1. Introduction
Osteosarcoma (OS) is the most common malignant bone-derived tumor and is particularly prevalent among children and adolescents. Although the long-term survival rate of OS patients has increased from 10% to nearly 80% within the last 25 years due to the use of neoadjuvant chemotherapy [
1], survival rates have plateaued [
2] and not risen over the last 15 years [
3]. Furthermore, despite recent advances in effective treatments, including surgery and chemotherapy, the 5-year survival rate of OS patients with lung metastasis is less than 30% [
4]. A better understanding of the molecular mechanisms involved in cancer progression is needed to reduce the financial burden and suffering of OS patients, to improve the treatment of OS, and to reduce the metastasis rate. Indeed, the mechanisms leading to OS metastasis remain unknown and represent an unmet challenge.
Cancer cell migration is a complex process that requires multiple steps and the participation of multiple signaling molecules. In general, these actions are controlled by a signaling cascade that is initiated by adhesion molecules (e.g., integrins) or growth factor receptors, such as the epidermal growth factor receptor (EGFR) [
5]. Several studies have shown that EGFR is expressed in bone and soft tissue tumors and that its expression is positively correlated with TNM (tumor-node-metastasis) stage in OS [
6,
7,
8,
9]. In vitro studies have also reported that EGFR inhibitors effectively reduce the growth of OS cells [
10,
11]. However, as one previous study found that the EGFR inhibitor gefitinib (20 μM) was unable to reduce the viability of several OS cell lines [
12], the role of EGFR in OS remains unresolved. Overall, a large-scale evaluation of EGFR mutations in OS is needed to determine the sensitivity of mutated EGFR to molecular targeting therapies [
13]. Moreover, novel downstream targets of EGFR that act via different mechanisms should be identified to further improve the survival rates of OS patients. The binding of EGF to its receptor induces EGFR dimerization at the plasma membrane, which results in trans-autophosphorylation and activation of the tyrosine kinase domain [
14,
15], ultimately leading to modulation of a variety of intracellular pathways, including the MAPK and PI3K/AKT pathways [
12].
Rho GTPases, small (21–25 kDa) GTP-binding proteins that belong to the Ras superfamily, play crucial roles in regulating cell migration, proliferation, polarity, apoptosis, cycle progression, adhesion, and motility [
16,
17,
18,
19]. EGF triggers cytoskeletal reorganization of actin microfilaments by regulating members of the Rho GTPase family, such as Rho A, Rac, and Cdc42 [
20]. Although EGFR stimulation has been reported to lead to reorganization of the cytoskeleton, membrane ruffling and focal adhesion through Rho protein activation [
21,
22], the mechanistic details by which Rho proteins function in EGF-induced OS cell migration remain unclear.
In this study, we hypothesized that the migration of OS cells induced by EGF might be regulated by the Rho family of GTPases. As our preliminary experiments indicated that Rho A has an important role in this process, the purpose of this study was to determine the detailed mechanism of Rho A’s involvement. We confirmed that EGF (10 ng/mL) stimulated MG63 cell migration and explored expression of Rho A and its downstream targets (ROCK1, LIMK2, Cofilin1) in OS cells. We applied different inhibitors and determined their effects on OS cell migration. This newly discovered signaling cascade described herein mediates EGF-induced actin microfilament formation in MG63 cells and may provide a new target for preventing and treating OS.
3. Discussion
EGF promotes OS migration through activation of EGFR, which positively correlates with TNM stage in OS. It is well known that most kinases activate their downstream effectors through phosphorylation; however, previous reports have shown that small molecule kinase inhibitors, such as Gefitinib and BIBW 2992, have little effect on OS development and no significant inhibitory effect on cell viability in vitro [
24]. These results suggest that EGFR is not a central regulator of OS cell proliferation under in vitro cell culture conditions. Thus, it is imperative to identify the common molecular alterations that drive OS proliferation and migration, and there is an urgent need to identify effective targets that are downstream from EGFR in OS.
In this study, MG63 cell migration was induced with 10 ng/mL EGF, which is in agreement with a previous study demonstrating that EGF is a cytokine with multifunctional activity, even in the same cell type [
25]. The results presented in
Figure 1F indicate that actin stress fiber formation was induced by EGF at different time points. Thus, we hypothesized that Rho GTPases are involved in cell cytoskeleton formation and organization. EGFR activation reportedly leads to membrane reorganization, ruffling of the cytoskeleton and focal adhesions through activation of Rho GTPases [
21], which are GTP-binding proteins involved in cell cytoskeleton formation and organization, as well as transcription, migration, and proliferation. Rho GTPases function as sensitive molecular switches existing either in an active GTP-bound form or in an inactive GDP-bound form, and the GTP hydrolytic activity of the former is responsible for their involvement in cytoskeleton rearrangement and cell motility [
26]. Jian Han et al. reported that Rho A and Rho C play important roles in the EGF-mediated migration of trophoblast cells and that Rho A regulates this migration through F-actin cytoskeleton reorganization [
27]. Taken together, we demonstrated that Rho A is clearly activated and in the GTP-bound form at 6 h after EGF stimulation. Additionally, we showed that Rho A plays a key role in MG63 cell migration when EGFR is overexpressed. To further clarify these findings, we inhibited GTP-Rho A with Rho A shRNA and the inhibitor C3. According to our results, Rho A inhibition suppresses MG63 cell migration and stress fiber formation. Furthermore, Rho A may be a downstream target of EGFR overexpression in OS. The downstream effectors of Rho A need to be examined and the signaling pathway dissected to reveal the mechanism of EGF-induced migration.
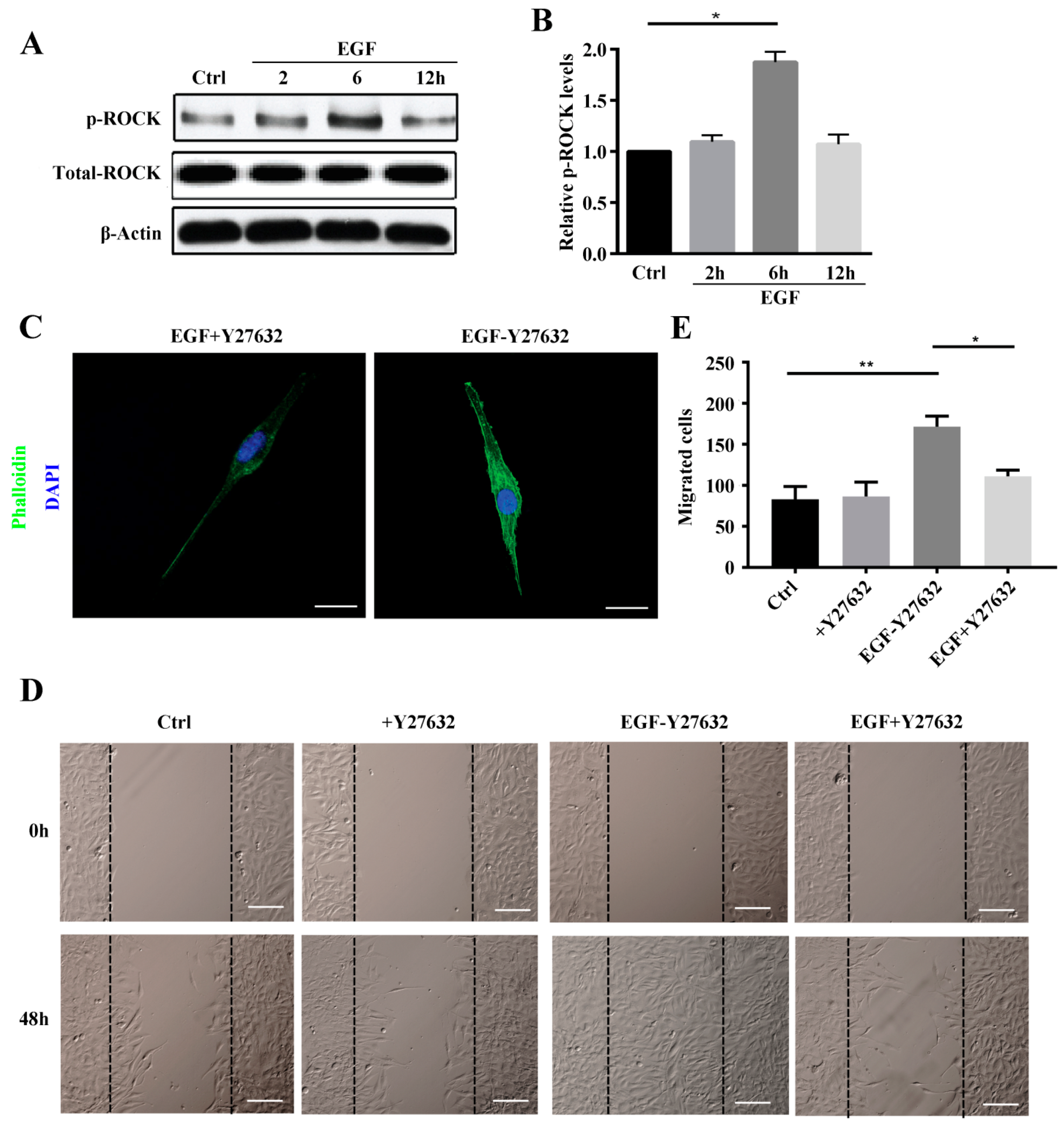
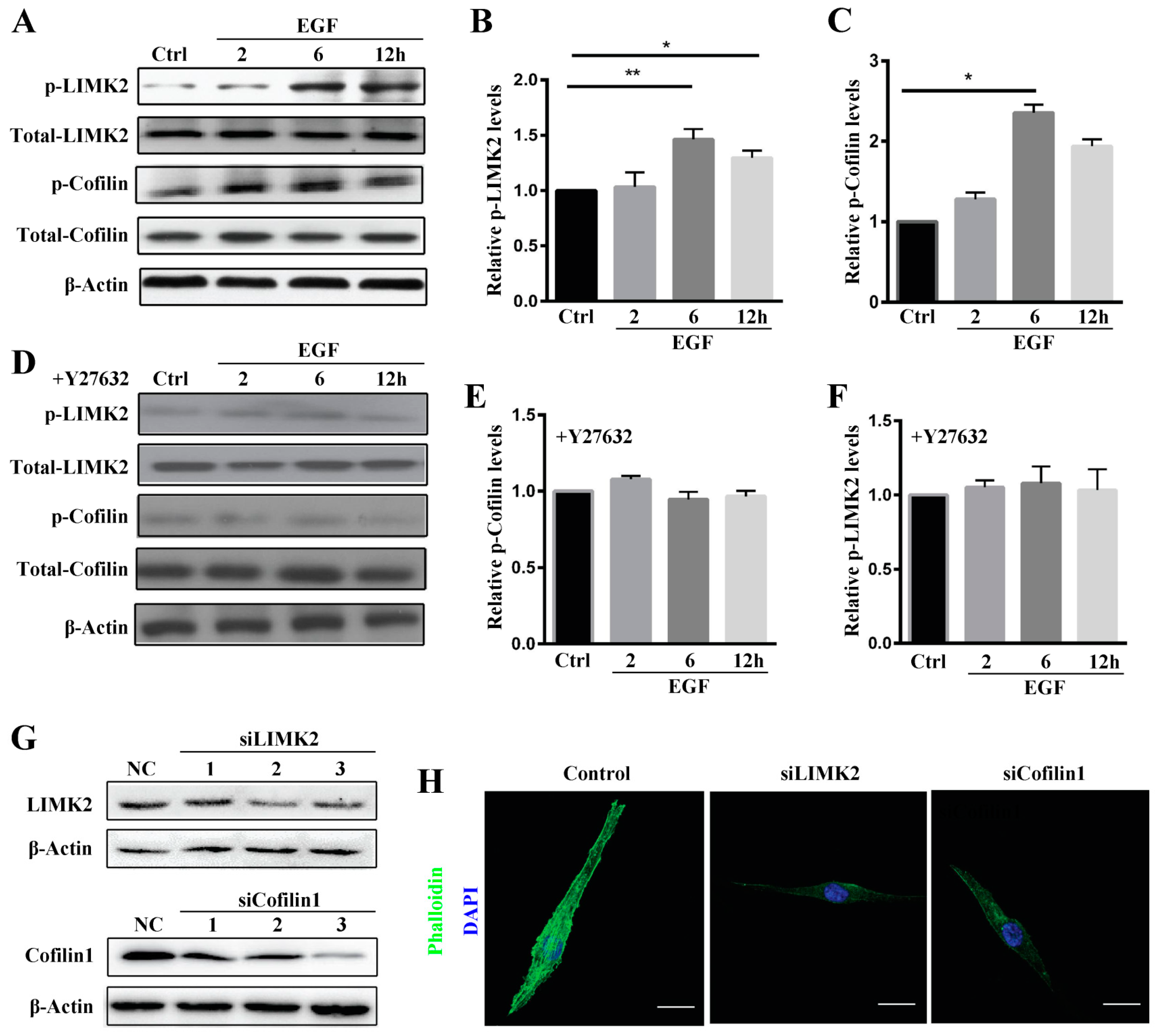
Rho-associated protein kinase (ROCK) belongs to the AGC (PKA/PKG/PKC) family of serine-threonine kinases, which are central regulators of the actin cytoskeleton that function downstream of the small GTPase Rho. ROCK signaling plays an essential role in a wide range of human diseases, and it has been recognized as a potential target and major modulator in the treatment of diseases involving abnormal cell motility, tumor cell invasion, and actin cytoskeleton organization [
28,
29]. Consistent with this, we found that ROCK inhibition reduced stress fiber formation. We also investigated ROCK downstream signaling to elucidate the mechanisms underlying ROCK participation in stress fiber formation. The data showed LIMK2 and Cofilin1 to be activated in MG63 cells after EGF treatment and that stress fiber formation was reduced by siRNA, which clearly demonstrates that the ROCK-LIMK2-Cofolin1 axis regulates stress fiber formation but does not inhibit cell migration.
In conclusion, our study shows that EGF (10 ng/mL) activates EGFR phosphorylation and stimulates MG63 cell migration, which can tailor the EGFR overexpression model. We herein provide evidence that MG63 cell migration and stress fiber formation occur through Rho A activation when EGFR is overexpressed. The downstream ROCK-LIMK2-Cofilin1 pathway also regulates the actin cytoskeleton formation in MG63 cells induced by EGF but not their migration. Our findings may have clinical implications that warrant further research using animal models to evaluate treatments for OS. This study has some limitations. First, we only used the MG63 cell line as a cell model, and additional OS cell lines, such as HOS, 143B, and Saos-2 cells, need to be explored. Second, primary OS cells from patients should be evaluated to assess the role of Rho A in regulating EGF-induced human OS cell migration. Third, evidence for the enhancement of migration and invasion after EGF treatment, as mediated through EGFR activation, is insufficient, and direct inhibition of EGFR activation by inhibitors or antibodies is required.
4. Materials and Methods
4.1. Materials and Reagents
The following antibodies served as primary antibodies: rabbit monoclonal antibodies against ROCK1 (Cell Signaling Technology, Danvers, MA, USA, #4035, 1:1000) and phosphorylated-ROCK (p-ROCK; Abcam, Cambridge, UK, #ab203273); rabbit polyclonal antibodies against LIMK2 (Cell Signaling Technology, Danvers, MA, USA, #3845) and phosphorylated-LIMK2 (p-Limk2; Sigma, St Louis, MO, USA, #SAB4300104); mouse monoclonal antibodies against Rho A (Santa Cruz Biotechnology, Dallas, TX, USA, #sc-418, 1:500). Antibodies against Cofilin (#5175) and p-Cofilin (#3311) were purchased from Cell Signaling Technology (Danvers, MA, USA).
EGF was obtained from Worthington Biochemical Corporation (Lakewood, NJ, USA). Regarding specific inhibitors, ROCK1 (Y-27632) was obtained from Selleck (Houston, TX, USA), and Rho Inhibitor I (CT04-A) was purchased from Cytoskeleton (Danvers, MA, USA). Whole Cell Lysis Protein Extraction Kit was purchased from Nanjing keyGEN BioTECH, Ltd. (Nanjing, China). Actin-Tracker Green was obtained from Shanghai Beyotime, Ltd. (Shanghai, China). UltraCruz® Hard-set Mounting Medium was purchased from Santa Cruz Biotechnology (TX, USA, #sc-359850).
4.2. Cell Culture
MG63 cells were obtained from the Department of Histology and Embryology, Southern Medical University (Guangzhou, China). MG63 cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Corning, Herndon, VA, USA) containing 10% fetal bovine serum (FBS, SeraBest, Aidenbach, Bavaria, Germany), 100 U/mL penicillin and 100 μg/mL streptomycin (Gibco, Grand Island, NY, USA) at 37 °C in a humidified atmosphere of 5% CO2. Unless otherwise noted, this medium was used as the standard medium. In some experiments, MG63 cells were pretreated with the ROCK1 inhibitor Y-27632 (10 mM; Selleck, TX, USA). EGF was added to the culture medium at the indicated time points (2, 6, and 12 h).
4.3. Pull Down Assay
Activity of Rho GTPases was assessed in OS cells using a pull-down assay for GTP-bound Rho according to the manufacturer’s protocol (Rho Assay Reagent (Rhotekin RBD, Agarose), Millipore, Burlington, MA, USA, cat. #14-383). GTP-bound Rho A was precipitated from cell lysates with PAK-GST Protein Beads ((human p21 activated kinase PBD) Cat. #PAK02). Briefly, MG63 cells were starved in serum-free medium for 24 h, rinsed with ice-cold Tris-buffered saline (TBS) and lysed in 600 µL of RIPA for 10 min. An aliquot (120 µg, 600 µL) of the extract was placed into two experimental tubes; GDP (1/100th of the aliquot volume) was then added to one tube, whereas GTPγS (1/100th of the aliquot volume) was added to the other tube. The tubes were incubated at room temperature for 15 min. The reaction was stopped by adding 1/10th the volume of stop buffer to each tube (final conc. 60 mM MgCl2). PAK-GST protein beads were resuspended, and 20 µg (20 µL) of the protein-bound beads was added to each reaction tube. The tubes were gently rotated at 4 °C for 1 h and then centrifuged at 5200× g at 4 °C for 1 min. The supernatant was removed and the beads were washed twice with 500 µL of wash buffer. The beads were pelleted and suspended in 25 µL of sodium dodecyl sulfate (SDS) sample buffer. The bound proteins were boiled in sample buffer at 99 °C for 10 min and then analyzed by western blotting to detect GTP-Rho A.
4.4. Lentiviral shRNA and Transduction In Vitro
Knockdown of Rho A was achieved via lentiviral transduction of human Rho A shRNA (sc-29471-V; Santa Cruz Biotechnology). According to the manufacturer’s instructions, scrambled shRNA (SC-108080) and the mCherry control (LPP-MCHR-LV105-025-C) were also included. The lentiviral particle solution was added to MG63 cells in fresh DMEM containing 5 mg/mL polybrene. The medium was changed after 12 h of infection. After successive 12-h intervals, the medium was changed to fresh DMEM, and the cells were cultured for 72 to 96 h. After transduction, quantitative reverse transcription-polymerase chain reaction (qRT-PCR) analysis was employed to determine the knockdown efficiency.
4.5. Real-Time Quantitative PCR
qRT-PCR of total RNA from MG63 cells was extracted using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions, and then 1 μg of RNA was reverse transcribed to generate cDNA using superscript reverse transcriptase (Invitrogen). qRT-PCR was carried out using a Real-time Quantitative PCR Detecting System. The primer sequences were as follows: Rho A shRNA (sc-29471-PR; Santa Cruz Biotechnology); GAPDH forward: 5′-TGACTTCAACAGCGACACCCA-3′; GAPDH reverse: 5′-CACCCTGTTGCTGTAGCCAAA-3′. The relative amounts of each transcript were calculated using the comparative Ct (2−ΔΔCt) method.
4.6. siRNA and Transfection In Vitro
The sequences of the human Cofilin1 and LIMK2 genes were obtained from GenBank (gene ID 1072 and 3985). Sequence-specific siRNAs targeting Cofilin1 and LIMK2 (Cofilin1 siRNA (5′-GGATCAAGCATGAATTGCA-3′) and LIMK2 siRNA (5′-GCTGCAAGGTGATCATTGA-3′)) were designed and synthesized by RiboBio Co., Ltd. (Guangzhou, China); a negative control siRNA (NC) was also designed by RiboBio Co., Ltd. All chemical transfections were performed using 60 nM of each siRNA and Lipofectamine™ 2000 (Invitrogen, Carlsbad, CA, USA), as described by the manufacturer. After 6 h at 37 °C, DMEM was replaced with complete growth medium (DMEM with 20% FBS). Cells not transfected with siRNA were used as an untreated control. Transcription expression was evaluated by western blotting.
4.7. Wound-Healing Assay
MG63 cells were seeded in 6-well plates at a density of 106 cells/well and allowed to reach adequate confluence over one day in 2.5 mL of DMEM supplemented with 10% FBS. Thereafter, the medium was removed and replaced with serum-free medium. On the next day (24 h), sterile 200 μL pipette tips were used to scrape the monolayer of MG63 cells across each well, creating a cell-free area. The cell monolayer was washed gently with phosphate-buffered saline (PBS) to remove detached cells and cell debris. At various time points (12, 24, 36, and 48 h), images were captured using an inverted microscope (Olympus, Tokyo, Japan) at 20× magnification to quantify the closure of the scratch. Differences between the wound area at time 0 and the other four time points were determined. The scratch area was measured using ImageJ software (National Institutes of Health, Bethesda, MD, USA).
4.8. Transwell Migration Assay
MG63 cells transfected with Rho A shRNA and the inhibitor Y27632 and negative control cells were starved overnight in serum-free medium. EGF (10 ng/mL) was added to the cells for 24 h before treating with trypsin. Cells (2 × 105) in 400 µL of medium were added to the top chamber of 24-well transwell plates (8 µm pore size, Corning, NY, USA). The bottom chambers were filled with 600 µL of medium containing EGF (10 ng/mL). Transwell cultures without EGF were used as a control. After 24 h under standard culture conditions, non-migrated cells on the upper side of the membrane were removed with a cotton swab. The cells at the bottom of the filter were fixed with Immunol Staining Fix Solution (Beyotime, Shanghai, China) and stained with 0.1% crystal violet at room temperature. After air-drying, the membrane was mounted on a glass slide and examined under a microscope. All experiments were repeated at least 3 times.
4.9. Fluorescent Staining of the Cytoskeleton
MG63 cells were washed three times with PBS and fixed with Immunol Staining Fix Solution (Beyotime) for 10 min, after which time they were washed with Immunol Staining Wash Buffer (Beyotime) 3 times for 5 min each. Actin-Tracker Green was diluted in Immunol Fluorescence Staining Secondary Antibody Dilution Buffer (Beyotime) and incubated with the cells for 40 min in the dark, after which they were washed with Immunol Staining Wash Buffer 3 times for 5 min each. Finally, nuclei were stained with UltraCruz® Hard-set Mounting Medium including 4′,6-diamidino-2-phenylindole (DAPI; sc-359850, Santa Cruz Biotechnology). Images were recorded using a Zeiss confocal laser-scanning microscope equipped with a Plan-Apochromat 60× oil-immersion objective lens (Zeiss LSM 880 Confocal Laser Scanning unit, Carl Zeiss AG, Oberkochen, Germany). Image processing, combining and analysis were performed using the ZEN software package (Carl Zeiss MicroImaging GmbH, Jena, Germany).
4.10. Western Blotting
To extract proteins, cultured MG63 cells were collected and lysed in 1X RIPA buffer according to the instructions. The protein content was determined using a BCA Protein Assay Kit (Beyotime, Shanghai, China) using BSA (bovine serum albumin) as the standard. Samples were boiled for 5 min at 99 °C and separated by 10% SDS-polyacrylamide gel electrophoresis (PAGE) using running buffer (190 mM glycine, 25 mM Tris, 0.1% SDS) and then transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, MA) using transfer buffer (190 mM glycine, 25 mM Tris, 30% methanol). The membranes were blocked for 2 h in 5% milk powder in TBS-Tween-20 (TBST) at room temperature, incubated overnight at 4 °C with the primary antibody diluted 1:1000 in 5% BSA. After washing three times with TBST, the membranes were incubated with a secondary antibody for 1 h at room temperature. Finally, the membranes were stained with ECL Western Blotting Substrate (Thermo, Waltham, MA, USA), scanned, and stored using a gel imaging system. The protein bands were analyzed using Image Lab software. Images shown in figures are representative of 3 independent experiments.
4.11. Statistical Analyses
All experiments were repeated three times. SPSS software (version 13.0, SPSS Inc., Chicago, IL, USA) was used for all statistical analyses, and the data are expressed as the mean ± SD. Statistical differences between groups were determined by one-way analysis of variance (ANOVA). A p-value < 0.05 was considered significant, and the significance level was defined as * p < 0.05, ** p < 0.01.