1. Introduction
The GLYATs (EC 2.3.1.13) present a family of proteins which play a physiologically important role in the detoxification of endogenous and xenobiotic acyl-CoA’s. In mammals, a variety of carboxylic acids xenobioties were conjugated with an amino acid, primarily glycine [
1], and the resulting peptides appear as excretory products in the urine. The conjugation, which occurs in both liver and kidney [
1], involves in a two step pathway: firstly, the carboxylic acid is ATP-dependent activated with CoA by carboxylic acid: CoA ligase to form an intermediate acyl-CoA product Gatley [
1–
3]; this acyl group is then transferred to the amino group of glycine by the action of an Glycine-N-acyltransferase(GLYAT). As a result, abnormality of GLYAT has been related to some pathological situations such as various organic acidemias [
4].
Up to now, the GLYAT proteins have been purified and biochemically identified from the liver or kidney of human, bovine, ovine, rabbit and rhesus monkey [
1,
5,
6]. It is clearly demonstrated that the enzyme has two kind of substrates in its functional pathway: the acyl-CoA donor and acyl-acceptor. A few members act as the role of acyl-donor: benzoyl-CoA, salicyl-CoA, isovaleryl-CoA, octanoyl-CoA etc. However, the acyl-CoA acceptor exhibits relatively high specifity for glycine [
1,
7].
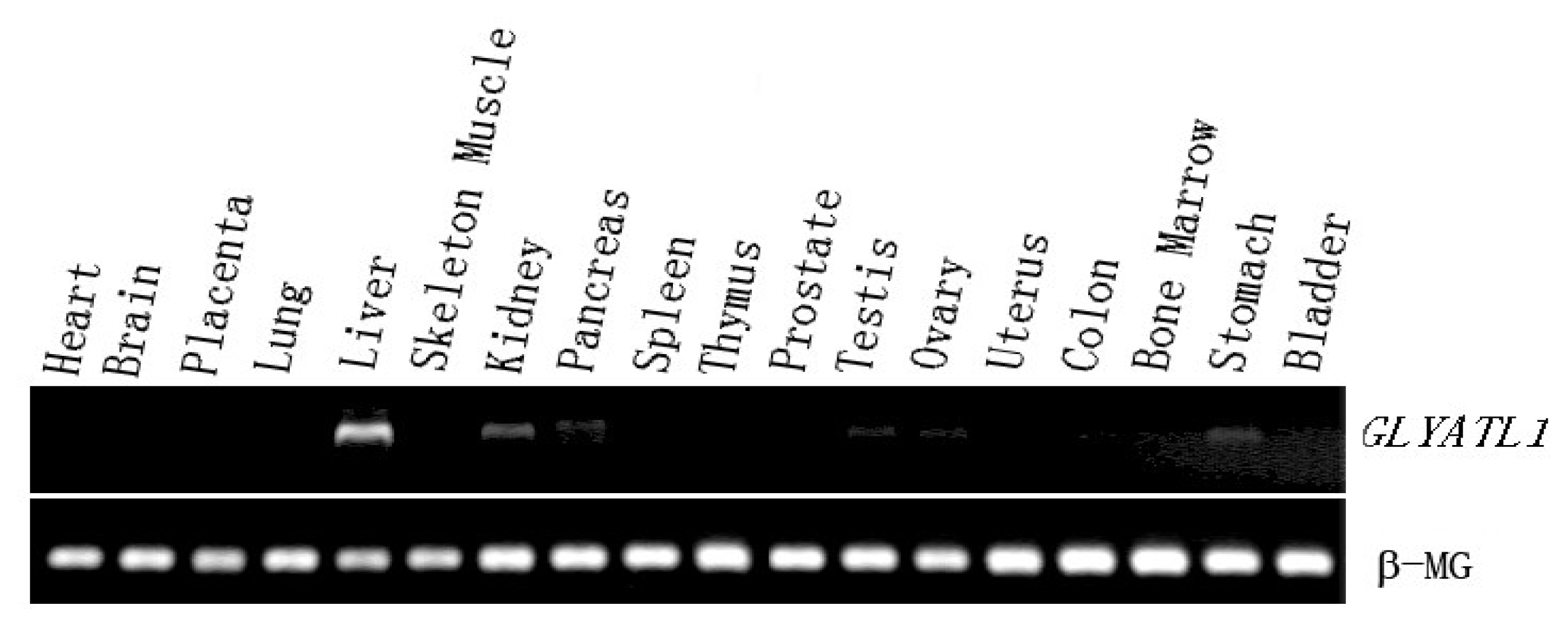
The completion of the Human Genome Project (HGP) makes it possible to investigate the gene nature of the GLYAT and it is helpful to obtain more comprehensive understanding for the function of this enzyme. In the present study, we identified a novel human acyl-CoA: glycine-N-acyltransferase gene, GLYATL1, which encoded a 302-amino acids protein. The RT-PCR analysis showed that GLYATL1 was expressed strongly in liver and weakly in kidney and stomach among 18 human tissues. The protein is subcellular localized in cytoplasm and overexpression of the protein activates the transcriptional activities of HSE, suggesting a potential role of this protein in HSE mediated transcriptional regulation.
2. Materials and methods
2.1 Molecular cloning and sequence analysis
Two primers (
GLYATL1-Forward and
GLYATL1-Reverse,
Table 1) were designed to amplify
GLYATL1 from the human liver cDNA library. PCR was performed at 94 °C, 4 min; 94 °C, 1 min; 57 °C, 1 min; 72 °C, 2 min for 32 cycles; then 72 °C, 10 min, using GeneAmp PCR system 9600 (Perkin-Elmer Cetus). PCR products were separated on 1 % (w/v) agarose gel. Then the product of interest was separated, cloned into pMD18-T vector (TaKaRa, Japan) and finally confirmed by sequencing. The full-length sequence of GLYATL1 was analyzed for homologies in GenBank by using the BLAST program and then submitted to GenBank. with the Accession No. DQ084381.
Protein sequence comparisons were carried out using BLAST2.0 at NCBI (
http://www.ncbi.nlm.nih.gov/blast). The gene structure analysis was performed by a BLAT search against the human genome database (
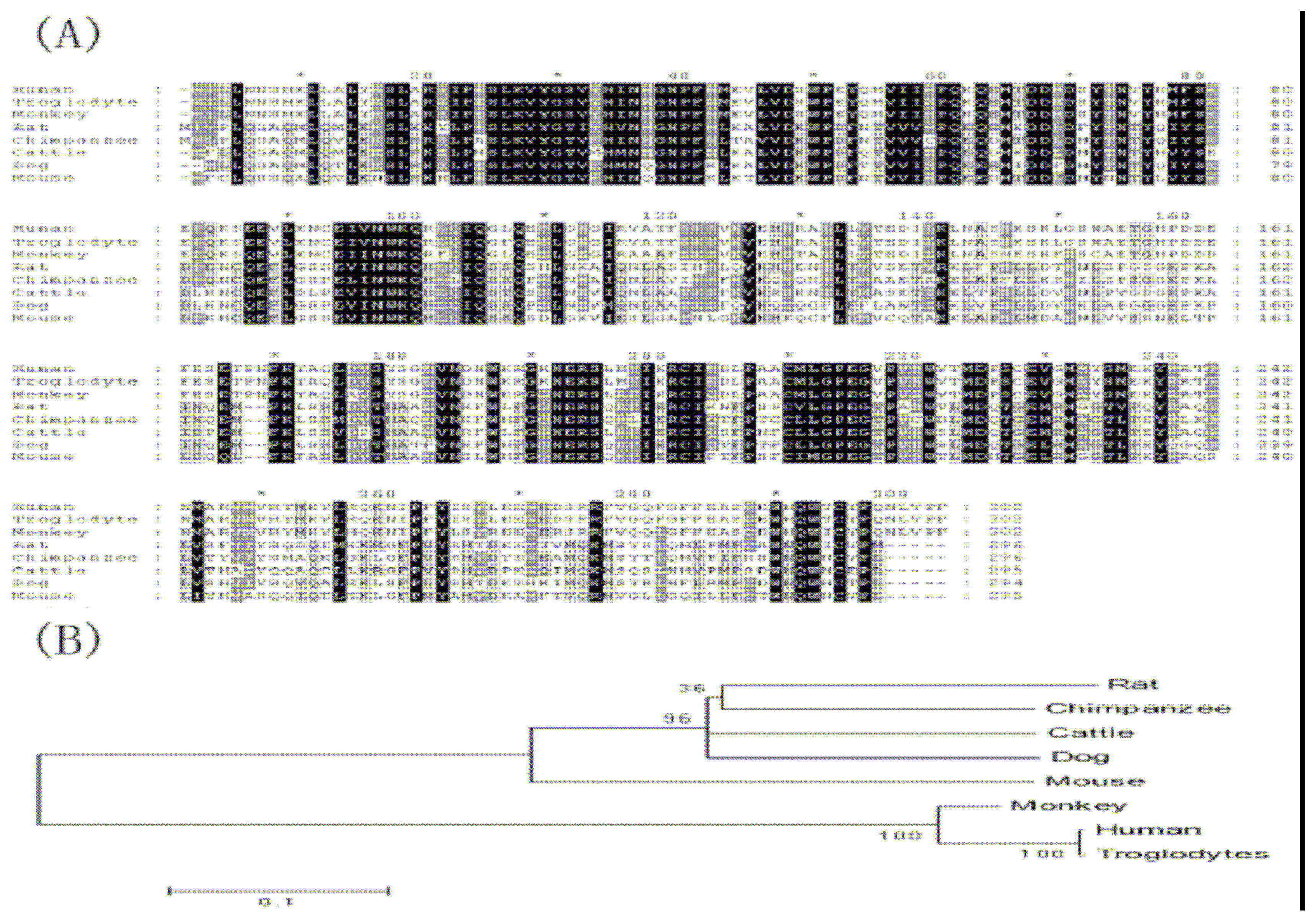
http://genome.ucsc.edu). The sequences were aligned by Clustal X software and viewed by GENEDOC software. Phylogenetic tree analysis of amino acid sequences deduced from
GLYATL1 DNA sequences was performed using Mega program.
2.2 Tissue distribution of GLYATL1
Human MTC (multiple-tissue cDNA) panels (Bio-Chain, USA) including bone marrow, kidney, stomach, bladder, lung, placenta, pancreas, heart, spleen, liver, thymus, testis, colon, uterus cervix, ovary, brain, skeletal muscle, and prostate served as templates to study the distribution of
GLYATL1 with the primers:
GLYATL1-RT-Forward and
GLYATL1-RT-Reverse (
Table 1). β-Macroglobin (β-MG) sense primer β-MG-F and antisense primer β-MG-R (
Table 1) were used to specifically amplify a 290 bp amplicon of C-terminal of β-MG, which served as the internal control. Products were separated by DNA electrophoresis in 2 % (w/v) agarose gel.
2.3 Cell culture, transfection and western blotting
HEK 293T cells were cultured in DMEM (Dulbecco’s modified Eagle’s medium; Gibco-BRL) which was supplemented with 10 % FBS, and kept in the CO2 incubator (37 °C, 5 % CO2). The cultured HEK 293T cells were transfected with expression vectors using Lipofectamine2000TM (Invitrogen, USA) according to the manufacturer’s protocol. The cells were harvested 48 h after transfection and washed twice with an ice cold PBS (phosphate-buffered saline) and then lysed on ice for 30 min in lysis buffer (Cell Signalling Technology, USA) supplemented with cocktail tablets and 0.1 mM PMSF with gentle shaking. The solution was then centrifuged at 12,000 g for 30 min at 4 °C to remove the debris, and supernatant was collected for use. Protein samples separated by SDS-PAGE were electro-transferred onto nitrocellulose membrane. The membrane was blocked at room temperature for 1 h with PBS containing 5 % (w/v) skim milk, and then washed with a mixture of PBS and 0.2 % Tween 20 (Sigma, USA). After blocking, incubated the membrane overnight at room temperature with mouse anti-Myc monoclonal antibody (CLONTECH, USA) diluted with PBS containing 1 % (w/v) BSA. Then the membrane was incubated with HRP-conjugated anti-mouse IgG antibody (Santa Cruz, USA) at room temperature for 1 h after washed with Tween-PBS. The membrane was then washed with Tween-PBS and then developed with the ECL system (Santa Cruz, USA).
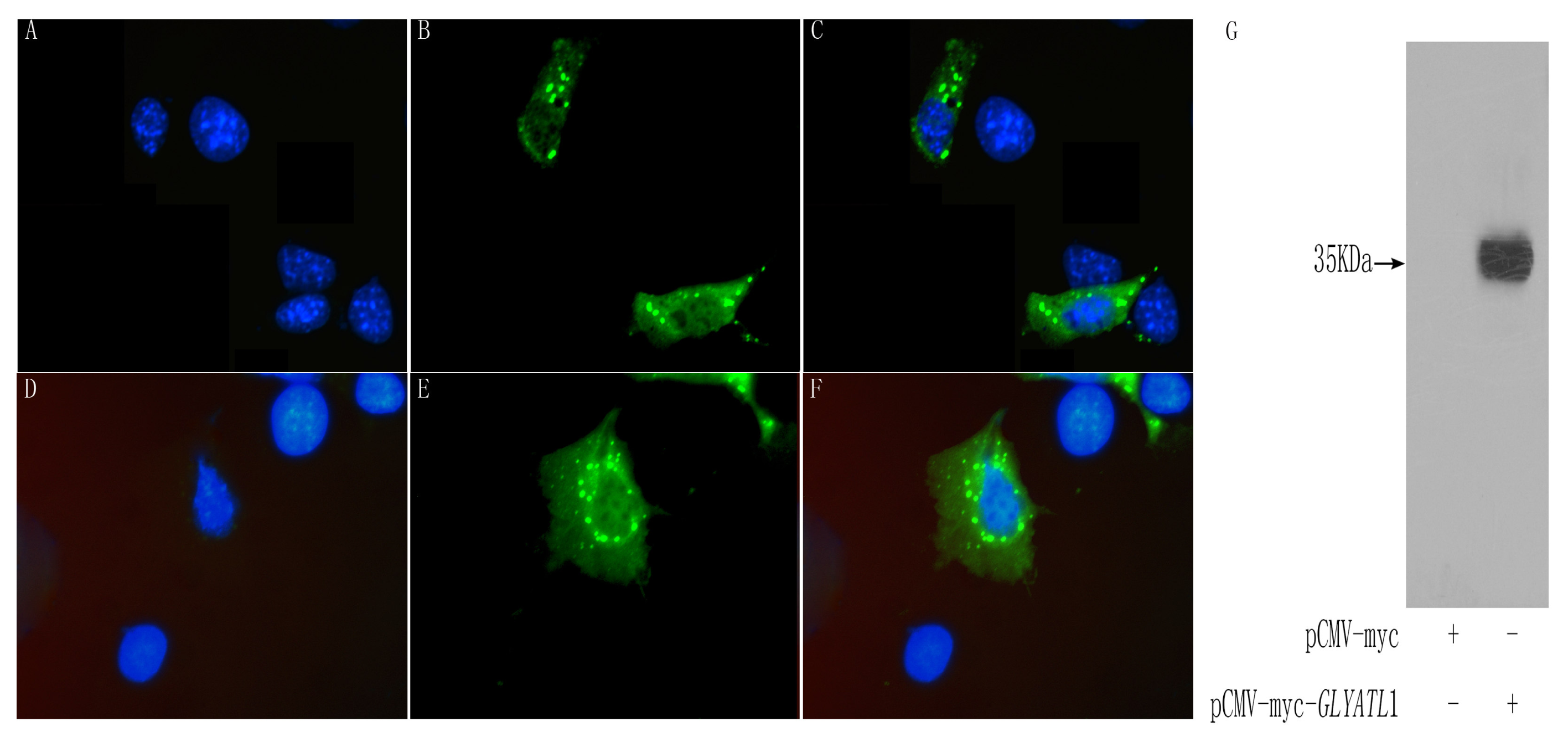
2.4 Subcellular localization analysis
To investigate the subcellular localization of the GLYATL1 protein, we proceeded in transient expression of Myc tagged vectors in COS-7 cells. Cells transfected with expression vectors were incubated on coverslips for 24–48 h. Cells were washed by PBS and then fixed in PBS containing 2–4 % paraformaldehyde, followed by 0.1 % Triton X-100/PBS treatment. After treatment with blocking reagent of 5 % goat serum for 20 min, cells were stained with mouse anti-Myc antibody, and stained with FITC-conjugated anti-mouse IgG goat antibody (ImmunoJackson, USA) for 1 h. Fluorochrome-labeled cells were visualized by using a microscope equipped with fluorescence optics (Leica DM RA2, Germany) and images were recorded with digital camera (Leica DC500). 2.5 μg/ml DAPI (4′,6-diamidino-2-phenylindole) in PBS was used to stain nuclei. In addition, pCMV-Myc was used as a negative control.
2.5 Pathway profiling assay
Mercury Pathway Profiling System (Clontech, USA) was used to investigate the potential roles of GLYAT1 in the signal pathways, and firefly luciferase as the reporter gene. HEK 293T cells were cultured in Dulbecco’s modified Eagle’s medium containing 10% fetal bovine serum. The cells were seeded on a 96-well plate at 1×104/well. After 24h, GLYAT1 expression vector, pCMV-Myc-GLYAT1, and each cisacting luciferase reporter vector of the Mercury Pathway Profiling System (Clontech, USA) were co-transfected into cells by the Lipofectamine2000 (Invitrogen, USA) according to the manufacturer’s protocol. After 48 h, the transfected cells were lysed and the activity of luciferase reporter gene was measured by dual-luciferase reporter assay system (Promega, USA). According to the primary results, another assay was performed to examine whether the activation of the HSE by the GLYATL1 was in a dose-dependent manner.
Discussion
The study of acyl-CoA: amino acid N-acyltransferases in liver or kidney appears important for several reasons. First, it may provide an explanation at the molecular level for the wide species diversity found in the excretion of amino acid conjugates of organic acids [
4]. Second, it should yield insights into the evolution and genetics of these important detoxifying systems. Third, it may relate to the pathogenesis and possibly the treatment of certain organic acidemias. Finally, it does provide a useful model for studying mechanisms of enzymatic transfer reactions.
During the past five decades, studies around the GLYAT mainly focused on the in vitro biochemical properties of the enzyme. However, the in vivo trait of the GLYAT, such as the genome structure, the cellular phenotype and etc, remained obscure. Although there are some other nucleotide acid sequences and correspondence amino acids sequence deposited in the GenBank database, they are mostly putative contigs which lack of support from experimental evidence. The major contribution of the present study lies in the first identification of a member of the GLYAT family in mRNA and cellular level.
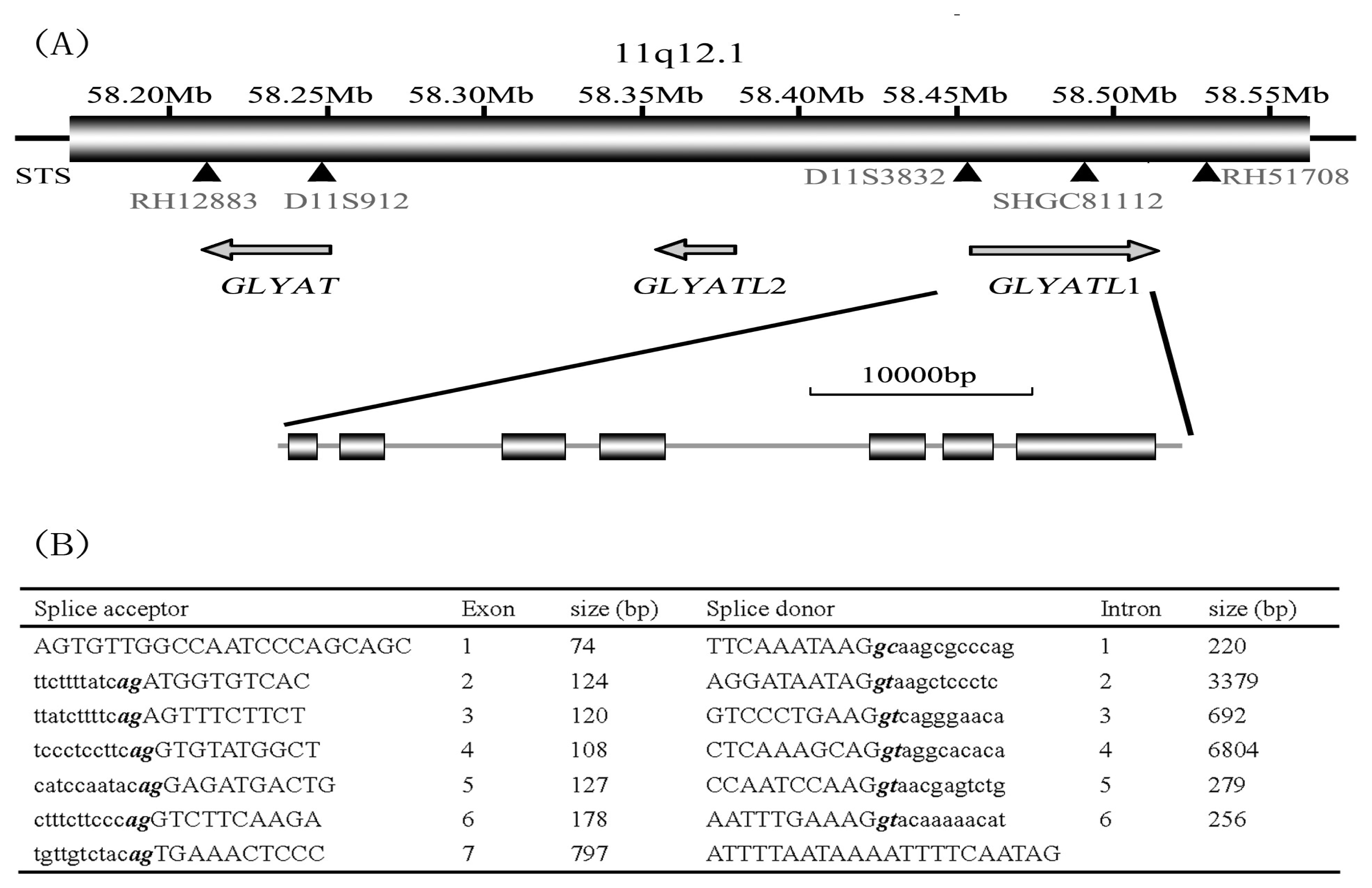
By the means of genome structure analysis, we isolated the human
GLYATL1 gene from human adult liver cDNA library. During the procedure of mapping the
GLYATL1 gene into the human chromosome 11q12.1, we interestingly found that another two members of this family localized in the neighborhood of the
GLYATL1 and there’s no other known gene between them (
Figure 3). And, a few partial mRNA with the sequence similarity to the nucleotide sequence of GLYATL domain also localized there. Further, through blast searching the GenBank database, we found that most human mRNA sequence similar to
GLYAT localized the 11q12.1 area. This finding implied: on one hand, this gene family might have some other unidentified members besides the
GLYAT,
GLYATL1,
GLYATL2, on the other hand, probably most the members belong to this family were cluster distributed in the 11q12.1 segment.
Previous works have reported the GLYAT protein exists in the liver and kidney tissues [
1,
6–
9], which is consistent with our experiment result. In the present study, we proved the GLYATL1 mainly expressed in the liver and kidney. Besides, we also detected a rather weak expression signal in some other adult tissue such as pancreas, testis, ovary and stomach. It was not reported before, which might suggest that these tissues, other than liver and kidney, could be the right place for GLYATL1 to execute its molecular function.
In response to elevated temperature, almost all organisms synthesize a certain set of proteins [
10], which are collectively referred to as heat shock proteins (HSPs) due to the method of induction. Induction of HSPs is regulated by the
trans-acting heat shock factors (HSFs) and
cis-acting heat shock element (HSE) present at the promoter region of each heat shock gene. Studies have revealed that many kinds of environmental stress, such as heavy metals, ethanol, amino acid analogues, anoxia and agents capable of perturbing protein structure, cause a similar response [
11]. Simultaneously, according to the previous research, the GLYAT was a component element of detoxification procedure in mammals by conjugating endogenous and exogenous carboxylic acids xenobiotics with amino acid. In our study, with the _help of the Mercury Pathway Profiling System, we found that GLYATL1 activated the transcriptional activity of the heat shock factor pathway (HSE) significantly. It was a clue about the correlation between the heat shock response pathway and the GLYATL1 involved detoxification mechanism. One of the possibilities lies in that, when executing its function in cells, GLYATL1 interacts with the heat shock factors (HSFs) and regulate the induction of HSPs consequently. Detail evidence for supporting our speculation is needed by further study.