Edges and Overlaps in Northwest Atlantic Phylogeography
Abstract
:1. Introduction
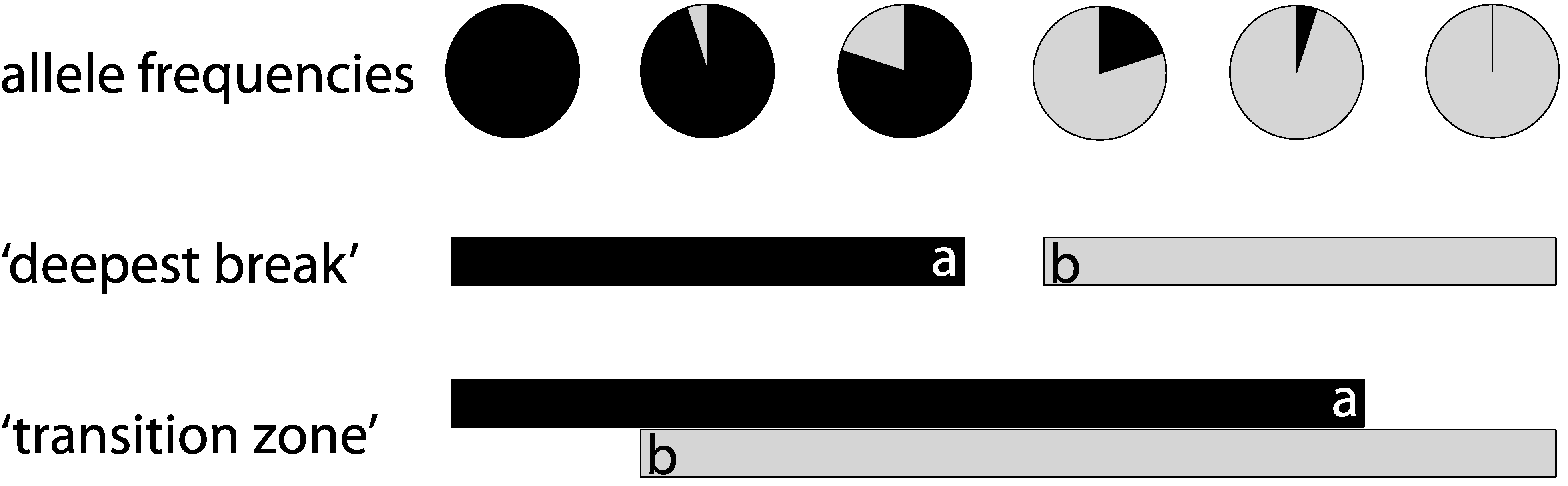
2. Methods
2.1. Literature Search/Study Compilation
2.2. Analysis of Phylogenetic Concordance
3. Results
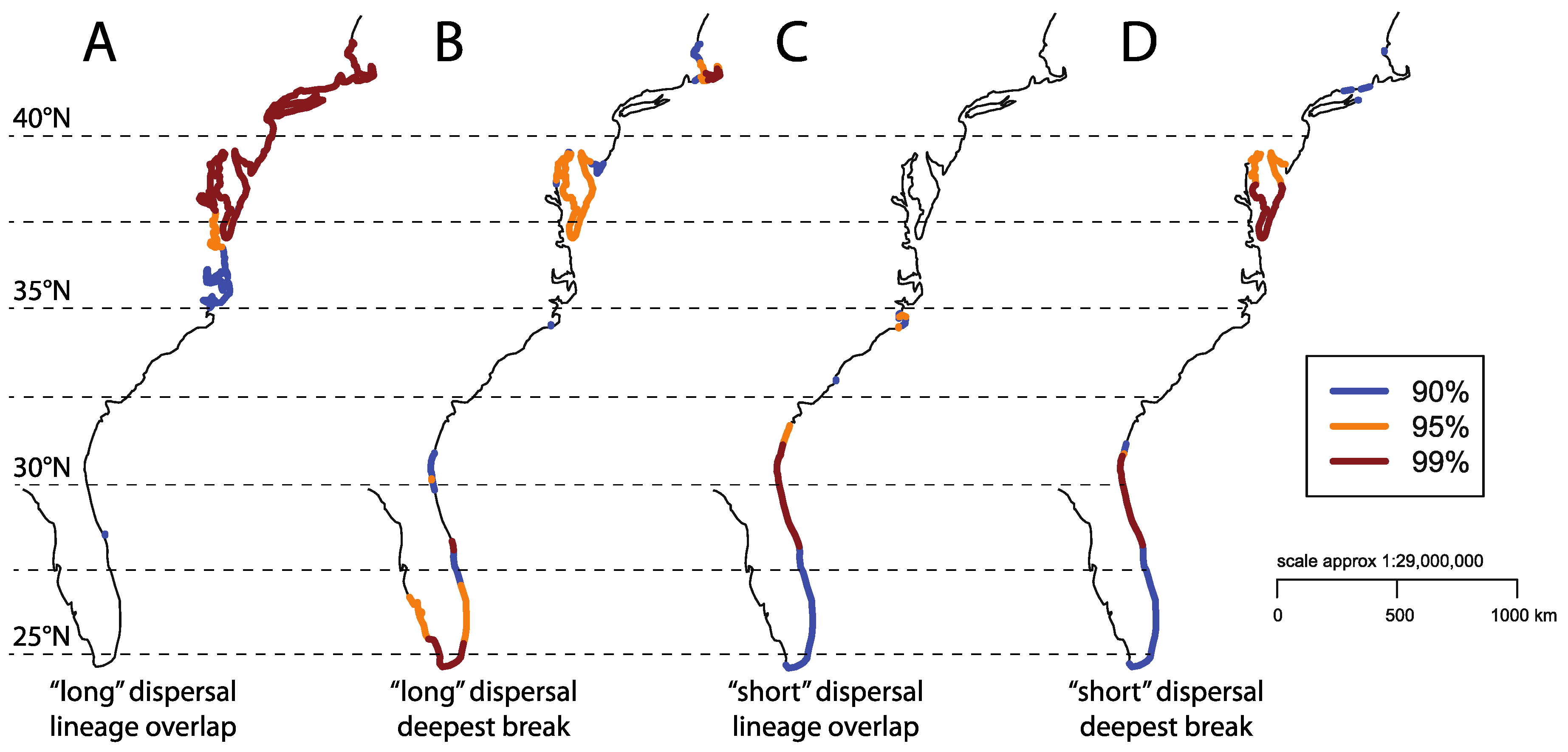
4. Discussion
Moving forward with Comparative Phylogeography
5. Conclusions
Supplementary Materials
Acknowledgments
References
- Burton, R.S. Intraspecific phylogeography across the Point Conception biogeographic boundary. Evolution 1998, 52, 734–745. [Google Scholar] [CrossRef]
- Dawson, M.N. Phylogeography in coastal marine animals: a solution from California? J. Biogeogr. 2001, 28, 723–736. [Google Scholar] [CrossRef]
- Wares, J.P. Community Genetics in the Northwestern Atlantic Intertidal. Mol. Ecol. 2002, 11, 1131–1144. [Google Scholar] [CrossRef]
- Pelc, R.A.; Warner, R.R.; Gaines, S.D. Geographical patterns of genetic structure in marine species with contrasting life histories. J. Biogeogr. 2009, 36, 1881–1890. [Google Scholar] [CrossRef]
- Marko, P.B. “What’s larvae got to do with it?” Disparate patterns of post-glacial population structure in two benthic marine gastropods with identical dispersal potential. Mol. Ecol. 2004, 13, 597–611. [Google Scholar] [CrossRef]
- Puritz, J.B.; Toonen, R.J. Coastal pollution limits pelagic larval dispersal. Nature Comm. 2011. [Google Scholar] [CrossRef]
- Pringle, J.M.; Wares, J.P. The maintenance of alongshore variation in allele frequency in a coastal ocean. Mar. Ecol. Prog. Series 2007, 335, 69–84. [Google Scholar] [CrossRef]
- Eckman, J.E. Closing the larval loop: linking larval ecology to the population dynamics of marine benthic invertebrates. J. Exp. Mar. Biol. Ecol. 1996, 200, 207–237. [Google Scholar] [CrossRef]
- Emlet, R.B. Developmental mode and species geographic range in regular sea urchins (Echinodermata: Echinoidea). Evolution 1995, 49, 476–489. [Google Scholar] [CrossRef]
- Grantham, B.A.; Eckert, G.L.; Shanks, A.L. Dispersal potential of marine invertebrates in diverse habitats. Ecol. Appl. 2003, 13, 108–116. [Google Scholar] [CrossRef]
- Havenhand, J.N. Evolutionary ecology of larval types. In Ecology of Marine Invertebrate Larvae; McEdward, L., Ed.; CRC Press: New York, NY, USA, 1995; pp. 79–122. [Google Scholar]
- Levin, L.A.; Bridges, T.S. Pattern and Diversity in Reproduction and Development; McEdward, L., Ed.; CRC Press: New York, NY, USA, 1995; pp. 1–48. [Google Scholar]
- McEdward, L. Ecology of Marine Invertebrate Larvae; CRC Press: New York, NY, USA, 1995; p. 464. [Google Scholar]
- Baums, I.B.; Paris, C.B.; Chérubin, L.M. A bio-oceanographic filter to larval dispersal in a reef-building coral. Limn. Ocean. 2006, 51, 1969–1981. [Google Scholar] [CrossRef]
- Connolly, S.R.; Roughgarden, J. A latitudinal gradient in northeast Pacific intertidal community structure: evidence for an oceanographically based synthesis of marine community theory. Amer. Nat. 1998, 151, 311–326. [Google Scholar]
- Dawson, M.N. Phylogeography in coastal marine animals: a solution from California? J. Biogeogr. 2001, 28, 723–736. [Google Scholar] [CrossRef]
- Galindo, H.M.; Olson, D.B.; Palumbi, S.R. Seascape genetics: A coupled oceanographic-genetic model predicts population structure of Caribbean corals. Curr. Biol. 2006, 16, 1622–1626. [Google Scholar] [CrossRef]
- Selkoe, K.A.; Watson, J.R.; White, C.; Horin, T.B.; Iacchei, M.; Mitarai, S.; Siegel, D.A.; Gaines, S.D.; Toonen, R.J. Taking the chaos out of genetic patchiness: seascape genetics reveals ecological and oceanographic drivers of genetic patterns in three temperate reef species. Mol. Ecol. 2010, 19, 3708–3726. [Google Scholar] [CrossRef]
- Zakas, C.; Binford, J.; Navarrete, S.A.; Wares, J.P. Restricted gene flow in Chilean barnacles reflects an oceanographic and biogeographic transition zone. Mar. Ecol. Prog. Ser. 2009, 394, 165–177. [Google Scholar] [CrossRef]
- Carlton, J.T. Community assembly and historical biogeography in the Atlantic Ocean: the potential role of human-mediated dispersal vectors. Hydrobiologia 2003, 503, 1–8. [Google Scholar] [CrossRef]
- Sunday, J.M.; Bates, A.E.; Dulvy, N.K. Thermal tolerance and the global redistribution of animals. Nature Clim. Change. 2012, 2, 686–690. [Google Scholar]
- Wiens, J.J. The niche, biogeography and species interactions. Phil. Trans. R. Soc. Lond. B 2011, 366, 2336–2350. [Google Scholar] [CrossRef]
- Sexton, J.P.; McIntyre, P.J.; Angert, A.L.; Rice, K.J. Evolution and ecology of species range limits. Ann. Rev. Ecol. Evol. Syst. 2009, 40, 415–436. [Google Scholar] [CrossRef]
- Dawson, M.N.; Grosberg, R.K.; Stuart, Y.E.; Sanford, E. Population genetic analysis of a recent range expansion: mechanisms regulating the poleward range limit in the volcano barnacle Tetraclita rubescens. Mol. Ecol. 2010, 19, 1585–1605. [Google Scholar] [CrossRef]
- Zacherl, D.; Gaines, S.D. The limits to biogeographical distributions: Insights from the northward range extension of the marine snail, Kelletia kelletia. J. Biogeography 2003, 30, 913–924. [Google Scholar] [CrossRef]
- Sanford, E.; Roth, M.S.; Johns, G.C.; Wares, J.P.; Somero, G.N. Local selection and latitudinal variation in a marine predator-prey interaction. Science 2003, 300, 1135–1137. [Google Scholar] [CrossRef]
- Harley, C.D.G.; Pankey, M.S.; Wares, J.P.; Grosberg, R.K.; Wonham, M.J. The impacts of climate change in coastal systems. Ecol. Lett. 2006, 9, 228. [Google Scholar] [CrossRef]
- Norberg, J.; Urban, M.C.; Vellend, M.; Klausmeier, C.A.; Loeuille, N. Eco-evolutionary responses of biodiversity to climate change. Nature Clim. Change 2012, 2, 747–751. [Google Scholar] [CrossRef]
- Wethey, D.S. Biogeography, competition, and microclimate: the barnacle Chthamalus fragilis in New England. Int. Comp. Biol. 2002, 42, 872–880. [Google Scholar] [CrossRef]
- Sanford, E.; Holzman, S.B.; Haney, R.A.; Rand, D.M.; Bertness, M.D. Larval tolerance, gene flow, and the northern geographic range limit of fiddler crabs. Ecology 2006, 87, 2882–2894. [Google Scholar] [CrossRef]
- Small, S.T.; Wares, J.P. Phylogeography and marine retention. Journal of Biogeography 2010, 37, 781–784. [Google Scholar] [CrossRef]
- Briggs, J.C. Marine zoogeography; McGraw Hill: New York, NY, USA, 1974. [Google Scholar]
- Briggs, J.C.; Bowen, B.W. A realignment of marine biogeographic provinces with particular reference to fish distributions. J. Biogeogr. 2012, 39, 12–30. [Google Scholar] [CrossRef]
- Olivero, J.; Márquez, A.L.; Real, R. Integrating fuzzy logic and statistics to improve the reliable delimitation of biogeographic regions and transition zones. Syst. Biol. 2012, 26, 1–21. [Google Scholar]
- Wares, J.P.; Gaines, S.D.; Cunningham, C.W. A Comparative Study of Asymmetric Migration Events across a Marine Biogeographic Boundary. Evolution 2001, 55, 295–306. [Google Scholar]
- Blanchette, C.A.; Miner, C.M.; Raimondi, P.T.; Lohse, D.; Heady, K.E.K.; Broitman, B.R. Biogeographical patterns of rocky intertidal communities along the Pacific coast of North America. J. Biogeogr. 2008, 35, 1593–1607. [Google Scholar] [CrossRef]
- Newman, W.A.; Abbott, D.P.; Haderlie, E.C. Cirripedia; Morris, R.H., Ed.; Stanford University Press: Stanford, CA, USA, 1980; pp. 504–535. [Google Scholar]
- Fortin, M.J.; Keitt, T.H.; Maurer, B.A.; Taper, M.L.; Kaufman, D.M.; Blackburn, T.M. Species’ geographic ranges and distributional limits: pattern analysis and statistical issues. Oikos 2005, 108, 7–17. [Google Scholar] [CrossRef]
- Kelly, M.W.; Sanford, E.; Grosberg, R.K. Limited potential for adaptation to climate change in a broadly distributed marine crustacean. Proc. Roy. Soc. B 2012, 279, 349–356. [Google Scholar] [CrossRef]
- Blackburn, T.M.; Cassey, P.; Gaston, K.J. Variations on a theme: sources of heterogeneity in the form of the interspecific relationship between abundance and distribution. J. Anim. Ecol. 2006, 75, 1426–1439. [Google Scholar] [CrossRef]
- Byers, J.E.; Pringle, J.M. Going against the flow: retention, range limits and invasions in advective environments. Mar. Ecol. Prog. Ser. 2006, 313, 27–41. [Google Scholar] [CrossRef]
- Case, T.J.; Taper, M.L. Interspecific competition, environmental gradients, gene flow, and the coevolution of species’ borders. Amer. Nat. 2000, 155, 583–605. [Google Scholar] [CrossRef]
- Gaines, S.D.; Lester, S.; Eckert, G.; Kinlan, B.P.; Sagarin, R.D.; Gaylord, B. Dispersal and geographic ranges in the sea. In Marine Macroecology; Witman, J.D., Roy, K., Eds.; University of Chicago Press: Chicago, IL, USA, 2009; pp. 227–249. [Google Scholar]
- Gaston, K.J. The structure and dynamics of geographic ranges. In Oxford Series Ecology and Evolution; Oxford University Press: Oxford, UK, 2003; p. 266. [Google Scholar]
- Gaylord, B.; Gaines, S.D. Temperature or transport? Range limits in marine species mediated solely by flow. Amer. Nat. 2000, 155, 769–789. [Google Scholar] [CrossRef]
- Gilman, S.E. Experimental manipulations at the northern geographic range limit of an intertidal snail. Ph.D. Thesis, University of California, Davis, CA, USA, 2003. [Google Scholar]
- Helmuth, B.; Mieszkowska, N.; Moore, P.; Hawkins, S.J. Living on the edge of two changing worlds: Forecasting the responses of rocky intertidal ecosystems to climate change. Ann. Rev. Ecol. Evol. Syst. 2006, 37, 373–404. [Google Scholar] [CrossRef]
- Holt, R.D. On the evolutionary ecology of species’ ranges. Evolutionary Ecology Research 2003, 5, 159–178. [Google Scholar]
- Jablonski, D.; Valentine, J.W. From regional to total geographic ranges: testing the relationship in Recent bivalves. Paleobiology 1990, 16, 126–142. [Google Scholar]
- Kirkpatrick, M.; Barton, N.H. Evolution of a species’ range. The American Naturalist 1997, 150, 1–23. [Google Scholar]
- Weersing, K.; Toonen, R.J. Population genetics, larval dispersal, and connectivity in marine systems. Mar. Ecol. Prog. Ser. 2009, 393, 1–12. [Google Scholar] [CrossRef]
- Strathman, R.R. Feeding and nonfeeding larval development and life-history evolution in marine invertebrates. Ann. Rev. Ecol. Syst. 1985, 16, 339–361. [Google Scholar]
- Strathman, R.R. What controls the type of larval development. Bull. Mar. Sci. 1986, 39, 616–622. [Google Scholar]
- Levin, L.A. Recent progress in understanding larval dispersal: new directions and digressions. Integr. Comp. Biol. 2006, 46, 282–297. [Google Scholar] [CrossRef]
- Shanks, A.L. Pelagic larval duration and dispersal distance revisited. Biol. Bull. 2009, 216, 373–385. [Google Scholar]
- Toews, D.P.L.; Brelsford, A. The biogeography of mitochondrial and nuclear discordance in animals. Mol. Ecol. 2012, 21, 3907–3930. [Google Scholar] [CrossRef]
- R_Development_Core_Team, R: A Language and Environment for Statistical Computing, R Foundation for Statistical Computing: Vienna, Austria, 2009.
- Irwin, D.E. Phylogeographic breaks without geographic barriers to gene flow. Evolution 2002, 56, 2383–2394. [Google Scholar]
- Gilman, S.E. Life at the edge: An experimental study of a poleward range boundary. Oecologia 2006, 148, 270–279. [Google Scholar] [CrossRef]
- Zakas, C.; Wares, J.P. Consequences of a poecilogonous life history for genetic structure in coastal populations of the polychaete Streblospio benedicti. Mol. Ecol. 2012, 21, 5447–5460. [Google Scholar] [CrossRef]
- Williams, P.H. Mapping variations in the strength and breadth of biogeographic transition zones using species turnover. Proc. R. Soc. Lond. B 1996, 263, 579–588. [Google Scholar] [CrossRef]
- Engle, V.D.; Summers, J.K. Latitudinal gradients in benthic community composition in Western Atlantic estuaries. J. Biogeogr. 1999, 26, 1007–1023. [Google Scholar] [CrossRef]
- Spalding, M.D.; Fox, H.E.; Halpern, B.S.; McManus, M.A.; Molnar, J.; Allen, G.R.; Davidson, N.; Jorge, Z.A.; Lombana, A.L.; Lourie, S.A.; et al. Marine ecoregions of the world: A bioregionalization of coastal and shelf areas. BioScience 2007, 57, 573–583. [Google Scholar] [CrossRef]
- Avise, J.C. Phylogeography; Harvard University Press: Cambridge, MA, USA, 2000. [Google Scholar]
- Hickerson, M.J.; Meyer, C.P. Testing comparative phylogeographic models of marine vicariance and dispersal using a hierarchical Bayesian approach. BMC Evol. Biol. 2008, 8, 322. [Google Scholar] [CrossRef]
- Nielsen, R.; Beaumont, M.A. Statistical inferences in phylogeography. Mol. Ecol. 2009, 18, 1034–1047. [Google Scholar]
- Huang, W.; Takebayashi, N.; Qi, Y.; Hickerson, M.J. MTML-msBayes: Approximate Bayesian comparative phylogeographic inference from multiple taxa and multiple loci with rate heterogeneity. BMC Bioinformatics 2011, 12, 1. [Google Scholar]
- Wares, J.P. Intraspecific variation and geographic isolation in Idotea balthica (Isopoda: Valvifera). J. Crust. Biol. 2001, 21, 1007–1013. [Google Scholar] [CrossRef]
- Bell, T.M. The maintenance of genetic diversity in the Atlantic isopod, Idotea balthica. Ph.D. Thesis, University of Georgia, Athens, GA, USA, 2009. [Google Scholar]
- Endler, J.A. Problems in distinguishing historical from ecological factors in Biogeography. Amer. Zoolo. 1982, 22, 441–452. [Google Scholar]
- Endler, J.A. Geographic Variation, Speciation, and Clines; Princeton University Press: Princeton, NJ, USA, 1977; p. 246. [Google Scholar]
- Barry, J.P.; Baxter, C.H.; Sagarin, R.D.; Gilman, S.E. Climate-related, long-term faunal changes in a California rocky intertidal community. Science 1995, 267, 672–675. [Google Scholar] [CrossRef]
- Southward, A.J. Forty years of changes in species composition and population density of barnacles on a rocky shore near Plymouth, England, UK. J. Mar. Biol. Assoc. UK 1991, 71, 495–514. [Google Scholar] [CrossRef]
- Perry, A.L.; Low, P.J.; Ellis, J.R.; Reynolds, J.D. Climate change and distribution shifts in marine fishes. Science 2005, 308, 1912–1915. [Google Scholar] [CrossRef]
- Hoffmann, A.A.; Daborn, P.J. Towards genetic markers in animal populations as biomonitors for human-induced environmental change. Ecol. Lett. 2007, 10, 63–76. [Google Scholar] [CrossRef]
- Meirmans, P.G. Using the AMOVA framework to estimate a standardized genetic differentiation measure. Evolution 2006, 60, 2399–2402. [Google Scholar]
- Dawson, M.N. Parallel phylogeographic structure in ecologically similar sympatric sister taxa. Mol. Ecol. 2012, 21, 987–1004. [Google Scholar] [CrossRef]
- Kelly, R.P.; Palumbi, S.R. Genetic Structure Among 50 Species of the Northeastern Pacific Rocky Intertidal Community. PloS One 2010, 5, e8594. [Google Scholar]
- Ilves, K.; Huang, W.; Wares, J.P.; Hickerson, M.J. Colonization and/or mitochondrial selective sweeps across the Atlantic intertidal assemblage revealed by multi-taxa approximate Bayesian computation. Mol. Ecol. 2010, 19, 4505–4519. [Google Scholar] [CrossRef]
- Ling, S.D.; Johnson, C.R.; Ridgway, K.; Hobday, A.J.; Haddon, M. Climate-driven range extension of a sea urchin: inferring future trends by analysis of recent population dynamics. Global Change Biol. 2009, 15, 719–731. [Google Scholar] [CrossRef]
- Wernberg, T.; Russell, B.D.; Moore, P.J.; Ling, S.D.; Smale, D.A.; Campbell, A.; Coleman, M.A.; Steinberg, P.D.; Kendrick, G.A.; Connell, S.D. Impacts of climate change in a global hotspot for temperate marine biodiversity and ocean warming. J. exp. mar. Biol. Ecol. 2011, 400, 7–16. [Google Scholar] [CrossRef]
- Williams, J.W.; Kharouba, H.M.; Veloz, S.; Vellend, M.; McLachlan, J.; Liu, Z.Y.; Otto-Bliesner, B.; He, F. The ice age ecologist: testing methods for reserve prioritization during the last global warming. Global Ecol. Biogeogr. 2012, 22, 289–301. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Altman, S.; Robinson, J.D.; Pringle, J.M.; Byers, J.E.; Wares, J.P. Edges and Overlaps in Northwest Atlantic Phylogeography. Diversity 2013, 5, 263-275. https://doi.org/10.3390/d5020263
Altman S, Robinson JD, Pringle JM, Byers JE, Wares JP. Edges and Overlaps in Northwest Atlantic Phylogeography. Diversity. 2013; 5(2):263-275. https://doi.org/10.3390/d5020263
Chicago/Turabian StyleAltman, Safra, John D. Robinson, James M. Pringle, James E. Byers, and John P. Wares. 2013. "Edges and Overlaps in Northwest Atlantic Phylogeography" Diversity 5, no. 2: 263-275. https://doi.org/10.3390/d5020263



