1. Introduction
Tissues of the musculoskeletal system (bone, cartilage, skeletal muscles, tendons, ligaments) are all comprised of various cell types that exhibit individual differential gene expression patterns. Together with their specific extracellular matrix (ECM), these cells are responsible for the mechanical and functional properties of their respective tissues and organs. Having knowledge of specific genes expressed solely in a given cell type of a particular tissue is not only helpful in identifying and isolating that particular cell type, but more importantly, such genes can be used as markers to track specific lineage specification and differentiation [
1], embryonic development [
2], and ultimately tissue regeneration [
3]. Such individual markers have been identified for musculoskeletal cell types such as osteoblasts, osteoclasts, osteocytes, chondroblasts, chondrocytes, skeletal myoblasts and myocytes, satellite cells, and tenocytes.
As an example, the various stages of osteoblast differentiation can be linked to marker genes to exactly correlate a cellular process, i.e., lineage commitment (i.e., Stro1), proliferation (i.e.,
CD44), maturation (i.e., bone sialoprotein), mineralization (i.e., osteocalcin) cell death (i.e.,
Bax), within a specific cell type such as osteoprogenitor, immature osteoblast, mature osteoblast, and osteocyte [
4]. The same is true for other cells types of the musculoskeletal system. Regardless, to date, there is no single marker that is expressed by all musculoskeletal cell types. Such a pan-musculoskeletal cell marker would complement the existing markers and be useful to study multiple cell types simultaneously.
Mustn1, originally discovered and termed
Mustang (
musculoskeletal
temporally
activated
novel
gene) [
5], represents such a pan-musculoskeletal cell marker.
The
Mustn1 gene which encodes a 9.6 kDa nuclear protein, was discovered during an expression screen for upregulated genes that play a role in the regeneration of a fractured bone [
6]. Bone fracture repair is a complex process that is defined by the interdependent phases of inflammation, angiogenesis, osteogenesis, chondrogenesis, endochondral ossification and remodeling and, thus, it serves as an excellent model of regeneration by enabling the experimenter to isolate individual genes that play a role in one or multiple phases [
7]. Following the screen,
Mustn1 was subsequently cloned and its temporal and spatial expression during bone regeneration (following a transverse fracture) was elucidated [
5]. Specifically, upregulated
Mustn1 expression during fracture repair was localized to multiple cell types within the callus, including periosteal osteoprogenitors, osteoblasts and proliferating chondrocytes. As
Mustn1 represented a novel gene, its expression was also investigated in multiple adult tissues and it was only found at high levels in skeletal muscle and tendon as well as lower amounts in bone and cartilage, making it a probable musculoskeletal specific gene. This review represents a timely attempt to summarize our knowledge of
Mustn1 in the area of phylogeny, genomic organization, promoter analyses, expression, functional perturbation, and disease states. Hopefully, this review may also induce other researchers to include
Mustn1 as a marker gene given the accrued evidence about its relevance in a substantial number of studies.
2. Phylogeny/Genomic Organization
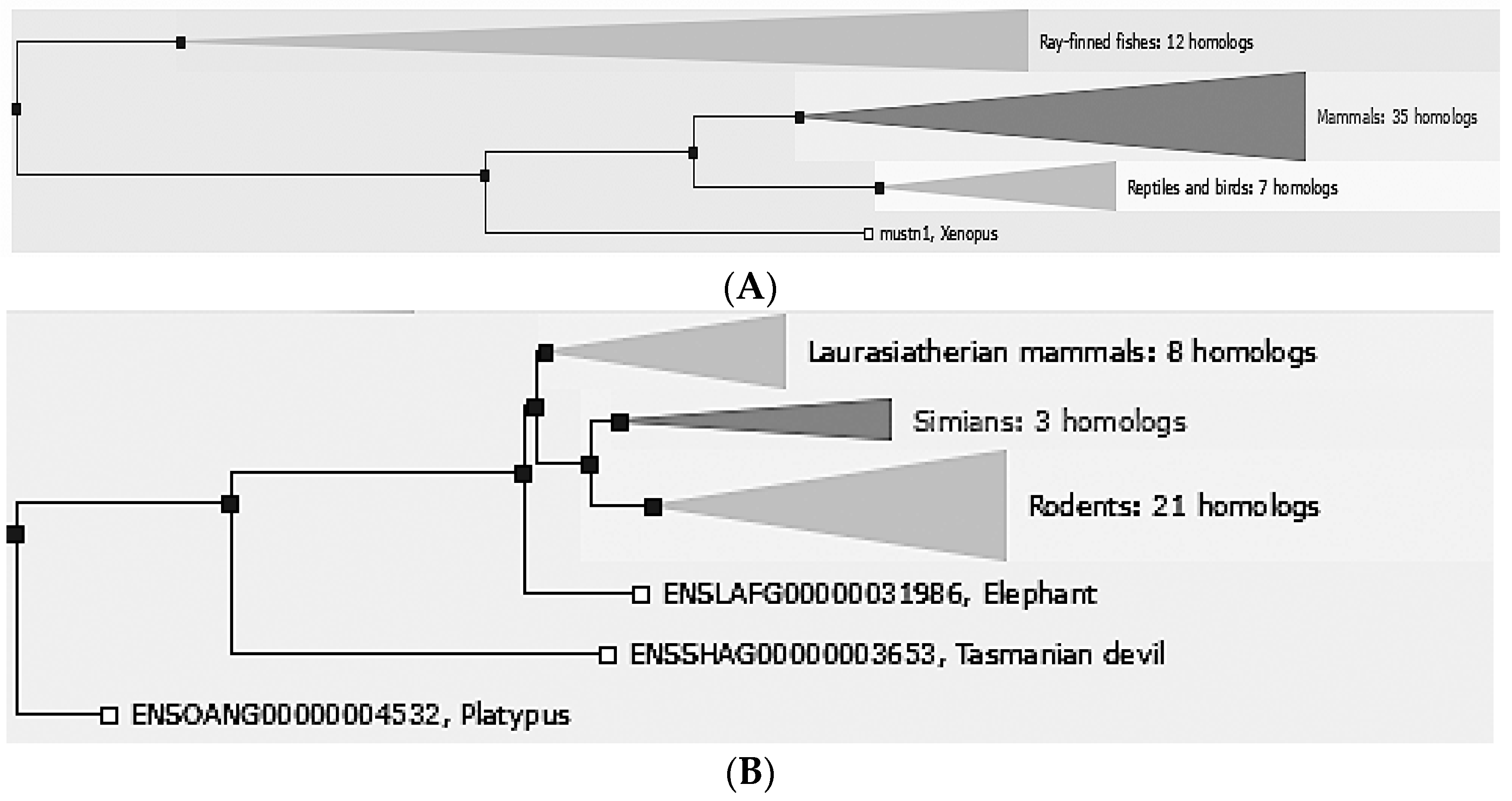
Mustn1 is only found in vertebrate organisms, ranging from fish to mammals. Comparative sequence analyses in e!Ensembl [
8], revealed that there are 55 different vertebrate homologs: 35 mammals, 7 reptiles and birds, 12 Ray-finned fishes. 1 amphibian (
Figure 1A). Further phylogenetic analyses of mammals include closely related laurasiatherian (placental, 8 homologs), simian (apes and monkeys, 3 homologs), and rodents (21 homologs) as well as the more distantly related ones; elephant, Tasmanian devil and platypus (
Figure 1B).
Mustn1 protein sequence homology between eight species that were identified as putative homologs of one another based on the MUSCLE algorithm [
9] and listed in HomoloGene (
https://www.ncbi.nlm.nih.gov/homologene/18744, access date: 12 January 2017) clearly shows their homolgy. For example, the
Homo sapiens protein amino acid sequence is 97.6%, 95.1%, 89.0%, 86.4%, 85.4%, 80.5%, and 70.0% identical to that of
Pan troglodytes,
Bos taurus,
Canis lupus,
Rattus norvegicus,
Mus musculus,
Gallus gallus,
and Xenopus tropicalis, respectively (
Figure 2A). Differences in amino acids between
H. sapiens Mustn1 and those of other species are highlighted in grey (
Figure 2A). In bold letters, the highly conserved nuclear localization signal (NLS) located at residues 10–18 is shown among all the species; the top six species show perfect conservation whereas the lower two show a slight divergence in amino acids, due to the overall lower homology (
Figure 2A).
Han et al. [
10] reported on the porcine
Mustn1 gene which is 78 amino acids and shares 92% and 89% homology with the human and mouse sequence, respectively. More recently, Xu et al. [
11] identified the duck
Mustn1 gene and showed that it is comprised of a 78-amino acid sequence with high similarity with that of other birds (96% with zebra finch and 94% with chicken) and lower with mammals (pig (85%), cow (83%), human (83%), rat (86%) and mouse (81%)). Lastly, we recently reported on the cloning of zebrafish
mustn1. Interestingly, we identified two orthologs,
mustn1a and
mustn1b, that shared 71% homology at the amino acid level and whose predicted proteins were highly related to other vertebrate members (63% to human, 61% to frog and chimp, 60% to dog, chicken and cow, 56% to mouse, and 54% to rat) [
12]. The discovery and comparative sequence analyses of the vertebrate
Mustn1 gene have given rise to a new protein family, labeled “
Mustang” (after the original name we assigned to this gene [
5]) as outlined by UnitPro (
http://www.uniprot.org/uniprot/?query=family:%22MUSTANG+family%22, access date: 12 January 2017).
The genomic organization of
Mustn1 is also conserved within these species. Specifically,
Mustn1 is comprised of 3 exons separated by two internal introns (
Figure 2B) in each of these eight vertebrate species shown in
Figure 2A. Moreover, in both
H. sapiens and
P. troglodytes, the closest of the eight species,
Mustn1 is located on chromosome 3. For the other species, it resides on other chromosomes, again indicating the divergence of this gene in more distantly related species. Interestingly, the genomic organization of the duck
Mustn1 also includes three exons, arranged as those of mammals [
11].
3. Promoter Analyses
The
Mustn1 promoter element was first isolated, cloned, sequenced and characterized in a myoblast cell line in vitro [
13]. The 1512-bp mouse
Mustn1 promoter representing the 5′-flanking region revealed the transcription start site, a TATA box, and multiple putative transcription factor binding sites, particularly AP-1 and AP-2. The activity of this promoter was detected in musculoskeletal cells and exceeded the levels of the control SV40 promoter in C2C12 myoblasts by ~45%. Promoter mutagenesis experiments indicated that one of four AP-1 sites present was required for robust transcriptional activation. The contribution of the AP-2 sites was found to be only marginal in promoter activity. Lastly, we determined that in both proliferating and differentiating C2C12 cells, the immediate early genes,
c-Fos,
Fra-2 and
JunD were required for transcriptional activation [
13].
Next, we utilized this mouse
Mustn1 promoter to drive the expression of Green Fluorescent Protein (GFP) and generated transgenic mice (
Mustn1PR0-GFP) so that GFP expression would serve as a surrogate for
Mustn1 during skeletal muscle development and regeneration [
14]. As we expected based on our previous studies with
Mustn1 expression during development (as described in
Section 4 and
Section 5),
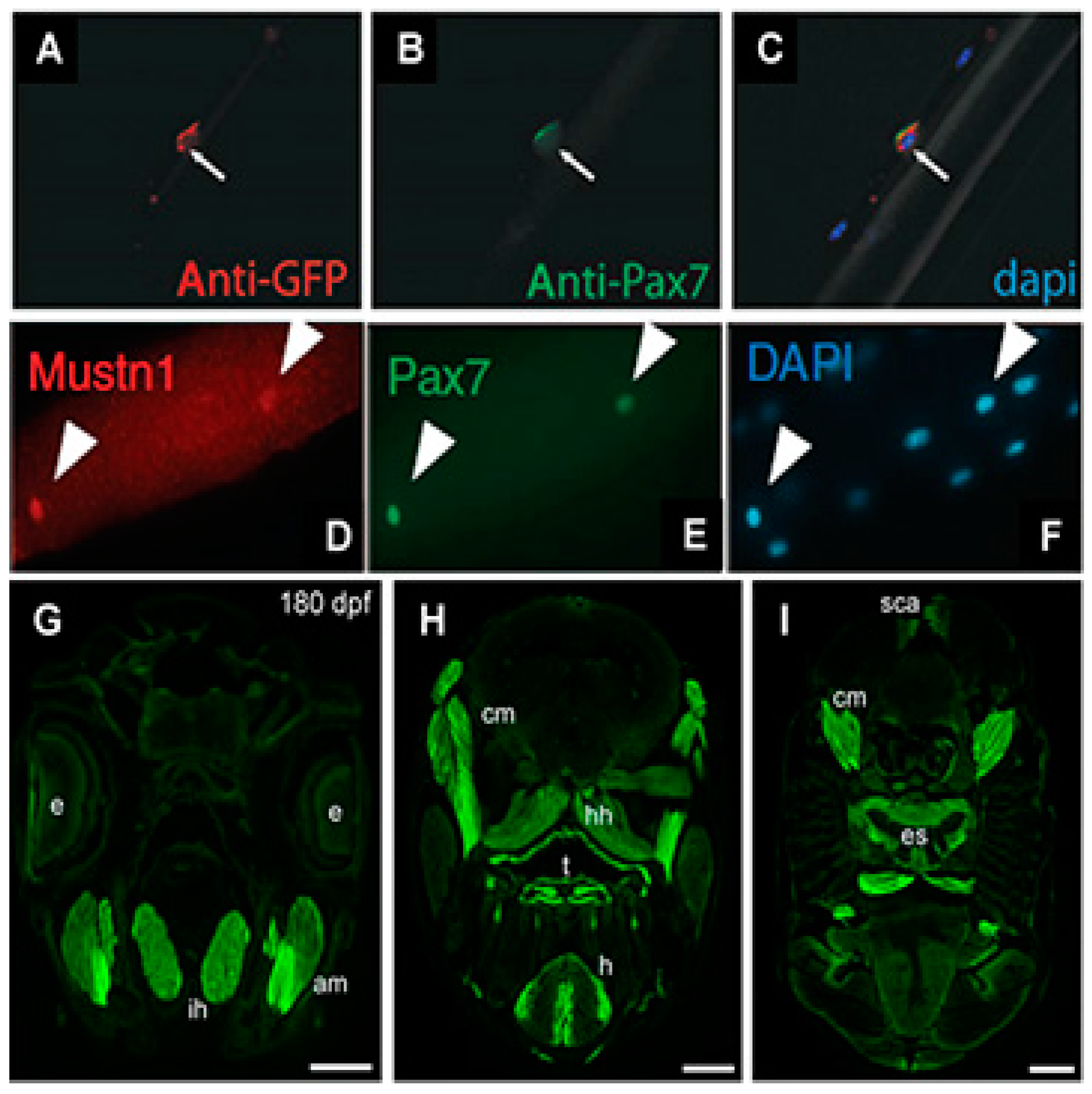
Mustn1PR0-GFP expression was observed within somites at embryonic day 12 and developing skeletal muscles at embryonic day 15 and 18. Cardiotoxin injury increased GFP expression at 3 days post-injury with decreasing levels observed thereafter. Moreover, GFP expression was detected in newly formed myotubes and satellite cells on freshly isolated, single myofibers which co-localized with
Pax7 (satellite cell marker) expression (
Figure 3A–C) [
14]. Collectively, these results indicated the expression GFP, as driven by the
Mustn1 promoter, is robust within both developing and regenerating skeletal muscle as well as satellite cells. Consistent with our data, Zhang et al. [
15] recently identified
Mustn1 as one of a group of novel regulators of satellite cell homeostasis and also showed that its expression co-localizes with
Pax7 in freshly isolated, single skeletal myofibers (
Figure 3D–E).
Very recently, Suarez-Bregua et al. [
16] isolated and characterized the zebrafish
mustn1b promoter by generating transgenic fish with this promoter driving eGFP expression. Specifically, the authors reported stable eGFP expression in a pattern that mirrors that of endogenous
mustn1b gene expression; in skeletal muscle pioneer cells and somites of embryos and in craniofacial and fin muscles of transgenic larvae. Some eGFP expression was also detected in embryonic cardiac muscle. In the adult fish, eGFP expression was detected in jaw (
Figure 3G), cranial muscles, tongue, heart, and esophagus (
Figure 3H,I). Light eGFP expression was also detected in the supracarinalis anterior, lateralis superficialis and hypoaxial muscles of the trunk (
Figure 3I). Lastly, functional analyses of the
mustn1b promoter also revealed that the MyoD binding site was crucial for
mustn1b expression in skeletal muscles [
16].
5. Cartilage Expression Analyses
As previously mentioned, upregulated
Mustn1 expression during bone regeneration was localized to multiple cell types within the fracture callus, including proliferating chondrocytes [
5]. More recently, we also detected
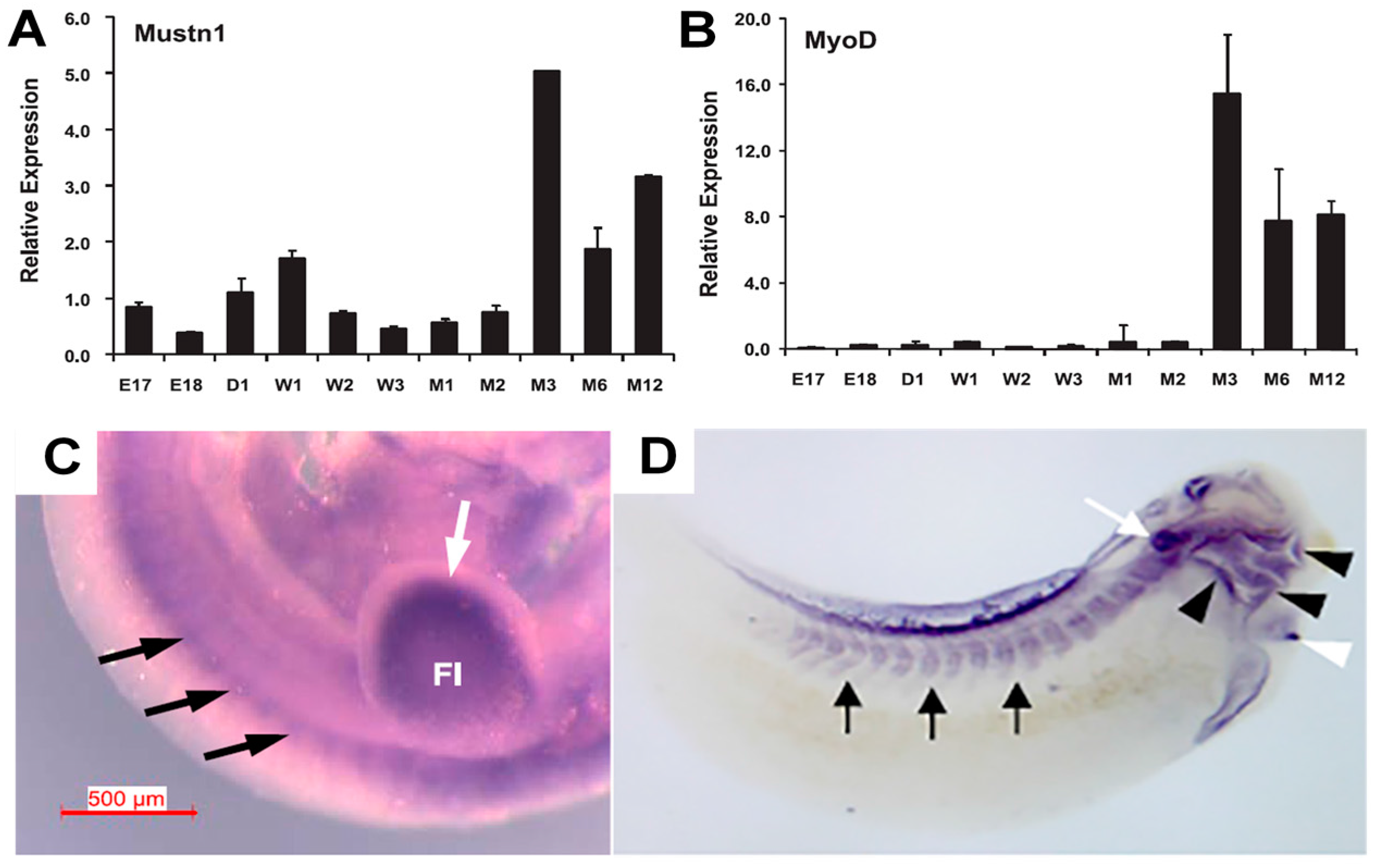
Mustn1 expression in articular cartilage chondrocytes as well as those in the germinal/reserve zone of the growth plate (unpublished observations). During mouse embryonic development,
Mustn1 expression was detected as early as 10.5 days post conception (dpc) in several areas of active cartilage and bone formation [
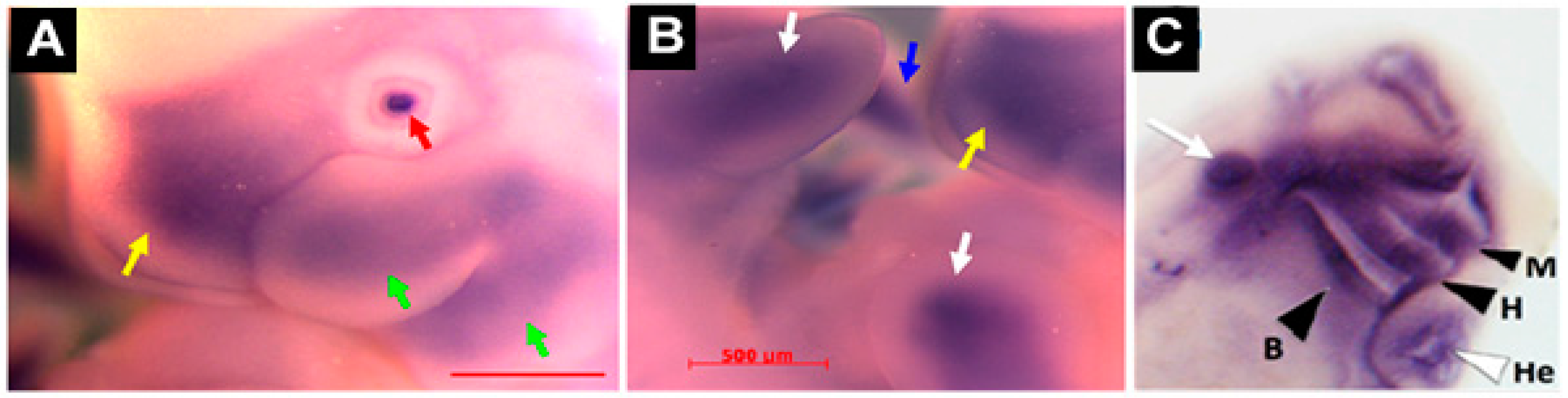
31]. For example, robust expression was present in the craniofacial region, especially the developing first branchial arch that begins to divide into the maxillary and mandibular components (
Figure 5A). In addition, hybridization was also detected in the frontonasal process (
Figure 5A). Similarly, the fore and hind limb buds also displayed robust
Mustn1 expression at 10.5 dpc and as development proceeded to 11.5 dpc, staining was again present along the entire length of both fore and hind limb buds and the posterior tip of the tail (
Figure 5B).
This is consistent with our findings in a rat model, where 16 dpc embryos showed
Mustn1 expression in mesenchymal condensations in developing digits and intervertebral discs, as well as the in the perichondrium of developing vertebral bodies [
5]. Similarly, we also observed intense
Mustn1 expression in the cranial region during
Xenopus development, specifically, at late tadpole stage 35, in anterior structures corresponding to mandibular, hyoid, branchial and other head cartilaginous tissues [
19] (
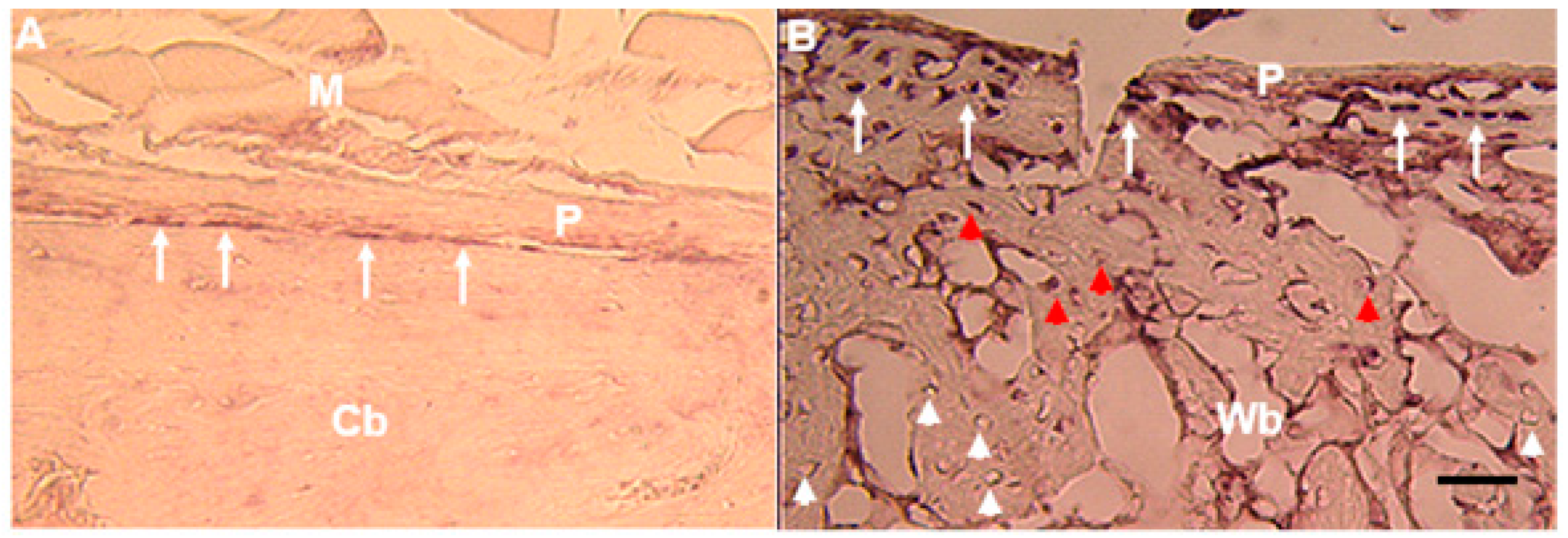
Figure 5C). In Zebrafish,
mustn1b mRNA was primary detected in the pharyngeal arches at 72 h post fertilization (hpf) and by five days, where the pharyngeal arch mesoderm begins to differentiate into the cartilage structures that will eventually form the jaw [
32], its expression was present in the ceratohyal and ceratobranchial elements of the pharyngeal skeleton. This pattern of expression was most likely a progression from the pharyngeal arch expression detected at 72 hpf [
12] consistent with the data using transgenic fish (
mustn1b: eGFP) (
Figure 3) [
16].
An interesting study was performed in rats in order to identify differentially expressed genes in mandibular condylar cartilage during natural growth and under mechanical strain as a result of mandibular advancement [
33]. To induce mandibular advancement, acrylic bite-jumping appliances were fitted to the upper incisors of the experimental rats to produce a continuous 3.5 mm anterior and 3mm inferior displacement of the mandible. The appliances were kept cemented in place and groups of rats were sacrificed on days 1, 3, 7, 9, 14, 21, 30, and 33 and compared to control (no appliances).
Mustn1 was one of five genes that were upregulated in the experimental rats and involved with different stages of chondrogenesis in mandibular condyle growth. Although
Mustn1 expression did not change in the control animals (natural growth), its expression was found at all days tested (1, 7, 9, 14, 30, 33), but was only significantly upregulated at days 7 (~2-fold), 9 (~2-fold), 14 (~3-fold), and 21 (~2-fold) during the advancement of the mandibular condyle in the experimental group, suggesting a role in the activation of mandibular condylar cartilage formation [
33].
More recently, the effect of pulsed and continuous ultrasound (US) exposure on chondrogenesis-related gene expression was tested in rat tibial articular cartilage [
34]; pulsed US is used clinically to accelerate fracture healing [
35,
36]. Three groups of rats were tested, a control group which was treated with sham sonication, a pulsed US group that received a pulse rate of 20%, at a frequency of 1 MHz, and an intensity of 1.5 W/cm
2 for 10 min and a continuous US group that was exposed continuously at a frequency of 1 MHz and an intensity of 1.5 W/cm
2 for 10 min. Each group received a single US treatment exposure. The two genes that were selected to monitor chondrogenesis were
Mustn1 and the classical chondrogenic marker, Sox9, a master regulatory gene encoding for a transcription factor [
37]. Results showed that mRNA expression of both
Mustn1 and Sox9, increased significantly in the continuous (~24% and 37%, respectively) and pulsed groups (44% and 52%, respectively), but the increase of
Mustn1 mRNA in the continuous US group was significantly more prominent than in the pulsed group (22% and 15%, respectively). Based on these results, the authors suggest that US stimulates chondrogenic gene expression in articular cartilage and may potentially serve as a therapeutic modality [
36].
6. Bone and Tendon Expression Analyses
In adult bone,
Mustn1 expression is localized to periosteal osteoprogenitor cells (
Figure 6A) and during fracture repair its expression is detected in osteoblasts as well osteocytes [
5,
38] (
Figure 6B). Similarly,
Mustn1 was also identified by microarray analysis in a fracture repair study utilizing the same transverse femoral fracture model as in our study, as the gene with the greatest fold increase from exposure to alcohol consumption as compared to control (no alcohol) at post-fracture day 3 [
39]. The upregulated expression of
Mustn1 was also verified by Q-PCR (~3-fold) and it coincided with the upregulated expression of Testin (~2-fold), a gene previously identified in the mouse to be expressed at embryonic day 10.5 in the mesenchyme of all branchial arches and also in the frontonasal processes, as well as in mesenchyme of the limb buds [
40]. Incidentally, Testin’s pattern of embryonic expression mirrors that of
Mustn1, as we previously described in the mouse [
31].
A well-designed study examined differential gene expression between pre-osteoblasts, osteoblasts and osteocytes using visual markers of bone lineage cells derived from dual GFP reporter mice. In these mice osteocytes expressed GFP (topaz) directed by the DMP1 promoter, while pre-osteoblasts and osteoblasts expressed GFP (cyan) expression driven by 2.3 kb of the Col1a1 promoter [
41]. Using this innovative approach,
Mustn1 was identified as a gene expressed by both Col2.3cyan+ (osteoblasts), and DMP1topaz+ (preosteocytes and osteocytes), again consistent with our previous observations [
5,
38].
The expression of
Mustn1 was also detected in an experiment where cultured primary osteoblasts, from calvarial and trabecular bone, isolated from
PTHrP+/+ and −/− mice were exposed to 1 g or simulated microgravity for 6 days with or without intermittent (2hr daily) PTHrP1–36 treatment [
42].
Mustn1 expression was upregulated ~2.2-fold in
PTHrP+/+ osteoblasts exposed to simulated microgravity as well as ~3.9-fold in
PTHrP−/− osteoblasts at 1 g. Additionally, it was found that
Mustn1 expression was down-regulated by ~0.6-fold in osteoblasts treated with PTHrP.
Mustn1 was also one of only 24 genes whose expression was common to all three conditions (up-regulated in simulated microgravity and
PTHrP ablation and down-regulated by PTHrP1–36 treatment). Lastly, and more importantly, cluster analysis of genes whose expression was modified by microgravity and similarly affected by
PTHrP ablation placed
Mustn1 in the same cluster as genes involved in bone growth, mineralization and bone morphogenetic protein (BMP) metabolism, suggesting a role of
Mustn1 in these key osteogenic processes [
42].
Although we previously demonstrated that
Mustn1 is expressed in tendon [
5], the only study that provided additional evidence that
Mustn1 is expressed in isolated tenocytes was reported recently from Mueller and colleagues [
43]. In this study, the authors isolated tenocytes cells from tail, Achilles, and hind limb deep digital flexor tendons from three-month-old male rats and cultured these into a monolayer as well as fibrin gels, respectively, for 7–10 days. RNA from both native tendons as well as from the three-dimensional tenocyte cultures were then used for gene expression profiling analyses. Results revealed that
Mustn1, together with other well-known tendon genes such as tenomodulin, elastin, keratocan, and lubricin, were more highly expressed (~3-fold) in native tendon than in monolayer or the three-dimensional tenocyte cultures. The data from all expression studies described in
Section 4,
Section 5 and
Section 6 are summarized in
Table 1.
7. Functional Perturbation and Regulation
The initial functional indication for
Mustn1 was derived from its amino acid sequence that reveled a NLS, thus making it a probable nuclear protein. We subsequently verified its subcellular localization by transfection experiments using a GFP-
Mustn1 fusion protein that labeled only the nuclei green, indicating active translation and nuclear import [
5]. The NLS was also demonstrated to be functional through nuclear localization of zebrafish
mustn1a fused to GFP [
44]. It is of interest to note that in our study as well as that of Cholski et al. [
44], nucleoli and the nuclear envelope were devoid of any staining suggesting that
Mustn1 is not involved in associated “housekeeping” processes (i.e., rRNA synthesis; nuclear import/export and/or structural functions). Although, these experiments suggested a nuclear function for
Mustn1, they did not address its function directly, but we hypothesized that
Mustn1 may function as a co-activator or co-regulator of transcription as part of a larger multi-protein transcription initiation complex [
5,
31].
To directly address the function of
Mustn1 in cells of the musculoskeletal system, myogenic and chondrogenic, we utilized the in vitro and in vivo approach of overexpression and silencing [
17]. Using the C2C12 pre-myoblastic cell line as a model for myogenic differentiation, we showed that silencing of
Mustn1 mRNA via RNA interference (RNAi) had no effect on the proliferation of these cells. In contrast,
Mustn1 silencing significantly impaired myoblast differentiation, preventing myofusion and ultimately myotube formation. Moreover,
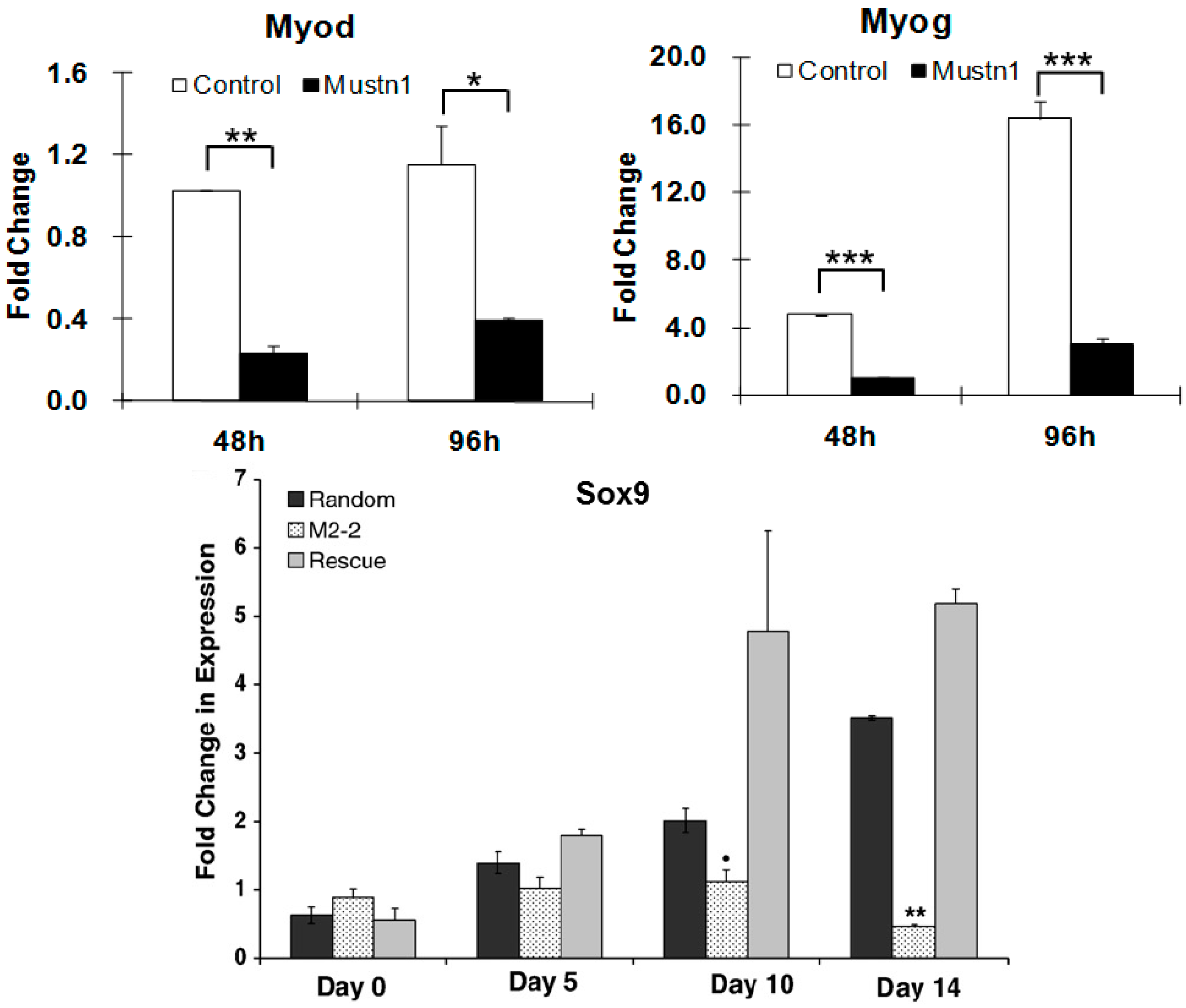
Mustn1-silenced myoblasts elongated poorly and were mono-nucleated as opposed to large, multi-nucleated myotubes present in the control cells, even after 6 days in the presence of myogenic differentiation medium. Additional immunocytochemical analyses of
Mustn1-silenced cells demonstrated significant reductions in both, the amount and timing of expression of the myogenic markers, myogenin (
Myog) and myosin heavy chain (
Myhc) at 4 and 6 days of differentiation. These decreases in
Myog and
Myhc protein expression in
Mustn1-silenced cells were also associated with robust decreases in
Myog,
MyoD (
Figure 7),
Myhc and desmin (
Des) mRNA expression, as well as those of myofusion markers, Calpain1 (
Capn1), Caveolin 3 (
Cav3), and Cadherin 15 (M-cadherin;
Cadh15). Taken together, the data indicates that
Mustn1 is an essential regulator of myogenic differentiation, myofusion and myotube formation [
17]. A more recent study also showed that knockdown of
Mustn1 via RNAi inhibited expansion of mouse primary skeletal muscle stem cells [
15].
Similar experiments were conducted with the pre-chondrocytic RCJ3.1C5.18 (RCJ) cell line [
31]. This cell line represents a heterogeneous cell population capable of differentiation from proliferating chondrocytes to terminally differentiated hypertrophic chondrocytes [
45]. We showed that
Mustn1 overexpression (~2–6-fold) had no statistically significant changes in either proliferation or chondrogenic differentiation. In contrast, both proliferation and differentiation (as assayed by proteoglycan production and cartilage specific gene expression) were significantly reduced in the
Mustn1 silenced cell lines. Specifically,
Mustn1 silencing led to an ~55–75% reduction in cell number. Similarly, an ~34–40% reduction in proteoglycans was observed as compared to parental and random control lines, which was also accompanied by significant downregulation of mRNA levels of the chondrogenic specific marker genes,
Sox9 (
Figure 7), Collagen type II (
Col II), and Collagen type X (
Col X), indicating that
Mustn1 is a necessary regulator of chondrocyte function [
31].
These results were further supported by in vivo experiments in
Xenopus where we utilized antisense morpholinos to downregulate
Mustn1 at the 4-cell stage [
19]. Specifically, the antisense morpholinos were injected into the dorsal and anterior tissues of the developing embryo, including the head and anterior paraxial mesoderm, as well as the anterior neural-ectodermal margin from which the cranial neural crest cells (NCCs) originate. Targeted knockdown of
Mustn1 resulted in phenotypes characterized by small or absent eyes (68% of injected tadpoles), a shortened body axis (49%), and tail kinks (45%) at both stages 37–38 and 40 that corresponded to early swimming/feeding tadpoles. Additionally, when we unilaterally injected the same
Mustn1 antisense morpholinos, we observed the same gross morphological defects on the injected side. More importantly,
Mustn1 knockdown reduced cranial
Sox9 mRNA expression and showed dramatic disruption in ~93% of the examined tadpoles that showed disrupted cartilage formation as detected by Alcian blue staining and in some cases, a complete absence of cartilaginous structures associated with the eye, as well as the ceratohyal cartilage and gill arches [
19] (
Figure 8). Such failure of cranial embryonic cartilage development observed with
Mustn1 downregulation, is consistent with
Xenopus phenotypes resulting from knockdown of other known chondrogenic regulators such as
Sox9 (depletion shows reduction in eye size and anterior Alcian blue-positive cartilaginous matrix, [
46]) and
Runx2 (ablation of cranial cartilage formation, [
47]).
Zebrafish
mustn1a was found as a gene whose expression was induced (~30-fold) by
Foxj1, a winged-helix transcription factor that serves as the master regulator of motile cilia biogenesis, through a systematic effort (microarrays and functional genomics) to discover novel ciliary genes [
44]. In the same study, morpholino knockdown of zebrafish
mustn1a resulted in a curved body axis, similar to what we observed with
Mustn1 knockdown in
Xenopus [
10] as described above. Lastly, zebrafish
mustn1a knockdown also caused defects in otolith and left-right asymmetry, as well as curling of cilia and disorganized γ-tubulin expression, a marker of the basal bodies [
44]. The data from all expression studies described in this section are summarized in
Table 2.
The observation that
Mustn1 knockdown both in vitro and in vivo, resulted in the downregulation of the master regulatory transcription factors,
MyoD,
Myog and
Sox9, suggest that
Mustn1 is involved in early stages during myogenic and chondrogenic differentiation, and at least in vivo, provides the first direct evidence that
Mustn1 is required for formation of embryonic cartilage. This is not surprising given that
Mustn1 expression is robust in these tissues during embryonic development (i.e., somites and limb buds), as previously reported [
5,
17,
19,
31] and outlined in
Section 4 and
Section 5. More importantly, these data also raise the question of whether
Mustn1 is a direct transcriptional regulator of
Myod,
Myog and
Sox9 expression. As
Mustn1 lacks a DNA binding motif, it precludes it from being a direct transcription factor and thus further supports the notion that
Mustn1 probably functions as a musculoskeletal co-activator or co-regulator. Specifically, we speculate that
Mustn1 is part of a transcriptional initiation complex responsible for the activation of these master regulatory genes during both myogenesis and chondrogenesis.
Although these data implicate
Mustn1 in the activation of
Myod,
Myog and
Sox9, they do not indicate which signaling pathway is involved with
Mustn1 activation. However, prior research into several important musculoskeletal signaling pathways implicates them in regulating
Mustn1 transcription. One such study involves Shh, Sonic hedgehog, which is a member of the well-known embryonic Hedgehog signaling pathway [
48]. It is well established that the downstream target genes regulated by morphogens such as Shh and other hedgehog proteins, Desert Hedgehog (Dhh) and Indian Hedgehog (Ihh), are ultimately responsible for processes such as cell proliferation, differentiation and skeletal development [
49]. For example, a murine multipotent mesodermal cell line (C3H10T1/2) was used to elucidate transcriptional targets of Shh. Following overexpression of Shh in C3H10T1/2 and a microarray screen,
Mustn1 was one of 141 genes showing a >1.5-fold increase in expression. Some of the additional genes identified include other transcriptional regulators, as well as those involved in developmental processes, including cellular proliferation and differentiation [
50].
C3H10T1/2 cells were also utilized as a model of osteoblastic differentiation in order to discover transcriptional targets of the Wnt signaling pathway, particularly, Wnt3A [
51]. Activation of signaling pathways by Wnts ultimately leads into a wide array of developmental processes that include cell proliferation, migration, differentiation, establishment of cell polarity, and specification of cell fate [
52]. Results from this experiment identified
Mustn1 as a Wnt3A signaling target gene, with an ~2–3-fold increase in its expression. Interestingly, many of the genes that showed Wnt3A-stimulated expression induction were previously identified as Wnt3A targets, which validates the authors’ experimental approach. Further, a subset of these Wnt3A target genes are already known to play a role in osteoblast function and include
Axin2,
Bmp4,
Cyr61,
Ctgf,
Hes1,
Igfbp2,
Omd,
Tgfb3,
Thbs1,
Twist1, and
Wisp1 [
51]. These results suggest that
Mustn1 expression in osteoblasts may be regulated by Wnt signaling. As such,
Mustn1 (along with other well-established genes) appears to be a target of multiple signaling pathways, including Hedgehog and Wnt, reinforcing the fact that
Mustn1 has a critical role in cellular processes that lead to the development of the various tissues of the musculoskeletal system.
8. Disease States
Beyond understanding the basics aspects of
Mustn1 biology (i.e., cloning, genomic structure, promoter analysis, expression, functional perturbations, etc.), we also want to investigate whether it is associated with any particular disorders or disease states. There are now several studies in the literature that have identified
Mustn1 in various disease states of the musculoskeletal system. A number of human genome-wide association studies (GWAS) searching for risk alleles for osteoarthritis, a musculoskeletal disease characterized by gradual loss of articular cartilage accompanied with physiological alterations in the subchondral bone and the synovium, have been conducted. Reynard and Loughlin [
53] reviewed a number of GWAS studies on osteoarthritis and summarized the data showing a number of potential risk genes for the disease. In addition to highly plausible candidate genes such as
RUNX2 and
CHST11, a transcription factor active in joint development and an enzyme that adds sulfate groups to cartilage proteoglycan, respectively, one of the studies reviewed (arcOGEN, with individuals of European and North American of European descent) also identified
Mustn1 as one of the signals. This led the authors to suggest that the arcOGEN study has provided very novel insights into the etiology of osteoarthritis by the fact that the majority of the genes identified, including
Mustn1, have not previously been suggested to have a role in osteoarthritis [
53]. The plausibility that
Mustn1 is linked to osteoarthritis is not surprising as
Mustn1 is known to be expressed in adult articular cartilage, especially by proliferating chondrocytes in the superficial/tangential zone (unpublished observations).
Aside from cartilage,
Mustn1 expression was also identified in skeletal muscle diseases. Van Lunteren and Moyer [
54], conducted an experiment searching for differentially expressed genes in the diaphragm muscle of streptozotocin-induced diabetic rats using microarrays. Data showed that 105 genes with at least 2-fold significantly changed expression (55 increased and 50 decreased) in the diaphragm of the diabetic rats.
Mustn1 was found to be one of the genes whose expression increased by ~3.2-fold in the diabetic diaphragm and following ontological analyses it was assigned to a group with nine other upregulated genes that are known to be involved in the formation and organization of tissue and organ structure (morphogenesis and organogenesis) [
54].
Another experimental study examined differential gene expression in broiler chickens that suffer from a muscle disorder characterized by palpably “hard” or tough breast muscle (referred to as “Wooden Breast”) [
55]. This myopathy predominantly affects the pectoralis major, and occasionally minor muscles and is associated with multifocal degeneration and necrosis of the muscle tissue with infiltration of inflammatory cells. Results from this study showed that
Mustn1 was upregulated ~4.9-fold in the affected birds and the authors speculate that this indicates compensatory hypertrophy or muscle repair secondary to muscle damage [
55]. Again, not surprising since
Mustn1 is expressed during skeletal muscle hypertrophy [
17,
24] and regeneration [
14].
Kennedy’s disease/Spinobulbar muscular atrophy (KD/SBMA) is a degenerative neuromuscular disease that affects males and is caused by polyglutamine expansion mutations of the androgen receptor (
AR) gene. Halievski and colleagues [
56] used a transgenic mouse model of KD/SBMA because it overexpresses wild-type
AR exclusively in myocytes and has a severe phenotype following acute androgen treatment in females, which reproduces the sex limited (male) and androgen dependent features of the KD/SBMA phenotype [
57]. Thus, treating non-symptomatic females with testosterone induces disease symptoms within 3 days and by 7 days these female mice develop severe symptoms that are typically seen in diseased males. Using microarray analysis of RNA from muscles from both transgenic females (treated for three or seven days with testosterone), it was shown that
Mustn1 expression increased in KD/SBMA muscles of treated female (~11-fold) and affected males (~3-fold), leading the authors to suggest that since
Mustn1 expression is found during skeletal muscle regeneration [
14], hypertrophy [
17,
24], and exercise [
26,
27,
28,
29,
30], its increase in KD/SBMA mice may contribute to their ability to recover following testosterone removal [
56].
Duchenne muscular dystrophy (DMD) is an inherited X-linked lethal muscle wasting disease caused by a mutation in the dystrophin gene that normally encodes for a protein that links the muscle cytoskeleton through a membrane complex to the extracellular matrix. The absence of dystrophin causes various structural and signaling defects in muscle, leading to dystrophic myofibers that are susceptible to damage during mechanical contractions. A recent study with dogs, investigated whether systemic delivery of skeletal muscle-resident stem (MuStem) cells isolated from a 10-week-old healthy dog could serve as a therapeutic modality for the treatment of the Golden Retriever muscular dystrophy (GRMD) dog model, which is characterized by rapid progressive clinical dysfunction and severe muscle tissue remodeling [
58]. These MuStem cells are early myogenic progenitors and uncommitted cells that can be induced to differentiate into myogenic cells. Specifically, the authors compared global gene expression profile in biceps femoris between healthy, GRMD and MuStem cell treated GRMD dogs four months after allogenic MuStem cell transplantation. Results showed that
Mustn1 was one of sixteen genes with significant upregulated expression (~2.5-fold as verified by Q-PCR) in MuStem GRMD dog muscle as compared to untreated control. Some of these 16 genes are also involved in processes such as muscle regeneration, cellular homeostasis, and metabolism. The authors concluded that their results clearly indicate that MuStem cells can positively affect many biological processes, even several months after their transplantation leading to an improvement in the treated GRMD dogs. Moreover, it is the actual gene expression that afforded the treated GRMD dogs the ability to maintain robust muscle fiber regeneration activity that probably led to the stability of the dystrophic muscles [
58].
Clubfoot is a malalignment of the bones and joints of the foot and ankle, and affects 1 in 1000 live births, however, little is known about its genetic or developmental basis. A missense mutation in the Pitx1, a bicoid homeodomain transcription factor, was previously identified in humans with a spectrum of lower extremity abnormalities, including clubfoot [
59]. Because this mutation reduces Pitx1 activity, the authors hypothesized that
Pitx1 haploinsufficiency could also cause clubfoot. Thus,
Pitx1+/− mice were generated and showed a clubfoot-like phenotype associated with deficits in vasculature and bone and muscle volume in the affected limbs [
60]. These observed morphological abnormalities suggested that the clubfoot phenotype results from changes during early embryonic limb development that affect all tissues in the limb. As such,
Pitx−/− mice were generated and skeletal muscle gene expression was analyzed via microarray using E12.5 hindlimb buds and compared to those of wild-type mice. Interestingly,
Mustn1 was one of 19 genes related to muscle development whose expression was downregulated (~2.3-fold) in the E12.5 hindlimb buds from the
Pitx−/− mice indicating that the muscle hypoplasia observed was due to abnormal early skeletal muscle development [
60].
9. Conclusions
It is abundantly clear that since our initial report on the cloning and expression studies of
Mustn1 [
5], a large number of studies implicate its expression with a role predominantly in tissues of the musculoskeletal system, as outlined herein. This was further demonstrated by the
Mustn1-specific functional perturbation studies described in
Section 7. And more recently,
Mustn1 expression has been linked to various disease states related to the musculoskeletal system as described above. Despite the wealth of
Mustn1-related information, there is still much that we do not know about this gene. For example, no one has conducted a general or conditional knockout (the gold standard of determining gene function) of
Mustn1 to show the consequences of its ablation on the developing musculoskeletal system. My laboratory is now in the process of generating a
Mustn1 conditional knockout in cartilage and we are hopeful that we will observe an interesting phenotype given the aforementioned functional knockdown data, both in vitro and in vivo, that clearly showed impairment of chondrogenic differentiation [
31] and cartilage formation in general [
19]. Moreover, generating a
Mustn1 specific knockout in bone, skeletal muscle, and tendon will also be an interesting avenue of research in order to be able to compare the outcome of these studies between all of the major tissues of the musculoskeletal system. In addition to directly determining
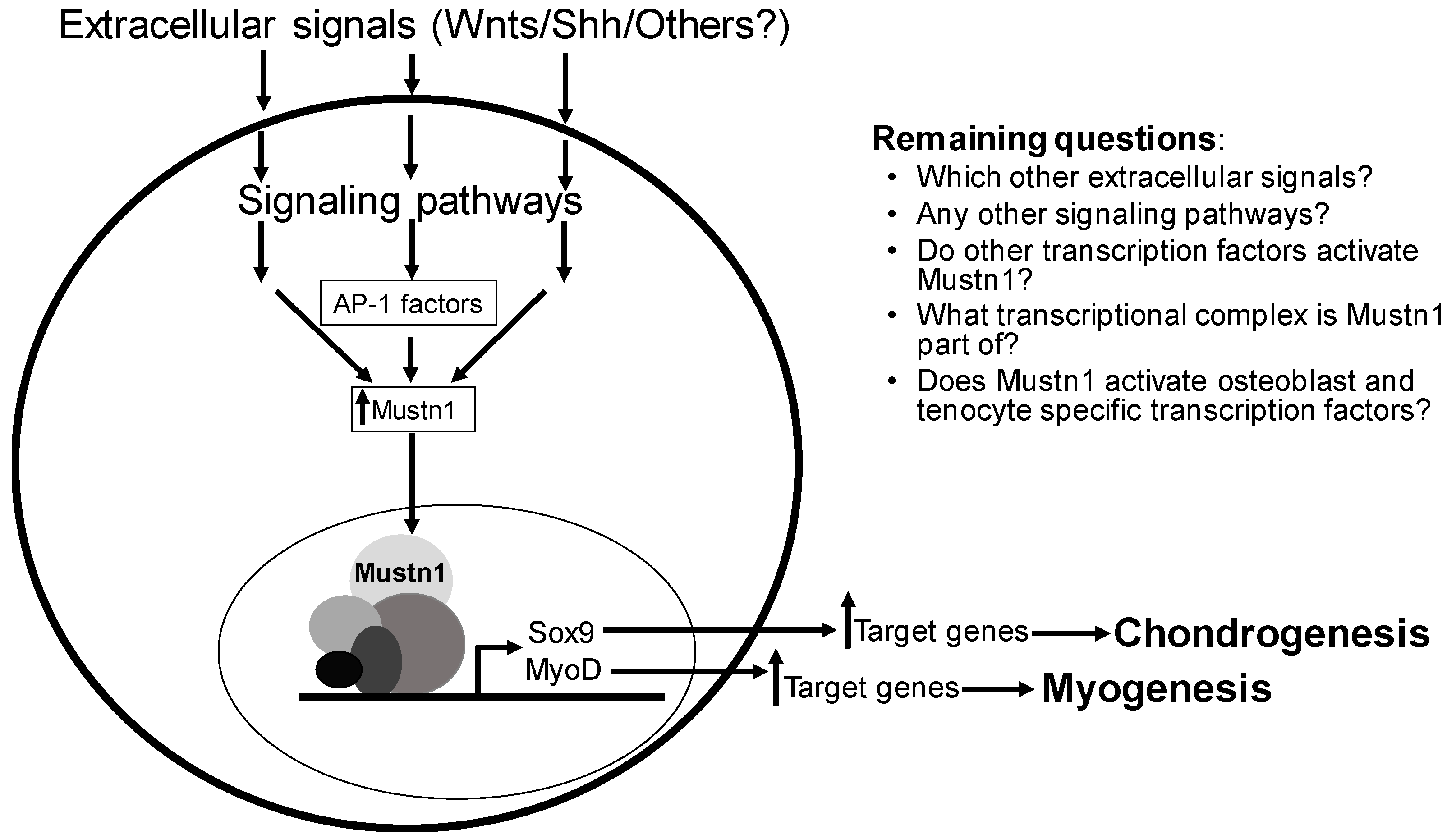
Mustn1 function, we also need to increase our knowledge of how its expression is regulated; by what specific signaling molecules and which signaling pathways (
Figure 9). Along with this, we should also seek to decipher which other transcription factor(s) are responsible for its direct expression.
Lastly, and more importantly, since we hypothesize that
Mustn1 functions as a cofactor of a transcriptional initiation complex, it would be a worthwhile effort to identify its interacting proteins. To this end, a study focusing on LNX1, Ligand of Numb, protein X 1, a RING (Really Interesting New Gene) domain-containing E3 ubiquitin ligase identified
Mustn1 as one of 62 potential interacting proteins [
61]. This search was based on the presence of PDZ (Post-synaptic density, 95 kDa, Discs large, Zona Occludens-1) domains, which are protein interaction domains that bind to the carboxy-terminal amino acids of binding partners. As LNX1 contains four PDZ domains the authors used a human protein array to identify direct LNX1 PDZ domain binding partners and
Mustn1 was one of 21 out of the original 62 proteins that had carboxy terminal tails that conform to PDZ domain binding motifs, though no direct experimental biochemical verification of a physical interaction between
Mustn1 and LNX1 was provided [
61].
Considering all of the current data together,
Mustn1 should be considered, not only as a pan-musculoskeletal cell/tissue marker, but more importantly as a regulatory protein whose expression precedes that of master regulatory genes such as
MyoD and
Myog in skeletal muscle and
Sox9 in cartilage (
Figure 9). Whether
Mustn1 is part of a multi-protein transcriptional complex responsible for activating these regulatory genes remains experimentally unknown. It is also very interesting to determine whether the same is true for critical transcription factors responsible for osteogenesis and tendogenesis; does
Mustn1 expression in osteoblasts precedes that of
Osx and
Runx2 and in tenocytes that of
Scx and
Mkh? Regardless, its prevalence in key cellular processes such as proliferation and differentiation and more complex tissue-based events as embryonic development, organogenesis and regeneration, leads me to believe that
Mustn1 will become an indispensable and critical early regulatory protein for all major cells of the musculoskeletal system. Hopefully, in the near future additional experimental evidence will emerge to support the suggested regulatory-based hypothesis for this important pan-musculoskeletal gene.