Functional Characterization of Soybean Glyma04g39610 as a Brassinosteroid Receptor Gene and Evolutionary Analysis of Soybean Brassinosteroid Receptors
Abstract
:1. Introduction
2. Results
2.1. Isolation of Glyma04g39610 (GmBRI1b)
2.2. Identification of Other BR Receptor Genes in Soybean
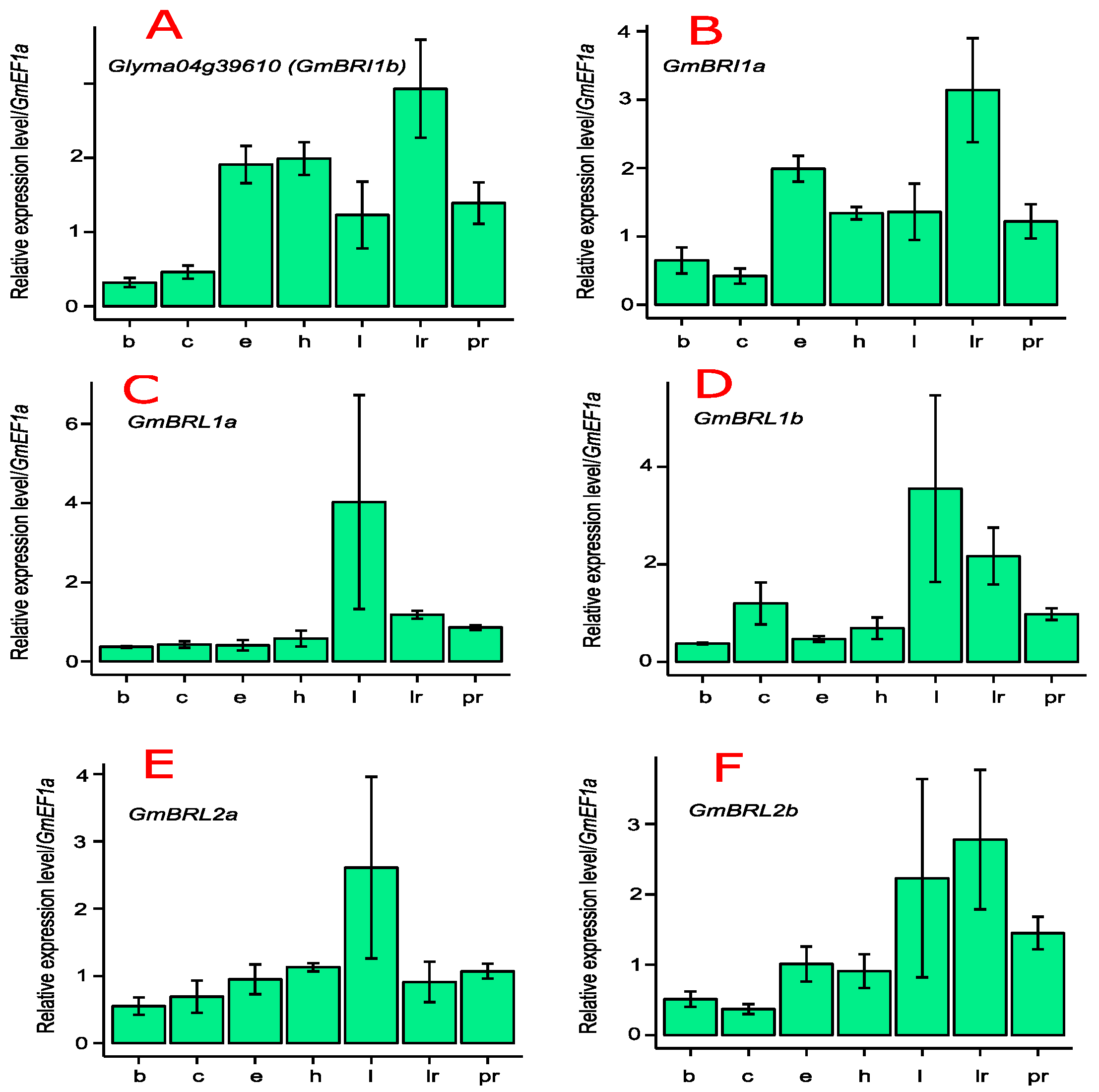
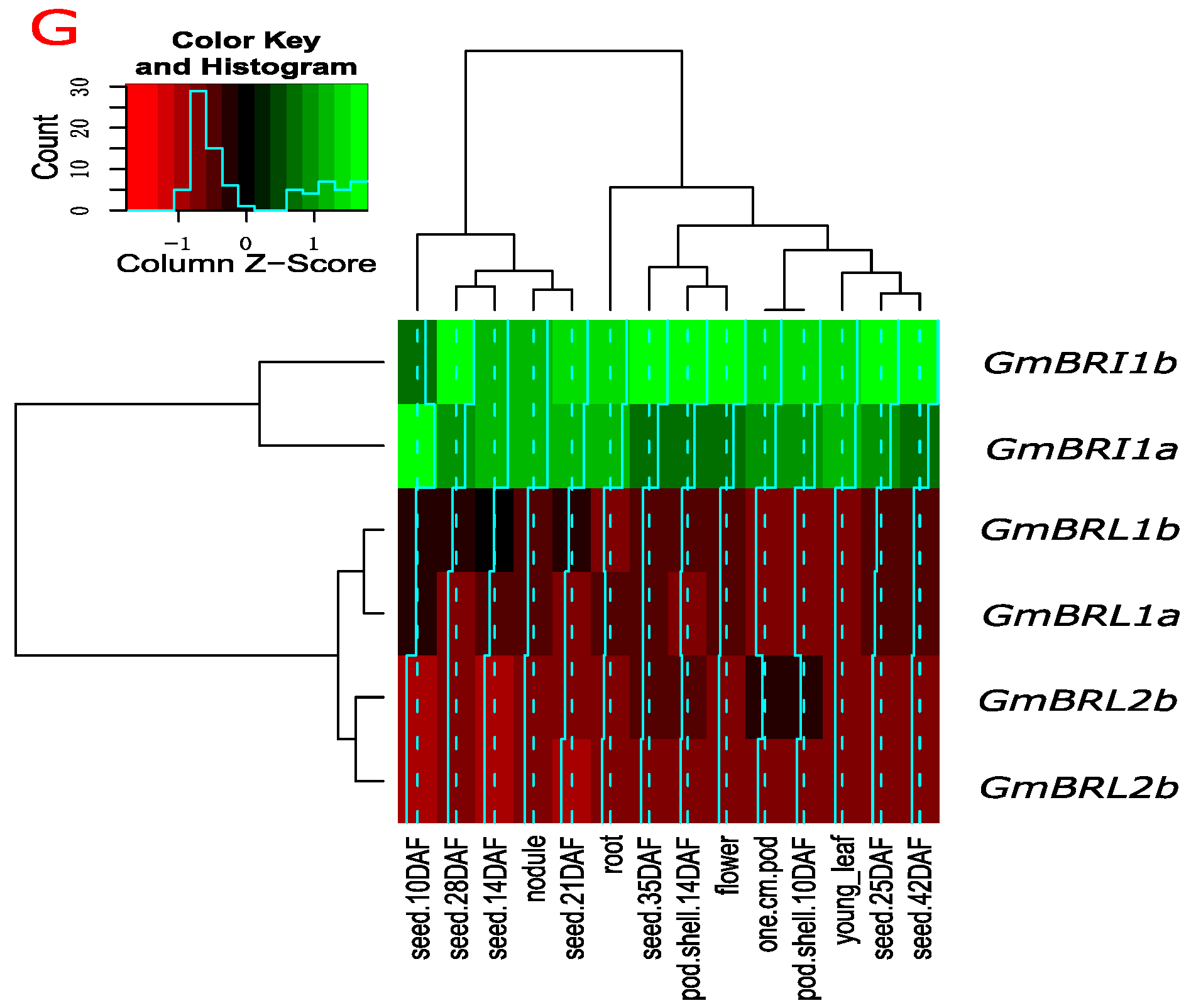
2.3. Transcript Levels of GmBRI1b and Other BR Receptor Genes in Soybean
2.4. Subcellular Localization of GmBRI1b
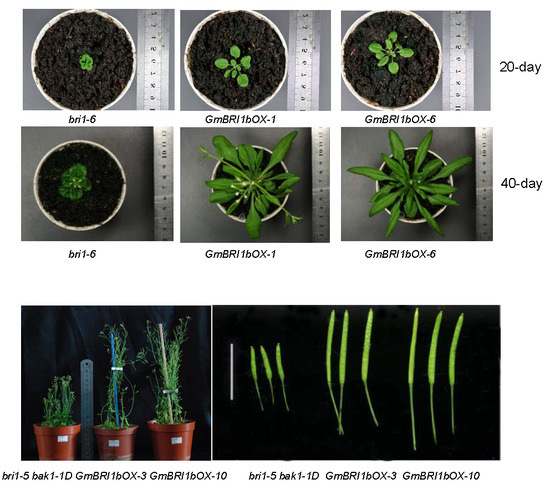
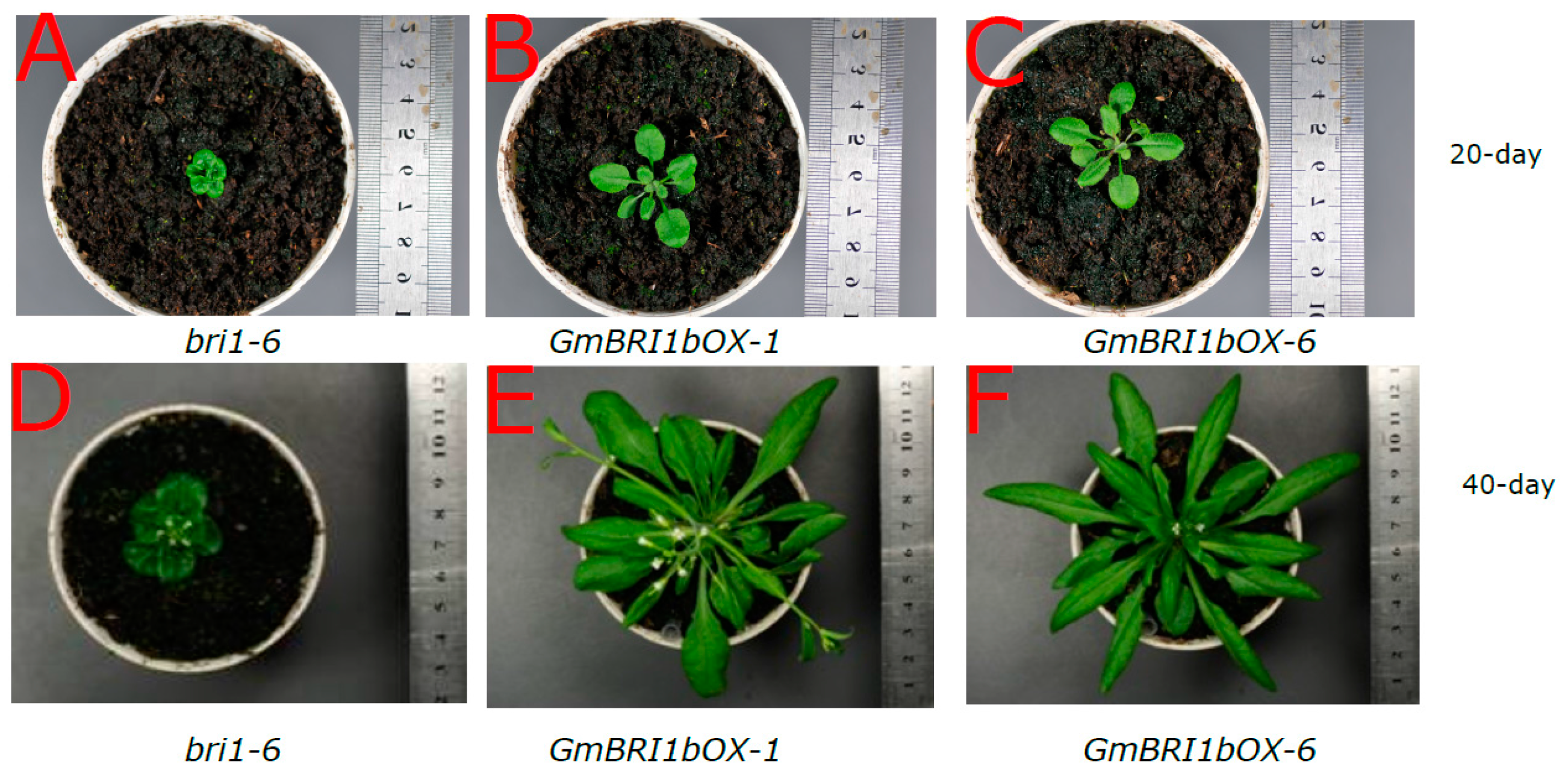
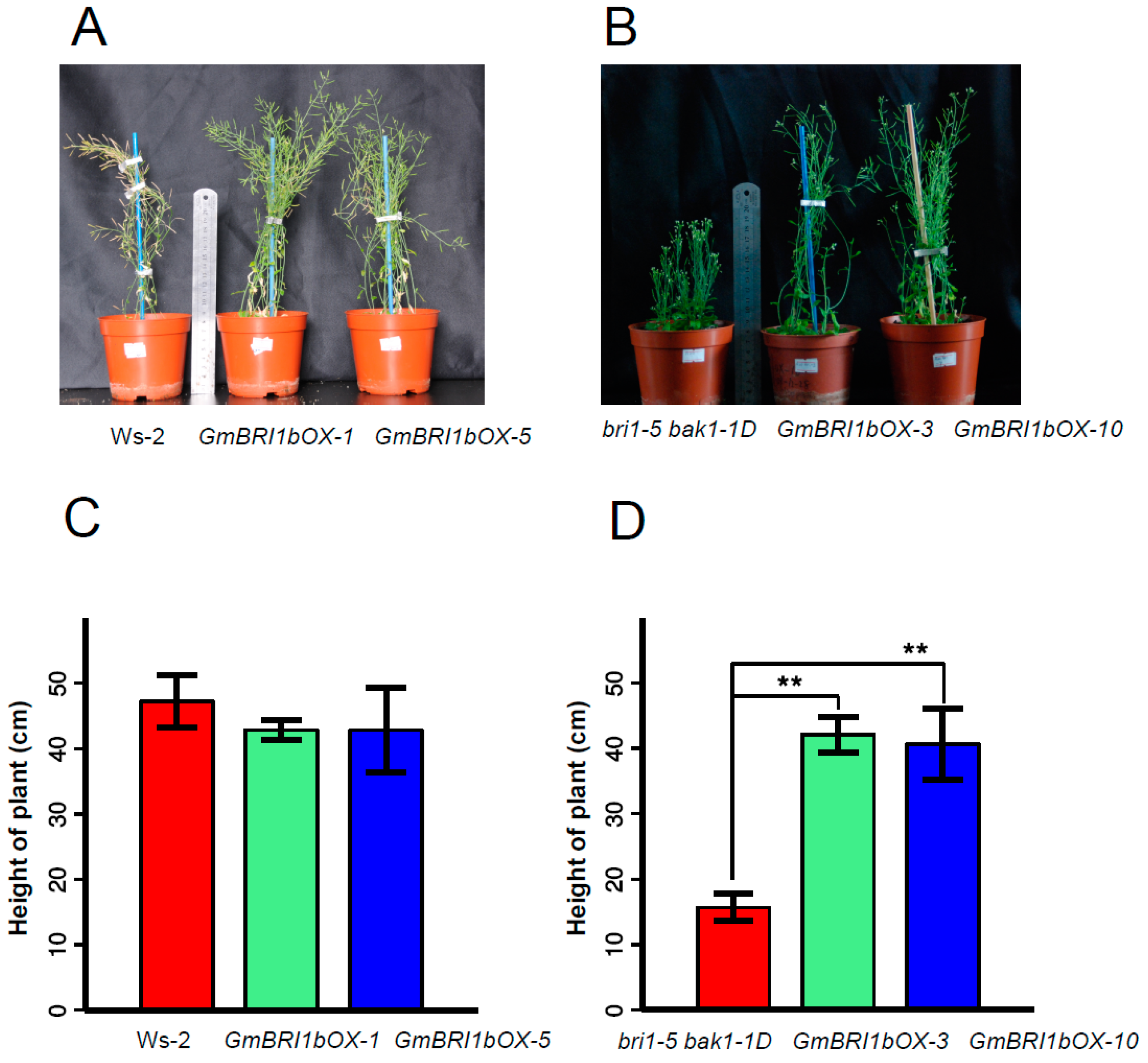
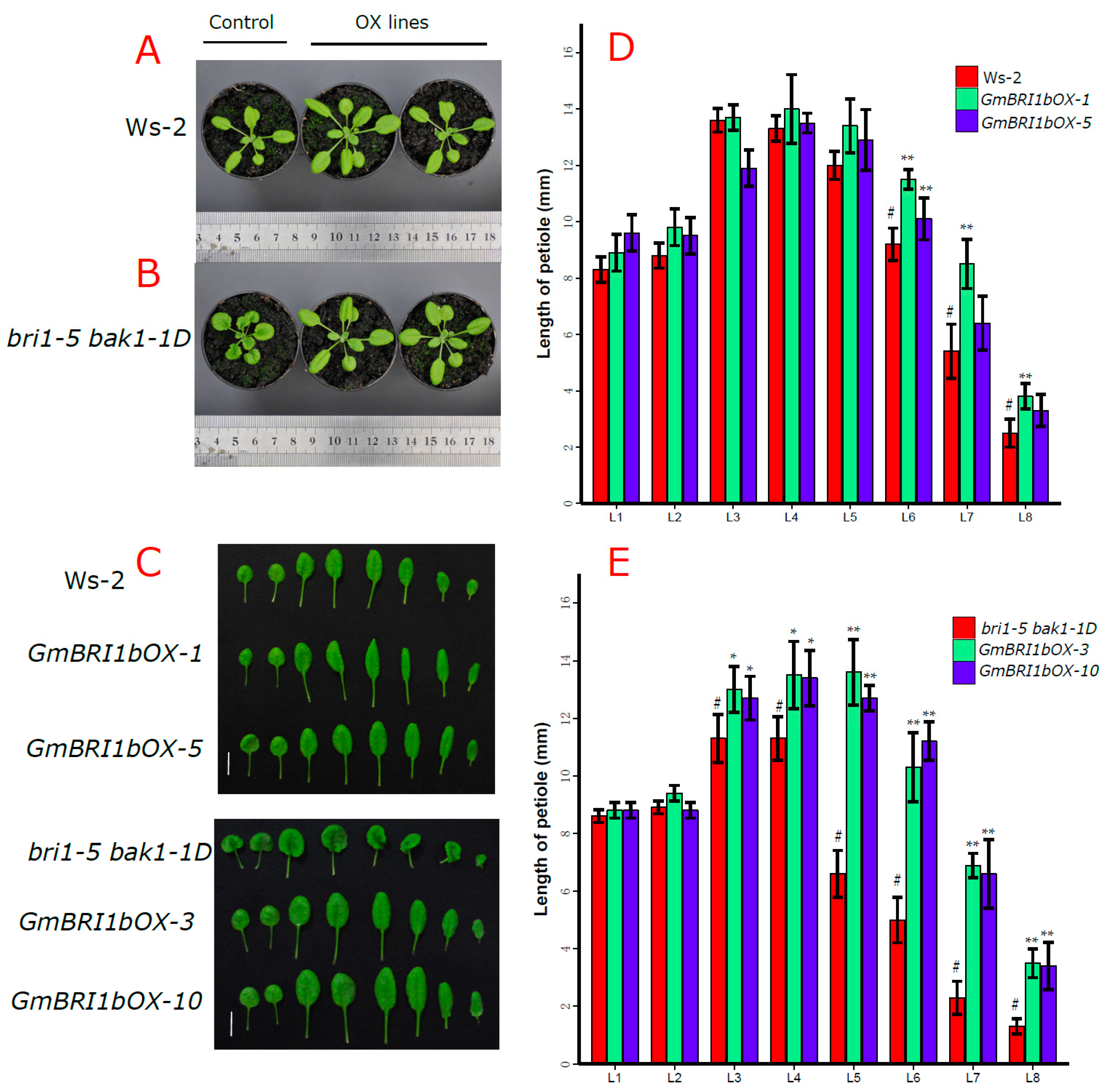
2.5. Functional Analysis of GmBRI1b in Arabidopsis
2.6. Ectopic Over-Expression of GmBRI1b Increased the Hypocotyl Length of the bri1-5 bak1-1D Mutant and Changed the Responses of the Wild Type and bri1-5 bak1-1D Mutant to Brassinazole
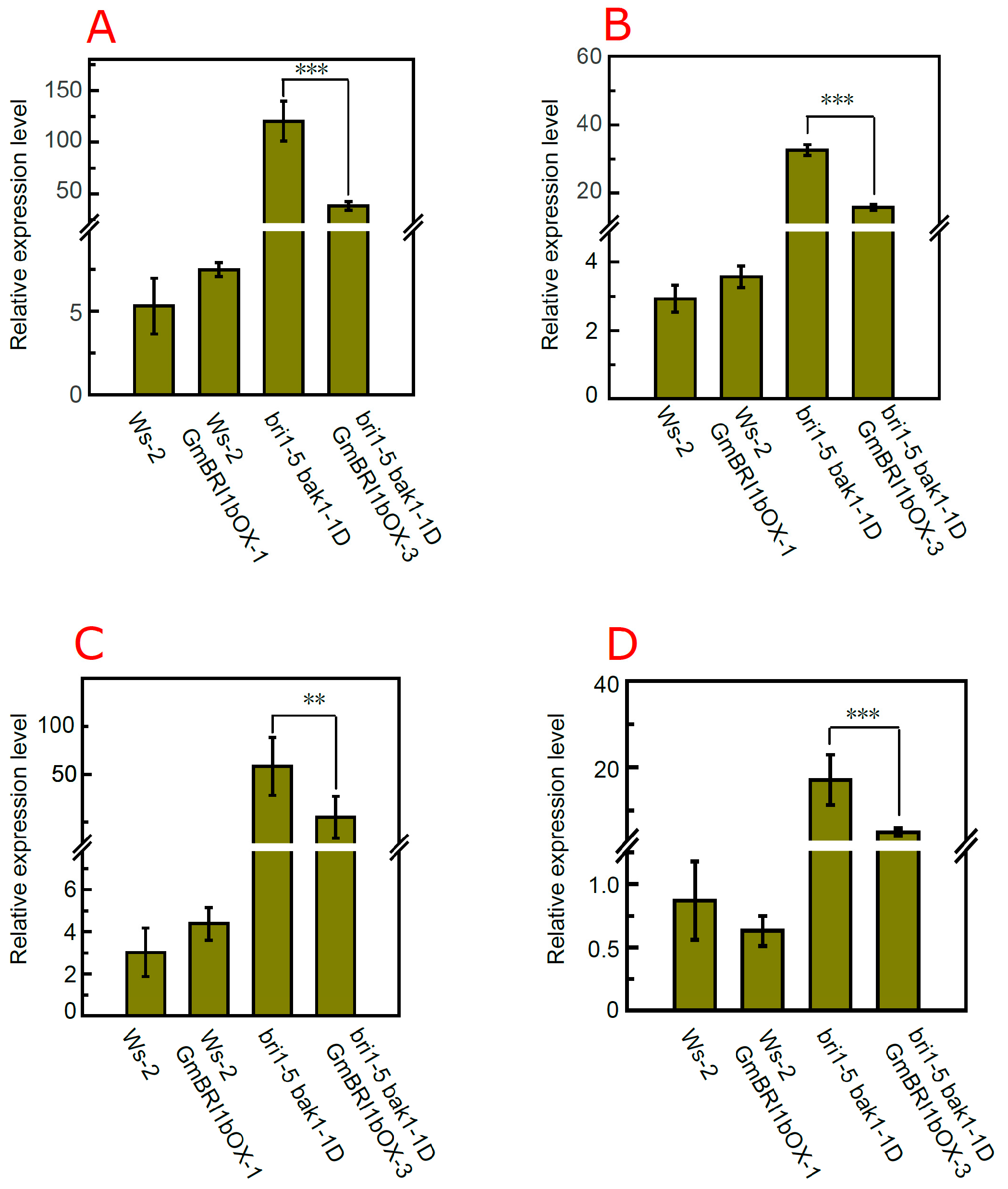
2.7. Over-Expression of GmBRI1b Altered the Expression Level of BR Biosynthesis-Related Genes in the bri1-5 bak1-1D Mutant
2.8. Structural Modeling of Soybean BR Receptors
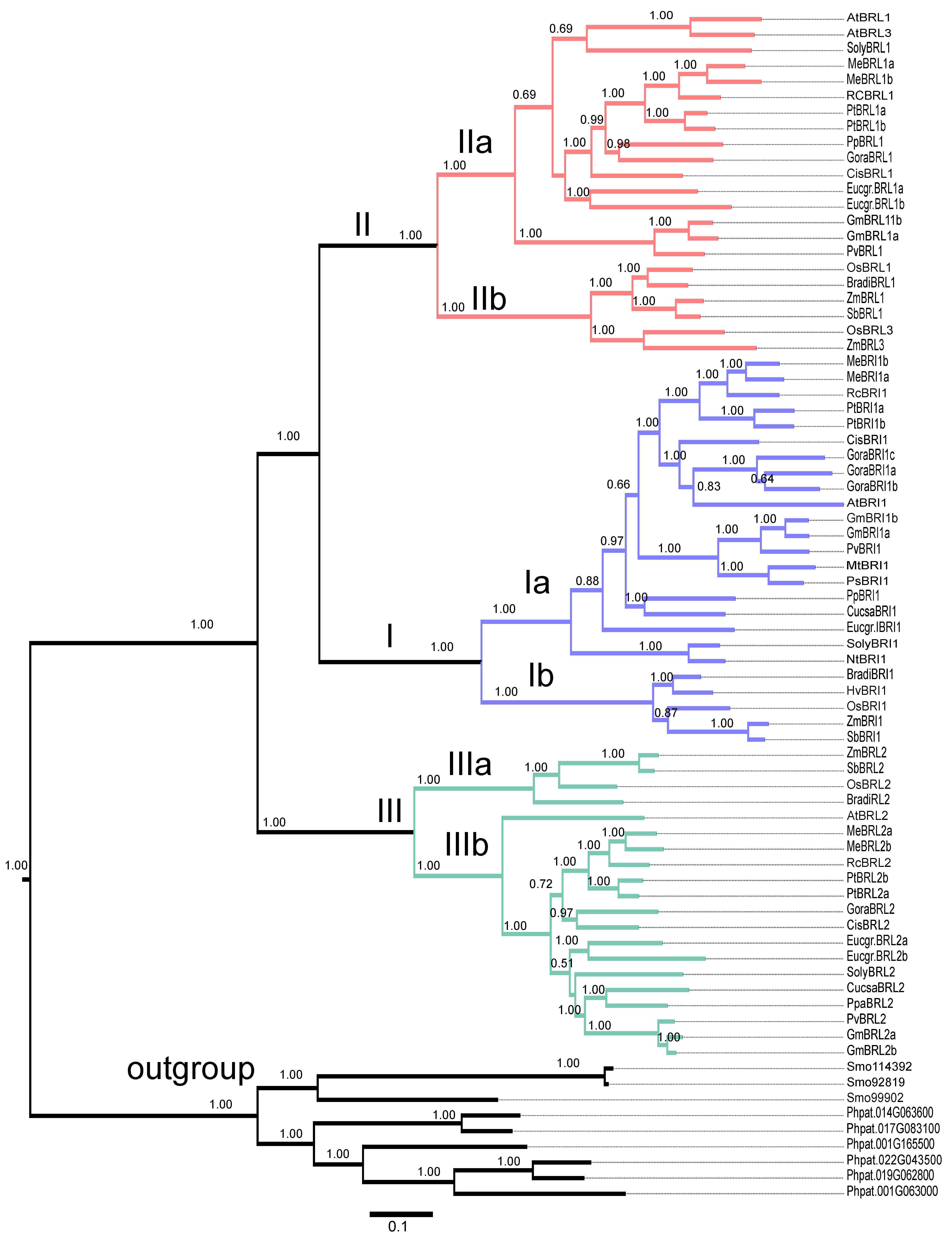
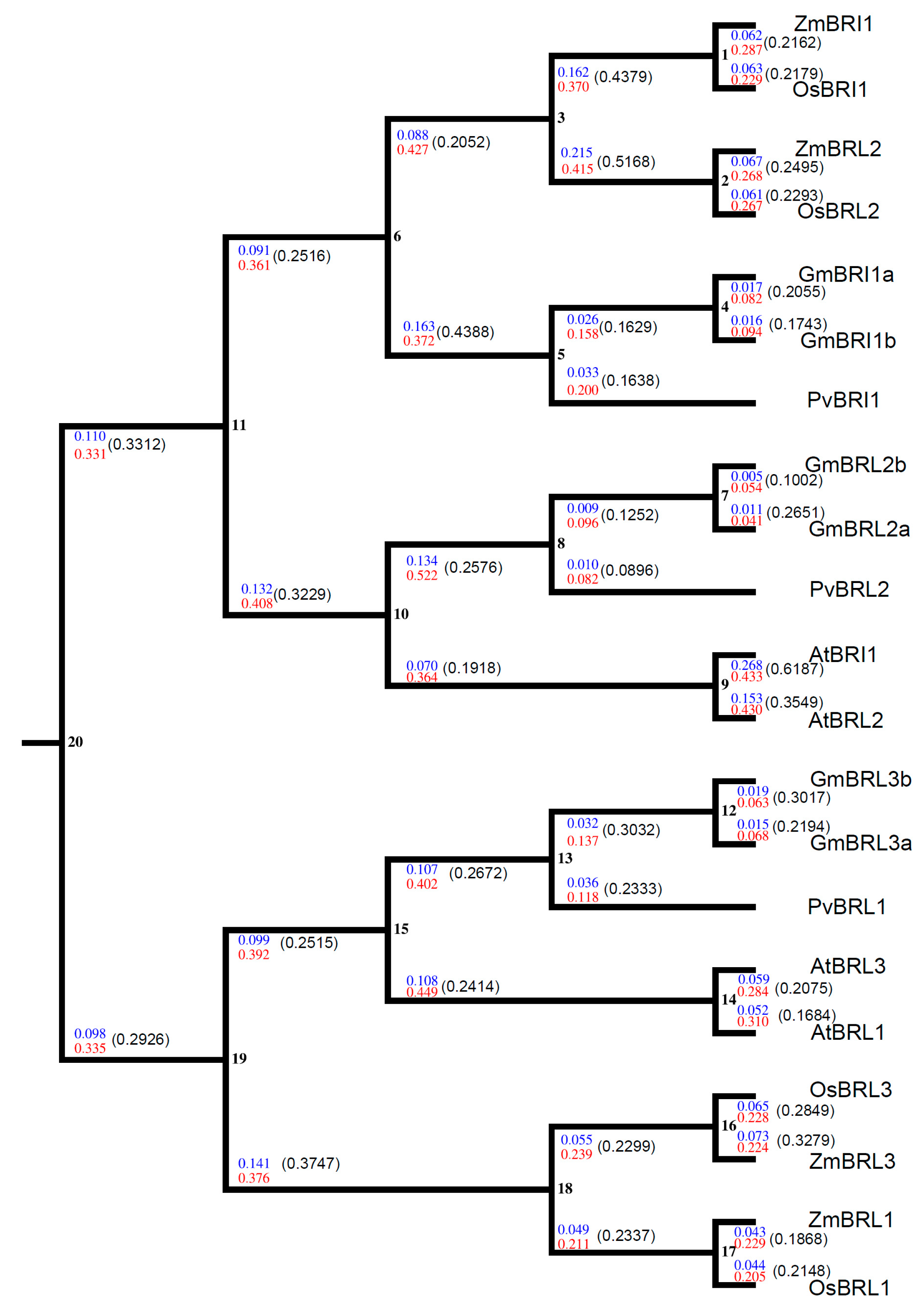
2.9. Evolutionary Analysis of BR Receptors in Plants
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth
4.2. Extraction of Genomic DNA and RNA and Reverse Transcription of mRNA
4.3. RACE Cloning of GmBRI1b
4.4. Analysis of Expression Patterns of Soybean BR Receptor Genes
4.5. Over-Expression of GmBRI1b in Arabidopsis
4.6. Subcellular Localization of GmBRI1b
4.7. Phenotypic Analysis of Transgenic Arabidopsis
4.8. Determination of the Expression Levels of BR Biosynthesis-Related Genes in the Wild Type, the Mutant, and Their Corresponding over-Expression Lines
4.9. Determination of the GmBRI1b Structure
4.10. Alignment of BR Receptors
4.11. Structural Modeling of Soybean BR Receptors
4.12. Estimation of Selection and Substitution Rates
4.13. Data Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| AA | Amino acid |
| BAK1 | BRI1 ASSOCIATED RECEPTOR KINASE 1 |
| BIN2 | BRASSINOSTEROID INSENSITIVE2 |
| BKI1 | BRI1 KINASE INHIBITOR 1 |
| BR | Brassinosteroid |
| BRI1 | BRASSINOSTEROID INSENSITIVE 1 |
| Brz | Brassinazole |
| BZR1 | BRASSINAZOLE-RESISTANT1 |
| ID | Island domain |
| ORF | Open reading frame |
| qRT-PCR | Quantitative real-time PCR |
References
- Müssig, C.; Shin, G.H.; Altmann, T. Brassinosteroids promote root growth in Arabidopsis. Plant Physiol. 2003, 133, 1261–1271. [Google Scholar] [CrossRef] [PubMed]
- Hacham, Y.; Holland, N.; Butterfield, C.; Ubeda-Tomas, S.; Bennett, M.J.; Chory, J.; Savaldi-Goldstein, S. Brassinosteroid perception in the epidermis controls root meristem size. Development 2011, 138, 839–848. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.W.; Michniewicz, M.; Bergmann, D.C.; Wang, Z.Y. Brassinosteroid regulates stomatal development by GSK3-mediated inhibition of a MAPK pathway. Nature 2012, 482, 419–422. [Google Scholar] [CrossRef] [PubMed]
- Leubner-Metzger, G. Brassinosteroids and gibberellins promote tobacco seed germination by distinct pathways. Planta 2001, 213, 758–763. [Google Scholar] [CrossRef] [PubMed]
- Chory, J.; Nagpal, P.; Peto, C.A. Phenotypic and genetic analysis of det2, a new mutant that affects light-regulated seedling development in Arabidopsis. Plant Cell 1991, 3, 445–459. [Google Scholar] [CrossRef] [PubMed]
- Szekeres, M.; Németh, K.; Koncz-Kálmán, Z.; Mathur, J.; Kauschmann, A.; Altmann, T.; Rédei, G.P.; Nagy, F.; Schell, J.; Koncz, C. Brassinosteroids rescue the deficiency of cyp90, a cytochrome p450, controlling cell elongation and de-etiolation in Arabidopsis. Cell 1996, 85, 171–182. [Google Scholar] [CrossRef]
- Bai, M.Y.; Fan, M.; Oh, E.; Wang, Z.Y. A triple helix-loop-helix/basic helix-loop-helix cascade controls cell elongation downstream of multiple hormonal and environmental signaling pathways in Arabidopsis. Plant Cell 2012, 24, 4917–4929. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, B.J.; Ross, J.J.; Reid, J.B. Nodulation phenotypes of gibberellin and brassinosteroid mutants of pea. Plant Physiol. 2005, 138, 2396–2405. [Google Scholar] [CrossRef] [PubMed]
- Nakashita, H.; Yasuda, M.; Nitta, T.; Asami, T.; Fujioka, S.; Arai, Y.; Sekimata, K.; Takatsuto, S.; Yamaguchi, I.; Yoshida, S. Brassinosteroid functions in a broad range of disease resistance in tobacco and rice. Plant J. 2003, 33, 887–898. [Google Scholar] [CrossRef] [PubMed]
- Dhaubhadel, S.; Browning, K.S.; Gallie, D.R.; Krishna, P. Brassinosteroid functions to protect the translational machinery and heat-shock protein synthesis following thermal stress. Plant J. 2002, 29, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Bajguz, A.; Hayat, S. Effects of brassinosteroids on the plant responses to environmental stresses. Plant Physiol. Biochem. 2009, 47, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sathiyamoorthy, P.; Nakamuracohen, S. In vitro root induction by 24-epibrassinolide on hypocotyl segments of soybean [Glycine max (l.) merr.]. Plant Growth Regul. 1990, 9, 73–76. [Google Scholar] [CrossRef]
- Zurek, D.M.; Rayle, D.L.; McMorris, T.C.; Clouse, S.D. Investigation of gene expression, growth kinetics, and wall extensibility during brassinosteroid-regulated stem elongation. Plant Physiol. 1994, 104, 505–513. [Google Scholar] [PubMed]
- Zhang, M.; Zhai, Z.; Tian, X.; Duan, L.; Li, Z. Brassinolide alleviated the adverse effect of water deficits on photosynthesis and the antioxidant of soybean (Glycine max L.). Plant Growth Regul. 2008, 56, 257–264. [Google Scholar] [CrossRef]
- Li, J.; Chory, J. A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 1997, 90, 929–938. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Nakano, T.; Gendron, J.; He, J.; Chen, M.; Vafeados, D.; Yang, Y.; Fujioka, S.; Yoshida, S.; Asami, T.; et al. Nuclear-localized BZR1 mediates brassinosteroid-induced growth and feedback suppression of brassinosteroid biosynthesis. Dev. Cell 2002, 2, 505–513. [Google Scholar] [CrossRef]
- Wang, X.; Chory, J. Brassinosteroids regulate dissociation of BKI1, a negative regulator of BRI1 signaling, from the plasma membrane. Science 2006, 313, 1118–1122. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wen, J.; Lease, K.A.; Doke, J.T.; Tax, F.E.; Walker, J.C. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 2002, 110, 213–222. [Google Scholar] [CrossRef]
- Li, J.; Nam, K.H.; Vafeados, D.; Chory, J. BIN2, a new brassinosteroid-insensitive locus in Arabidopsis. Plant Physiol. 2001, 127, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Nam, K.H. Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science 2002, 295, 1299–1301. [Google Scholar] [PubMed]
- He, J.X.; Gendron, J.M.; Yang, Y.; Li, J.; Wang, Z.Y. The GSK3-like kinase BIN2 phosphorylates and destabilizes BZR1, a positive regulator of the brassinosteroid signaling pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 2002, 99, 10185–10190. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Peng, P.; Schmitz, R.J.; Decker, A.D.; Tax, F.E.; Li, J. Two putative BIN2 substrates are nuclear components of brassinosteroid signaling. Plant Physiol. 2002, 130, 1221–1229. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Yuan, M.; Wang, R.; Yang, Y.; Wang, C.; Oses-Prieto, J.A.; Kim, T.W.; Zhou, H.W.; Deng, Z.; Gampala, S.S.; et al. PP2A activates brassinosteroid-responsive gene expression and plant growth by dephosphorylating BZR1. Nat. Cell Biol. 2001, 13, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Vafeados, D.; Tao, Y.; Yoshida, S.; Asami, T.; Chory, J. A new class of transcription factors mediates brassinosteroid-regulated gene expression in Arabidopsis. Cell 2005, 120, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Peng, P.; Yan, Z.; Zhu, Y.; Li, J. Regulation of the Arabidopsis GSK3-like kinase BRASSINOSTEROID-INSENSITIVE 2 through proteasome-mediated protein degradation. Mol. Plant 2008, 1, 338–346. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, X.; Meisenhelder, J.; Hunter, T.; Yoshida, S.; Asami, T.; Chory, J. Autoregulation and homodimerization are involved in the activation of the plant steroid receptor BRI1. Dev. Cell 2005, 8, 855–865. [Google Scholar] [CrossRef] [PubMed]
- Hothorn, M.; Belkhadir, Y.; Dreux, M.; Dabi, T.; Noel, J.P.; Wilson, I.A.; Chory, J. Structural basis of steroidhormone perception by the receptor kinase BRI1. Nature 2011, 474, 467–471. [Google Scholar] [CrossRef] [PubMed]
- She, J.; Han, Z.; Kim, T.W.; Wang, J.; Cheng, W.; Chang, J.; Shi, S.; Wang, J.; Yang, M.; Wang, Z.Y.; et al. Structural insight into brassinosteroid perception by BRI1. Nature 2011, 474, 472–476. [Google Scholar] [CrossRef] [PubMed]
- Cano-Delgado, A.; Yin, Y.; Yu, C.; Vafeados, D.; Mora-Garcia, S.; Cheng, J.C.; Nam, K.H.; Li, J.; Chory, J. BRL1 and BRL3 are novel brassinosteroid receptors that function in vascular differentiation in Arabidopsis. Development 2004, 131, 5341–5351. [Google Scholar] [CrossRef] [PubMed]
- Zhou, A.; Wang, H.; Walker, J.C.; Li, J. BRL1, a leucine-rich repeat receptor-like protein kinase, is functionally redundant with BRI1 in regulating Arabidopsis brassinosteroid signaling. Plant J. 2004, 40, 399–409. [Google Scholar] [CrossRef] [PubMed]
- Yamamuro, C.; Ihara, Y.; Wu, X.; Noguchi, T.; Fujioka, S.; Takatsuto, S.; Ashikari, M.; Kitano, H.; Matsuoka, M. Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 2000, 12, 1591–1606. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, A.; Fujioka, S.; Sunohara, H.; Kamiya, N.; Hong, Z.; Inukai, Y.; Miura, K.; Takatsuto, S.; Yoshida, S.; Ueguchi-Tanaka, M.; et al. The role of OsBRI1 and its homologous genes, OsBRL1 and OsBRL3, in rice. Plant Physiol. 2006, 140, 580–590. [Google Scholar] [CrossRef] [PubMed]
- Montoya, T.; Nomura, T.; Farrar, K.; Kaneta, T.; Yokota, T.; Bishop, G.J. Cloning the tomato Curl3 gene highlights the putative dual role of the leucine-rich repeat receptor kinase tBRI1/SR160 in plant steroid hormone and peptide hormone signaling. Plant Cell 2002, 14, 3163–3176. [Google Scholar] [CrossRef] [PubMed]
- Nomura, T.; Bishop, G.J.; Kaneta, T.; Reid, J.B.; Chory, J.; Yokota, T. The LKA gene is BRASSINOSTEROID INSENSITIVE 1 homolog of pea. Plant J. 2003, 36, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Chono, M.; Honda, I.; Zeniya, H.; Yoneyama, K.; Saisho, D.; Takeda, K.; Takatsuto, S.; Hoshino, T.; Watanabe, Y. A semidwarf phenotype of barley uzu results from a nucleotide substitution in the gene encoding a putative brassinosteroid receptor. Plant Physiol. 2003, 133, 1209–1219. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Fokar, M.; Asami, T.; Yoshida, S.; Allen, R.D. Characterization of the brassinosteroid insensitive 1genes of cotton. Plant Mol. Biol. 2004, 54, 221–232. [Google Scholar] [CrossRef] [PubMed]
- Kir, G.; Ye, H.; Nelissen, H.; Neelakandan, A.K.; Kusnandar, A.S.; Luo, A.; Inzé, D.; Sylvester, A.W.; Yin, Y.; Becraft, P.W. RNA interference knockdown of BRASSINOSTEROID INSENSITIVE1 in maize reveals novel functions for brassinosteroid signaling in controlling plant architecture. Plant Physiol. 2015, 169, 826–839. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Navarro, C.; Moore, J.; Ott, A.; Baumert, E.; Mohan, A.; Gill, K.S.; Sandhu, D. Evolutionary, comparative and functional analyses of the brassinosteroid receptor gene, BRI1, in wheat and its relation to other plant genomes. PLoS ONE 2015, 10, e0127544. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Sun, S.; Wu, C.; Han, T.; Wang, Q. Isolation and characterization of the brassinosteroid receptor gene (GmBRI1) from Glycine max. Int. J. Mol. Sci. 2014, 15, 3871–3888. [Google Scholar] [CrossRef] [PubMed]
- Schmutz, J.; Cannon, S.; Schlueter, J.; Ma, J.; Mitros, T. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef] [PubMed]
- Childs, K.L.; Hamilton, J.P.; Zhu, W.; Ly, E.; Cheung, F.; Wu, H.; Rabinowicz, P.D.; Town, C.D.; Buell, C.R.; Chan, A.P. The TIGR plant transcript assemblies database. Nucleic Acids Res. 2007, 35, D846–D851. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, D.L.; Chappey, C.; Lash, A.E.; Leipe, D.D.; Madden, T.L.; Schuler, G.D.; Tatusova, T.A.; Rapp, B.A. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2000, 28, 10–14. [Google Scholar] [CrossRef] [PubMed]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Doerks, T.; Bork, P. SMART: Recent updates, new developments and status. Nucleic Acids Res. 2015, 43, D257–D260. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.H.; Tang, H.; Wang, X.; Paterson, A.H. PGDD: A database of gene and genome duplication in plants. Nucleic Acids Res. 2013, 41, D1152–D1158. [Google Scholar] [CrossRef] [PubMed]
- Severin, A.J.; Woody, J.L.; Bolon, Y.T.; Joseph, B.; Diers, B.W.; Farmer, A.D.; Muehlbauer, G.J.; Nelson, R.T.; Grant, D.; Specht, J.E.; et al. RNA-Seq atlas of Glycine max: A guide to the soybean transcriptome. BMC Plant Biol. 2010, 10, 160. [Google Scholar] [CrossRef] [PubMed]
- Nelson, B.K.; Cai, X.; Nebenführ, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 2007, 51, 1126–1136. [Google Scholar] [CrossRef] [PubMed]
- Noguchi, T.; Fujioka, S.; Choe, S.; Takatsuto, S.; Yoshida, S.; Yuan, H.; Feldmann, K.A.; Tax, F.E. Brassinosteroid-insensitive dwarf mutants of Arabidopsis accumulate brassinosteroids. Plant Physiol. 1999, 121, 743–752. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Jin, H. Regulation of brassinosteroid signaling. Trends Plant Sci. 2007, 12, 37–41. [Google Scholar] [CrossRef] [PubMed]
- Nagata, N.; Min, Y.K.; Nakano, T.; Asami, T.; Yoshida, S. Treatment of dark-grown Arabidopsis thaliana with a brassinosteroid-biosynthesis inhibitor, brassinazole, induces some characteristics of light-grown plants. Planta 2000, 211, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Kota, U.; He, K.; Blackburn, K.; Li, J.; Goshe, M.B.; Huber, S.C.; Clouse, S.D. Sequential transphosphorylation of the BRI1/BAK1 receptor kinase complex impacts early events in brassinosteroid signaling. Dev. Cell 2008, 15, 220–235. [Google Scholar] [CrossRef] [PubMed]
- Mathur, J.; Molnár, G.; Fujioka, S.; Takatsuto, S.; Sakurai, A.; Yokota, T.; Adam, G.; Voigt, B.; Nagy, F.; Maas, C.; et al. Transcription of the Arabidopsis CPD gene, encoding a steroidogenic cytochrome p450, is negatively controlled by brassinosteroids. Plant J. 1998, 14, 593–602. [Google Scholar] [CrossRef] [PubMed]
- Bancos, S.; Nomura, T.; Sato, T.; Molnár, G.; Bishop, G.J.; Koncz, C.; Yokota, T.; Nagy, F.; Szekeres, M. Regulation of transcript levels of the Arabidopsis cytochrome p450 genes involved in brassinosteroid biosynthesis. Plant Physiol. 2002, 130, 504–513. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Asami, T.; Yoshida, S.; Nakamura, Y.; Matsuo, T.; Okamoto, S. Brassinosteroid homeostasis in Arabidopsis is ensured by feedback expressions of multiple genes involved in its metabolism. Plant Physiol. 2005, 138, 1117–1125. [Google Scholar] [CrossRef] [PubMed]
- Pieper, U.; Webb, B.M.; Barkan, D.T.; Schneidman-Duhovny, D.; Schlessinger, A.; Braberg, H.; Yang, Z.; Meng, E.C.; Pettersen, E.F.; Huang, C.C.; et al. ModBase, a database of annotated comparative protein structure models, and associated resources. Nucleic Acids Res. 2011, 39, D465–D474. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Fiser, A.; Do, R.K.; Sali, A. Modeling of loops in protein structures. Protein Sci. 2000, 9, 1753–1773. [Google Scholar] [CrossRef] [PubMed]
- PyMOL. The PyMOL molecular graphics system, version 1.5.0. Available online: https://www.pymol.org (accessed on 3 July 2012).
- Huelsenbeck, J.P.; Ronquist, F. Mrbayes: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Suyama, M.; Torrents, D.; Bork, P. PAL2NAL: Robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 2006, 34, W609–W612. [Google Scholar] [CrossRef] [PubMed]
- Kutschera, U.; Wang, Z.Y. Brassinosteroid action in flowering plants: A Darwinian perspective. J. Exp. Bot. 2012, 63, 3511–3522. [Google Scholar] [CrossRef] [PubMed]
- Hayat, S.; Ahmad, A. Brassinosteroids: A Class of Plant Hormone; Springer: Heidelberg, Germany, 2011. [Google Scholar]
- Friedrichsen, D.M.; Joazeiro, C.A.; Li, J.; Hunter, T.; Chory, J. Brassinosteroid-insensitive-1 is a ubiquitously expressed leucine-rich repeat receptor serine/threonine kinase. Plant Physiol. 2000, 123, 1247–1256. [Google Scholar] [CrossRef] [PubMed]
- Terakado, J.; Fujihara, S.; Goto, S.; Kuratani, R.; Suzuki, Y.; Yoshida, S.; Yoneyama, T. Systemic effect of a brassinosteroid on root nodule formation in soybean as revealed by the application of brassinolide and brassinazole. Soil Sci. Plant Nutr. 2005, 51, 389–395. [Google Scholar] [CrossRef]
- Libault, M.; Joshi, T.; Takahashi, K.; Hurley-Sommer, A.; Puricelli, K.; Blake, S.; Finger, R.E.; Taylor, C.G.; Xu, D.; Nguyen, H.T.; et al. Large-scale analysis of putative soybean regulatory gene expression identifies a Myb gene involved in soybean nodule development. Plant Physiol. 2009, 151, 1207–1220. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Nagpal, P.; Vitart, V.; McMorris, T.C.; Chory, J. A role for brassinosteroids in light-dependent development of Arabidopsis. Science 1996, 272, 398–401. [Google Scholar] [CrossRef] [PubMed]
- Choe, S.; Dilkes, B.P.; Fujioka, S.; Takatsuto, S.; Sakurai, A.; Feldmann, K.A. The DWF4 gene of Arabidopsis encodes a cytochrome p450 that mediates multiple 22α-hydroxylation steps in brassinosteroid biosynthesis. Plant Cell 1998, 10, 231–243. [Google Scholar] [PubMed]
- Bishop, G.J.; Harrison, K.; Jones, J.D. The tomato Dwarf gene isolated by heterologous transposon tagging encodes the first member of a new cytochrome p450 family. Plant Cell 1996, 8, 959–969. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Mao, H. On the origin and evolution of plant brassinosteroid receptor kinases. J. Mol. Evol. 2014, 78, 118–129. [Google Scholar] [CrossRef] [PubMed]
- Cheon, J.; Fujioka, S.; Dilkes, B.P.; Choe, S. Brassinosteroids regulate plant growth through distinct signaling pathways in Selaginella and Arabidopsis. PLoS ONE 2013, 12, e81938. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.P.; Pandey, B.K.; Deveshwar, P.; Narnoliya, L.; Parida, S.K.; Giri, J. JAZ repressors: Potential involvement in nutrients deficiency response in rice and chickpea. Front. Plant Sci. 2015, 6, 975. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.D.; Ling, L.Z.; Yi, T.S. Evolution and divergence of SBP-box genes in land plants. BMC Genom. 2015, 16, 787. [Google Scholar] [CrossRef] [PubMed]
- Victoria, F.C.; Bervald, C.M.; daMaia, L.C.; deSousa, R.O.; Panaud, O.; deOliveira, A.C. Phylogenetic relationships and selective pressure on gene families related to iron homeostasis in land plants. Genome 2012, 55, 883–900. [Google Scholar] [CrossRef] [PubMed]
- Marshall, O.J. PerlPrimer: Cross-platform, graphical primer design for standard, bisulphite and real-time PCR. Bioinformatics 2004, 20, 2471–2472. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Zakrajsek, B.A.; Mills, A.G.; Gorn, V.; Singer, M.J.; Reed, M.W. Quantitative reverse transcription-polymerase chain reaction to study mRNA decay: Comparison of endpoint and real-time methods. Anal. Biochem. 2000, 285, 194–204. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.Y.; Huang, T.K.; Tseng, C.Y.; Lai, Y.S.; Lin, S.I.; Lin, W.Y.; Chen, J.W.; Chiou, T.J. PHO2-dependent degradation of PHO1 modulates phosphate homeostasis in Arabidopsis. Plant Cell 2012, 24, 2168–2183. [Google Scholar] [CrossRef] [PubMed]
- Florea, L.; Hartzell, G.; Zhang, Z.; Rubin, G. M.; Miller, W. A computer program for aligning a cDNA sequence with a genomic DNA sequence. Genome Res. 1998, 8, 967–974. [Google Scholar] [PubMed]
- Notredame, C.; Higgins, D.G.; Heringa, J. T-coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 2000, 302, 205–217. [Google Scholar] [CrossRef] [PubMed]
- Sali, A.; Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 1993, 234, 779–815. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.Y.; Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 2006, 15, 2507–2524. [Google Scholar] [CrossRef] [PubMed]
- Liberles, D.A. Evaluation of methods for determination of a reconstructed history of gene sequence evolution. Mol. Biol. Evol. 2001, 18, 2040–2047. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing. Available online: http://www.R-project.org/ (accessed on 10 September 2012).
- Warnes, G.R.; Bolker, B.; Bonebakker, L.; Gentleman, R.; Huber, W.; Liaw, W.; Lumley, T.; Maechler, M.; Magnusson, A.; Moeller, S.; et al. gplots: Various R programming tools for plotting data. Available online: http://CRAN.R-project.org/package=gplots (accessed on 12 January 2014).
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2009. [Google Scholar]





| Gene | Locus | EST | TM | SP | KD | Length (AA) | Localization | Int/Ext |
|---|---|---|---|---|---|---|---|---|
| GmBRI1a | Glyma06g15270 | Yes | 784..806 | 1..20 | 871..1143 | 1184 | Plas | 0/1 |
| GmBRI1b | Glyma04g39610 | Yes | 787..809 | 1..22 | 874..1146 | 1187 | Plas | 0/1 |
| GmBRL1a | Glyma04g12860 | Yes | 827..849 | 1..43 | 928..1202 | 1207 | Cyto | 0/1 |
| GmBRL1b | Glyma06g47870 | Yes | 843..865 | No | 912..1186 | 1211 | Plas | 0/1 |
| GmBRL2a | Glyma05g26771 | Yes | 752..774 | 1..32 | 756..1038 | 1053 | Nucl | 1/2 |
| GmBRL2b | Glyma08g09750 | Yes | no | 1..29 | 837..1121 | 1136 | Plas | 0/1 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peng, S.; Tao, P.; Xu, F.; Wu, A.; Huo, W.; Wang, J. Functional Characterization of Soybean Glyma04g39610 as a Brassinosteroid Receptor Gene and Evolutionary Analysis of Soybean Brassinosteroid Receptors. Int. J. Mol. Sci. 2016, 17, 897. https://doi.org/10.3390/ijms17060897
Peng S, Tao P, Xu F, Wu A, Huo W, Wang J. Functional Characterization of Soybean Glyma04g39610 as a Brassinosteroid Receptor Gene and Evolutionary Analysis of Soybean Brassinosteroid Receptors. International Journal of Molecular Sciences. 2016; 17(6):897. https://doi.org/10.3390/ijms17060897
Chicago/Turabian StylePeng, Suna, Ping Tao, Feng Xu, Aiping Wu, Weige Huo, and Jinxiang Wang. 2016. "Functional Characterization of Soybean Glyma04g39610 as a Brassinosteroid Receptor Gene and Evolutionary Analysis of Soybean Brassinosteroid Receptors" International Journal of Molecular Sciences 17, no. 6: 897. https://doi.org/10.3390/ijms17060897







