The Roles of Long Non-Protein-Coding RNAs in Osteo-Adipogenic Lineage Commitment
Abstract
:1. Introduction
2. Epigenetics and ncRNAs
3. Epigenetics in Osteo-Adipogenesis
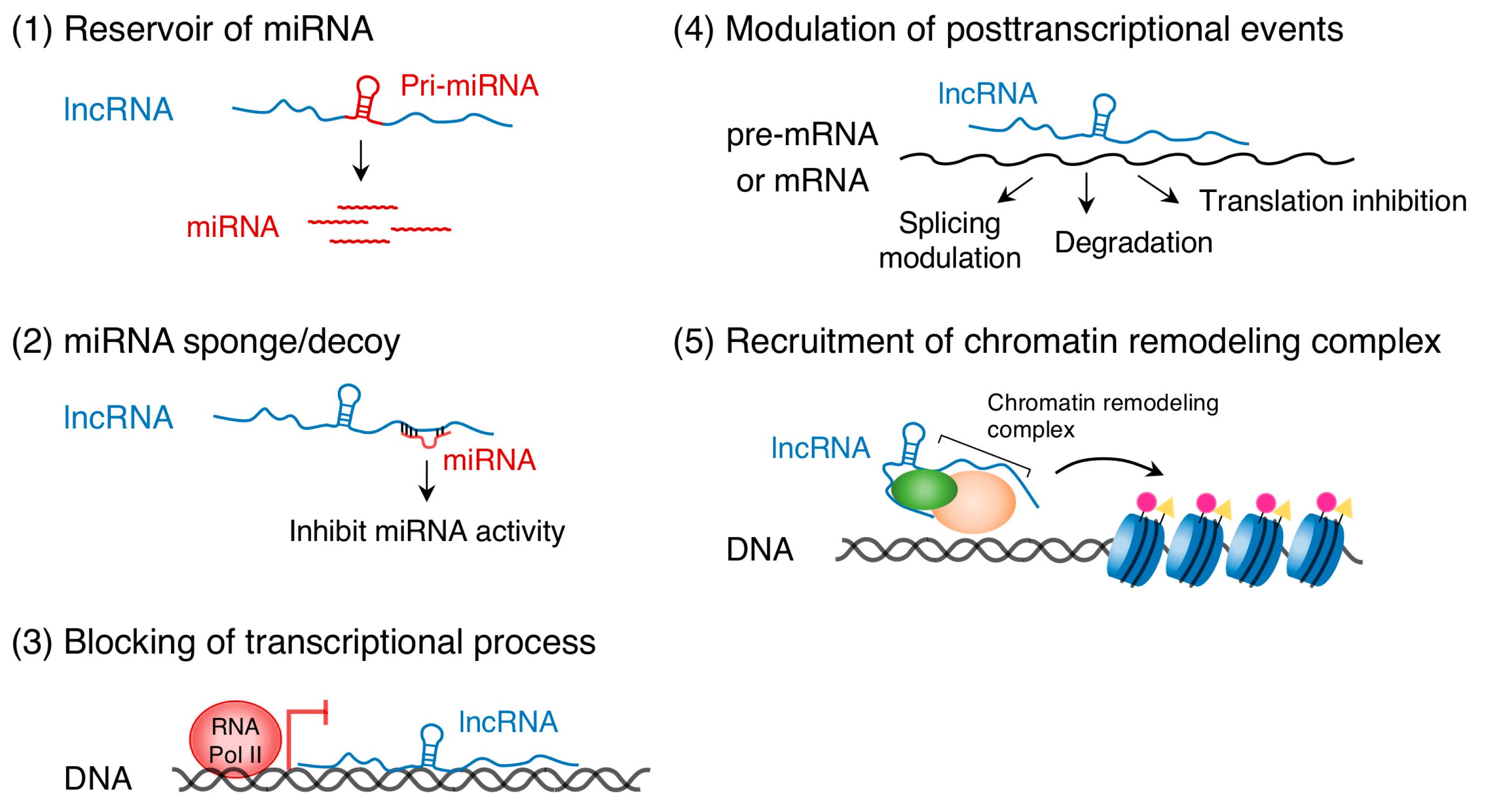
4. Functional Roles of lncRNAs in TGF-β- and WNT-Dependent Osteo-Adipogenesis
5. Epigenetics in Osteo-Adipogenic Transdifferentiation
6. Concluding Remarks and Outlook
Conflicts of Interest
Abbreviations
| lncRNAs | Long non-coding RNAs |
| BMSCs | Bone marrow mesenchymal stem cells |
| MAT | Marrow adipose tissue |
| ncRNAs | Non-protein-coding RNAs |
| miRNAs | MicroRNAs |
| HDACs | Histone deacetylases |
| TGF-β | Transforming growth factor-β |
| MAPK | Mitogen-activated protein kinase |
| ERK | Extracellular signal-regulated kinase |
| H3K4me3 | Trimethylation of histone H3 at lysine 4 |
| H3K36me3 | Trimethylation of histone H3 at lysine 36 |
| MSCs | Mesenchymal stem cells |
| H3K9me3 | Trimethylation of histone H3 at lysine 9 |
| H3K27me3 | Trimethylation of histone H3 at lysine 27 |
| CAR cells | CXC chemokine ligand 12-abundant reticular cells |
References
- Prockop, D.J. Marrow stromal cells as stem cells for nonhematopoietic tissues. Science 1997, 276, 71–74. [Google Scholar] [CrossRef] [PubMed]
- Nombela-Arrieta, C.; Ritz, J.; Silberstein, L.E. The elusive nature and function of mesenchymal stem cells. Nat. Rev. Mol. Cell Biol. 2011, 12, 126–131. [Google Scholar] [CrossRef] [PubMed]
- Scheller, E.L.; Cawthorn, W.P.; Burr, A.A.; Horowitz, M.C.; MacDougald, O.A. Marrow adipose tissue: Trimming the fat. Trends Endocrinol. Metab. 2016, 27, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Georgiou, K.R.; Hui, S.K.; Xian, C.J. Regulatory pathways associated with bone loss and bone marrow adiposity caused by aging, chemotherapy, glucocorticoid therapy and radiotherapy. Am. J. Stem Cells 2012, 1, 205–224. [Google Scholar] [PubMed]
- Scheller, E.L.; Rosen, C.J. What’s the matter with MAT? Marrow adipose tissue, metabolism, and skeletal health. Ann. N. Y. Acad. Sci. 2014, 1311, 14–30. [Google Scholar] [CrossRef] [PubMed]
- Rosen, E.D.; Sarraf, P.; Troy, A.E.; Bradwin, G.; Moore, K.; Milstone, D.S.; Spiegelman, B.M.; Mortensen, R.M. PPARγ is required for the differentiation of adipose tissue in vivo and in vitro. Mol. Cell 1999, 4, 611–617. [Google Scholar] [CrossRef]
- Cock, T.-A.; Back, J.; Elefteriou, F.; Karsenty, G.; Kastner, P.; Chan, S.; Auwerx, J. Enhanced bone formation in lipodystrophic PPARγhyp/hyp mice relocates haematopoiesis to the spleen. EMBO Rep. 2004, 5, 1007–1012. [Google Scholar] [CrossRef] [PubMed]
- Moerman, E.J.; Teng, K.; Lipschitz, D.A.; Lecka-Czernik, B. Aging activates adipogenic and suppresses osteogenic programs in mesenchymal marrow stroma/stem cells: The role of PPAR-γ2 transcription factor and TGF-β/BMP signaling pathways. Aging Cell 2004, 3, 379–389. [Google Scholar] [CrossRef] [PubMed]
- Singh, L.; Brennan, T.A.; Russell, E.; Kim, J.-H.; Chen, Q.; Johnson, F.B.; Pignolo, R.J. Aging alters bone-fat reciprocity by shifting in vivo mesenchymal precursor cell fate towards an adipogenic lineage. Bone 2016, 85, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Kaikkonen, M.; Lam, M.; Glass, C. Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc. Res. 2011, 90, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Peschansky, V.J.; Wahlestedt, C. Non-coding RNAs as direct and indirect modulators of epigenetic regulation. Epigenetics 2013, 9, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Panning, B.; Jaenisch, R. RNA and the epigenetic regulation of X chromosome inactivation. Cell 1998, 93, 305–308. [Google Scholar] [CrossRef]
- Mohn, F.; Schübeler, D. Genetics and epigenetics: Stability and plasticity during cellular differentiation. Trends Genet. 2009, 25, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Smith, Z.D.; Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Zentner, G.E.; Henikoff, S. Regulation of nucleosome dynamics by histone modifications. Nat. Struct. Mol. Biol. 2013, 20, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Cedar, H.; Bergman, Y. Linking DNA methylation and histone modification: Patterns and paradigms. Nat. Rev. Genet. 2009, 10, 295–304. [Google Scholar] [CrossRef] [PubMed]
- Ivey, K.; Srivastava, D. MicroRNAs as regulators of differentiation and cell fate decisions. Cell Stem Cell 2010, 7, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Taipaleenmäki, H.; Hokland, L.; Chen, L.; Kauppinen, S.; Kassem, M. Micro-RNAs: Targets for enhancing osteoblast differentiation and bone formation. Eur. J. Endocrinol. 2012, 166, 359–371. [Google Scholar] [CrossRef] [PubMed]
- Ha, M.; Kim, N.V. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Hamam, D.; Ali, D.; Kassem, M.; Aldahmash, A.; Alajez, N. MicroRNAs as regulators of adipogenic differentiation of mesenchymal stem cells. Stem Cells Dev. 2015, 24, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Liang, H.; Zhang, C.-Y.; Zen, K. miRNA regulates noncoding RNA: A noncanonical function model. Trends Biochem. Sci. 2012, 37, 457–459. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Witmer, P.D.; Casey, E.; Valle, D.; Sukumar, S. DNA methylation regulates microRNA expression. Cancer Biol. Ther. 2007, 6, 1290–1294. [Google Scholar] [CrossRef]
- Sinkkonen, L.; Hugenschmidt, T.; Berninger, P.; Gaidatzis, D.; Mohn, F.; Artus-Revel, C.; Zavolan, M.; Svoboda, P.; Filipowicz, W. MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 2008, 15, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zheng, Y.; Jia, L.; Li, W. Long noncoding RNA H19 promotes osteoblast differentiation via TGF-β1/Smad3/HDAC signaling pathway by deriving miR-675. Stem Cells 2015, 33, 3481–3492. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zheng, Y.; Jin, C.; Li, X.; Jia, L.; Li, W. Long non-coding RNA H19 inhibits adipocyte differentiation of bone marrow mesenchymal stem cells through epigenetic modulation of histone deacetylases. Sci. Rep. 2016, 6, 28897. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.-C.; Fu, W.-M.; Wang, Y.-B.; Sun, Y.-X.; Xu, L.-L.; Wong, C.-W.; Chan, K.-M.; Li, G.; Waye, M.M.; Zhang, J.-F. H19 activates Wnt signaling and promotes osteoblast differentiation by functioning as a competing endogenous RNA. Sci. Rep. 2016, 6, 20121. [Google Scholar] [CrossRef] [PubMed]
- Martianov, I.; Ramadass, A.; Barros, A.; Chow, N.; Akoulitchev, A. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature 2007, 445, 666–670. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.-H.; Abdelmohsen, K.; Gorospe, M. Posttranscriptional gene regulation by long noncoding RNA. J. Mol. Biol. 2013, 425, 3723–3730. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.; Mondal, T.; Mohammad, F.; Enroth, S.; Redrup, L.; Komorowski, J.; Nagano, T.; Mancini-DiNardo, D.; Kanduri, C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 2008, 32, 232–246. [Google Scholar] [CrossRef] [PubMed]
- Mondal, T.; Subhash, S.; Vaid, R.; Enroth, S.; Uday, S.; Reinius, B.; Mitra, S.; Mohammed, A.; James, A.; Hoberg, E.; et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA-DNA triplex structures. Nat. Commun. 2015, 6, 7743. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Chang, H. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Tay, Y.; Rinn, J.; Pandolfi, P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014, 505, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.L. Linking long noncoding RNA localization and function. Trends Biochem. Sci. 2016, 41, 761–772. [Google Scholar] [CrossRef] [PubMed]
- James, A. Review of signaling pathways governing MSC osteogenic and adipogenic differentiation. Scientifica 2013, 2013, 684736. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, A.D.; Olsen, B.R. Osteoblast-adipocyte lineage plasticity in tissue development, maintenance and pathology. Cell. Mol. Life Sci. 2014, 71, 493–497. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Shou, P.; Zheng, C.; Jiang, M.; Cao, G.; Yang, Q.; Cao, J.; Xie, N.; Velletri, T.; Zhang, X.; et al. Fate decision of mesenchymal stem cells: Adipocytes or osteoblasts? Cell Death Differ. 2016, 23, 1128–1139. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Chen, G.; Li, Y.-P. TGF-β and BMP signaling in osteoblast, skeletal development, and bone formation, homeostasis and disease. Bone Res. 2016, 4, 16009. [Google Scholar] [CrossRef] [PubMed]
- Komori, T.; Yagi, H.; Nomura, S.; Yamaguchi, A.; Sasaki, K.; Deguchi, K.; Shimizu, Y.; Bronson, R.T.; Gao, Y.-H.; Inada, M.; et al. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 1997, 89, 755–764. [Google Scholar] [CrossRef]
- Kang, J.; Alliston, T.; Delston, R.; Derynck, R. Repression of Runx2 function by TGF-β through recruitment of class II histone deacetylases by Smad3. EMBO J. 2005, 24, 2543–2555. [Google Scholar] [CrossRef] [PubMed]
- Ge, C.; Yang, Q.; Zhao, G.; Yu, H.; Kirkwood, K.; Franceschi, R. Interactions between extracellular signal-regulated kinase 1/2 and P38 Map kinase pathways in the control of RUNX2 phosphorylation and transcriptional activity. J. Bone Miner. Res. 2012, 27, 538–551. [Google Scholar] [CrossRef] [PubMed]
- Rahimi, N.; Tremblay, E.; McAdam, L.; Roberts, A.; Elliott, B. Autocrine secretion of TGF-β1 and TGF-β2 by pre-adipocytes and adipocytes: A potent negative regulator of adipocyte differentiation and proliferation of mammary carcinoma cells. Vitro Cell. Dev. Biol. 1998, 34, 412–420. [Google Scholar] [CrossRef]
- Choy, L.; Skillington, J.; Derynck, R. Roles of autocrine TGF-β receptor and Smad signaling in adipocyte differentiation. J. Cell Biol. 2000, 149, 667–682. [Google Scholar] [CrossRef] [PubMed]
- Choy, L.; Derynck, R. Transforming growth factor-β inhibits adipocyte differentiation by Smad3 interacting with CCAAT/enhancer-binding protein (C/EBP) and repressing C/EBP transactivation function. J. Biol. Chem. 2003, 278, 9609–9619. [Google Scholar] [CrossRef] [PubMed]
- Clouthier, D.E.; Comerford, S.A.; Hammer, R.E. Hepatic fibrosis, glomerulosclerosis, and a lipodystrophy-like syndrome in PEPCK-TGF-β1 transgenic mice. J. Clin. Investig. 1997, 100, 2697–2713. [Google Scholar] [CrossRef] [PubMed]
- Hu, E.; Kim, J.B.; Sarraf, P.; Spiegelman, B.M. Inhibition of adipogenesis through MAP kinase–mediated phosphorylation of PPARγ. Science 1999, 274, 2100–2103. [Google Scholar] [CrossRef]
- Beekum, O.; Fleskens, V.; Kalkhoven, E. Posttranslational modifications of PPAR-γ: Fine-tuning the metabolic master regulator. Obesity 2009, 17, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Ge, C.; Cawthorn, W.P.; Li, Y.; Zhao, G.; MacDougald, O.A.; Franceschi, R.T. Reciprocal control of osteogenic and adipogenic differentiation by ERK/MAP kinase phosphorylation of Runx2 and PPARγ transcription factors. J. Cell. Physiol. 2016, 231, 587–596. [Google Scholar] [CrossRef] [PubMed]
- Bennett, C.N.; Ross, S.E.; Longo, K.A.; Bajnok, L.; Hemati, N.; Johnson, K.W.; Harrison, S.D.; MacDougald, O.A. Regulation of Wnt signaling during adipogenesis. J. Biol. Chem. 2002, 277, 30998–31004. [Google Scholar] [CrossRef] [PubMed]
- Gaur, T.; Lengner, C.J.; Hovhannisyan, H.; Bhat, R.A.; Bodine, P.V.N.; Komm, B.S.; Javed, A.; van Wijnen, A.J.; Stein, J.L.; Stein, G.S.; et al. Canonical WNT signaling promotes osteogenesis by directly stimulating Runx2 gene expression. J. Biol. Chem. 2005, 280, 33132–33140. [Google Scholar] [CrossRef] [PubMed]
- Kennell, J.A.; MacDougald, O.A. Wnt signaling inhibits adipogenesis through β-catenin-dependent and -independent mechanisms. J. Biol. Chem. 2005, 280, 24004–24010. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Bennett, C.N.; Gerin, I.; Rapp, L.A.; Hankenson, K.D.; MacDougald, O.A. Wnt signaling stimulates osteoblastogenesis of mesenchymal precursors by suppressing CCAAT/enhancer-binding protein α and peroxisome proliferator-activated receptor γ. J. Biol. Chem. 2007, 282, 14515–14524. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Gao, Y.; Zhang, Z.; Cao, Q.; Zhang, X.; Zou, J.; Cao, Y. Kdm2a/b lysine demethylases regulate canonical Wnt signaling by modulating the stability of nuclear β-catenin. Dev. Cell 2015, 33, 660–674. [Google Scholar] [CrossRef] [PubMed]
- Fan, Z.; Yamaza, T.; Lee, J.; Yu, J.; Wang, S.; Fan, G.; Shi, S.; Wang, C.-Y. BCOR regulates mesenchymal stem cell function by epigenetic mechanisms. Nat. Cell Biol. 2009, 11, 1002–1009. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Ma, Y.; Ma, P.; Wang, S.; Fan, Z. Demethylation of epiregulin gene by histone demethylase FBXL11 and BCL6 corepressor inhibits osteo/dentinogenic differentiation. Stem Cells 2013, 31, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Kanazawa, A.; Tsukada, S.; Kamiyama, M.; Yanagimoto, T.; Nakajima, M.; Maeda, S. Wnt5b partially inhibits canonical Wnt/beta-catenin signaling pathway and promotes adipogenesis in 3T3-L1 preadipocytes. Biochem. Biophys. Res. Commun. 2005, 330, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Nishizuka, M.; Koyanagi, A.; Osada, S.; Imagawa, M. Wnt4 and Wnt5a promote adipocyte differentiation. FEBS Lett. 2008, 582, 3201–3205. [Google Scholar] [CrossRef] [PubMed]
- Van Tienen, F.H.J.; Laeremans, H.; Kallen, C.J.H.; Smeets, H.J.M. Wnt5b stimulates adipogenesis by activating PPARγ, and inhibiting the β-catenin dependent Wnt signaling pathway together with Wnt5a. Biochem. Biophys. Res. Commun. 2009, 387, 207–211. [Google Scholar] [CrossRef] [PubMed]
- Inui, M.; Martello, G.; Piccolo, S. MicroRNA control of signal transduction. Nat. Rev. Mol. Cell Biol. 2010, 11, 252–263. [Google Scholar] [CrossRef] [PubMed]
- Bhushan, R.; Grünhagen, J.; Becker, J.; Robinson, P.; Ott, C.-E.; Knaus, P. miR-181a promotes osteoblastic differentiation through repression of TGF-β signaling molecules. Int. J. Biochem. Cell Biol. 2013, 45, 696–705. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Cai, J.; Cai, X.; Chen, L. MiR-346 regulates osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by targeting the Wnt/β-catenin pathway. PLoS ONE 2013, 8, e72266. [Google Scholar] [CrossRef] [PubMed]
- Jeong, B.-C.; Kim, T.; Kim, H.; Lee, S.-H.; Choi, Y. Transmembrane protein 64 reciprocally regulates osteoblast and adipocyte differentiation by modulating Wnt/β-catenin signaling. Bone 2015, 78, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Chen, Y.; Guo, L.; Jiang, T.; Lin, Z. miR-23a/b regulates the balance between osteoblast and adipocyte differentiation in bone marrow mesenchymal stem cells. Bone Res 2016, 4, 16022. [Google Scholar] [CrossRef] [PubMed]
- Lecka-Czernik, B.; Gubrij, I.; Moerman, E.J.; Kajkenova, O.; Lipschitz, D.A.; Manolagas, S.C.; Jilka, R.L. Inhibition of Osf2/Cbfa1 expression and terminal osteoblast differentiation by PPARγ2. J. Cell. Biochem. 1999, 74, 357–371. [Google Scholar] [CrossRef]
- Jeon, M.J.; Kim, J.A.; Kwon, S.H.; Kim, S.W.; Park, K.S.; Park, S.W.; Kim, S.Y.; Shin, C.S. Activation of peroxisome proliferator-activated receptor-γ inhibits the Runx2-mediated transcription of osteocalcin in osteoblasts. J. Biol. Chem. 2003, 278, 23270–23277. [Google Scholar] [CrossRef] [PubMed]
- Akune, T.; Ohba, S.; Kamekura, S.; Yamaguchi, M.; Chung, U.; Kubota, N.; Terauchi, Y.; Harada, Y.; Azuma, Y.; Nakamura, K.; et al. PPARγ insufficiency enhances osteogenesis through osteoblast formation from bone marrow progenitors. J. Clin. Investig. 2004, 113, 846–855. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, M.; Lin, L.; Chen, P.; Ma, K.; Zhou, C.; Ao, Y. Runx2 overexpression enhances osteoblastic differentiation and mineralization in adipose-derived stem cells in vitro and in vivo. Calc. Tissue Int. 2006, 79, 169–178. [Google Scholar] [CrossRef] [PubMed]
- McGee-Lawrence, M.; Westendorf, J. Histone deacetylases in skeletal development and bone mass maintenance. Gene 2011, 474, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Jeon, E.-J.; Lee, K.-Y.; Choi, N.-S.; Lee, M.-H.; Kim, H.-N.; Jin, Y.-H.; Ryoo, H.-M.; Choi, J.-Y.; Yoshida, M.; Nishino, N.; et al. Bone morphogenetic protein-2 stimulates Runx2 acetylation. J. Biol. Chem. 2006, 281, 16502–16511. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, T.; Kahler, R.; Li, X.; Westendorf, J. Histone deacetylase 3 interacts with Runx2 to repress the osteocalcin promoter and regulate osteoblast differentiation. J. Biol. Chem. 2004, 279, 41998–42007. [Google Scholar] [CrossRef] [PubMed]
- Jensen, E.; Schroeder, T.; Bailey, J.; Gopalakrishnan, R.; Westendorf, J. Histone deacetylase 7 associates with Runx2 and represses its activity during osteoblast maturation in a deacetylation-independent manner. J. Bone Miner. Res. 2008, 23, 361–372. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Zhang, P.; Ge, J.; Cheng, J.; Dong, W.; Yuan, H.; Du, Y.; Yang, M.; Sun, R.; Jiang, H. Histone deacetylase 8 suppresses osteogenic differentiation of bone marrow stromal cells by inhibiting histone H3K9 acetylation and RUNX2 activity. Int. J. Biochem. Cell Biol. 2014, 54, 68–77. [Google Scholar] [CrossRef] [PubMed]
- Razidlo, D.; Whitney, T.; Casper, M.; McGee-Lawrence, M.; Stensgard, B.; Li, X.; Secreto, F.; Knutson, S.; Hiebert, S.; Westendorf, J. Histone deacetylase 3 depletion in osteo/chondroprogenitor cells decreases bone density and increases marrow fat. PLoS ONE 2010, 5, e11492. [Google Scholar] [CrossRef] [PubMed]
- Haberland, M.; Mokalled, M.; Montgomery, R.; Olson, E. Epigenetic control of skull morphogenesis by histone deacetylase 8. Genes Dev. 2009, 23, 1625–1630. [Google Scholar] [CrossRef] [PubMed]
- Yoo, E.; Chung, J.-J.; Choe, S.; Kim, K.; Kim, J. Down-regulation of histone deacetylases stimulates adipocyte differentiation. J. Biol. Chem. 2006, 281, 6608–6615. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, T.; Idelman, G.; Blanco, V.; Blomkalns, A.; Piegore, M.; Weintraub, D.; Kumar, S.; Rajsheker, S.; Manka, D.; Rudich, S.; et al. Histone deacetylase 9 is a negative regulator of adipogenic differentiation. J. Biol. Chem. 2011, 286, 27836–27847. [Google Scholar] [CrossRef] [PubMed]
- Ye, L.; Fan, Z.; Yu, B.; Chang, J.; Hezaimi, K.; Zhou, X.; Park, N.-H.; Wang, C.-Y. Histone demethylases KDM4B and KDM6B promotes osteogenic differentiation of human MSCs. Cell Stem Cell 2012, 11, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Okamura, H.; Nakashima, Y.; Haneji, T. Histone demethylase Jmjd3 regulates osteoblast differentiation via transcription factors Runx2 and Osterix. J. Biol. Chem. 2013, 288, 33530–33541. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Guo, F.; Wang, G.; Wang, J.; Zheng, F.; Guan, X.; Chang, A.; Zhang, X.; Dai, C.; Li, S.; et al. MiR-20a regulates adipocyte differentiation by targeting lysine-specific demethylase 6b and transforming growth factor-β signaling. Int. J. Obes. 2015, 39, 1282–1291. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Jin, Q.; Lee, J.-E.; Su, I.; Ge, K. Histone H3K27 methyltransferase Ezh2 represses Wnt genes to facilitate adipogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 7317–7322. [Google Scholar] [CrossRef] [PubMed]
- Dudakovic, A.; Camilleri, E.; Xu, F.; Riester, S.; McGee-Lawrence, M.; Bradley, E.; Paradise, C.; Lewallen, E.; Thaler, R.; Deyle, D.; et al. Epigenetic control of skeletal development by the histone methyltransferase Ezh2. J. Biol. Chem. 2015, 290, 27604–27617. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.; Chen, Q.-M.; Hong, C.; Wang, C.-Y. Histone methyltransferases and demethylases: Regulators in balancing osteogenic and adipogenic differentiation of mesenchymal stem cells. Int. J. Oral Sci. 2015, 7, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Nebbioso, A.; Dell’Aversana, C.; Bugge, A.; Sarno, R.; Valente, S.; Rotili, D.; Manzo, F.; Teti, D.; Mandrup, S.; Ciana, P.; et al. HDACs class II-selective inhibition alters nuclear receptor-dependent differentiation. J. Mol. Endocrinol. 2010, 45, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Kallen, A.N.; Zhou, X.-B.; Xu, J.; Qiao, C.; Ma, J.; Yan, L.; Lu, L.; Liu, C.; Yi, J.-S.; Zhang, H.; et al. The Imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol. Cell 2013, 52, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.-C.; Fu, W.-M.; Wong, C.-W.; Wang, Y.; Wang, W.-M.; Hu, G.-X.; Zhang, L.; Xiao, L.-J.; Wan, D.; Zhang, J.-F.; et al. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget 2015, 6, 22513–22525. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Zhang, A.; Shi, Z.; Ma, F.; Pu, P.; Wang, T.; Zhang, J.; Kang, C.; Zhang, Q. MicroRNA-200a suppresses the WNT/β-catenin signaling pathway by interacting with β-catenin. Int. J. Oncol. 2012, 40, 1162–1170. [Google Scholar] [PubMed]
- Lee, H.; Woo, K.; Ryoo, H.; Baek, J. Distal-less homeobox 5 inhibits adipogenic differentiation through the down-regulation of peroxisome proliferator-activated receptor γ expression. J. Cell. Physiol. 2013, 228, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Itoh, T.; Nozawa, Y.; Akao, Y. MicroRNA-141 and -200a are involved in bone morphogenetic protein-2-induced mouse pre-osteoblast differentiation by targeting distal-less homeobox 5. J. Biol. Chem. 2009, 284, 19272–19279. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Fu, M.; Bookout, A.; Kliewer, S.; Mangelsdorf, D. MicroRNA let-7 regulates 3T3-L1 adipogenesis. Mol. Endocrinol. 2009, 23, 925–931. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Sun, X.; Cai, H.; Sun, Y.; Plath, M.; Li, C.; Lan, X.; Lei, C.; Lin, F.; Bai, Y.; et al. Long non-coding RNA ADNCR suppresses adipogenic differentiation by targeting miR-204. Biochim. Biophys. Acta 2016, 1859, 871–882. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhao, L.; Xing, L.; Chen, D. MicroRNA-204 regulates Runx2 protein expression and mesenchymal progenitor cell differentiation. Stem Cells 2010, 28, 357–364. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Chen, K.; Wang, F.; Zhao, L.; Wan, X.; Wang, L.; Mo, Z. miR-204-5p promotes the adipogenic differentiation of human adipose-derived mesenchymal stem cells by modulating DVL3 expression and suppressing Wnt/β-catenin signaling. Int. J. Mol. Med. 2015, 35, 1587–1595. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.-X.; Yan, Y.-W.; Chen, D.; Ai, C.-Z.; Lu, X.; Xu, S.-S.; Jiang, S.; Zhong, G.-S.; Chen, D.-B.; Jiang, Y.-Z. Long non-coding RNA HoxA-AS3 interacts with EZH2 to regulate lineage commitment of mesenchymal stem cells. Oncotarget 2016, 7, 63561–63570. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, W.; Ge, X.; Yang, S.; Huang, M.; Zhuang, W.; Chen, P.; Zhang, X.; Fu, J.; Qu, J.; Li, B. Upregulation of lncRNA MEG3 promotes osteogenic differentiation of mesenchymal stem cells from multiple myeloma patients by targeting BMP4 transcription. Stem Cells 2015, 33, 1985–1997. [Google Scholar] [CrossRef] [PubMed]
- Kagami, M.; O’Sullivan, M.; Green, A.; Watabe, Y.; Arisaka, O.; Masawa, N.; Matsuoka, K.; Fukami, M.; Matsubara, K.; Kato, F.; et al. The IG-DMR and the MEG3-DMR at human chromosome 14q32.2: Hierarchical interaction and distinct functional properties as imprinting control centers. PLoS Genet. 2010, 6, e1000992. [Google Scholar] [CrossRef] [PubMed]
- Braconi, C.; Kogure, T.; Valeri, N.; Huang, N.; Nuovo, G.; Costinean, S.; Negrini, M.; Miotto, E.; Croce, C.; Patel, T. microRNA-29 can regulate expression of the long non-coding RNA gene MEG3 in hepatocellular cancer. Oncogene 2011, 30, 4750–4756. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Wang, L.; Lu, L.; Jiang, P.; Sun, H.; Wang, H. Inhibition of miR-29 by TGF-β-Smad3 signaling through dual mechanisms promotes transdifferentiation of mouse myoblasts into myofibroblasts. PLoS ONE 2012, 7, e33766. [Google Scholar] [CrossRef] [PubMed]
- Kwon, J.; Nabinger, S.; Vega, Z.; Sahu, S.; Alluri, R.; Abdul-Sater, Z.; Yu, Z.; Gore, J.; Nalepa, G.; Saxena, R.; et al. Pathophysiological role of microRNA-29 in pancreatic cancer stroma. Sci. Rep. 2015, 5, 11450. [Google Scholar] [CrossRef] [PubMed]
- Kapinas, K.; Kessler, C.; Ricks, T.; Gronowicz, G.; Delany, A. MiR-29 modulates Wnt signaling in human osteoblasts through a positive feedback loop. J. Biol. Chem. 2010, 285, 25221–25231. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Hassan, M.; Jafferji, M.; Aqeilan, R.; Garzon, R.; Croce, C.; Wijnen, A.; Stein, J.; Stein, G.; Lian, J. Biological functions of miR-29b contribute to positive regulation of osteoblast differentiation. J. Biol. Chem. 2009, 284, 15676–15684. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, X.; Qian, S.; Guo, L.; Huang, H.; He, Q.; Liu, Y.; Ma, C.; Tang, Q.-Q. Down-regulation of type I Runx2 mediated by dexamethasone is required for 3T3-L1 adipogenesis. Mol. Endocrinol. 2012, 26, 798–808. [Google Scholar] [CrossRef] [PubMed]
- Bäckesjö, C.; Li, Y.; Lindgren, U.; Haldosén, L. Activation of Sirt1 decreases adipocyte formation during osteoblast differentiation of mesenchymal stem cells. J. Bone Miner. Res. 2006, 21, 993–1002. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Merrell, A.J.; Stanger, B.Z. Adult cell plasticity in vivo: De-differentiation and transdifferentiation are back in style. Nat. Rev. Mol. Cell Biol. 2016, 17, 413–425. [Google Scholar] [CrossRef] [PubMed]
- Geurtzen, K.; Knopf, F.; Wehner, D.; Huitema, L.F.; Schulte-Merker, S.; Weidinger, G. Mature osteoblasts dedifferentiate in response to traumatic bone injury in the zebrafish fin and skull. Development 2014, 141, 2225–2234. [Google Scholar] [CrossRef] [PubMed]
- Park, D.; Spencer, J.A.; Koh, B.; Kobayashi, T.; Fujisaki, J.; Clemens, T.L.; Lin, C.P.; Kronenberg, H.M.; Scadden, D.T. Endogenous bone marrow MSCs are dynamic, fate-restricted participants in bone maintenance and regeneration. Cell Stem Cell 2012, 10, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Malaval, L.; Aubin, J. The mature osteoblast phenotype is characterized by extensive plasticity. Exp. Cell Res. 1997, 232, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Schiller, P.C.; D’Ippolito, G.; Brambilla, R.; Roos, B.A.; Howard, G.A. Inhibition of gap-junctional communication induces the trans-differentiation of osteoblasts to an adipocytic phenotype in vitro. J. Biol. Chem. 2001, 276, 14133–14138. [Google Scholar] [CrossRef] [PubMed]
- Yoshiko, Y.; Oizumi, K.; Hasegawa, T.; Minamizaki, T.; Tanne, K.; Maeda, N.; Aubin, J.E. A subset of osteoblasts expressing high endogenous levels of PPARγ switches fate to adipocytes in the rat calvaria cell culture model. PLoS ONE 2010, 5, e11782. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Liu, M.; Ono, N.; Bringhurst, R.; Kronenberg, H.; Guo, J. Loss of Wnt/β-catenin signaling causes cell fate shift of preosteoblasts from osteoblasts to adipocytes. J. Bone Miner. Res. 2012, 27, 2344–2358. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Seong, S.; Kim, K.; Kim, I.; Jeong, B.-C.C.; Kim, N. Downregulation of Runx2 by 1,25-dihydroxyvitamin D3 induces the transdifferentiation of osteoblasts to adipocytes. Int. J. Mol. Sci. 2016, 17, 770. [Google Scholar] [CrossRef] [PubMed]
- Meyer, M.B.; Benkusky, N.A.; Sen, B.; Rubin, J.; Pike, J.W. Epigenetic plasticity drives adipogenic and osteogenic differentiation of marrow-derived mesenchymal stem cells. J. Biol. Chem. 2016, 291, 17829–17847. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Link, D. Targeting of mesenchymal stromal cells by cre-recombinase transgenes commonly used to target osteoblast lineage cells. J. Bone Miner. Res. 2016, 31, 2001–2007. [Google Scholar] [CrossRef] [PubMed]
- Omatsu, Y.; Sugiyama, T.; Kohara, H.; Kondoh, G.; Fujii, N.; Kohno, K.; Nagasawa, T. The essential functions of adipo-osteogenic progenitors as the hematopoietic stem and progenitor cell niche. Immunity 2010, 33, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Perino, M.; Veenstra, G.J.C. Chromatin control of developmental dynamics and plasticity. Dev. Cell 2016, 38, 610–620. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, B.E.; Mikkelsen, T.S.; Xie, X.; Kamal, M.; Huebert, D.J.; Cuff, J.; Fry, B.; Meissner, A.; Wernig, M.; Plath, K. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 2006, 125, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, T.; Ku, M.; Jaffe, D.; Issac, B.; Lieberman, E.; Giannoukos, G.; Alvarez, P.; Brockman, W.; Kim, T.-K.; Koche, R.; et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 2007, 448, 553–560. [Google Scholar] [CrossRef] [PubMed]
- Rugg-Gunn, P.J.; Cox, B.J.; Ralston, A.; Rossant, J. Distinct histone modifications in stem cell lines and tissue lineages from the early mouse embryo. Proc. Natl. Acad. Sci. USA 2010, 107, 10783–10790. [Google Scholar] [CrossRef] [PubMed]
- Voigt, P.; Tee, W.-W.; Reinberg, D. A double take on bivalent promoters. Genes Dev. 2013, 27, 1318–1338. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Gordon, J.; Whitfield, T.; Tai, P.; Wijnen, A.; Stein, J.; Stein, G.; Lian, J. Chromatin dynamics regulate mesenchymal stem cell lineage specification and differentiation to osteogenesis. Biochim. Biophys. Acta 2017, 1860, 438–449. [Google Scholar] [CrossRef] [PubMed]
- Hemming, S.; Cakouros, D.; Isenmann, S.; Cooper, L.; Menicanin, D.; Zannettino, A.; Gronthos, S. EZH2 and KDM6A act as an epigenetic switch to regulate mesenchymal stem cell lineage specification. Stem Cells 2014, 32, 802–815. [Google Scholar] [CrossRef] [PubMed]
- Rojas, A.; Aguilar, R.; Henriquez, B.; Lian, J.; Stein, J.; Stein, G.; Wijnen, A.; Zundert, B.; Allende, M.; Montecino, M. Epigenetic control of the bone-master Runx2 gene during osteoblast-lineage commitment by the histone demethylase JARID1B/KDM5B. J. Biol. Chem. 2015, 290, 28329–28342. [Google Scholar] [CrossRef] [PubMed]
- Santoro, S.; Dulac, C. Histone variants and cellular plasticity. Trends Genet. 2015, 31, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Wilson, V.L.; Jones, P.A. DNA methylation decreases in aging but not in immortal cells. Science 1983, 220, 1055–1057. [Google Scholar] [CrossRef] [PubMed]
- Wilson, V.L.; Smith, R.A.; Ma, S.; Cutler, R.G. Genomic 5-methyldeoxycytidine decreases with age. J. Biol. Chem. 1987, 262, 9948–9951. [Google Scholar] [PubMed]
- Fuke, C.; Shimabukuro, M.; Petronis, A.; Sugimoto, J.; Oda, T.; Miura, K.; Miyazaki, T.; Ogura, C.; Okazaki, Y.; Jinno, Y. Age related changes in 5-methylcytosine content in human peripheral leukocytes and placentas: An HPLC-based study. Ann. Hum. Genet. 2004, 68, 196–204. [Google Scholar] [CrossRef] [PubMed]
- Maegawa, S.; Hinkal, G.; Kim, H.; Shen, L.; Zhang, L.; Zhang, J.; Zhang, N.; Liang, S.; Donehower, L.; Issa, J.-P. Widespread and tissue specific age-related DNA methylation changes in mice. Genome Res. 2010, 20, 332–340. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Groen, T.; Kadish, I.; Li, Y.; Wang, D.; James, S.; Karpf, A.; Tollefsbol, T. Insufficient DNA methylation affects healthy aging and promotes age-related health problems. Clin. Epigenet. 2011, 2, 349–360. [Google Scholar] [CrossRef] [PubMed]
- Pal, S.; Tyler, J. Epigenetics and aging. Sci. Adv. 2016, 2, e1600584. [Google Scholar] [CrossRef] [PubMed]
- Feser, J.; Tyler, J. Chromatin structure as a mediator of aging. FEBS Lett. 2011, 585, 2041–2048. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.-H.; Zhao, T.; Cui, K.; Hu, G.; Chen, Q.; Chen, W.; Wang, X.-W.; Soto-Gutierrez, A.; Zhao, K.; Deng, C.-X. Negative reciprocal regulation between Sirt1 and Per2 modulates the circadian clock and aging. Sci. Rep. 2016, 6, 28633. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.; Perrien, D.; Fleming, N.; Nyman, J.; Ono, K.; Connelly, L.; Moore, M.; Lwin, S.; Yull, F.; Mundy, G.; et al. Silent information regulator (Sir) T1 inhibits NF-κB signaling to maintain normal skeletal remodeling. J. Bone Miner. Res. 2013, 28, 960–969. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.-W.; Lee, S.; Seo, M.-S.; Park, S.-B.; Kurtz, A.; Kang, S.-K.; Kang, K.-S. Histone deacetylase controls adult stem cell aging by balancing the expression of polycomb genes and jumonji domain containing 3. Cell. Mol. Life Sci. 2010, 67, 1165–1176. [Google Scholar] [CrossRef] [PubMed]
- Dimmeler, S.; Nicotera, P. MicroRNAs in age-related diseases. EMBO Mol. Med. 2013, 5, 180–190. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kim, K.; Noh, J.; Yoon, J.-H.; Abdelmohsen, K.; Gorospe, M. Long noncoding RNAs in diseases of aging. Biochim. Biophys. Acta 2016, 1859, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Issa, J.-P. Aging and epigenetic drift: A vicious cycle. J. Clin. Investig. 2014, 124, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Mosteiro, L.; Pantoja, C.; Alcazar, N.; Marión, R.; Chondronasiou, D.; Rovira, M.; Fernandez-Marcos, P.; Muñoz-Martin, M.; Blanco-Aparicio, C.; Pastor, J.; et al. Tissue damage and senescence provide critical signals for cellular reprogramming in vivo. Science 2016, 354, aaf4445. [Google Scholar] [CrossRef] [PubMed]
- Li, C.-J.; Cheng, P.; Liang, M.-K.; Chen, Y.-S.; Lu, Q.; Wang, J.-Y.; Xia, Z.-Y.; Zhou, H.-D.; Cao, X.; Xie, H. MicroRNA-188 regulates age-related switch between osteoblast and adipocyte differentiation. J. Clin. Investig. 2015, 125, 1509–1522. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Tsang, K.; Tang, H.; Chan, D.; Cheah, K. Hypertrophic chondrocytes can become osteoblasts and osteocytes in endochondral bone formation. Proc. Natl. Acad. Sci. USA 2014, 111, 12097–12102. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; von der Mark, K.; Henry, S.; Norton, W.; Adams, H.; de Crombrugghe, B. Chondrocytes transdifferentiate into osteoblasts in endochondral bone during development, postnatal growth and fracture healing in mice. PLoS Genet. 2014, 10, e1004820. [Google Scholar] [CrossRef] [PubMed]
- Guan, D.; Zhang, W.; Zhang, W.; Liu, G.-H.; Belmonte, J. Switching cell fate, ncRNAs coming to play. Cell Death Dis. 2013, 4, e464. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, D.; DeWitt, N. In vivo cellular reprogramming: The next generation. Cell 2016, 166, 1386–1396. [Google Scholar] [CrossRef] [PubMed]
| LncRNAs | Expression Profiles | Functions | Effector Molecules | Target Genes | References |
|---|---|---|---|---|---|
| H19 | Osteogenic ↑ | miRNA precursor | miR-675 | TGFB1, HDAC4, HDAC5, HAC6 | [24,25] |
| Adipogenic ↓ | miRNA sponge | miR-200a, miR-141, miR-22, let-7 | DLX5, CTNNB1 | [26,83,84,85,87] | |
| MEG3 | Osteogenic ↑ | protein recruiter | PRC2 | TGFB2, TGFBR1, SMAD2 | [30] |
| Adipogenic ↓ | |||||
| ADNCR | Osteogenic ? | miRNA sponge | miR-204 | RUNX2, SIRT1, DVL3 | [89,90,91] |
| Adipogenic ↓ | |||||
| HOXA-AS3 | Osteogenic → | protein recruiter | EZH2 | RUNX2 | [92] |
| Adipogenic ↑ |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yoshioka, H.; Yoshiko, Y. The Roles of Long Non-Protein-Coding RNAs in Osteo-Adipogenic Lineage Commitment. Int. J. Mol. Sci. 2017, 18, 1236. https://doi.org/10.3390/ijms18061236
Yoshioka H, Yoshiko Y. The Roles of Long Non-Protein-Coding RNAs in Osteo-Adipogenic Lineage Commitment. International Journal of Molecular Sciences. 2017; 18(6):1236. https://doi.org/10.3390/ijms18061236
Chicago/Turabian StyleYoshioka, Hirotaka, and Yuji Yoshiko. 2017. "The Roles of Long Non-Protein-Coding RNAs in Osteo-Adipogenic Lineage Commitment" International Journal of Molecular Sciences 18, no. 6: 1236. https://doi.org/10.3390/ijms18061236




