1. Introduction
Ankylosing spondylitis (AS), a prototypical chronic immune-mediated inflammatory arthritis disorder of spondyloarthritides (SpA), is a highly familial, heritable disease as shown by strong evidence from genetic association twin and family studies [
1,
2,
3]. Multiple genetic interactions have been implicated in the aetiology of AS disease and present with heterogeneous manifestations [
4,
5,
6]. The genetic transmission and patterns of inheritance in AS are highly complex. Therefore, there is immense interest for identifying the genetic factors in the AS complex disease entity.
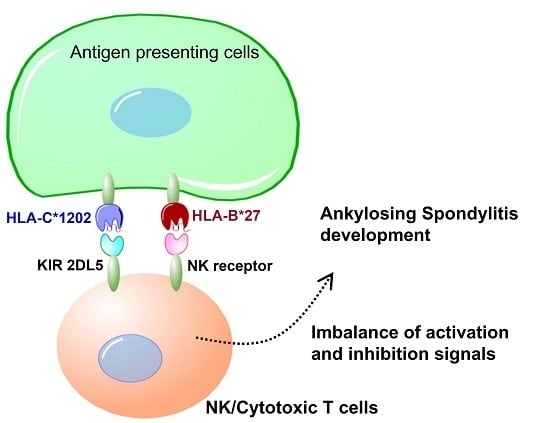
Natural killer (NK) cells dynamically interact via inhibitory surface receptors with human leukocyte antigen (HLA) ligands to acquire a “licensing” process, and express functional competence during development [
7]. Licensed NK cells with self-MHC (Major Histocompatibility Complex) specific receptors are more readily activated and more responsive than unlicensed NK cells without self-MHC-specific receptors [
8]. NK cell activity reflects the balance between inhibitory receptors specific for MHC class I molecules and activating receptors with diverse specificities. The elevated frequency of killer immunoglobulin-like receptor (KIR) 3DL1 (Three Ig Domains and Long cytoplasmic tail 1) expressing NK cells in SpA patients may contribute to a reduction in IFN-γ production [
9]. NK cells interact with fibroblast-like synoviocytes (FLS), leading to proinflammatory responses with increased IL-15 expression by FLS followed by the production of proinflammatory chemokines, cytokines, and matrix metalloproteinases (MMPs) in SpA and rheumatoid arthritis (RA) patients [
10]. However, the precise role of NK cells in the pathogenesis of SpA remains unclear, as conflicting data require further clarification.
KIRs (killer immunoglobulin-like receptors) regulate the cytolytic activity of NK cells and certain T cells through binding to HLA class I ligands [
11]. KIRs comprise critical inhibitory receptors to ensure self-recognition, which dampens NK cell activation upon interaction with cognate MHC class I ligands [
11]. Human NK cells subjected to NK cell licensing highlight the potential functional influence of KIR and HLA genes in disease as well as inter-individual differences in NK cell potency [
12,
13]. Specific HLA Class I molecules may bind and trigger cell surface receptors specified by KIR genes that regulate the physiological functions of NK cells [
13]. Thus, it is important to identify relevant associations between the KIR functional genotype that contributes to AS susceptibility and the development of clinical characteristics.
The
KIR gene clusters show extensive genetic diversity, as do the HLA Class I loci, which encode ligands for KIR molecules. The activating
KIR genes and their corresponding HLA ligand groups show strong negative correlations across populations with distinct evidence for coevolution [
8,
14]. The inhibitory receptor and autologous HLA interactions impact cell function and demonstrate that the resting human NK repertoire is highly attuned but variegated in the immune response [
14]. Increasing evidence from epidemiological studies reveals that particular
KIR and HLA genotypes are associated with certain human disease outcomes, although the functional explanation for these associations is poorly understood. The interaction of HLA-C with KIR plays the dominant role to control human NK cell response [
15]. In addition, HLA-C alleles appear to associate with AS susceptibility [
16,
17,
18]. Herein, we aimed to investigate the association of HLA-C alleles, KIR functional genes and their interaction in the genetic predisposition to disease susceptibility and clinical phenotypes in Taiwanese patients with AS.
3. Discussion
Genetic differences in MHC alleles, especially HLA-B27, confer the greatest risk to AS but are not the only significant genetic factor driving the disease development process. The highly polymorphic KIRs modulate NK and T cell actions against target cells through their interactions with HLA-C ligands. The current study shows that specific
KIRs, HLA Class I genes and their combinations under natural selection result in the diverse characteristics of individuals and populations, and contribute to AS pathogenesis in the Taiwanese population [
19,
20]. As a whole, we observed that in the Taiwanese population, the
KIR AA haplotype, HLA-C1202 allele and HLA-C1 epitope confer risk for AS development, while
KIR2DL5,
KIR Bx and HLA-C2 have a protective effect.
KIRs are involved in the activation/inhibition of NK cells through their interaction with MHC class I, particularly on target cells. KIRs and MHC class I ligands, the HLA-B group (particularly HLA-B27) and possibly other HLA molecules have been associated with differential NK cell activity and function, adding to the growing evidence for the involvement of KIRs in AS disease development [
21]. Previous studies have demonstrated that
KIR3DL1/
KIR3DS1, 2DL5 and
KIR2DS1 are risk factors in the pathogenesis of AS [
21,
22]. A positive association of KIR3DS1 (activating receptor) and a negative association of KIR3DL1 (inhibitory receptor) with AS development have been suggested. Nevertheless, neither the
KIR gene content of particular
KIR haplotypes nor
KIR3DL2 polymorphisms contribute to AS in Caucasian (UK) patients [
23,
24].
KIR3DS1 segregates as an allele with a short cytoplasmic tail and characterizes activation receptors that are expressed by a higher percentage of NK cells in
KIR3DS1 homozygous donors than in heterozygous donors [
25].
KIR3DS1 shows an increased frequency association in combination with HLA-B alleles carrying Bw4-I80 in the trans position in Spanish and Azoreans populations with AS, compared with B27 controls, whereas KIR3DL1 was decreased in AS patients [
26,
27]. KIR3DS1 or KIR3DL1 in combination with the HLA-B27s/HLA-B Bw4-I80 genotypes may modulate the disease development of AS [
28]. The activation of either NK or T cells via the KIR3DS1 receptor may represent a critical event in AS development, while the presence of the functional KIR3DL1 receptor confers a protective effect. In Asians, studies with small sample sizes have demonstrated that the frequency of 3DL1/3DL1 is decreased, while that of 3DL1/3DS1 is increased in Chinese and Thai AS populations [
21,
29]. As a consequence, a general imbalance mediated by protective/inhibitory and risk/activating allotypes from specific
KIR genotypes is responsible for AS susceptibility. With the genetic background of HLA-B27, variation at the KIRs and their corresponding specific HLA-C ligands may contribute to the pathogenesis of AS. Both HLA-C and HLA-B are located at the MHC cluster on Chromosome 6. We found HLA-C1202 allele had strong linkage with HLA-B27 positivity. However, their influence on the ability of NK cells and T cells to recognize and lyse targets in the immune response and the functional effects of KIR polymorphisms remain largely unknown [
26,
27,
28,
29,
30].
KIR gene variations with activating effects might associate with susceptibility to AS by influencing NK cell activity once HLA-C2 epitope ligands are present [
16,
17,
18,
21]. It has been proposed that the orientations or conformations of peptides, even the conformational flexibility of HLA proteins, might affect the association of HLA subtypes with AS [
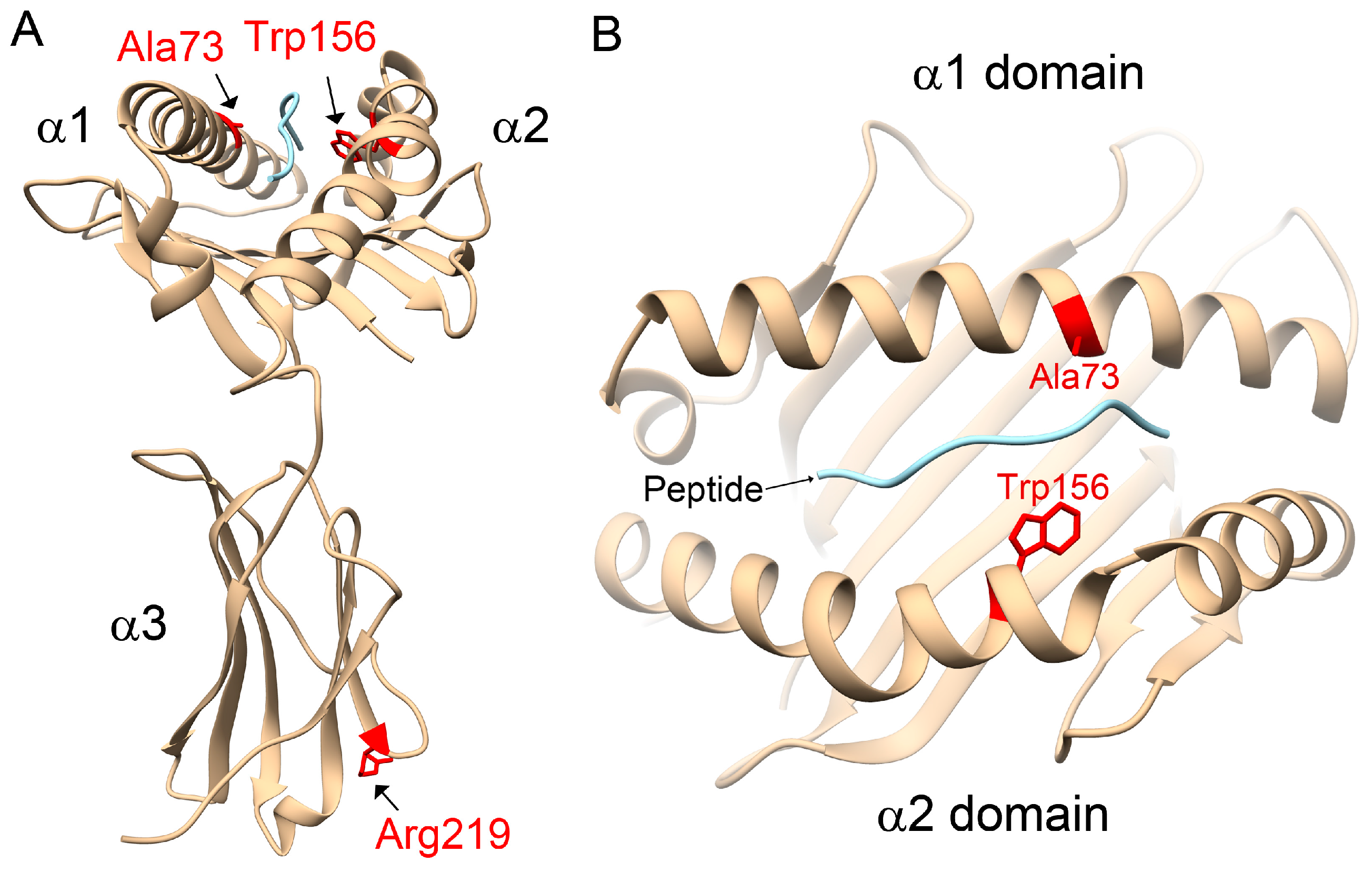
31]. However, in opposition to the literature, our study observed that the HLA-C1 epitope was a risk factor for AS, especially the HLA-C*12:02:02 allele containing the HLA-C1 epitope. Compared with other HLA-C alleles, 3D structural modelling of the highest-frequency allele in AS patients, HLA-C*12:02:02, showed different protein-peptide interactions resulting from the binding affinities, orientations, or conformations during peptide presentation on Ala73 and Trp156. Although the precise underlying mechanisms remain to be determined, we speculate that it may be related to differential NK cell activity.
KIR2DL5 exhibits distinct inhibitory capacities through recruiting both SHP-1 and SHP-2, and its inhibitory capacity is more similar to that of the cytoplasmic domain of KIR2DL4 than that of KIR3DL1 [
32]. Notably, our study comprised a large cohort of samples and paradoxically identified that KIR2DL5 was significantly less prevalent in the Taiwanese AS patient group than the control group (
pfishier = 0.004833).
KIR2DL5 belongs to the
KIR B haplotypes and shows associations with activating receptors related to high viral loads in primary human cytomegalovirus (HCMV) infection following HCMV-positive renal transplant, severe pandemic influenza A (H1N1) and dengue virus infections [
32,
33,
34,
35].
KIR2DL5 was shown to decrease the risk of systemic lupus erythaematosus (SLE), but increased the overall risk for viral infections in Japanese subjects [
36]. However, our study had limitations, such as lacking genotyping for KIR2DL5A and KIR2DL5B (non-allelic expression form) at seven nucleotide positions that differentiate two functional genotypes characterizing KIR2DL5 protein expression [
37,
38].
KIR2DL5A is the predominant genotype in Asian populations, which is consistent with the results from our 282 commercially genotyped individuals [
37].
KIR B haplotypes also showed a protective role for AS in our study, although the combinatory effects of activating KIRs are difficult to explain. Our study suggests that KIR2DL5 may serve as an independent KIR protective marker for Taiwanese AS, although the ligand is unknown [
20].
KIR2DS4 was identified as a risk factor for AS based on a previous meta-analysis [
22]. KIR
2DS4 presents as either a fully functional (KIR2DS4-
f) or deleted non-functional (KIR2DS4/1D) variant with a 22-bp deletion in exon 5, due to a frame shift mutation and premature stop codon yielding a truncated KIR2DS4 protein with loss of the transmembrane and cytoplasmic domains [
39]. Our data revealed that
KIR2DS4 deletion carriers with low KIR2DS4 functional expression were at risk for AS. Additionally, our study showed that
KIR A haplotypes with multiple
KIR inhibitory genes represented risk factors, suggesting that this inhibitory regulatory receptor is critical for AS susceptibility.
These discrepant results indicate the complex diversification of
KIR gene structure and human evolution by genetic selection over long-term environmental stimulation. The remarkable polymorphism of
KIR and HLA genes warrants descriptive gene frequency studies in different populations. The diversity distribution of Asians, especially Taiwanese citizens, demonstrates a similar high frequency (>98.6%) in framework-specific
KIR genes (2DL1, 2DL3, 2DL4, 3DL1, 3DL2) among normal subjects and AS patients, although we did not apply these results to functional analysis. Nevertheless, the present study extends the association and confirms the contribution of the
KIR genes to AS susceptibility, with an imbalance between activating and inhibitory
KIR genes seeming to influence susceptibility to AS [
40].
Combinatorial analyses of KIR genes and their HLA-C ligands may reveal the interaction effects contributing to AS pathogenesis. We observed protective effects for KIR2DL5 with the heterozygous HLA-C1C2 ligand combination genotype in a multivariate analysis of Taiwanese AS development. Our results indicated that HLA-C was beneficial in KIR2DL5 AS carriers possessing at least one C2 allele, while the opposite was not true for homozygous C1 carriers (C1/C1), resulting in risk outcomes. In contrast, the KIR2DS4 deletion and HLA-C1C1 ligand combination was identified as a risk factor for AS pathogenesis. These results suggest that the KIR gene content in combination with the specific ligand may enhance the influence on AS development.
4. Materials and Methods
4.1. Characteristics of the Study Populations
The present study recruited patients who fulfilled the 1984 revised New York diagnostic criteria for AS. Radiographs of the cervical, thoracic and lumbar spine were examined by rheumatology specialists to evaluate syndesmophyte formations according to the modified Stoke’s Ankylosing Spondylitis Spinal Score (mSASSS). To ensure the accuracy of evaluation, two rheumatology specialists independently scored the syndensmophyte formations by blindly reading radiographs of AS patients to establish inter- and intra-reader reliability. The X-ray observations were further classified into three groups: Group 1 patients showed no spine erosion, sclerosis or syndesmophyte formation; Group 2 patients showed <4 fused syndesmophyte formations (mSASSS < 24); and Group 3 patients showed >4 syndesmophyte formations (mSASSS ≥ 24), as described previously [
41]. For the comparisons in this study, 952 healthy blood donors (568 males and 384 females, mean age: 47.98 ± 9.97) were recruited. This study was approved by the ethics committees of Chang Gung Memorial Hospital (Institutional Review Board (IRB) approval number 104-7310B; 26 November 2015). All experiments were performed in accordance with relevant guidelines and regulations. All patients provided written informed consent according to the Declaration of Helsinki.
4.2. HLA-B27 Determination
HLA-B27 antigen positivity was determined by flow cytometry analysis and/or polymerase chain reaction (PCR) assays. Briefly, the whole blood samples were stained with BDTM HLA-B27 kit with fluorescein-conjugated anti-HLA-B27 phycoerythrin-conjugated anti-CD3 antibodies that specifically bind to leucocyte antigen. Samples were then analyzed using a FACSCalibur flow cytometer and HLA-B27 software (Becton Dickinson, San Jose, CA, USA). In PCR assays, two sets of primers were used to amplify HLA-B gene with genomic DNA. The first set of primers detect human HLA-B27/B40/B55 while the second set of primer specifically target HLA-B27. Amplification of human β-globin gene in the same PCR reaction was included as an internal control.
4.3. Nucleic Acid Isolation
Genomic DNA was isolated from Ethylenediaminetetraacetic acid (EDTA) anti-coagulated peripheral blood using the Puregene DNA isolation kit (Gentra Systems, Minneapolis, MN, USA).
4.4. Genotyping of KIRs
The presence and absence of KIR genes (KIR gene profiles) were established using the PCR-SSP KIR genotyping kit (KIR Genotyping SSP Kit). KIR2DL1 (279), 2DL3 (281), 2DL4 (282), 3DL1 (278), 3DL2 (282), and 3DL3 (282) showed positive rates over 98.5%. Thereafter, nine KIR genes (KIR2DL2, 2DL3, 2DL5, 2DS1, 2DS2, 2DS3, 2DS4-full, 2DS4-deleted, 2DS5, and 3DS1) in the remainder of the samples were genotyped using an in-house PCR-SSP primer set, the results of which were consistent with commercial kits applied to the same 282 samples. PCR reactions were prepared at a volume of 10 µL containing 0.08 µL Taq DNA polymerase (5 U/L), 9 µL Ready PCR buffer and 1 µL DNA (40 ng/L). All of the KIR genes were amplified using an ABI 9700 PCR Cycler (Applied Biosystems, Forster City, CA, USA) under the following conditions: initial denaturation at 96 °C for 2 min, followed by 10 cycles of 15 s at 96 °C and 1 min at 65 °C, and 20 cycles of 15 s at 96 °C, 50 s at 61 °C, and 30 s at 72 °C, with a 4 °C hold. The PCR products were electrophoresed on a 2% agarose gel and then visualized under ultraviolet light.
4.5. HLA-C Genotyping and DNA Sequencing
Genotyping of the HLA-C allele was performed using a commercial sequencing-based typing (SBT) kit (Applied Biosystems, Forster City, CA, USA). HLA-C genotypes were assigned by the HLA SBT uTYPE 6.0 software (Applied Biosystems, Forster City, CA, USA).
4.6. Homology Modelling of Protein Structures for HLA-C Alleles
The amino acid sequences, residue numbers and the sequence alignment of HLA-C alleles were searched for on the HLA nomenclature website (
http://hla.alleles.org). The α chain of HLA-C*12:02:02 is a protein of 342 amino acids. Based on a sequence search using Basic Local Alignment Search Tool (BLAST) (
https://blast.ncbi.nlm.nih.gov), the structural information for HLA-Cw3 (Protein Date Bank codes: 1EFX) [
42] was selected as a template for homology modelling of HLA-C alleles, in which the identity between HLA-C*02:02:02 and HLA-Cw was 96% within 277 aligned amino acids (A.A. 2–278). A human self-peptide of sequence RRKWRRWHL derived from vasoactive intestinal peptide receptor type 1 (pVIPR) is a known peptide displayed by AS-associated HLA-B*27 [
31]. This RRKWRRWHL peptide was also modelled in complex with the HLA-C alleles to indicate the peptide-binding site. Subsequently, the 3D structures of human HLA-C alleles with peptides were simulated by Modeller 9.12 [
43], with the python scripts using the functions of the AUTOMODEL class. The Discrete Optimized Protein Energy (DOPE) method [
43] was used to select the best model from the 10 initially generated models. The protein-peptide interactions were analyzed using HotLig [
44]. The molecular models were rendered using Chimera [
45].
4.7. Statistical Analysis
Frequency comparisons of HLA-C alleles, individual
KIR genes,
KIR haplotypes (group A and group B) and KIR/HLA-C pairs between patients and controls were made using logistic regression or the
χ2-test, and Fisher’s exact test was applied when appropriate. Based on the risk allele identified,
p-values, odds ratios (ORs), and 95% confidence intervals (CIs) were then calculated. To account for multiple testing corrections, false discovery rate (FDR)-corrected
p-values were generated using an FDR correction method in the modified version of QVALUE software (
http://genomics.princeton.edu/storeylab/qvalue/). To investigate the genetic association with clinical characteristics, we controlled for each of the clinical characteristics and performed stepwise logistic regression analyses.