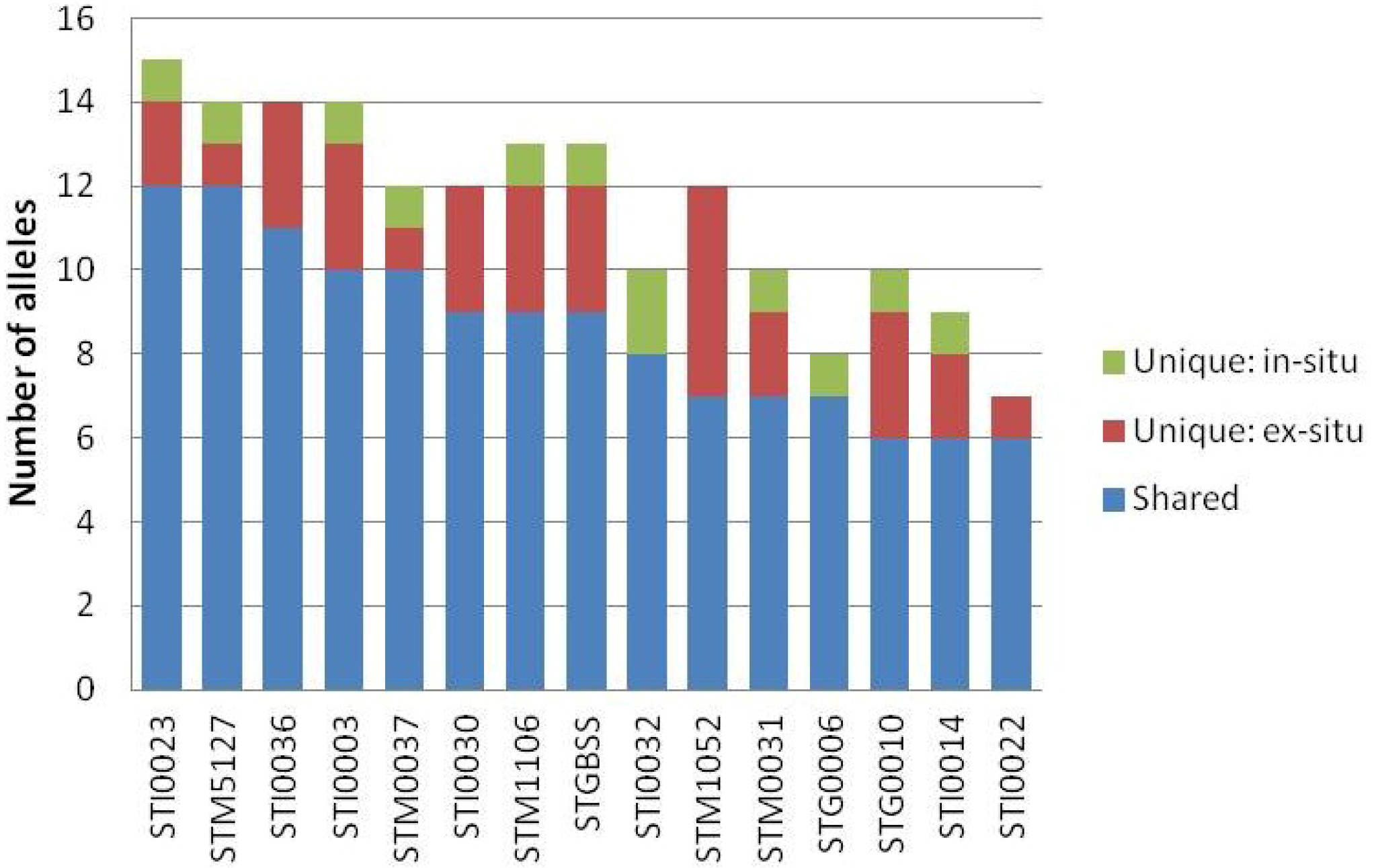
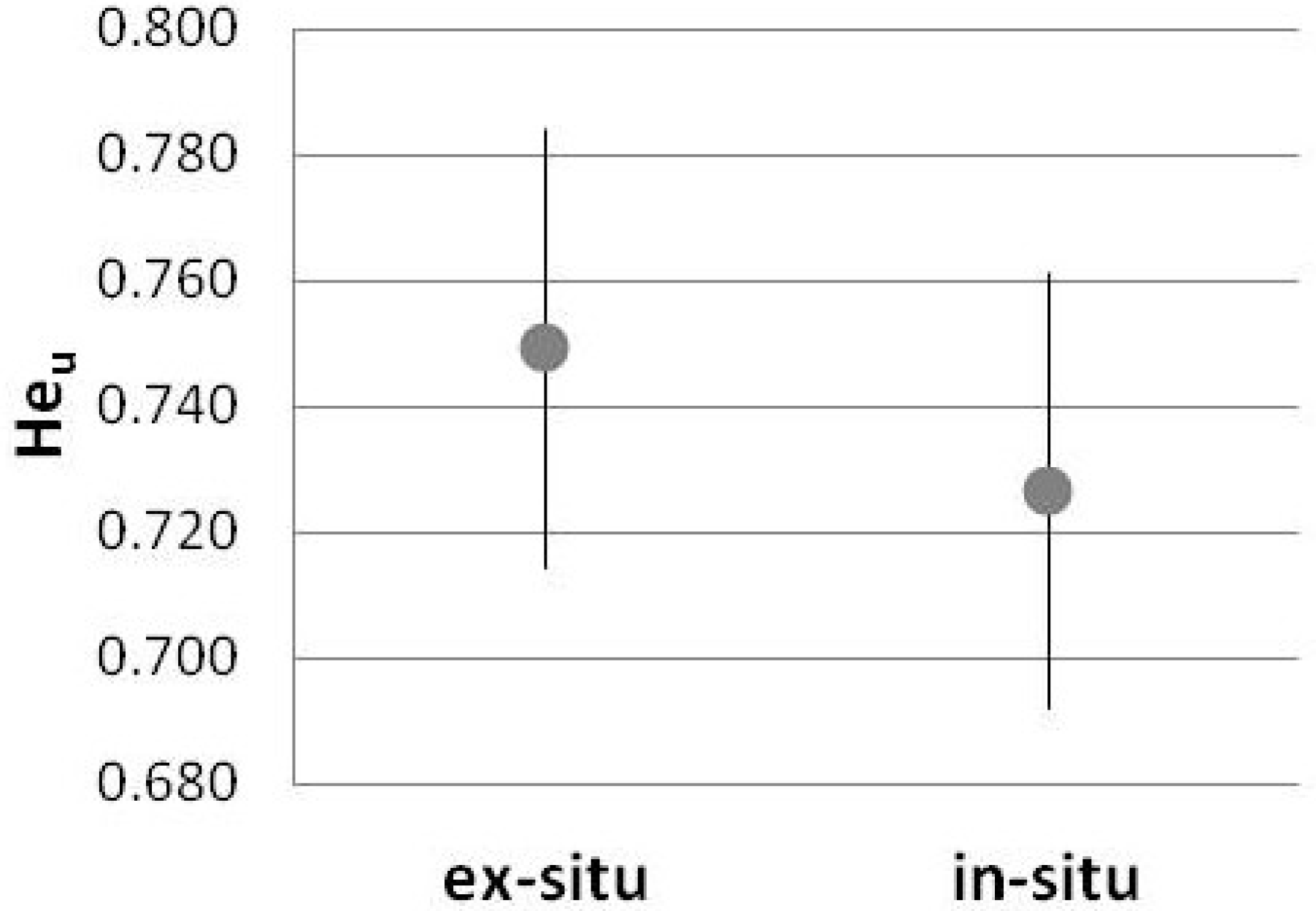
3. Results
| Population | Landrace cultivars (n) | Frequent (f > 10%) | Moderate (f < 10%) | Scarce (f < 5%) | Rare (f < 1%) | ||||
|---|---|---|---|---|---|---|---|---|---|
| n | % | n | % | n | % | n | % | ||
| Ex situ | 173 | 72 | 44.7 | 13 | 8.1 | 52 | 32.3 | 24 | 14.9 |
| In situ | 989 | 68 | 48.2 | 16 | 11.3 | 24 | 17.0 | 33 | 23.4 |
| Source of variation | d.f. | Sum of squares | Variance components | Percentage of variation |
|---|---|---|---|---|
| Among the in situ and ex situ populations | 1 | 64.3 | 0.17184 Va | 1.24 |
| Within the in situ and ex situ populations | 1,160 | 15,922.2 | 13.72606 Vb | 98.76 |
| Total | 1,161 | 15,986.6 | 13.90 | 100 |
| Fixation Index: | Significance test (1023 permutations): | |||
| Fst: 0.01236 | Va and Fst, p is <0.001 | |||

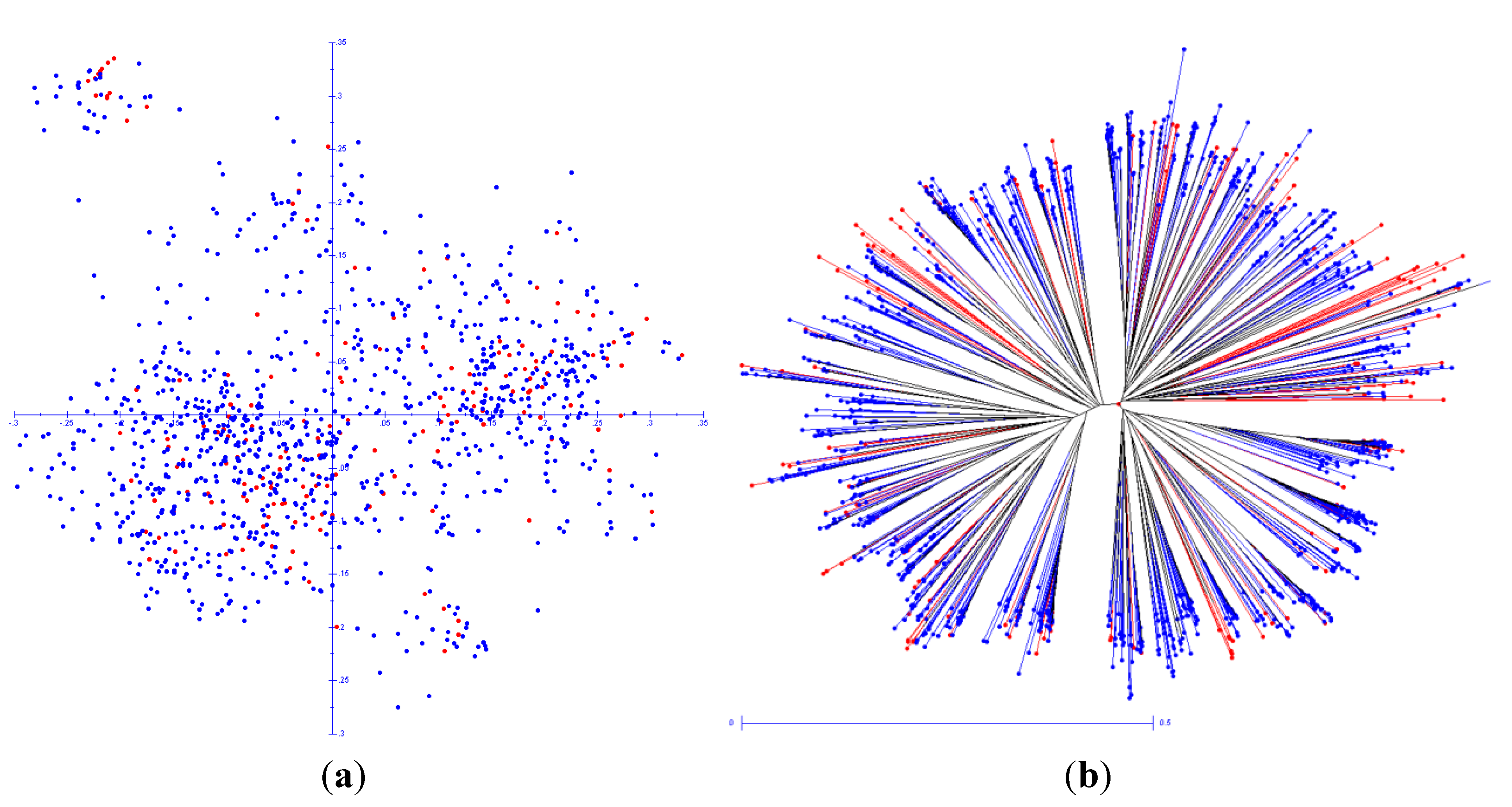
4. Discussion
4.1. SSR-Based Population Genetics
4.2. SSR-Based Overlap of Landraces
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Ovchinnikova, A.; Krylova, E.; Gavrilenko, T.; Smekalova, T.; Zhuk, M.; Knapp, S.; Spooner, D.M. Taxonomy of cultivated potatoes (Solanum section Petota: Solanaceae). Bot. J. Linn. Soc. 2011, 165, 107–155. [Google Scholar] [CrossRef]
- Spooner, D.M.; Núñez, J.; Trujillo, G.; Del Rosario Herrera, M.; Guzmán, F; Ghislain, M. Extensive simple sequence repeat genotyping of potato landraces support a major reevaluation of their gene pool structure and classification. Proc. Natl. Acad. Sci. USA 2007, 104, 19398–19403. [Google Scholar]
- Zeven, A.C. Landraces: A review of definitions and classifications. Euphytica 1998, 104, 127–139. [Google Scholar] [CrossRef]
- Brickell, C.D.; Baum, B.R.; Hetterscheid, W.L.A.; Leslie, A.C.; McNeill, J.; Trehane, P.; Vrugtman, F.; Wiersema, J.H. International Code of Nomenclature for Cultivated Plants; International Society for Horticultural Science: Leuven, Belgium, 2004. [Google Scholar]
- Correl, D.S. The Potato and its Wild Relatives; Texas Research Foundation: Renner, TX, USA, 1962. [Google Scholar]
- Hermsen, J.H. Schatgraven in het genencentrum van de aardappel (Solanum tuberosum L.). Vakbl. voor Biol. 1974, 14, 204–208. [Google Scholar]
- Hawkes, J.G. Hunting the Wild Potatoes in the South American Andes: Memories of the British Empire Potato Collecting Expedition to South America 1938–1939; Nijmegen University: Nijmegen, The Netherlands, 2003. [Google Scholar]
- Ochoa, C.M. Las Papas del Perú: base de datos 1947–1997 (In Spanish); UNALM, COSUDE, CIP: Lima, Peru, 2003. [Google Scholar]
- Vargas, C. Las Papas Sudperuanas, Parte I; Universidad Nacional del Cusco: Cusco, Peru, 1949. [Google Scholar]
- Vargas, C. Las Papas Sudperuanas, Parte II; Universidad Nacional del Cusco: Cusco, Peru, 1956. [Google Scholar]
- Loskutov, I.G. Vavilov and His Institute: A History of the World Collection of Plant Genetic Resources in Russia; IPGRI: Rome, Italy, 1999. [Google Scholar]
- Scott, G.J. Plants, people, and the conservation of biodiversity of potatoes in Peru. Nat. Conserv. 2011, 9, 1–18. [Google Scholar] [CrossRef]
- De Haan, S.; Núñez, J.; Bonierbale, M.; Ghislain, M. Multilevel agrobiodiversity and conservation of Andean potatoes in central Peru. Mt. Res. Dev. 2010, 30, 222–231. [Google Scholar] [CrossRef]
- Monteros, A.R. Potato landraces: Description and dynamics in three areas in Ecuador. Ph.D. thesis, Wageningen Agricultural University, Wageningen, The Netherlands, 2011. [Google Scholar]
- Vigouroux, Y.; Mariac, C.; De Mita, S.; Pham, J.L.; Gérard, B.; Kapran, I.; Sagnard, F.; Deu, M.; Chantereau, J.; Ali, A.; et al. Selection for earlier flowering crop associated with climatic variations in the Sahel. PLoS One 2011, 6, e19563. [Google Scholar] [CrossRef] [Green Version]
- Hijmans, R.J.; Garrett, K.A.; Huamán, Z.; Zhang, D.P.; Schreuder, M.; Bonierbale, M. Assessing the geographical representativeness of genebank collections: The case of Bolivian wild potatoes. Conserv. Biol. 2000, 14, 1755–1765. [Google Scholar] [CrossRef]
- Johns, T.; Keen, S.L. Ongoing evolution of the potato on the Altiplano of Western Bolivia. Econ. Bot. 1986, 40, 409–424. [Google Scholar] [CrossRef]
- Rabinowitz, D.; Linder, C.R.; Douches, D.S.; Quiros, C.F. Determining natural hybridization between S. sparsipilum and S. stenotomum in the Andes. Am. Potato J. 1988, 65, 496. [Google Scholar]
- Scarcelli, N.; Tostain, S.; Vigouroux, Y.; Luong, V.; Baco, M.N.; Agbangla, C.; Daïnou, O.; Pham, J.L. Genetic structure of farmer-managed varieties in clonally-propagated crops. Genetica 2011, 139, 1055–1064. [Google Scholar] [CrossRef]
- Brush, S.B. Farmers’ Bounty: Locating Crop Diversity in the Contemporary World; Yale University Press: New Haven, CT, USA, 2004. [Google Scholar]
- Etisham-Ul-Haq, M.; Allnutt, T.R.; Smith-Ramirez, C.; Gardner, M.F.; Armesto, J.J.; Newton, A.C. Patterns of genetic variation in in and ex situ populations of the threatened Chilean vine Berberidopsis corallina, detected using RAPD markers. Ann. Bot. 2001, 87, 813–821. [Google Scholar] [CrossRef]
- Lautherbach, D.; Burkart, M.; Gemeinholzer, B. Rapid genetic differentiation between ex situ and their in situ source populations: An example of the endangered Silene otites (Caryophyllaceae). Bot. J. Linn. Soc. 2012, 168, 64–75. [Google Scholar]
- Malice, M.; Martin, N.; Pissard, A.; Rojas-Beltran, J.A.; Gandarillas, A.; Bertin, P.; Baudoin, J.P. A preliminary study of the genetic diversity of Bolivian oca (Oxalis tuberosa Mol.) varieties maintained in situ and ex situ through the utilization of ISSR molecular markers. Genet. Resour. Crop. Evol. 2007, 54, 685–690. [Google Scholar] [CrossRef]
- Rice, E.B.; Smith, M.E.; Mitchell, S.E.; Kresovich, S. Conservation and change: A comparison of in situ and ex situ conservation of Jala maize germplasm. Crop Sci. 2006, 46, 428–436. [Google Scholar] [CrossRef]
- Gomez, O.J.; Blair, M.W.; Frankow-Lindberg, B.E.; Gullberg, U. Comparative study of common bean (Phaseolus vulgaris L.) landraces conserved ex situ in genebanks and in situ by farmers. Genet. Resour. Crop Evol. 2005, 52, 371–380. [Google Scholar] [CrossRef]
- Che, Y.H.; Yang, Y.P.; Yang, X.M.; Li, X.Q.; Li, L.H. Genetic diversity between ex situ and in situ samples of Agropyron cristatum (L.) Gaertn based on simple sequence repeat molecular markers. Crop Pasture Sci. 2011, 62, 639–644. [Google Scholar] [CrossRef]
- Hamblin, M.T.; Warburton, M.L.; Buckler, E.S. Empirical comparison of simple sequence repeats and single nucleotide polymorphisms in assessment of maize diversity and relatedness. PLoS One 2007, 2, e1367. [Google Scholar] [CrossRef]
- Van Inghelandt, D.; Melchinger, A.E.; Lebreton, C.; Stich, B. Population structure and genetic diversity in a commercial maize breeding program assessed with SSR and SNP markers. Theor. Appl. Genet. 2010, 120, 1289–1299. [Google Scholar] [CrossRef]
- Yang, X.; Xu, Y.; Shah, T.; Li, H.; Han, Z.; Li, J.; Yan, J. Comparison of SSRs and SNPs in assessment of genetic relatedness in maize. Genetica 2011, 139, 1045–1054. [Google Scholar] [CrossRef]
- Fregene, M.A.; Suarez, M.; Mkumbira, J.; Kulembeka, H.; Ndedya, E.; Kulaya, A.; Mitchel, S.; Gullberg, U.; Rosling, H.; Dixon, A.G.O.; et al. Simple sequence repeat marker diversity in cassava landraces: Genetic diversity and differentiation in a asexually propagated crop. Theor. Appl. Genet. 2003, 107, 1083–1093. [Google Scholar] [CrossRef]
- Tenzer, I.; Degli Ivanissevich, S.; Morgante, M.; Gessler, C. Identification of microsatellite markers and their application to population genetics of Venturia inaequalis. Phytopathology 1999, 89, 748–753. [Google Scholar] [CrossRef]
- Carputo, D.; Alioto, D.; Aversano, R.; Garramone, R.; Miraglia, V.; Villano, C.; Frusciante, L. Genetic diversity among potato species as revealed by phenotypic resistances and SSR markers. Plant Genet. Resour. 2013. [Google Scholar] [CrossRef]
- Ruiz de Galarreta, J.I.; Barandalla, L.; Lorenzo, R.; Gonzalez, J.; Rios, D.J.; Ritter, E. Microsatellite variation in potato landraces from the island of La Palma. Span. J. Agric. Res. 2007, 5, 186–192. [Google Scholar]
- Ashkenazi, V.; Chani, E.; Lavi, U.; Levy, D.; Hillel, J.; Veilleux, R.E. Development of microsatellites markers in potato and their use in phylogenetic and fingerprinting analysis. Genome 2001, 44, 50–62. [Google Scholar] [CrossRef]
- Ghislain, M.; Andrade, D.; Rodriguez, F.; Hijmans, R.J.; Spooner, D.M. Genetic analysis of the cultivated potato Solanum tuberosum L. Phureja Group using RAPDs and nuclear SSRs. Theor. Appl. Genet. 2006, 113, 1515–1527. [Google Scholar] [CrossRef]
- Ghislain, M.; Nunez, J.; Herrera, M.R.; Pignataro, J.; Guzman, F.; Bonierbale, M.; Spooner, D.M. Robust and highly informative microsatellite-based genetic identity kit for potato. Mol. Breed. 2009, 23, 377–388. [Google Scholar] [CrossRef]
- Rubina, A.; Barreda, J. Atlas del Departamento de Huancavelica (In Spanish); Centro de Estudios y Promoción del Desarrollo (DESCO), Buena Ventura: Lima, Peru, 2000. [Google Scholar]
- CIP. In Catálogo de Variedades de Papa Nativa de Huancavelica–Perú (In Spanish); CIP, FEDECCH: Lima, Peru, 2006.
- Nei, M. Molecular Evolutionary Genetics; Columbia University Press: New York, NY, USA, 1987. [Google Scholar]
- CIP. Supplementary dataset. Available online: Supplementary dataset. Available online: https://research.cip.cgiar.org/confluence/x/_aogAw (accessed on 24 April 2013).
- Rohlf, F.J. NTSYSpc: Numerical Taxonomy System, version 2.20; Exeter Publishing: Setauket, NY, USA, 2008. [Google Scholar]
- Perrier, X.; Jacquemoud-Collet, J.P. DARwin Software, version 5.0.158; CIRAD: Montpellier, France, 2006. [Google Scholar]
- Excoffier, L.; Lischer, H.E. Arlequin suite version 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef]
- Hawkes, J.G. The Potato: Evolution, Biodiversity & Genetic Resources; Smithsonian Institution Press: Washington, DC, USA, 1990. [Google Scholar]
- Ochoa, C.M. Las Papas de Sudamerica: Peru (In Spanish); Centro Internacional de la Papa: Lima, Peru, 1999. [Google Scholar]
- Zimmerer, K.S.; Douches, D.S. Geographical approaches to crop conservation: The partitioning of genetic diversity in Andean potatoes. Econ. Bot. 1991, 45, 176–189. [Google Scholar] [CrossRef]
- Rodriguez, F.; Ghislain, M.; Clausen, A.M.; Jansky, S.H.; Spooner, D.M. Hybrid origins of cultivated potatoes. Theor. Appl. Genet. 2010, 121, 1187–1198. [Google Scholar] [CrossRef]
- Spooner, D.M.; McLean, K.; Ramsay, G.; Waugh, R.; Bryan, G.J. A single domestication for potato based on multilocus amplified fragment length polymorphism genotyping. Proc. Natl. Acad. Sci. USA 2005, 102, 14694–14699. [Google Scholar] [CrossRef]
- Thiele, G. Informal potato seed systems in the Andes: Why are they important and what should we do with them? World Dev. 1999, 27, 83–99. [Google Scholar] [CrossRef]
- Brush, S.B. Ethnoecology, biodiversity, and modernization in Andean potato agriculture. J. Ethnobiol. 1992, 12, 161–185. [Google Scholar]
- Johns, T.; Huaman, Z.; Ochoa, C.; Schmiediche, P.E. Relationships among wild, weed, and cultivated potatoes in the Solanum ajanhuiri complex. Syst. Bot. 1987, 12, 541–552. [Google Scholar] [CrossRef]
- Quiros, C.F.; Ortega, R.; Van Raamsdonk, L.; Herrera-Montoya, M.; Cisneros, P.; Schmidt, E.; Brush, S.B. Increase of potato genetic resources in their center of diversity: The role of natural outcrossing and selection by Andean farmers. Genet. Resour. Crop Evol. 1992, 39, 107–113. [Google Scholar]
- Rabinowitz, D.; Linder, C.R.; Ortega, R.; Begazo, D.; Murguia, H.; Douches, D.S.; Quiros, C.F. High levels of interspecific hybridization between Solanum sparsipilum and S. stenotomum in experimental plots in the Andes. Am. Potato J. 1990, 67, 73–82. [Google Scholar] [CrossRef]
- Scurrah, M.; Celis-Gamboa, C.; Chumbiauca, S.; Salas, A.; Visser, R.G.F. Hybridization between wild and cultivated potato species in the Peruvian Andes and biosafety implications for the deployment of GM potatoes. Euphytica 2008, 164, 881–892. [Google Scholar]
- Wilson, F. Towards a political economy of roads: Experiences from Peru. Dev. Change 2004, 35, 525–546. [Google Scholar]
- Zimmerer, K.S. Geographies of seed networks for food plants (potato, ulluco) and approaches to agrobiodiversity conservation in the Andes. Soc. Nat. Resour. 2003, 16, 583–601. [Google Scholar] [CrossRef]
- Bellon, M.R. Conceptualizing interventions to support on-farm genetic resource conservation. World Dev. 2004, 32, 159–172. [Google Scholar] [CrossRef]
- Community Biodiversity Management: Promoting Resilience and the Conservation of Plant Genetic Resources; De Boef, W.; De Boef, W.; Subedi, A.; Peroni, N.; Thijssen, M.; O’Keeffe, E. (Eds.) Routledge: Oxford, UK, 2013.
- Brush, S.B.; Taylor, J.E.; Bellon, M.R. Technology adoption and biological diversity in Andean potato agriculture. J. Dev. Econ. 1992, 39, 365–387. [Google Scholar] [CrossRef]
- Zimmerer, K.S. The loss and maintenance of native crops in mountain agriculture. GeoJournal 1992, 27, 61–72. [Google Scholar] [CrossRef]
- De Haan, S.; Almekinders, C.; Thiele, G.; Scurrah, M. Farmer seed systems and infraspecific diversity of potato in Peru’s central Highlands. In Potato Diversity at Height: Multiple Dimensions of Farmer-driven In-situ Conservation in the Andes; Wageningen Agricultural University: Wageningen, The Netherlands, 2009; pp. 133–160. [Google Scholar]
- Pedersen, S.; Tremblay, J.; Errázuriz, C.; Gamarra, J. The sequelae of political violence: Assessing trauma, suffering and dislocation in the Peruvian highlands. Soc. Sci. Med. 2008, 67, 205–217. [Google Scholar] [CrossRef]
- Zimmerer, K.S. Labor shortages and crop diversity in the southern Peruvian Sierra. Geogr. Rev. 1991, 82, 414–432. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
De Haan, S.; Núñez, J.; Bonierbale, M.; Ghislain, M.; Van der Maesen, J. A Simple Sequence Repeat (SSR) Marker Comparison of a Large In- and Ex-situ Potato Landrace Cultivar Collection from Peru Reaffirms the Complementary Nature of both Conservation Strategies. Diversity 2013, 5, 505-521. https://doi.org/10.3390/d5030505
De Haan S, Núñez J, Bonierbale M, Ghislain M, Van der Maesen J. A Simple Sequence Repeat (SSR) Marker Comparison of a Large In- and Ex-situ Potato Landrace Cultivar Collection from Peru Reaffirms the Complementary Nature of both Conservation Strategies. Diversity. 2013; 5(3):505-521. https://doi.org/10.3390/d5030505
Chicago/Turabian StyleDe Haan, Stef, Jorge Núñez, Merideth Bonierbale, Marc Ghislain, and Jos Van der Maesen. 2013. "A Simple Sequence Repeat (SSR) Marker Comparison of a Large In- and Ex-situ Potato Landrace Cultivar Collection from Peru Reaffirms the Complementary Nature of both Conservation Strategies" Diversity 5, no. 3: 505-521. https://doi.org/10.3390/d5030505




