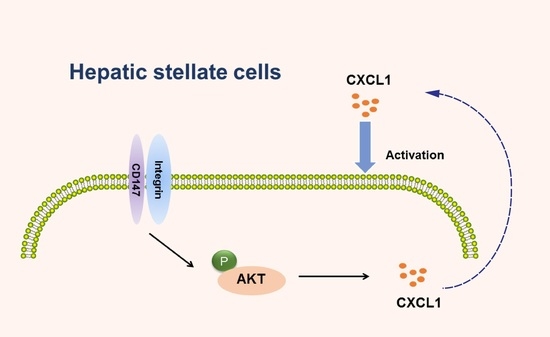
CD147 Promotes CXCL1 Expression and Modulates Liver Fibrogenesis
Abstract
:1. Introduction
2. Results
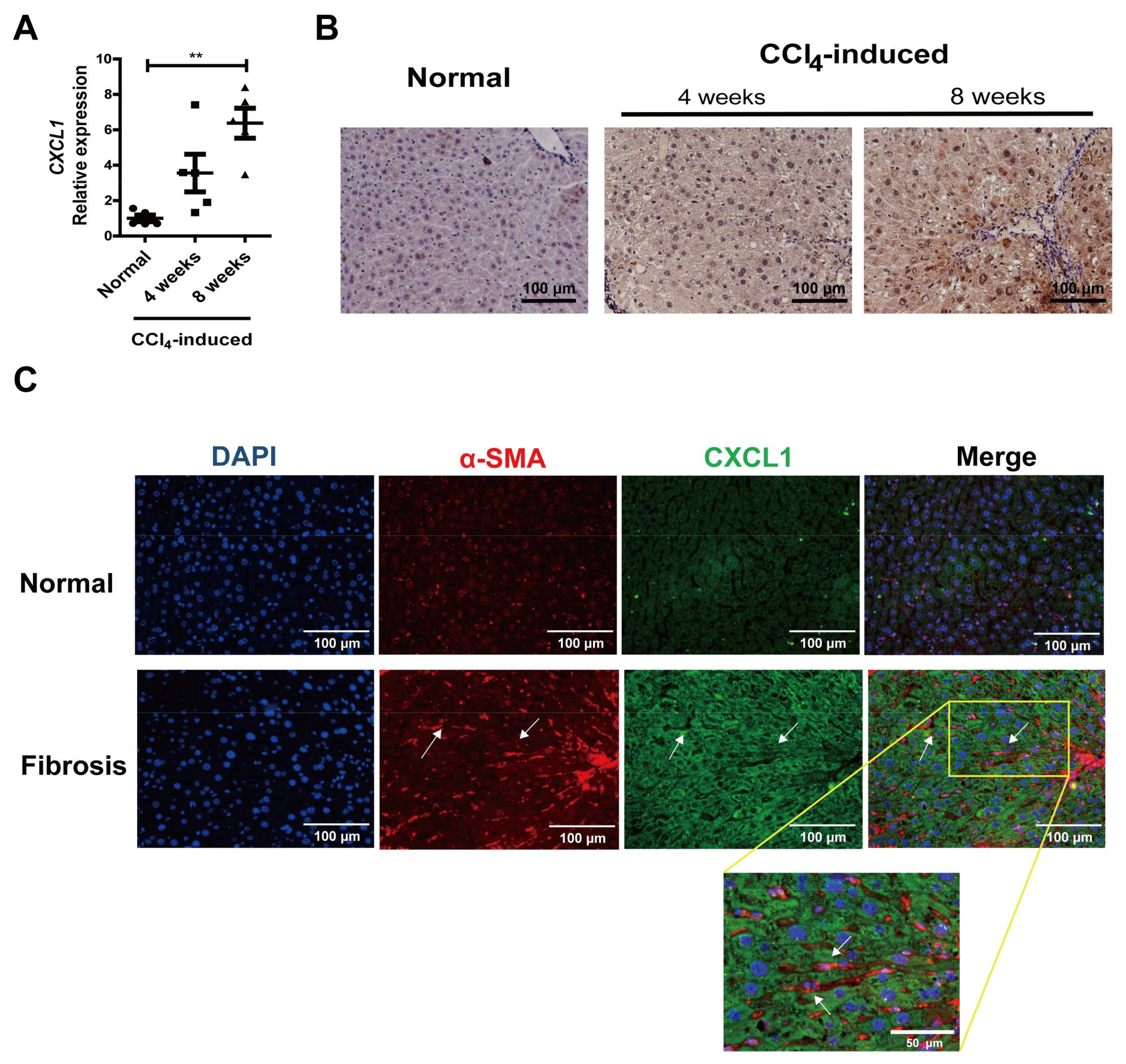
2.1. CXCL1 Expression Was Increased in Activated HSCs
2.2. CXCL1 Promoted HSCs Activation and Co-Localized with CD147 in HSCs
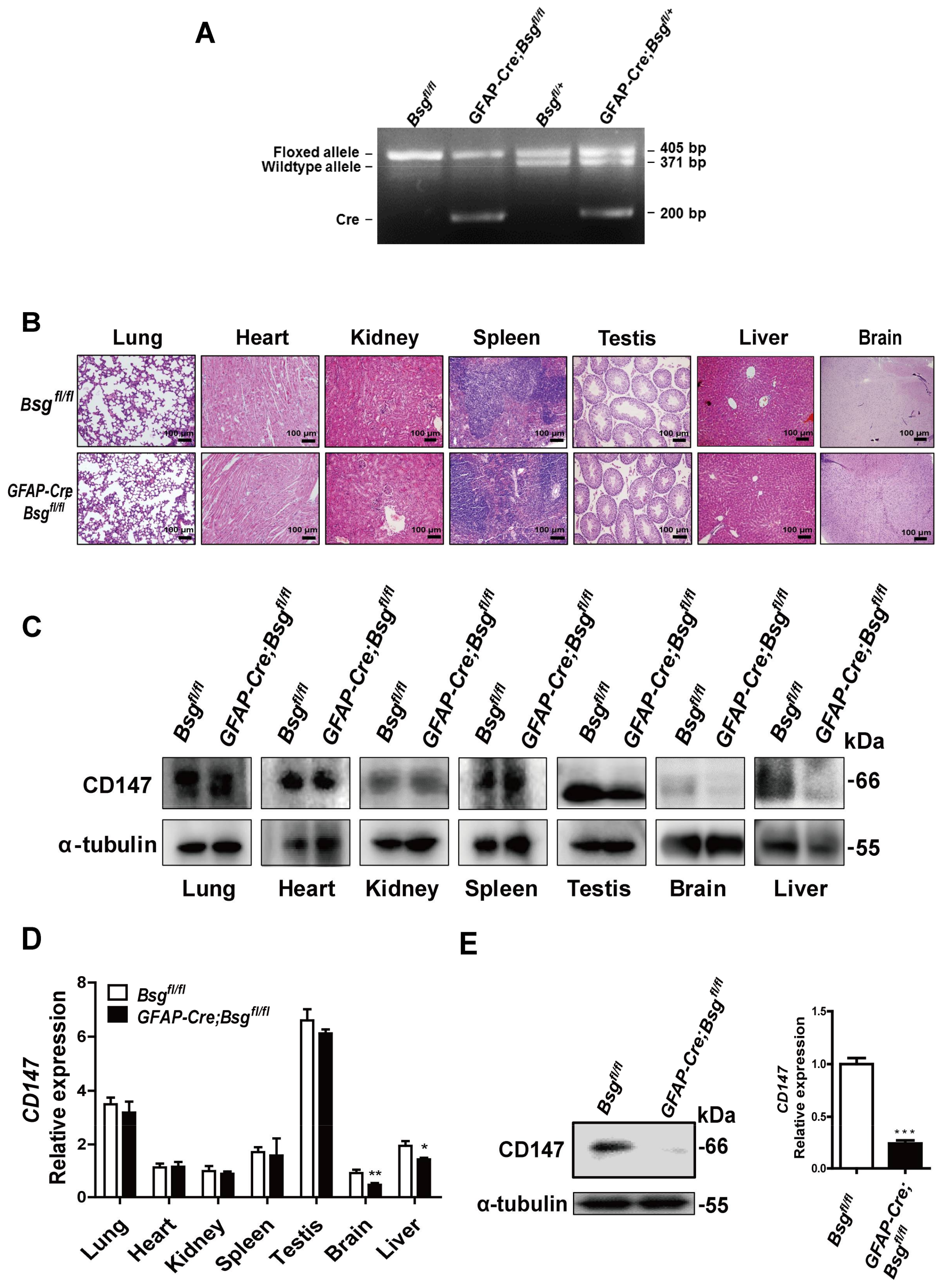
2.3. Generation of HSCs-Specific CD147-Knockout Mice
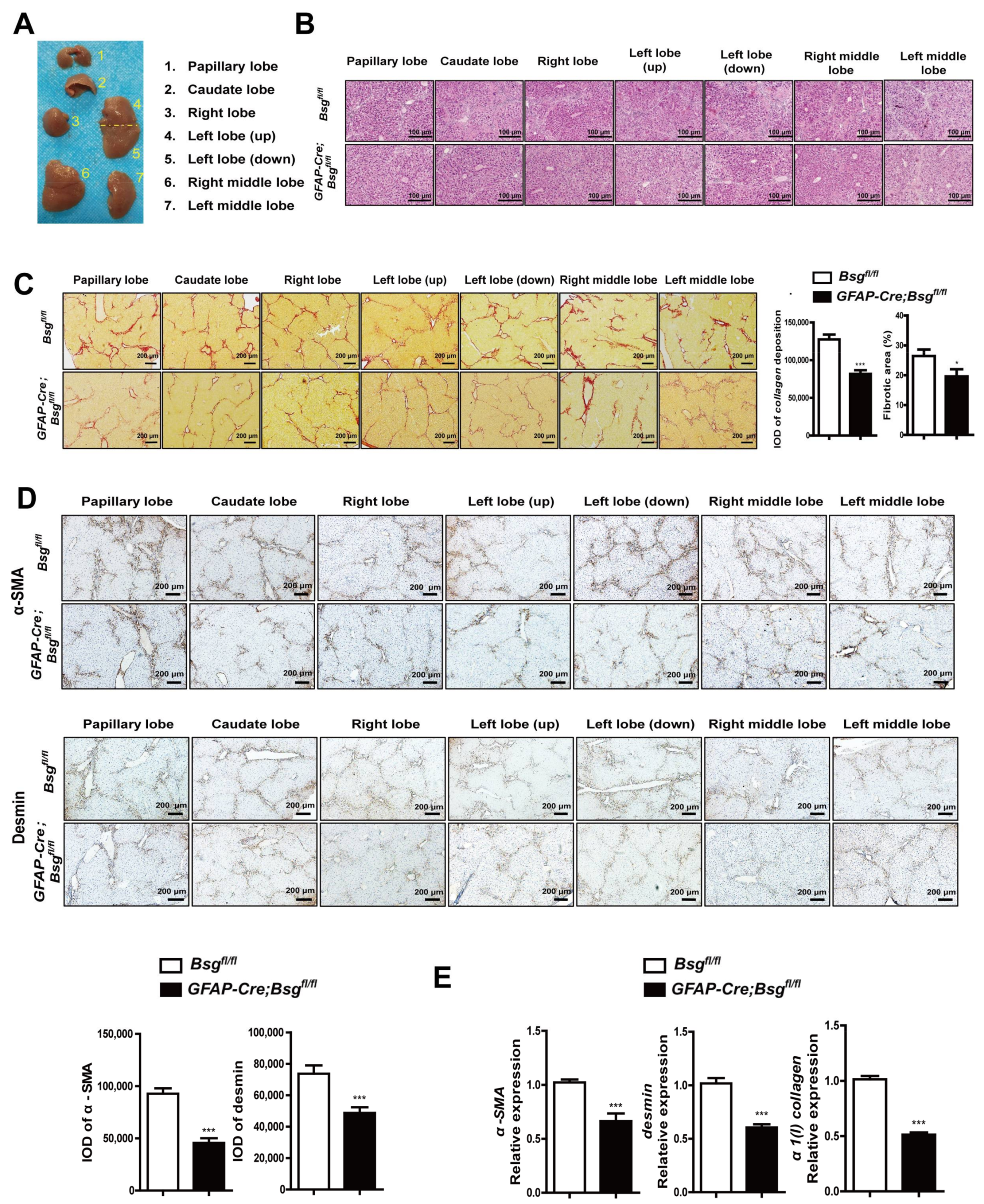
2.4. CD147 Deletion in HSCs Alleviated CCl4-Induced Liver Fibrosis and Deregulated CXCL1 Expression
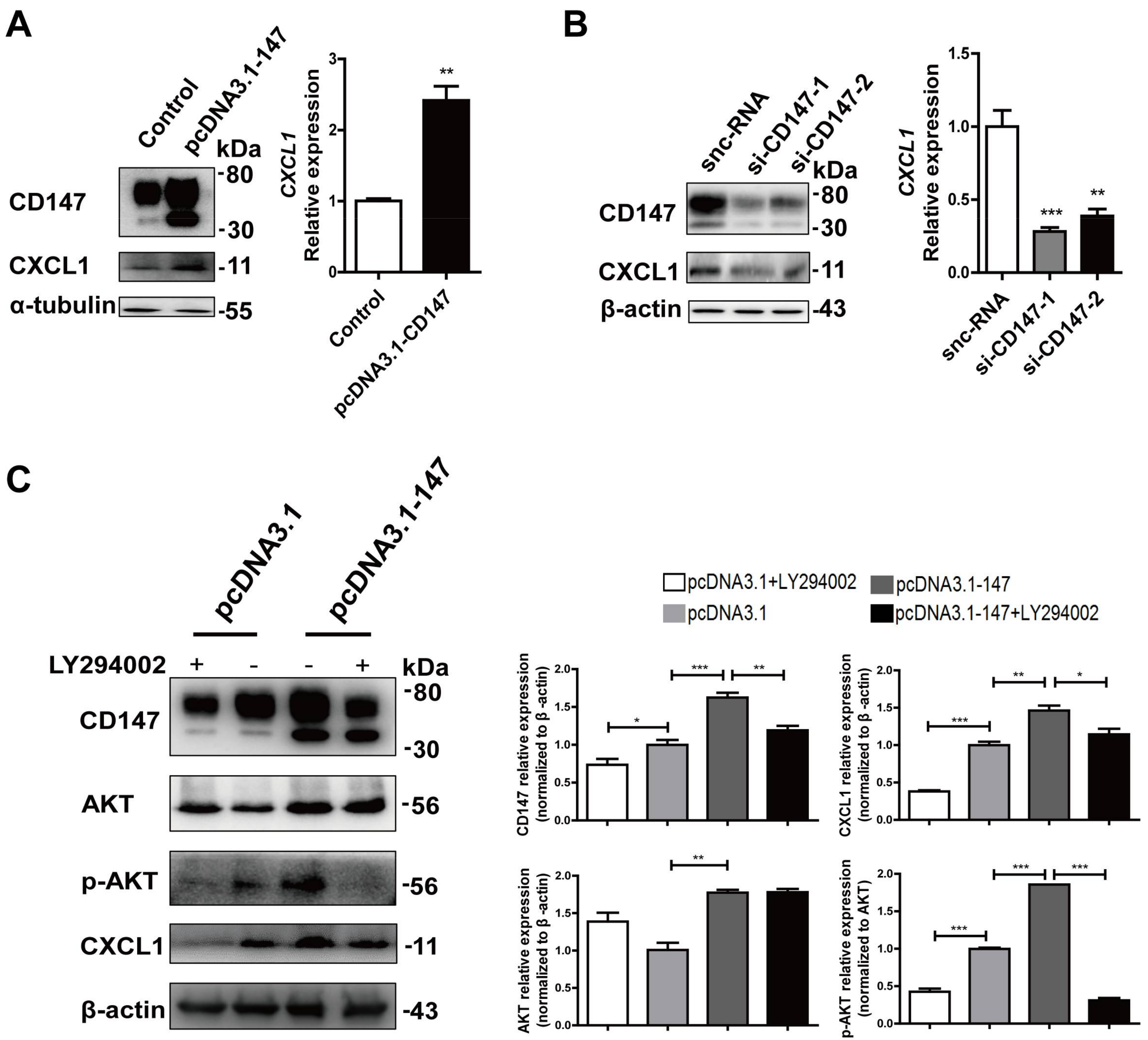
2.5. CD147-Regulated CXCL1 Expression in HSCs via the PI3K/AKT Pathway
3. Discussion
4. Materials and Methods
4.1. Cells Culture and Reagents
4.2. Mice
4.3. siRNA Transfection
4.4. Western Blot
4.5. Real-Time PCR
4.6. Enzyme-Linked Immunosorbent Assay (ELISA)
4.7. Cell Contraction Assay
4.8. Collagen Staining
4.9. Immunohistochemistry
4.10. Immunofluorescence
4.11. Fluorescence Activated Cell Sorting (FACS)
4.12. CCK-8
4.13. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| α-SMA | α-smooth muscle actin |
| CXCL1 | CXC chemokine-ligand-1 |
| CCl4 | Carbon tetrachloride |
| ECM | Extracellular matrix |
| ELISA | Enzyme-linked immunosorbent assay |
| FACS | Fluorescence activated cell sorting |
| GFAP | Glial fibrillary acidic protein |
| HSC | Hepatic stellate cell |
| MFI | Median fluorescence intensity |
| snc-RNA | Silencer-negative control siRNA |
References
- Higashi, T.; Friedman, S.L.; Hoshida, Y. Hepatic stellate cells as key target in liver fibrosis. Adv. Drug Deliv. Rev. 2017, 121, 27–42. [Google Scholar] [CrossRef] [PubMed]
- Pellicoro, A.; Ramachandran, P.; Iredale, J.P.; Fallowfield, J.A. Liver fibrosis and repair: Immune regulation of wound healing in a solid organ. Nat. Rev. Immunol. 2014, 14, 181–194. [Google Scholar] [CrossRef] [PubMed]
- Sahin, H.; Trautwein, C.; Wasmuth, H.E. Functional role of chemokines in liver disease models. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 682–690. [Google Scholar] [CrossRef] [PubMed]
- Nischalke, H.D.; Berger, C.; Luda, C.; Muller, T.; Berg, T.; Coenen, M.; Kramer, B.; Korner, C.; Trebicka, J.; Grunhage, F.; et al. The CXCL1 rs4074 A allele is associated with enhanced CXCL1 responses to TLR2 ligands and predisposes to cirrhosis in HCV genotype 1-infected Caucasian patients. J. Hepatol. 2012, 56, 758–764. [Google Scholar] [CrossRef] [PubMed]
- Stefanovic, L.; Brenner, D.A.; Stefanovic, B. Direct hepatotoxic effect of KC chemokine in the liver without infiltration of neutrophils. Exp. Biol. Med. 2005, 230, 573–586. [Google Scholar] [CrossRef]
- Strieter, R.M.; Burdick, M.D.; Mestas, J.; Gomperts, B.; Keane, M.P.; Belperio, J.A. Cancer CXC chemokine networks and tumour angiogenesis. Eur. J. Cancer 2006, 42, 768–778. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.X.; Wang, Q.; Zhang, Z.M.; Huang, S.; Yuan, W.P.; Liu, J.Y.; Ban, K.C.; Zhao, Y.N. Identifying serological biomarkers of hepatocellular carcinoma using surface-enhanced laser desorption/ionization-time-of-flight mass spectroscopy. Cancer Lett. 2009, 279, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Grass, G.D.; Dai, L.; Qin, Z.; Parsons, C.; Toole, B.P. CD147: Regulator of hyaluronan signaling in invasiveness and chemoresistance. Adv. Cancer Res. 2014, 123, 351–373. [Google Scholar] [PubMed]
- Kanekura, T.; Chen, X. CD147/basigin promotes progression of malignant melanoma and other cancers. J. Dermatol. Sci. 2010, 57, 149–154. [Google Scholar] [CrossRef] [PubMed]
- Partridge, E.A.; Le Roy, C.; Di Guglielmo, G.M.; Pawling, J.; Cheung, P.; Granovsky, M.; Nabi, I.R.; Wrana, J.L.; Dennis, J.W. Regulation of cytokine receptors by Golgi N-glycan processing and endocytosis. Science 2004, 306, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Li, H.Y.; Ju, D.; Zhang, D.W.; Li, H.; Kong, L.M.; Guo, Y.; Li, C.; Wang, X.L.; Chen, Z.N.; Bian, H. Activation of TGF-beta1-CD147 positive feedback loop in hepatic stellate cells promotes liver fibrosis. Sci. Rep. 2015, 5, 16552. [Google Scholar] [CrossRef]
- Zhang, D.W.; Zhao, Y.X.; Wei, D.; Li, Y.L.; Zhang, Y.; Wu, J.; Xu, J.; Chen, C.; Tang, H.; Zhang, W.; et al. HAb18G/CD147 promotes activation of hepatic stellate cells and is a target for antibody therapy of liver fibrosis. J. Hepatol. 2012, 57, 1283–1291. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Teng, Y.; Sun, Q.; Xu, J.; Chen, Y.T.; Hou, N.; Cheng, X.; Yang, X.; Chen, Z.N. Important functional roles of basigin in thymocyte development and T cell activation. Int. J. Biol. Sci. 2013, 10, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Maubach, G.; Lim, M.C.; Zhang, C.Y.; Zhuo, L. GFAP promoter directs lacZ expression specifically in a rat hepatic stellate cell line. World J. Gastroenterol. 2006, 12, 723–730. [Google Scholar] [CrossRef]
- Neubauer, K.; Knittel, T.; Aurisch, S.; Fellmer, P.; Ramadori, G. Glial fibrillary acidic protein—A cell type specific marker for Ito cells in vivo and in vitro. J. Hepatol. 1996, 24, 719–730. [Google Scholar] [CrossRef]
- Puche, J.E.; Lee, Y.A.; Jiao, J.; Aloman, C.; Fiel, M.I.; Munoz, U.; Kraus, T.; Lee, T.; Yee, H.F., Jr.; Friedman, S.L. A novel murine model to deplete hepatic stellate cells uncovers their role in amplifying liver damage in mice. Hepatology 2013, 57, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Iimuro, Y.; Nishio, T.; Morimoto, T.; Nitta, T.; Stefanovic, B.; Choi, S.K.; Brenner, D.A.; Yamaoka, Y. Delivery of matrix metalloproteinase-1 attenuates established liver fibrosis in the rat. Gastroenterology 2003, 124, 445–458. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, P.; Iredale, J.P. Reversibility of liver fibrosis. Ann. Hepatol. 2009, 8, 283–291. [Google Scholar] [PubMed]
- Sun, M.; Kisseleva, T. Reversibility of liver fibrosis. Clin. Res. Hepatol. Gastroenterol. 2015, 39, S60–S63. [Google Scholar] [CrossRef] [PubMed]
- Atta, H.M. Reversibility and heritability of liver fibrosis: Implications for research and therapy. World J. Gastroenterol. 2015, 21, 5138–5148. [Google Scholar] [CrossRef]
- Huang, Y.; Deng, X.; Liang, J. Modulation of hepatic stellate cells and reversibility of hepatic fibrosis. Exp. Cell. Res. 2017, 352, 420–426. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Xu, J.; Brenner, D.A.; Kisseleva, T. Reversibility of liver fibrosis and inactivation of fibrogenic myofibroblasts. Curr. Pathobiol. Rep. 2013, 1, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Wasmuth, H.E.; Tacke, F.; Trautwein, C. Chemokines in liver inflammation and fibrosis. Semin. Liver Dis. 2010, 30, 215–225. [Google Scholar] [CrossRef] [PubMed]
- Morrison, M.C.; Liang, W.; Mulder, P.; Verschuren, L.; Pieterman, E.; Toet, K.; Heeringa, P.; Wielinga, P.Y.; Kooistra, T.; Kleemann, R. Mirtoselect, an anthocyanin-rich bilberry extract, attenuates non-alcoholic steatohepatitis and associated fibrosis in ApoE∗3Leiden mice. J. Hepatol. 2015, 62, 1180–1186. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Sadanandam, A.; Varney, M.L.; Nannuru, K.C.; Singh, R.K. Small interfering RNA-mediated CXCR1 or CXCR2 knock-down inhibits melanoma tumor growth and invasion. Int. J. Cancer 2010, 126, 328–336. [Google Scholar] [CrossRef] [PubMed]
- Saintigny, P.; Massarelli, E.; Lin, S.; Ahn, Y.H.; Chen, Y.; Goswami, S.; Erez, B.; O'Reilly, M.S.; Liu, D.; Lee, J.J.; et al. CXCR2 expression in tumor cells is a poor prognostic factor and promotes invasion and metastasis in lung adenocarcinoma. Cancer Res. 2013, 73, 571–582. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, Y.; Raimondo, M.; Woodward, T.A.; Wallace, M.B.; Gill, K.R.; Tong, Z.; Burdick, M.D.; Yang, Z.; Strieter, R.M.; Hoffman, R.M.; et al. CXC-chemokine/CXCR2 biological axis promotes angiogenesis in vitro and in vivo in pancreatic cancer. Int. J. Cancer 2009, 125, 1027–1037. [Google Scholar] [CrossRef] [PubMed]
- Nischalke, H.D.; Berger, C.; Lutz, P.; Langhans, B.; Wolter, F.; Eisenhardt, M.; Kramer, B.; Kokordelis, P.; Glassner, A.; Muller, T.; et al. Influence of the CXCL1 rs4074 A allele on alcohol induced cirrhosis and HCC in patients of European descent. PLoS ONE 2013, 8, e80848. [Google Scholar] [CrossRef] [PubMed]
- Chang, B.; Xu, M.J.; Zhou, Z.; Cai, Y.; Li, M.; Wang, W.; Feng, D.; Bertola, A.; Wang, H.; Kunos, G.; et al. Short- or long-term high-fat diet feeding plus acute ethanol binge synergistically induce acute liver injury in mice: An important role for CXCL1. Hepatology 2015, 62, 1070–1085. [Google Scholar] [CrossRef] [PubMed]
- Bataller, R.; Brenner, D.A. Liver fibrosis. J. Clin. Investig. 2005, 115, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Forbes, S.J.; Russo, F.P.; Rey, V.; Burra, P.; Rugge, M.; Wright, N.A.; Alison, M.R. A significant proportion of myofibroblasts are of bone marrow origin in human liver fibrosis. Gastroenterology 2004, 126, 955–963. [Google Scholar] [CrossRef] [PubMed]
- Troeger, J.S.; Mederacke, I.; Gwak, G.Y.; Dapito, D.H.; Mu, X.; Hsu, C.C.; Pradere, J.P.; Friedman, R.A.; Schwabe, R.F. Deactivation of hepatic stellate cells during liver fibrosis resolution in mice. Gastroenterology 2012, 143, 1073-83.e22. [Google Scholar] [CrossRef] [PubMed]
- Pulli, B.; Ali, M.; Iwamoto, Y.; Zeller, M.W.; Schob, S.; Linnoila, J.J.; Chen, J.W. Myeloperoxidase-hepatocyte-stellate cell cross talk promotes hepatocyte injury and fibrosis in experimental nonalcoholic steatohepatitis. Antioxid. Redox Signal. 2015, 23, 1255–1269. [Google Scholar] [CrossRef] [PubMed]
- Henderson, N.C.; Arnold, T.D.; Katamura, Y.; Giacomini, M.M.; Rodriguez, J.D.; McCarty, J.H.; Pellicoro, A.; Raschperger, E.; Betsholtz, C.; Ruminski, P.G.; et al. Targeting of alphav integrin identifies a core molecular pathway that regulates fibrosis in several organs. Nat. Med. 2013, 19, 1617–1624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Legate, K.R.; Wickstrom, S.A.; Fassler, R. Genetic and cell biological analysis of integrin outside-in signaling. Genes Dev. 2009, 23, 397–418. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wu, J.; Song, F.; Tang, J.; Wang, S.J.; Yu, X.L.; Chen, Z.N.; Jiang, J.L. Extracellular membrane-proximal domain of HAb18G/CD147 binds to metal ion-dependent adhesion site (MIDAS) motif of integrin beta1 to modulate malignant properties of hepatoma cells. J. Biol. Chem. 2012, 287, 4759–4772. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.L.; Wu, J.; Wei, D.; Zhang, D.W.; Feng, H.; Chen, Z.N.; Bian, H. Newcastle disease virus represses the activation of human hepatic stellate cells and reverses the development of hepatic fibrosis in mice. Liver Int. 2009, 29, 593–602. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xu, J.; Chen, L.; Zhong, W.D.; Zhang, Z.; Mi, L.; Zhang, Y.; Liao, C.G.; Bian, H.J.; Jiang, J.L.; et al. HAb18G (CD147), a cancer-associated biomarker and its role in cancer detection. Histopathology 2009, 54, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Ru, N.Y.; Zhang, Y.; Li, Y.; Wei, D.; Ren, Z.; Huang, X.F.; Chen, Z.N.; Bian, H. HAb18G/CD147 promotes epithelial-mesenchymal transition through TGF-beta signaling and is transcriptionally regulated by Slug. Oncogene 2011, 30, 4410–4427. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Cui, J.; Yang, X.M.; Liu, Z.Y.; Song, F.; Li, L.; Jiang, J.L.; Chen, Z.N. Cytoplasmic fragment of CD147 generated by regulated intramembrane proteolysis contributes to HCC by promoting autophagy. Cell Death Dis. 2017, 8, e2925. [Google Scholar] [CrossRef] [PubMed]


| Gene | Forward Sequence (5′–3′) | Reverse Sequence (5′–3′) |
|---|---|---|
| Mouse GFAP Cre | ACTCCTTCATAAAGCCCTCG | ATCACTCGTTGCATCGACCG |
| Mouse CD147 Loxp | ATAGAAATGGGGGATGCTCTG | GGCTCTGTCTTCACTTGGGTT |
| Human CXCL1 | ATGGCCCGCGCTGCTCTCTCC | GTTGGATTTGTCACTGTTCAG |
| Human CD147 | ACTCCTCACCTGCTCCTTGA | GCCTCCATGTTCAGGTTCTC |
| Human α1(I) collagen | AACATGACCAAAAACCAAAAGT | CATTGTTTCCTGTGTCTTCTGG |
| Human GAPDH | GCACCGTCAAGGCTGAGAAC | ATGGTGGTGAAGACGCCAGT |
| Mouse CD147 | TGGCAAGTATGTGGTGGTAT | GTGAGATGGTTTCCCGAGT |
| Mouse α-SMA | GTCCCAGACATCAGGGAGTAA | TCGGATACTTCAGCGTCAGGA |
| Mouse desmin | AACAGCCTCGGTTCCTTGAG | GACCTGAGGCTAAACAGGCG |
| Mouse α1(I) collagen | GCTCCTCTTAGGGGCCACT | CCACGTCTCACCATTGGGG |
| Mouse CXCL1 | CACAGGGGCGCCTATCGCCAA | CAAGGCAAGCCTCGCGACCAT |
| Mouse GAPDH | AGGTCGGTGTGAACGGATTTG | TGTAGACCATGTAGTTGAGGTCA |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, W.-P.; Ju, D.; Li, H.; Yuan, L.; Cui, J.; Luo, D.; Chen, Z.-N.; Bian, H. CD147 Promotes CXCL1 Expression and Modulates Liver Fibrogenesis. Int. J. Mol. Sci. 2018, 19, 1145. https://doi.org/10.3390/ijms19041145
Shi W-P, Ju D, Li H, Yuan L, Cui J, Luo D, Chen Z-N, Bian H. CD147 Promotes CXCL1 Expression and Modulates Liver Fibrogenesis. International Journal of Molecular Sciences. 2018; 19(4):1145. https://doi.org/10.3390/ijms19041145
Chicago/Turabian StyleShi, Wen-Pu, Di Ju, Hao Li, Lin Yuan, Jian Cui, Dan Luo, Zhi-Nan Chen, and Huijie Bian. 2018. "CD147 Promotes CXCL1 Expression and Modulates Liver Fibrogenesis" International Journal of Molecular Sciences 19, no. 4: 1145. https://doi.org/10.3390/ijms19041145
APA StyleShi, W.-P., Ju, D., Li, H., Yuan, L., Cui, J., Luo, D., Chen, Z.-N., & Bian, H. (2018). CD147 Promotes CXCL1 Expression and Modulates Liver Fibrogenesis. International Journal of Molecular Sciences, 19(4), 1145. https://doi.org/10.3390/ijms19041145